Abstract
SARS-CoV-2 (severe acute respiratory syndrome coronavirus 2) pandemic represents the primary public health concern nowadays, and great efforts are made worldwide for efficient management of this crisis. Considerable scientific progress was recorded regarding SARS-CoV-2 infection in terms of genomic structure, diagnostic tools, viral transmission, mechanism of viral infection, symptomatology, clinical impact, and complications, but these data evolve constantly. Up to date, neither an effective vaccine nor SARS-CoV-2 specific antiviral agents have been approved, but significant advances were enlisted in this direction by investigating repurposed approved drugs (ongoing clinical trials) or developing innovative antiviral drugs (preclinical and clinical studies). This review presents a thorough analysis of repurposed drug admitted for compassionate use from a chemical structure—biological activity perspective highlighting the ADME (absorption, distribution, metabolism, and excretion) properties and the toxicophore groups linked to potential adverse effects. A detailed pharmacological description of the novel potential anti-COVID-19 therapeutics was also included. In addition, a comprehensible overview of SARS-CoV-2 infection in terms of general description and structure, mechanism of viral infection, and clinical impact was portrayed.
1. Introduction
Since the beginning of the new decade of the 21st century—2020, humanity has been burdened by the emergence of a novel coronavirus known as SARS-CoV-2 (severe acute respiratory syndrome coronavirus 2), which caused a deadly outbreak of coronavirus disease (COVID-19) [1]. The first mention of this novel virus dates from the end of December 2019, and it is linked to a cluster of atypical pneumonia cases (27 cases) recorded in Wuhan, Hubei province, China [2,3]. The infection was declared a pandemic on 11 March 2020 by the World Health Organization (WHO) [4] and became an imperious public health concern.
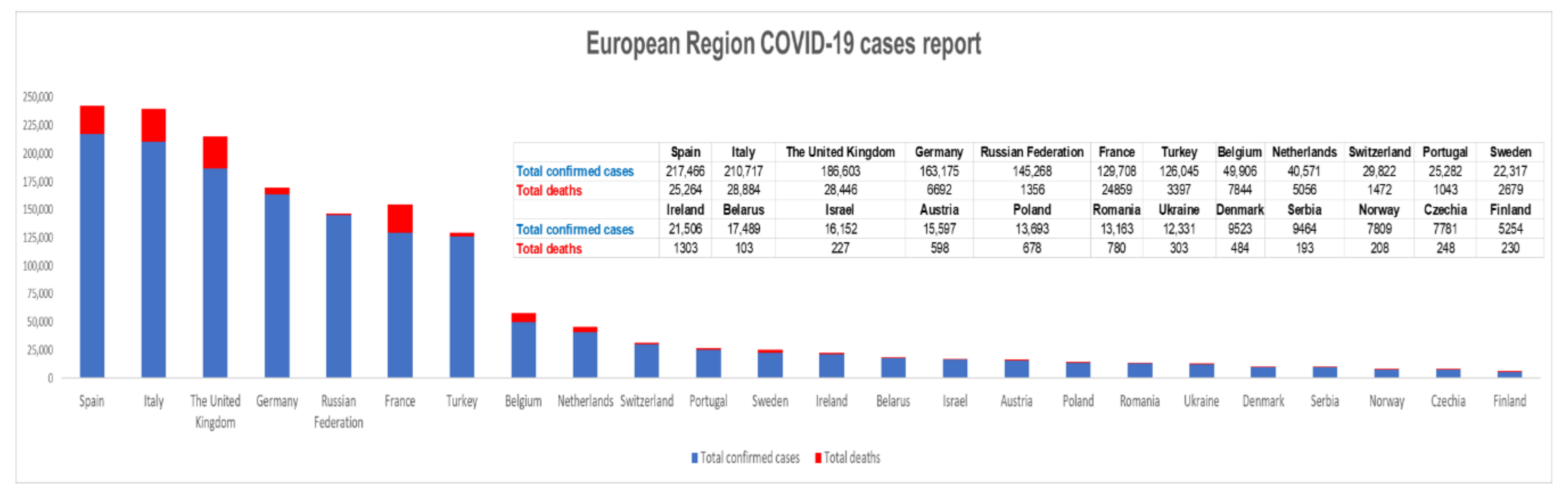
SARS-CoV-2 is the third highly pathogenic coronavirus that crossed the species barrier to cause fatal pneumonia in humans, after the SARS and MERS viruses causing the “severe acute respiratory syndrome” in 2002–2003 and the “Middle East respiratory syndrome” in 2012, respectively [1]. It has been reported that the SARS-CoV pandemic potentially led to up to 8000 cases of infection with an approximately 10% fatality rate in the early 2000s, while MERS-CoV produced over 1700 cases and an approximately 36% fatality rate later on [5]. Still, the recently discovered coronavirus elicits a greater rate of transmission, being already spread on all continents and encountering over 3,430,000 cases of infection up to the date of writing this article [6]. Therefore, to date of writing this article, the number of confirmed COVID-19 patients worldwide [7] was 3,435,894 with 239,604 reported deaths (Figure 1) [8] and, based on the trends observed in the last days, these numbers might be underestimated in the upcoming period.

Figure 1.
The European Region coronavirus disease (COVID-19) cases situation reported on 4 May 2020 according to World Health Organization (WHO) reports (in the graphic are presented only the countries with more than 5000 cases at that date) [8].
SARS-CoV-2 pandemic represents the topic in focus in these latter days, and although notable progress was recorded in gaining knowledge about this fatal disease, there still are multiple gaps to fill for a full perspective. This review plans to offer a comprehensible overview of SARS-CoV-2 infection in terms of general description and structure, mechanism of viral infection, clinical impact, and investigational anti-COVID-19 therapy.
3. Drugs Recommended in COVID-19 Therapeutic Guidelines
Up to date, neither an effective vaccine nor SARS-CoV-2-specific antiviral agents have been approved, and the management of coronavirus disease (COVID-19) focuses mainly on supportive care and symptomatic treatment [17,35,48].
Taking into consideration the global threat posed by the SARS-CoV-2 pandemic, imperative collaborative solutions were proposed in order to find a treatment against this unpredictable virus, such as the “SOLIDARITY” clinical trial for COVID-19 treatment, a study of great ampleness launched by WHO on 20th of March [49] and the scientific collaboration of a considerable number of researchers from different countries for the development of a COVID-19 vaccine [50]. At present, there are over 1300 studies recorded as clinical trials for COVID-19 in the ClinicalTrials.gov database [51], information that can also be accessed via the WHO Trial Registry Network on the International Clinical Trial Registry Platform (ICTRP) [52].
Considering the urgent need to find a treatment for COVID-19, two approaches were nominated: (i) a short term approach with immediate results—administration of approved drugs (originally introduced for other pathologies) with a high safety profile that showed promising in vitro results against SARS-CoV, MERS-CoV, and SARS-CoV-2 combined with convalescent plasma treatment (agents tested in ongoing clinical trials); and (ii) a long term approach—development of novel antiviral agents and vaccines against SARS-CoV-2.
On the basis of the literature data and the Chinese observational notes, interim COVID-19 treatment guidelines were created, guidelines that were adapted to each country and are constantly updated (Table 1) [53,54,55].

Table 1.
Interim coronavirus disease (COVID-19) therapeutic guidelines in different countries dependent on clinical stage.
The therapeutic drugs mentioned in the interim guidelines for COVID-19 treatment (Table 1) were also selected for the SOLIDARITY trial: remdesivir (RDV), an experimental antiviral; chloroquine (CQ), an antimalarial agent (or hydroxychloroquine—HCQ); a combination of lopinavir and ritonavir (LPV/r), an anti-HIV medication; and lopinavir + ritonavir + interferon-beta (an antiviral and modulator of the immune system that acts by suppressing pro-inflammatory cytokines) (see Table 2) [56].

Table 2.
Brief description of COVID-19 therapeutic options recommended by World Health Organization (WHO) guidelines [117,118,119,120].
3.1. Remdesivir—RDV (GS-5734)
Remdesivir (2-Ethylbutyl (2S)-2-{[(S)-{[(2R,3S,4R,5R)-5-(4aminopyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-dihydroxytetrahydrofuran-2-yl] methoxy}(phenoxy) phosphoryl] amino} propanoate) (Figure 5, see brief description in Table 2) is considered at this point the most promising treatment for SARS-CoV-2 infection on the basis of the latest outcomes acquired within the phase III clinical trials sponsored by the manufacturer [57] and the National Institute of Allergy and Infectious Diseases (NIAID) in the Adaptive COVID-19 Treatment Trial (ACTT) [58].
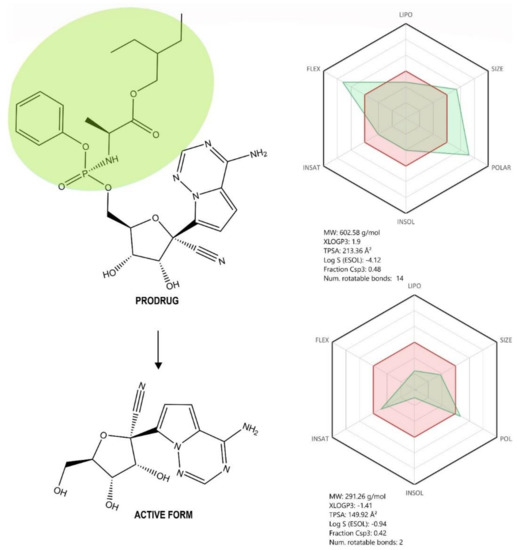
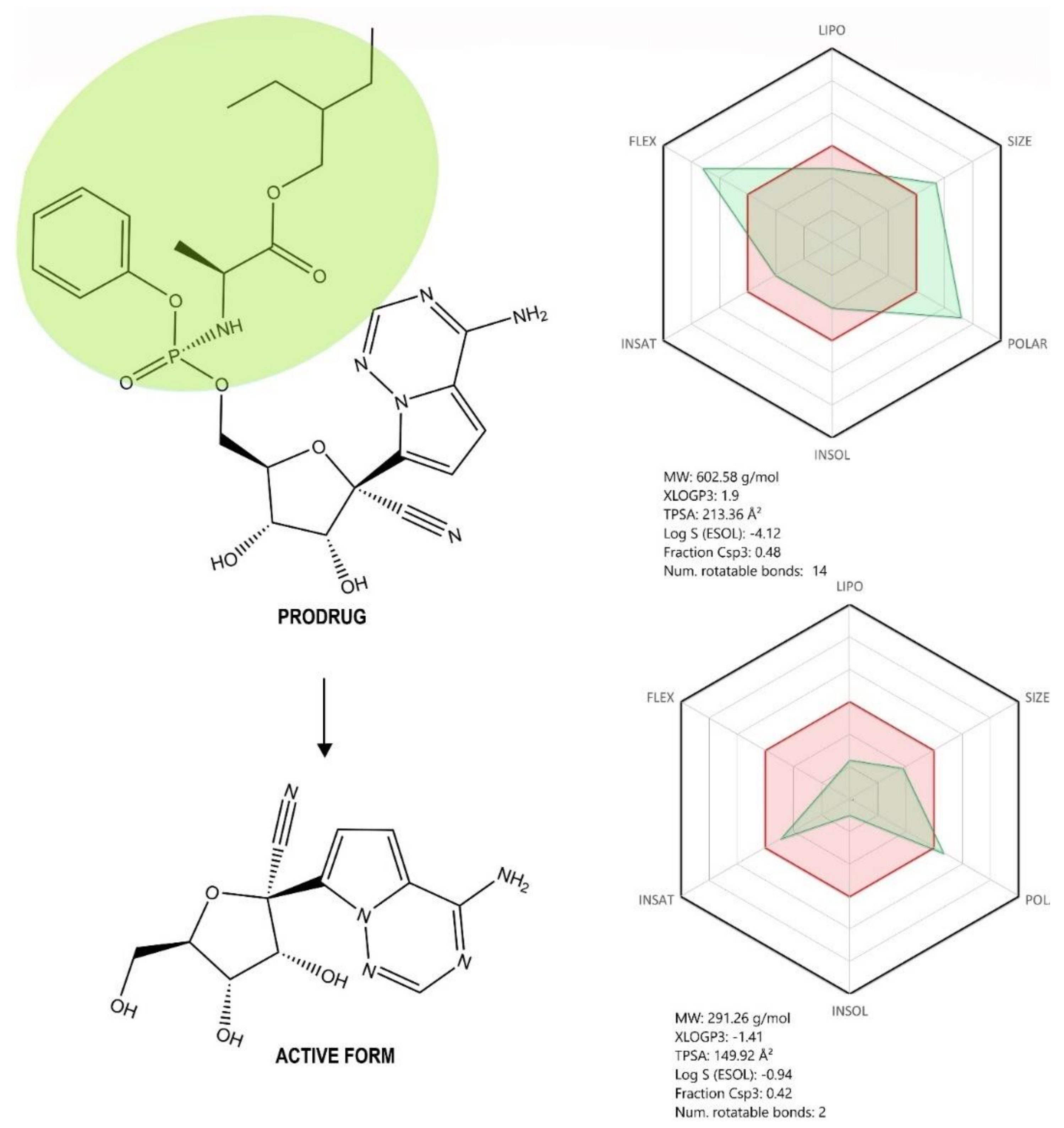
Figure 5.
Chemical structure of the prodrug Remdesivir (RDV) and its active form GS-441524. Absorption, distribution, metabolism, and excretion (ADME) profile was achieved using the free web tool SwissADME; the red highlighted area represents the suitable physicochemical space for oral bioavailability covering value intervals for LIPO (lipophility): −0.7 < XLOGP3 < +5.0, SIZE: 150 g/mol < MV < 500 g/mol, POLAR (polarity): 20Å2 < TPSA < 130Å2, INSOLU (insolubility): 0 < Log S (ESOL) < 6, INSATU (insaturation): 0.25 < Fraction Csp3 < 1, FLEX (flexibility): 0 < Num. rotatable bonds < 9, whereas the overlapped green highlighted area shows the calculated ADME profile for the molecule [63].
RDV is the monophosphoramidate prodrug of C-adenosine nucleoside analogue GS-441524 (1′-cyano 4-aza-7,9-dideazaadenosine C-nucleoside—a compound that is prescribed as therapy against infectious peritonitis in cats and felines, a disease determined by a feline coronavirus) [59,60]. This drug was developed by Gilead Sciences, Inc. as an antiviral candidate against the Ebola virus, but it also proved to be highly efficient in vitro against multiple RNA viruses, both negative sense, paramyxoviridae (parainfluenza type 3 virus, measles and mumps viruses, and nipah virus, among others) and pneumoviridae (respiratory syncytial virus), and positive sense viruses, coronaviridae (SARS-CoV, MERS-CoV, mouse hepatitis virus (MHV), and HCoV-OC43, among others) [60,61,62].
Structurally, RDV is a nucleoside analogue, which closely resembles AMP (adenosine monophosphate); important structural differences occur both in the nucleobase region, where the purine ring is replaced by a pyrrolo[2,1-f][1,2,4]triazine, and the ribose region, where a cyanide group is present in the fifth position. Moreover, the phosphate group is attached to a phenoxi group and an α-Alanine-2-ethyl-butyl ester, thus increasing the molecule’s lipophilicity as well as its degree of structural flexibility (Figure 5). Given its molecular shape, RDV acts as ATP (adenosine triphosphate) competitive inhibitor, targeting an enzyme involved in the viral genome replication, namely RNA-dependent RNA polymerase (RdRp) [31,48,61].
RDV is actually a prodrug metabolized intracellularly in different human cells [31,60,61] following the subsequent action of various enzymes (esterases and phosphoramidases), and is converted to the monophosphate form of RDV’s active metabolite, GS-441525, which is further phosphorylated by nucleoside-phosphate kinases to its active triphosphate form, interfering with the action of the viral RdRp of Ebola virus via insertion into the newly nascent viral RNA chain and disruption of the virus transcription [61,64,65,66]. RDV exerts a similar mechanism of action in coronaviruses by triggering the inhibition of RdRp complex and the premature termination of the viral RNA chain [31,48]. Moreover, RDV was able to bypass the proofreading 3′-5′ exoribonuclease (ExoN), a unique feature of CoVs, responsible for the decreased activity of various nucleoside analogues used as antivirals [59]. Recent studies identified the architecture of RdRp complex in SARS-CoV-2, the target site for RDV, an important step for a better understanding of the antiviral mechanism of action of RDV in SARS-CoV-2 infection [67,68,69].
In order to investigate whether this mechanism of action is also responsible for the antiviral effect on SARS-CoV-2, a recent study, using computational methods, constructed a homology model of SARS-CoV-2 NSP12 RdRp (nonstructural protein 12 of the RdRp complex) [70]. On the basis of this model, the authors assessed ATP/RDV binding with SARS-CoV-2 NSP12 by means of molecular docking, using a previous co-crystal structure of the poliovirus RdRp. The study showed that the difference between the calculated binding energy of RDV triphosphate and that of ATP was significantly large, concluding that ATP can be blocked out of the binding site when RDV triphosphate is locally present, deeming that RDV could act as an effective SARS-CoV-2 RNA-chain terminator, stopping its RNA reproduction [70].
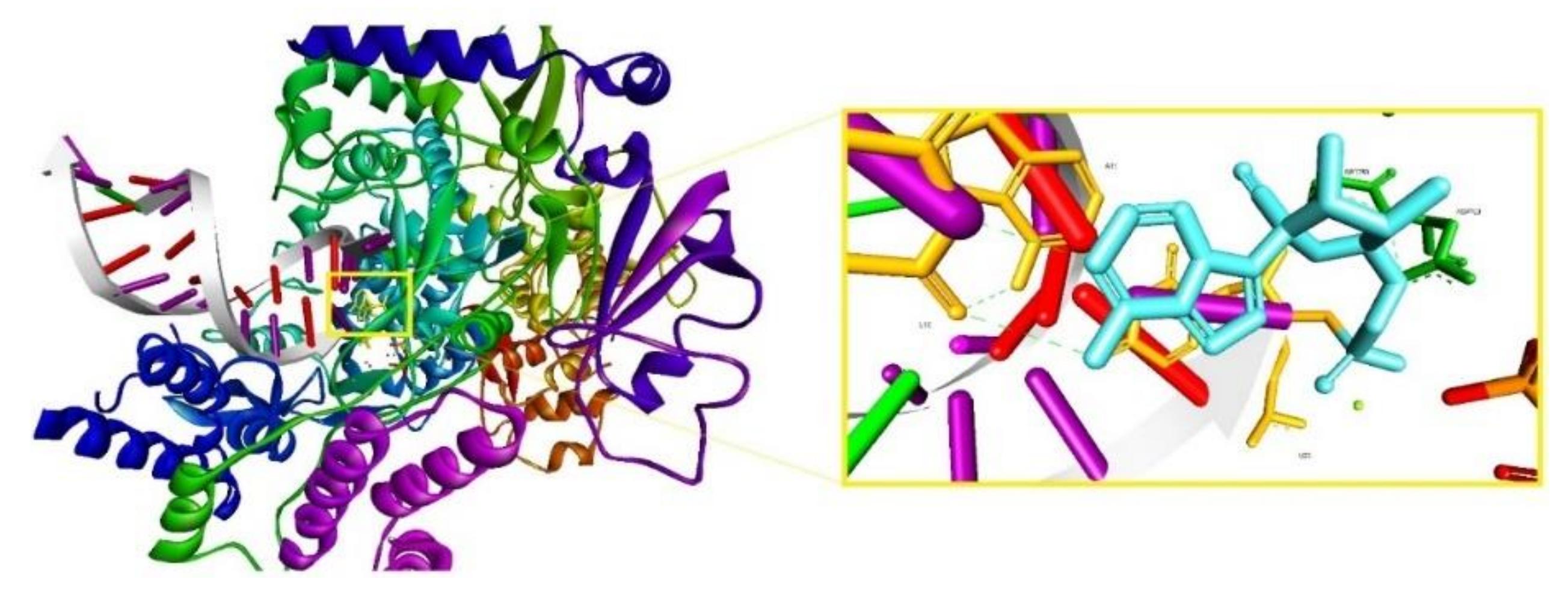
Following the same research direction, to evaluate the mechanism of action of RDV, a study published the 3D cryo-EM structure of SARS-CoV-2 RdRp in the apo form at 2.8 Å resolution and in complex with a template-primer RNA (50-base long) and RDV at 2.5 Å resolution [69]. According to this study, the characterized complex structure revealed that the partial double-stranded RNA template was inserted into the central channel of the RdRp, where RDV is covalently incorporated into the primer strand at the first replicated base pair, inhibiting the RdRp via non-obligated RNA chain termination. The complex structure is depicted in Figure 6 and is available in the RCSB Protein Data Base under the ID 7BV2 [69]. According to the authors, the structure comparison and sequence alignment analysis carried out on the obtained complex suggest that RNA recognition and RDV inhibition of RdRp is highly preserved in various RNA viruses, providing a basis for broad spectrum nucleotide analog antiviral-based drug design [69].
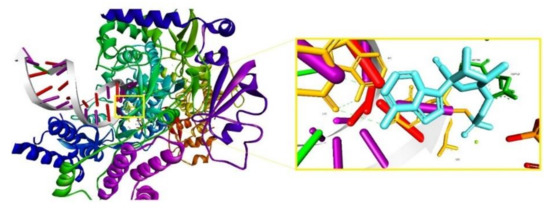
Figure 6.
The nsp12-nsp7-nsp8 complex bound to the template-primer RNA and triphosphate form of RDV, Protein Data Base (PDB) ID: 7BV2 (left); RDV covalently bound to U20; hydrogen bonds are depicted as light green dotted lines, amino acid residues in dark green, and nucleotides in orange.
At the same time, in line with the reported findings related to RDV’s antiviral mechanism of action described by computational methods, several studies aimed to validate the presumed antiviral effect of RDV on SARS-CoV-2 through biological assessment.
Considerable proof has been gathered lately supporting the effectiveness of RDV against SARS-CoV-2 virus: (i) in vitro, Wang et al. proved that RDV efficiently inhibited the viral infection at low-micromolar concentrations in two different cell lines (Vero E6 and Huh-7) with a notable half-maximal effective concentration (EC50) value of 0.77 μM [56,61]; (ii) in vivo studies on animal models of SARS-CoV and MERS-CoV confirmed the RDV antiviral potential by reducing clinical symptoms of the infection [31]; and (iii) in clinical efficacy trials of RDV on severe COVID-19 patients, an improved clinical outcome was noticed, but several adverse reactions observed in the RDV-treated group were also reported [71,72].
Phase I, phase II, and phase III clinical trials on healthy volunteers and patients with Ebola virus infection asserted the pharmacokinetic properties and a safety profile of this compound (high human tolerance, no cytotoxicity, hepato- or renal toxicities, and no/reduced severe adverse effects were detected) [31].
Currently, RDV is tested in multiple ongoing phase 3 clinical trials for COVID-19 treatment (NCT04252664, NCT04257656, NCT04292730, NCT04292899, NCT04280705, Solidarity trial (WHO); DisCoVeRy trial (INSERM) in Belgium, and so on) and The European Medicine Agency (EMA) adopted Article 83, which provides recommendations on compassionate use of RDV for COVID-19 treatment in the European Union [73].
The manufacturer of Remdesivir, Gilead Sciences, Inc., sponsored two randomized, open-label, multi-center phase 3 clinical trials (also known as SIMPLE studies) that were developed in countries with a large number of cases: (1) SIMPLE trial 1 was developed on severe COVID-19 patients in order to assess the effectiveness and the safety of a 5-day versus 10-day treatment with RDV (first dose—200 mg/day intravenously administration, followed by 100 mg/day for 4 or 9 days, respectively) in addition to standard care, and (2) SIMPLE trial 2 performed on patients with moderate COVID-19 symptomatology aiming to evaluate the effectiveness and the safety of a 5-day or 10-day treatment with RDV versus standard care—the results will be available by the end of May [57].
On 29 April 2020, Gilead Sciences, Inc. provided in a press release the preliminary results obtained within SIMPLE trial 1 on severe COVID-19 patients, as follows: (i) a similar clinical improved outcome after a 5-day or 10-day treatment with RDV; (ii) an earlier start of RDV treatment (within 10 days of symptoms onset) reduces the hospitalization duration; and (iii) a high tolerance of RDV treatment (in both experimental settings: 5-day or 10-day treatment) in most patients, but some adverse effects occasionally occurred, such as nausea (10% of the patients), acute respiratory failure (a higher percentage in the 10-day group: 10.7% versus 6%), elevated liver enzymes (ALT) values (7.3% of the patients), and stopping the treatment in 3% of the patients owing to liver damage (high enzymes values) [57].
On 1 June 2020, Gilead Sciences, Inc. announced the results obtained from SIMPLE trial 2, as follows: (i) the 5-day treatment with RDV led to a significant improved clinical status in patients with moderate COVID-19 symptomatology at day 11 as compared with standard care group (in 65% of patients); and (ii) in terms of safety and adverse effects noticed, RDV treatment was well tolerated and the most common adverse effects observed were nausea (10% of the patients), diarrhea (5%), and headache (5% of the patients), and no deaths were recorded as compared with the standard care group that enlisted four deaths [74]. In addition, since 7 May 2020, Remdesivir (Veklury®) was approved in Japan as a treatment for patients with severe COVID-19 pathology [75].
Positive results regarding the effectiveness of RDV as a treatment for COVID-19 patients were also achieved in the Adaptive COVID-19 Treatment Trial (ACTT) financed by the National Institute of Allergy and Infectious Diseases (NIAID—part of the National Institutes of Health), a clinical trial that included 1063 patients: (i) patients treated with RDV (10 days—first day 200 mg/day intravenously, followed by 100 mg/day for 9 days) recovered faster (31%) as compared with patients that received placebo; in addition, the mortality rate was lower in the RDV-treated group compared with placebo (8% versus 11.6%, respectively) [57]. The preliminary report results of the ACTT study were published by Beigel et al. in The New England Journal of Medicine on 22 May 2020 [76]. On the basis of these preliminary promising data, European Medicine Agency (EMA) initiated the “rolling review” for RDV, resulting in the acceleration of the assessment process of RDV for marketing authorization [77], while the FDA (Food and Drug Administration) authorized RDV for emergency use as a treatment for hospitalized COVID-19 patients [78].
These latest data regarding RDV efficacy against SARS-CoV-2 infection are stimulating, although the gaps concerning its safety profile are rather wide at present, and the forthcoming results from the ongoing clinical trials are essential to fill them. On the basis of these considerations, it is recommended that, during the treatment, the physicians should be well-aware of the considerable number of factors/conditions (mainly in the case of severely ill patients with comorbidities—diabetes, cardiovascular pathology, and aged people) that might interfere with this compound and lead to adverse events. To prevent or to minimize any kind of undesired effects that might aggravate patients’ status, some key information about RDV pharmacokinetics, pharmacodynamics, and drug-drug interactions should be reviewed, data that will be provided in the following paragraphs.
According to the summary on compassionate use of RDV, this drug is formulated in two pharmaceutical forms (a solution—5 mg/mL and a lyophilized formulation—100 mg RDV powder) and is recommended to be administered intravenously (30–120 min) after reconstitution in 0.9% saline or 5% glucose solutions, with the therapeutic dosage being as follows: 200 mg on day 1 and 100 mg/day for the following 9 days. The recommended formulation of RDV for compassionate use is the lyophilized powder that must be reconstituted prior to use and administered intravenously, as stated above [73,79].
RDV is conditioned as a prodrug of the nucleoside analog GS-441524 and displays the following pharmacokinetic profile: 100% absorption after iv. administration (after oral administration RDV suffers an almost complete first-pass hepatic clearance), distribution as a free fraction in a proportion of 12.1% (moderate affinity for plasmatic proteins binding), and t1/2 (plasmatic half-life) of approximatively 1 h. Given its nucleotide analog structural features and high lipophilicity compared with that of the active form (Figure 5), RDV undergoes intense hepatic metabolization by cytochrome P450 enzymes (CYP2C8, CYP2D6, and CYP3A4) and is predominantly excreted in the urine as more hydrosoluble metabolites. The potential for interactions with other drugs is considered to be low for RDV (inhibitor of CYP3A4 and OATPs hepatic uptake transporter, according to several in vitro studies; no in vivo studies were performed) but is currently unknown for its metabolites.
In terms of the safety profile, it can be stated that this aspect is incompletely characterized, although several data were provided by the studies conducted by the manufacturer regarding the possible side effects (single doses ranging from 3 to 225 mg; multiple doses of 150 mg/daily for 7 to 14 days), such as phlebitis, constipation, headache, ecchymosis, nausea, pain in the extremities, as well as an increase of the hepatic enzymes values. The augmented values of hepatic transaminases are the only adverse effects directly associated with RDV according to manufacturer data; hypersensitivity reactions and renal events are also recommended to be monitored [79].
Nevertheless, considering that RDV as well as its active metabolite are both ATP competitors, potential future side effects may emerge owing to the fact that multiple kinase (which all have ATP binding sites) functions could be altered by possible RDV inhibition.
In addition to the side effects mentioned in the summary on compassionate use of RDV, a cohort study conducted by Grein and collaborators on severe hospitalized COVID-19 patients treated with RDV for 10 days (the dosage recommended for compassionate use: 200 mg on day 1 and 100 mg/day on day 2 to day 10) and supportive care reported other adverse effects, such as increased hepatic enzymes (most common), diarrhea, rash, renal disturbance, hypotension, acute kidney injury, atrial fibrillation, multiple-organ dysfunction syndrome, hypernatremia, deep-vein thrombosis, acute respiratory distress syndrome, pneumothorax, hematuria, delirium, septic shock, and pyrexia (most of these effects were noticed in patients on invasive ventilation). Taking into consideration the limitations of the study (small cohort size, a relatively short duration of the follow-up, lack of a randomized control group) and the paucity of information regarding COVID-19, it is questionable if these side effects are directly related to RDV administration or are in fact consequences/complications determined by the SARS-CoV-2 infection [71].
One of the randomized, double-blind, placebo-controlled, multicentre clinical trials developed in China (NCT04257656) in order to verify RDV efficacy as a COVID-19 treatment (the dosage recommended for compassionate use was applied: 200 mg on day 1 followed by 100 mg/daily from day 2 to day 10 co-administered with lopinavir-ritonavir/interferons/corticosteroids) indicated an improved outcome in patients treated with RDV, with a decreased duration of invasive mechanical ventilation; however, these benefits were not statistically significant. Several adverse effects were recorded within the RDV-treated group and were classified as follows: most common—constipation, hypoalbuminemia, hypokalemia, thrombocytopenia, anemia, and an augmented total bilirubin; common—increased blood glucose, rash, augmented blood lipids, high white blood cell count, hyperlipidemia, increased neutrophils number, high blood urea nitrogen, augmented aspartate aminotransferase, nausea, diarrhea, reduced sodium level, and increased serum potassium, as well as rare severe adverse events as respiratory failure, acute respiratory distress syndrome, and cardiopulmonary failure. Even though a plethora of side-effects in the RDV-treated group were reported, these reactions could not be directly correlated to RDV, and the conclusion of the authors in terms of RDV safety profile was that this drug “was adequately tolerated and no new safety concerns were identified”. Moreover, the average proportion of severe adverse events was lower in the RDV-treated group compared with the placebo group [72].
The in vitro metabolism studies (for the identification of drug interactions) for RDV revealed an inhibitory effect on CYP3A4 enzyme; as this enzyme catalyzes multiple drug metabolism pathways, it is important to determine if RDV could interfere with these drugs and initiate different side effects. An ample study in this direction was conducted by the Liverpool Drug Interactions Group (University of Liverpool), and revealed the following findings: (i) co-administration of RDV with drugs from different classes as analgesics (metamizole), antibacterials (rifabutin), anticonvulsants (eslicarbazepine, oxcarbazepine, rufinamide), anti-hypertensives for pulmonary hypertension (bosentan), and steroids (betamethasone, dexamethasone) can lead to potential interaction that might require an adjustment of the RDV dose or a close patient monitoring; and (ii) co-administration of RDV with drugs from the classes of antibacterials (rifampicin, rifapentine), anticonvulsants (carbamazepine, phenobarbital, phenytoin, primidone), and antidepressants (St John’s wort) is not recommended.
Key information is represented by the lack of clinically significant interaction between RDV and other classes of drugs used as comedication for COVID-19 patients as analgesics, antiarrhythmics, antibacterials, anticoagulants, anti-platelets and fibrinolytics, antidiabetics, antifungals, anti-hypertensives (ACE inhibitors, angiotensin antagonists, diuretics, other agents), neuroleptics/antipsychotics, other antivirals for COVID-19 treatment, anxiolytics/hypnotics/sedatives, beta blockers, bronchodilators, calcium channel blockers, contraceptives, gastrointestinal agents, anti-emetics, hormone replacement therapy, immunosuppressants, inotropes and vasopressors, lipid lowering agents, and steroids [80].
In view of all the data stated above about RDV, several recommendations and cautions are suggested:
- The duration of RDV treatment should be settled at 5 days (200 mg day 1 followed by 100 mg/daily for the following 4 days) in severe COVID-19 cases—on the basis of the results obtained within SIMPLE trial 1, this measure represents a major step in reducing the risk of drug–drug interactions/side-effects/complications/aggravation of patient status;
- As the RDV directly-linked adverse effect is hepatic injury, a possible preventive approach would consist of the co-administration of hepatoprotective medication (for example, essentials phospholipids or albumin solution—it should be a patient-dependent decision after analyzing the risk/benefit ratio) during the RDV treatment (in severely ill COVID-19 patients, liver dysfunction was reported, but until now, it is unclear what causes it—the medication administered or the SARS-CoV-2 infection, thus liver protection would do no harm);
- As RDV is predominantly eliminated through urine (mostly as metabolites), the renal function should be monitored during the treatment and, in the case of renal impairment, the treatment should be stopped
- Co-administration of other anti-COVID-19 drugs (lopinavir/ritonavir, chloroquine, hydroxychloroquine, interferon) is possible because no drug–drug interactions were reported; surveillance is recommended
- Co-administration of antidiabetics and cardiovascular medication (anti-hypertensives, anticoagulants, beta blockers, and so on) is possible because no drug–drug interactions were reported; surveillance is recommended
- Co-administration of RDV with other hepatotoxic drugs is not recommended
- To remedy/to prevent the mild adverse reactions reported in the clinical trials, such as nausea, diarrhea, hyperlipidemia, and so on, symptomatic medication could be administered during RDV treatment because no interactions were described between these classes of drugs and RDV; strict surveillance is recommended
- To follow the updates on other clinical trials that tested RDV.
Every medical decision regarding the treatment of COVID-19 patients should be taken after a thorough analysis of the risk/benefit ratio and all adverse events should be reported in order to fill the gaps in the RDV toxicological profile.
3.2. Chloroquine and Hydroxychloroquine
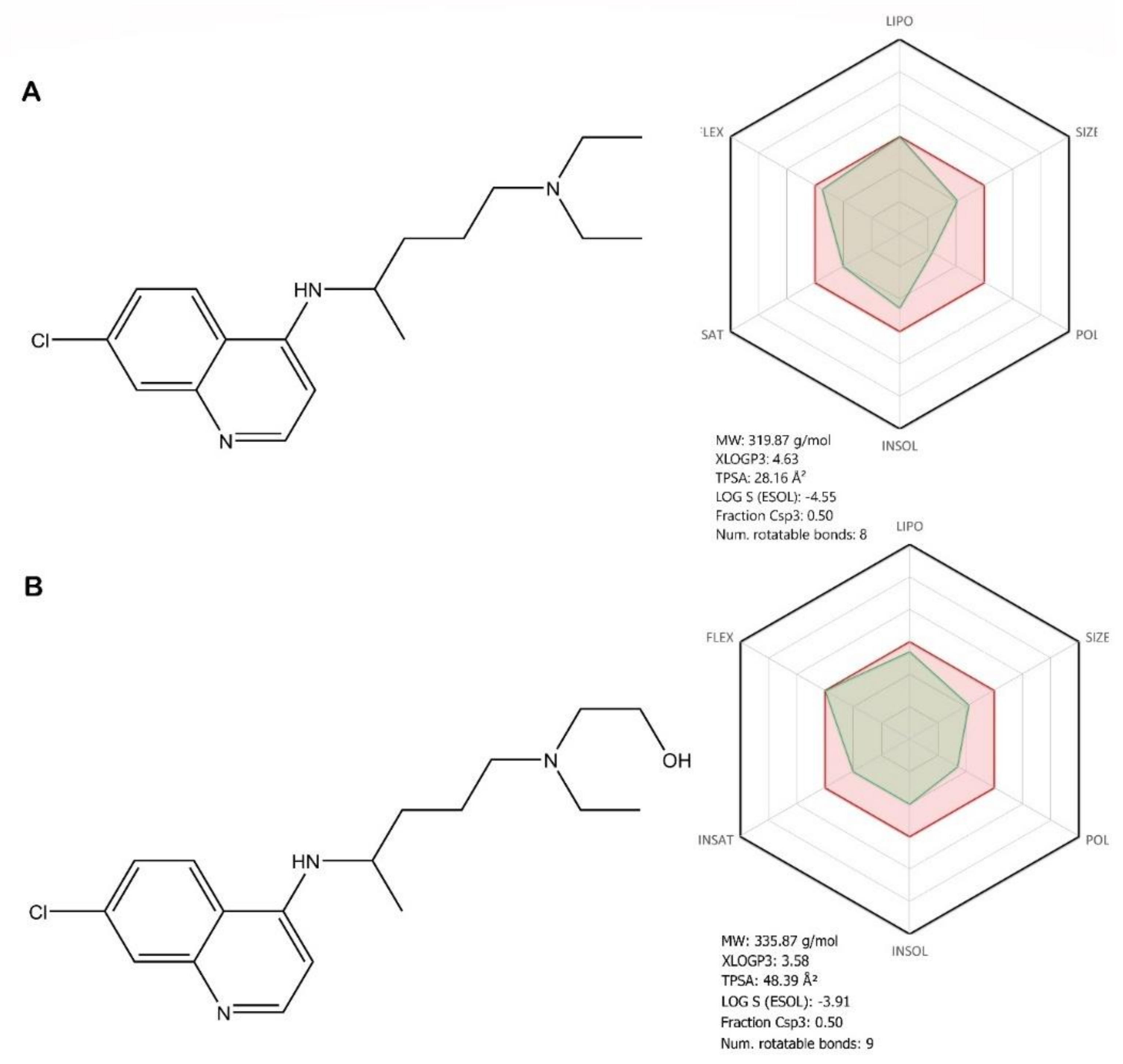
Chloroquine (CQ—Figure 7A, see brief description in Table 2), a 70-year-old antimalaria drug, currently one of the agents for amoebiasis and other protozoal diseases and antimalarials associated with irreversible retinal damage and life-threatening and fatal cardiomyopathy, has been recently reported as a potential broad-spectrum antiviral drug [61,81].
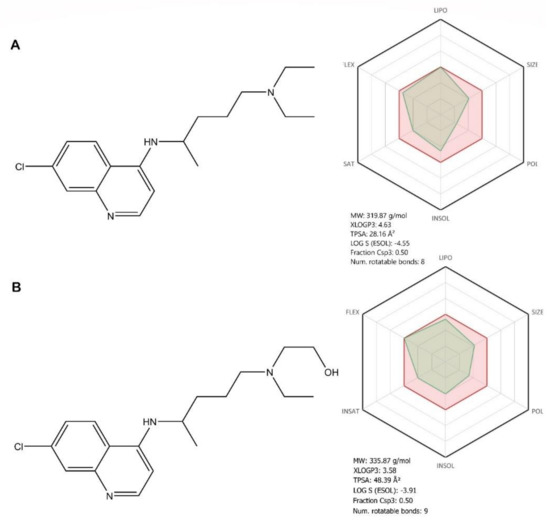
Figure 7.
Chemical structure of chloroquine (A) and hydroxychloroquine (B) ADME profile was achieved using the free web tool SwissADME; the red highlighted area represents the suitable physicochemical space for oral bioavailability, covering value intervals for the following: LIPO (lipophility): −0.7 < XLOGP3 < +5.0, SIZE: 150 g/mol < MV < 500 g/mol, POLAR (polarity): 20Å2 < TPSA < 130Å2, INSOLU (insolubility): 0 < Log S (ESOL) < 6, INSATU (insaturation): 0.25 < Fraction Csp3 < 1, FLEX (flexibility): 0 < Num. rotatable bonds < 9, whereas the overlapped green highlighted area shows the calculated ADME profile for the molecule [63].
Hydroxychloroquine (HCQ—Figure 7B, see brief description in Table 2) is a chloroquine analogue [82], one of the antimalarials and other anti-inflammatory and antirheumatic agents associated with ocular toxicity and cardiomyopathy, is currently recommended in the treatment of immune diseases, such as systemic lupus erythematosus and rheumatoid arthritis [83]. HCQ has been used for the last 30 years to treat intracellular bacterium Coxiella burnetii, which produces the Q fever, being the only effective agent that kills intracellular pathogens. Another important therapeutic activity is against the intracellular bacterium Tropheryma wippley [84].
Structurally, both compounds are 7-chloro-quinoline derivatives, with a novaldiamine substituent in the fourth position, where HCQ has an additional hydroxyl group grafted at the end of the chain. In terms of bioavailability, the additional hydroxyl group, according to the absorption, distribution, metabolism, and excretion (ADME) profile of HCQ, leads, as expected, to an improvement in hydrosolubility, flexibility, and polarity, as well as to a decrease in lipophilicity compared with CQ (Figure 7). These differences in the ADME profiles of the two molecules, owing to the presence of the hydroxyl group, can lead to different pharmacological behaviors, in terms of therapeutic efficacy and outcome, but also in the occurrence of toxic effects. These pharmacokinetic aspects will be further discussed.
The reposition of CQ and HCQ as antiviral candidates for COVID-19 treatment was based on several in vitro and in vivo studies that reported their therapeutic effect against several coronaviruses, such as human OC43, SARS-CoV, and MERS-CoV [81,85]. The molecular mechanism of action of CQ and HCQ has not been fully elucidated yet [82].
Previous studies have explored the mechanism of the antiviral action of CQ against SARS-CoV. The authors concluded that a possible mode of action of the drug, in post-infection treatment, would be the increase of the endosomal pH value, owing to the presence of the three nitrogen atoms within the CQ molecule, which give its basic properties, leading to abrogation of virus-endosome fusion [86,87]. These findings also suggest that pre-infection treatment with CQ is responsible for cell surface expression of under-glycosylated ACE2, leading to a decrease in the viral spike protein–cell receptor affinity [86]. In line with these reports, a recent study also showed that CQ/HCQ induced pH elevation, within acidic intracellular organelles, such as early endosomes (EEs) and endolysosomes (ELs), and also caused the disruption of SARS-CoV-2 transport between EE and EL [83]—a phase that seems to be required in viral genome release, in SARS-like coronavirus infections [88].
At the same time, regarding the molecular mechanism of action of CQ/HCQ, the possibility to find a potential target, until experimental validation, was the subject of in silico determinations. Wu et al. screened two compound libraries (ZINC and a natural compound library of their own) against 19 SARS-CoV-2 protein targets. Among the results, the authors showed that chloroquine can target nonstructural proteins such as Nsp3b, exhibiting suitable docking scores [89].
In terms of anti-inflammatory and immunomodulatory activities, a number of mechanisms have been proposed, for both CQ and HCQ, which involve the following: decreased cytokine production, suppression of the immune effector cells and platelet function, protection of the cell surface from external disorders, competitive binding to nucleic acid ligands or toll-like receptors (TLRs), interference with lysosomal function, reduction of lysosomal enzyme leakage, and interference with endosomal NADPH oxidase (NOX) [90].
Depending on their activity against SARS-CoV-2, the possible mechanisms of action can be divided into two main categories: (1) inhibition of viral enzymes/processes (viral DNA and/or RNA polymerase), glycosylation of viral proteins, assemblage of virus, new virus particle transport, and virus release; and (2) ACE2 cellular receptor inhibition, acidification at the surface of the cell membrane thus inhibiting virus fusion, and immunomodulation of cytokine release [90].
The promising in vitro results of CQ and HCQ against CoVs led to an early clinical interest in the use of these two compounds as therapy for COVID-19 and multiple clinical trials (over 50, most of them evaluating the HCQ effects [51], were set in motion [91]). The data obtained from the clinical studies (final results or pre-print texts) present methodological flaws [91] and are inconclusive: (i) an improved clinical outcome was observed in HCQ-treated group, but was not statistically relevant [92]; (ii) coadministration of HCQ with azithromycin determined the decrease of viral load in COVID-19 patients [93]; (iii) CQ inhibited pneumonia exacerbation and shortened the infection course (improved lung imaging and increased viral clearance) [94]; (iv) HCQ proved to be better in terms of efficacy compared with CQ [95,96]; and (v) HCQ apparently offered no protection against SARS-CoV-2 infection (results of a large healthcare Israelian database analysis) [97]. The association of HCQ and azithromycin for the treatment COVID-19 patients was based on several premises: azithromycin proved in vitro activity against Ebola and Zika viruses, as well as a preventive effect against severe respiratory tract infections to patients presenting viral infection [93], but further studies confirmed the efficacy of HCQ and azithromycin combination [98].
CQ and HCQ are also evaluated in three ample clinical trials; that is, the SOLIDARITY trial sponsored by WHO that evaluates these compounds as possible therapy against SARS-CoV-2 infection; the HERO-HCQ, a study sponsored by the National Institutes of Health (NIH) that assesses their preventive potential; and the DisCoVeRy trial, launched by INSERM (French Institut National de la Santé Et de la Recherche Médicale) [99,100]. Even though the two compounds share similar chemical structures, it has been reported that HCQ presents some therapeutic benefits when compared with CQ such as a lower toxicity in animals [83]. It is important to note that both CQ and HCQ interfere with the cytochrome P450 isoenzymes and drug transporters: CYP2C8, CYP2D6, CYP3A4, P-gp being primarily metabolized by CYP2C8 and CYP3A4, and known inhibitors of the drug transporter P-glycoprotein (P-gp), thus explaining the interactions (increasing/decreasing) with associated antiviral drugs. Both CQ and HCQ show favorable pharmacokinetic properties: efficient oral absorption and tissue distribution patterns, leading to high concentrations in the liver, spleen, kidney, and lungs for CQ and the bone marrow, liver, kidneys, lungs, adrenal gland, and pituitary gland for HCQ. It should be noted that melanin-containing cells bind strongly to the chloroquine process, explaining the retinal toxicity of both compounds [82,90].
Moreover, given the fact that CQ and HCQ are both quinine/quinidine analogs, one major concern in the therapeutically use of these drugs is related to the QTc interval prolongation and the increased incidence of ventricular arrhythmias [101].
From a toxicological point of view, both substances show significant adverse reactions and drug–drug interactions, as follows: most of the antiarrhythmics (amiodarone, bepridil, flecainide, mexiletine) are not recommended to be coadministered with CQ and HCQ owing to the increased risk of QT development; several antibacterials (rifampicin, rifapentine) and anticonvulsants (phenobarbital, phenytoin, primidone, carbamazepine) are also not recommended as co-medication.
The most common adverse effects associated with CQ and HCQ use are gastrointestinal disorders, hypoglycemia, and QT interval prolongation (after short term treatment), as well as cardiomyopathy and retinal toxicity (after long-term treatment) [102]. Several concerns were raised regarding the safety profile of HCQ administered for COVID-19 treatment (the doses recommended in several guidelines for COVID-19 treatment—800 mg/day on day 1, followed by 400 mg/day for 4 days —are higher as compared with those administered conventionally for chronic therapies) in terms of cardiac toxicity and retinopathy [101,103]. Moreover, a retrospective study conducted on 95 hospitalized COVID-19 patients treated with CQ showed that 23% of the patients presented QT interval prolongation [104]. A case report stated that CQ treatment induced Torsade de Pointes in a COVID-19 female patient [105].
Coadministration of CQ and HCQ with the other investigational anti-COVID-19 agents (LPV/r, atazanavir) should be performed under strict monitoring, as LPV/r increases their concentrations and, subsequently, the risk for adverse effects. Moreover, when the coadministration is decided, ECG (electrocardiogram) monitoring is required, and the treatment period should be as short as possible. No interactions were reported so far between RDV and CQ or HCQ; therefore, their coadministration can be considered safe, at least until further notice [80].
The current preclinical and clinical data regarding the use of CQ and HCQ in SARS-CoV-2 infection are not considered robust [102], and EMA did not approve chloroquine for standard use in COVID-19 pathology and restricted its use exclusively in clinical trials or through national emergency use programs [101,106].
At present, there are more than 200 ongoing clinical studies enlisted regarding the use of CQ/HCQ alone or in combination with macrolides (azithromycin or clarithromycin) as treatment or as pre- or post-exposure prophylaxis in COVID-19 patients. Although positive results were obtained in small size clinical studies, the recent data collected from larger size studies indicate that these drugs have no significant benefit in clinical outcome; moreover, their association with macrolides might augment the risk of cardiac adverse effects (arrhythmia) and mortality of hospitalized patients [98].
In addition, the latest update provided by WHO on 17 June 2020 regarding the assessment of hydroxychloroquine as a treatment for COVID-19 infection notifies that this part of the SOLIDARITY trial was suspended based on the evidence gathered according to which hydroxychloroquine had no impact on decreasing the mortality of hospitalized COVID-19 patients as compared with standard care. Similar results were furnished by the United Kingdom’s Recovery Trial, DisCoVeRy trial, and a Cochrane review, which also stopped their study [107].
3.3. Lopinavir/Ritonavir
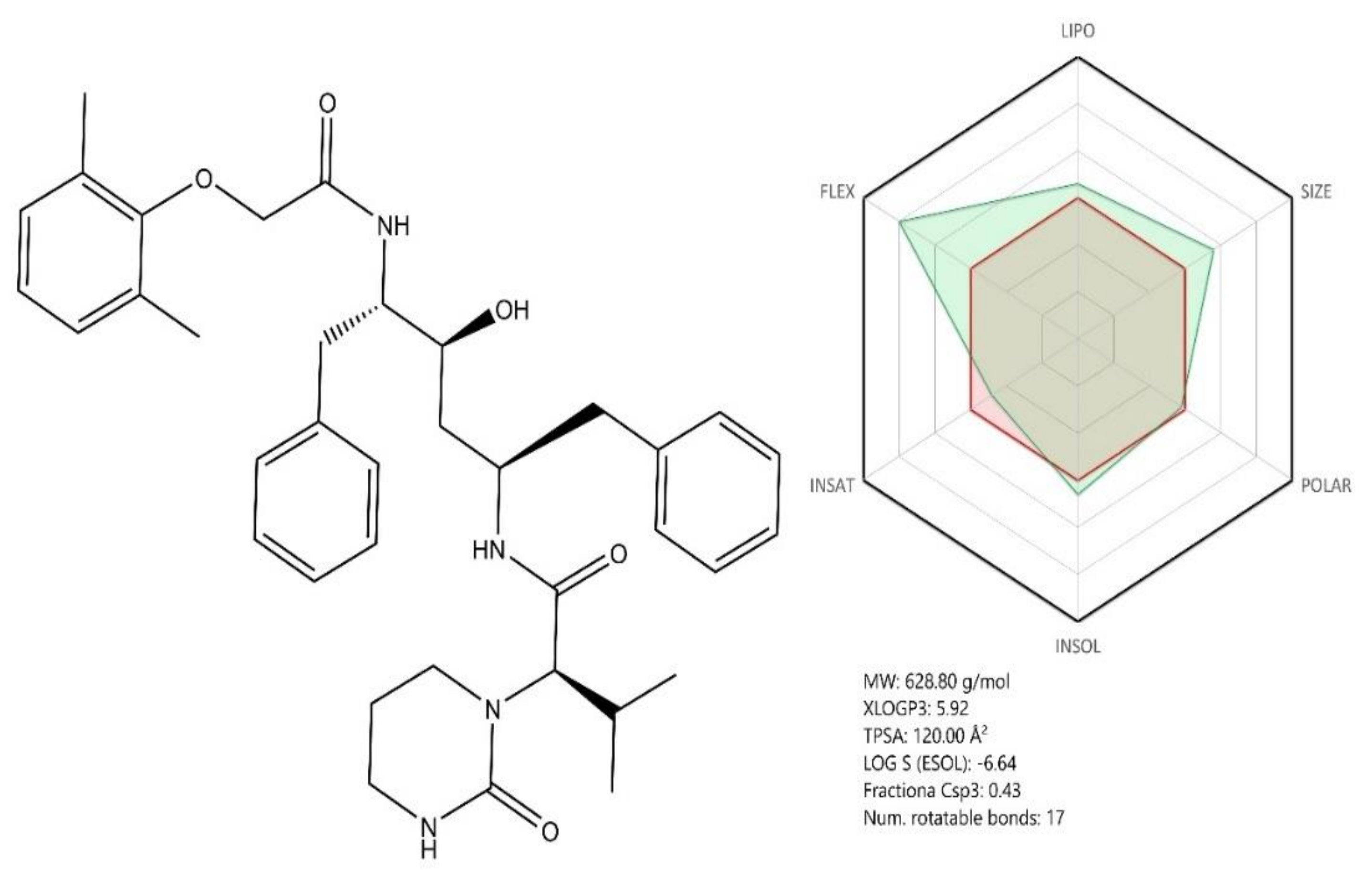
Lopinavir and ritonavir (LPV/r) (Figure 8, see brief description in Table 2) are two antiviral drugs acting as viral protease inhibitors, currently used against HIV (human immunodeficiency virus) infection [108]. Structurally, the two compounds are 1,6-diphenyl-4-hydroxy-2,5-diaminohexane derivatives whose amino groups contain side chains with various substituted cyclic moieties such as phenoxy/3,5-diazinan (Lopinavir) or 1,3-thiazole (Figure 8). Given their molecular structure, both compounds share similar pharmacokinetic profiles based on their calculated ADME profile (Figure 8 and Figure 9). Previous studies showed that the combination of the two compounds had a beneficial effect against CoVs infections such as SARS-CoV, which triggered the investigation of their potential antiviral effect against SARS-CoV-2 as well [109].
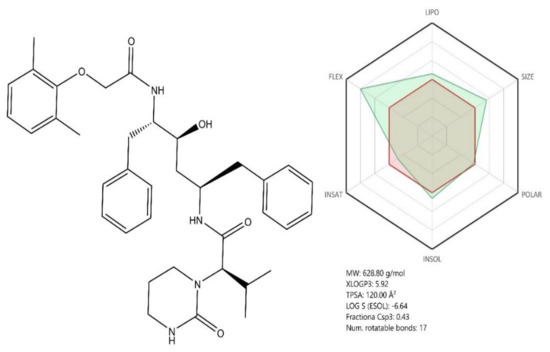
Figure 8.
Chemical structure of Lopinavir; ADME profile was achieved using the free web tool SwissADME; the red highlighted area represents the suitable physicochemical space for oral bioavailability covering value intervals for the following: LIPO (lipophility): −0.7 < XLOGP3 < +5.0, SIZE: 150 g/mol < MV < 500 g/mol, POLAR (polarity): 20Å2 < TPSA < 130Å2, INSOLU (insolubility): 0 < Log S (ESOL) < 6, INSATU (insaturation): 0.25 < Fraction Csp3 < 1, FLEX (flexibility): 0 < Num. rotatable bonds < 9, whereas the overlapped green highlighted area shows the calculated ADME profile for the molecule [63].
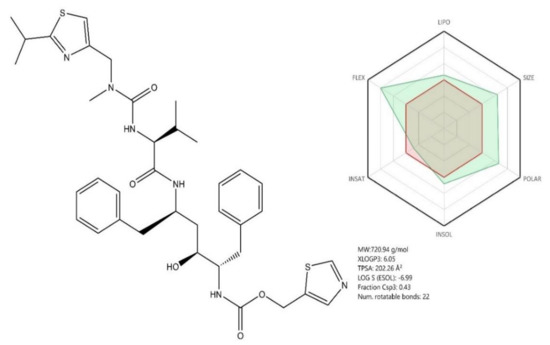
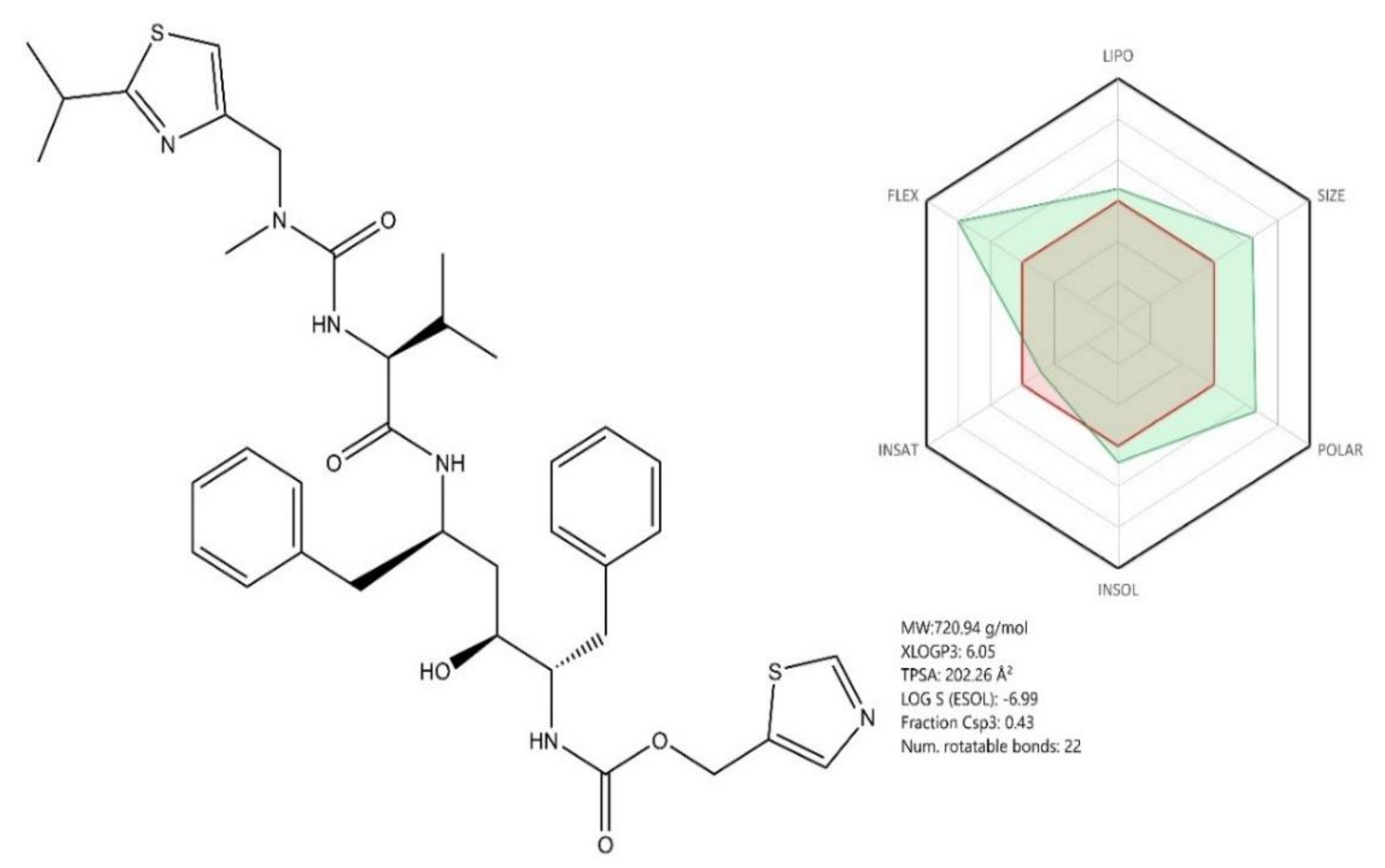
Figure 9.
Chemical structure of Ritonavir; ADME profile was achieved using the free web tool SwissADME; the red highlighted area represents the suitable physicochemical space for oral bioavailability covering value intervals for the following: LIPO (lipophility): −0.7 < XLOGP3 < +5.0, SIZE: 150 g/mol < MV < 500 g/mol, POLAR (polarity): 20Å2 < TPSA < 130Å2, INSOLU (insolubility): 0 < Log S (ESOL) < 6, INSATU (insaturation): 0.25 < Fraction Csp3 < 1, FLEX (flexibility): 0 < Num. rotatable bonds < 9, whereas the overlapped green highlighted area shows the calculated ADME profile for the molecule [63].
CoVs encode specific proteases such as papain—like protease (PLpro) and main protease (Mpro or 3CLpro) [48,108]. Both enzymes are involved in the control of the CoVs’ gene expression and replication [108] by inducing the proteolysis of viral polyproteins into functional and individual units [48] consisting of spike proteins, membrane proteins, envelope proteins, nucleoproteins, replicases, and other polymerases [110]. Mpro, a highly conserved homodimer among Coronaviridae members such as SARS-CoV and MERS-CoV, has emerged as a potential target for lopinavir/ritonavir owing to its role in self and polyprotein maturation [108]. A previous study assessed the ability of several antivirals to act as inhibitors of the SARS-CoV specific viral protease Mpro, by means of molecular docking and molecular dynamic simulations. The conclusion was that the molecules with the best scores for potential viral protease inhibition were lopinavir and ritonavir [111]. In accordance with previous studies, to investigate a possible mechanism for the antiviral activity of LPV/r, Muralidharan et al. performed molecular docking and molecular dynamics simulations to explore the affinity of these antiretroviral drugs to the SARS-CoV-2 protease, and their results showed a stronger binding energy of the two drugs combined compared with that exerted by each drug separately [108]. Their study also revealed that Lopinavir was able to interact with the viral protease predominantly through hydrophobic interactions, while in the Ritonavir–protease complexes, both hydrogen bonding and hydrophobic interactions were deemed as key features for enzyme inhibition [108].
However, the LPV/r combination in the treatment of COVID-19 is not justified or, at least, needs further investigations. Cao B and co-workers conducted a randomized, controlled, open-label trial in adult hospitalized patients with COVID-19 to test the efficacy and safety of the oral lopinavir (400 mg) associated with ritonavir (200 mg) in SARS-CoV-2 infection. At the end of their study, there was no evidence that LPV/r combination exerted any significant antiviral effect against SARS-CoV-2, as the viral RNA was still detected in 40.7% of the patients belonging to the LPV/r group [31,112]. Moreover, European Society of Intensive Care Medicine (ESICM) does not currently recommend LPV/r for routine use [31,113].
The pharmacokinetic profile of LPV/r consists of the following parameters: (1) a good absorption rate after oral administration, (2) lopinavir presents a high affinity for serum proteins (98–99% bound to serum proteins, mainly of alpha-1-acid glycoprotein (AAG)), (3) hepatic biotransformation of lopinavir is almost exclusively by isozyme CYP3A (ritonavir is a strong inhibitor of CYP3A, increasing the levels of lopinavir), and (4) elimination in urine and feces both untransformed and metabolized. Taking into consideration that LPV/r (Kaletra) was approved as anti-HIV therapy for almost 20 years ago (2001), a long list of adverse effects was ascribed to this product, classified as follows: (i) very common: upper respiratory tract infection, diarrhea, nausea; (ii) common: lower respiratory tract infection, skin infections (including cellulitis, folliculitis, and furuncle), anemia, leucopenia, neutropenia, lymphadenopathy, urticaria, angioedema, blood glucose disorders including diabetes mellitus, hypertriglyceridemia, hypercholesterolemia, weight decrease, decreased appetite, anxiety, headache (migraine), insomnia, dizziness, hypertension, pancreatitis, abdominal pain, hepatitis (with increased AST (aspartate aminotransferase), ALT, and GGT (Gamma-glutamyl transferase) values), skin disorders, myalgia, and erectile dysfunction, among others; (iii) uncommon: immune reconstitution inflammatory syndrome, hypogonadism, weight increase, abnormal dreams, cerebrovascular accident, convulsion, dysgeusia, ageusia, tremor, visual impairment, tinnitus, deep vein thrombosis, gastrointestinal hemorrhage, gastrointestinal ulcer, jaundice, hepatic steatosis, alopecia, rhabdomyolysis, osteonecrosis, and nephritis, among others; and (iv) rare: Stevens–Johnson syndrome and erythema multiforme [114].
As both lopinavir and ritonavir are CYP3A enzyme inhibitors, multiple drug–drug interactions were described, thus increased attention is recommended when this combination is co-administered with other substances metabolized by the same enzyme. Among the long list of interactions (for the detailed list, visit [80]), we will present several contraindicated drugs to be co-administered with LPV/r in the COVID-19 actual context: dextropropoxyphene, amiodarone, bepril, dofetilide, rifampicin, apixaban, clopidogrel, ticagrelor, St John’s wort, aliskiren, ivabradine, lercanidipine, ranolazine, sildenafil, quetiapine, antivirals such as hepatitis C virus (HCV) DDAs (direct-acting antiviral agents), midazolam, triazolam, cisapride, domperidone, sirolimus, lovastatin, simvastatin, budesonide, fluticasone, mometasone, and triamcinolone [114].
In light of the toxicological profile of LPV/r and its high potential for drug–drug interactions, China National Center for Adverse Drug Reaction Monitoring collected and analyzed data from hospitals concerning medication safety for COVID-19 patients; their report included the following findings: 179 patients (82.5%) developed adverse effects after the lopinavir/ritonavir treatment and 37 patients (17.1%) after chloroquine use. The adverse effects related to LPV/r use consisted of the following: hypertriglyceridemia, hypercholesterolemia, gastrointestinal reactions (diarrhea), liver injury, and rush [115].
In the opinion of the Infectious Diseases Society of America (IDSA), the published clinical data concerning LPV/r efficacy as COVID-19 treatment are not sufficient to recommend this medicine as treatment, only in a clinical trial context [116].
5. Closing Remarks and Future Perspectives
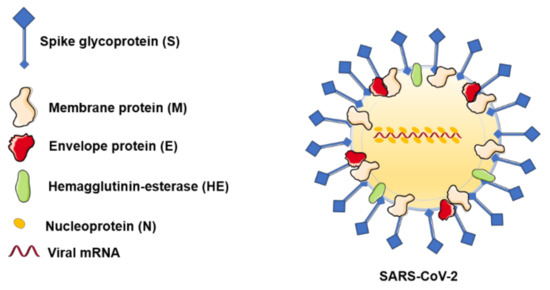
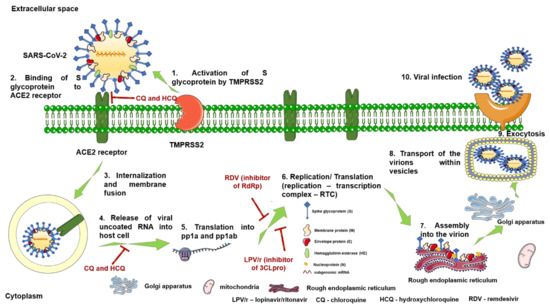
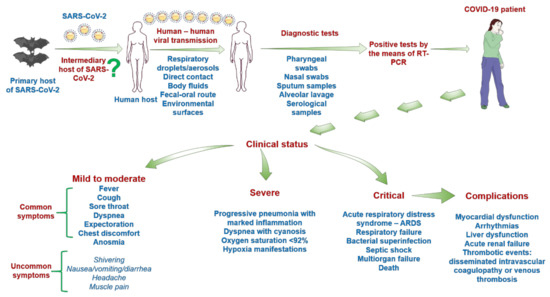
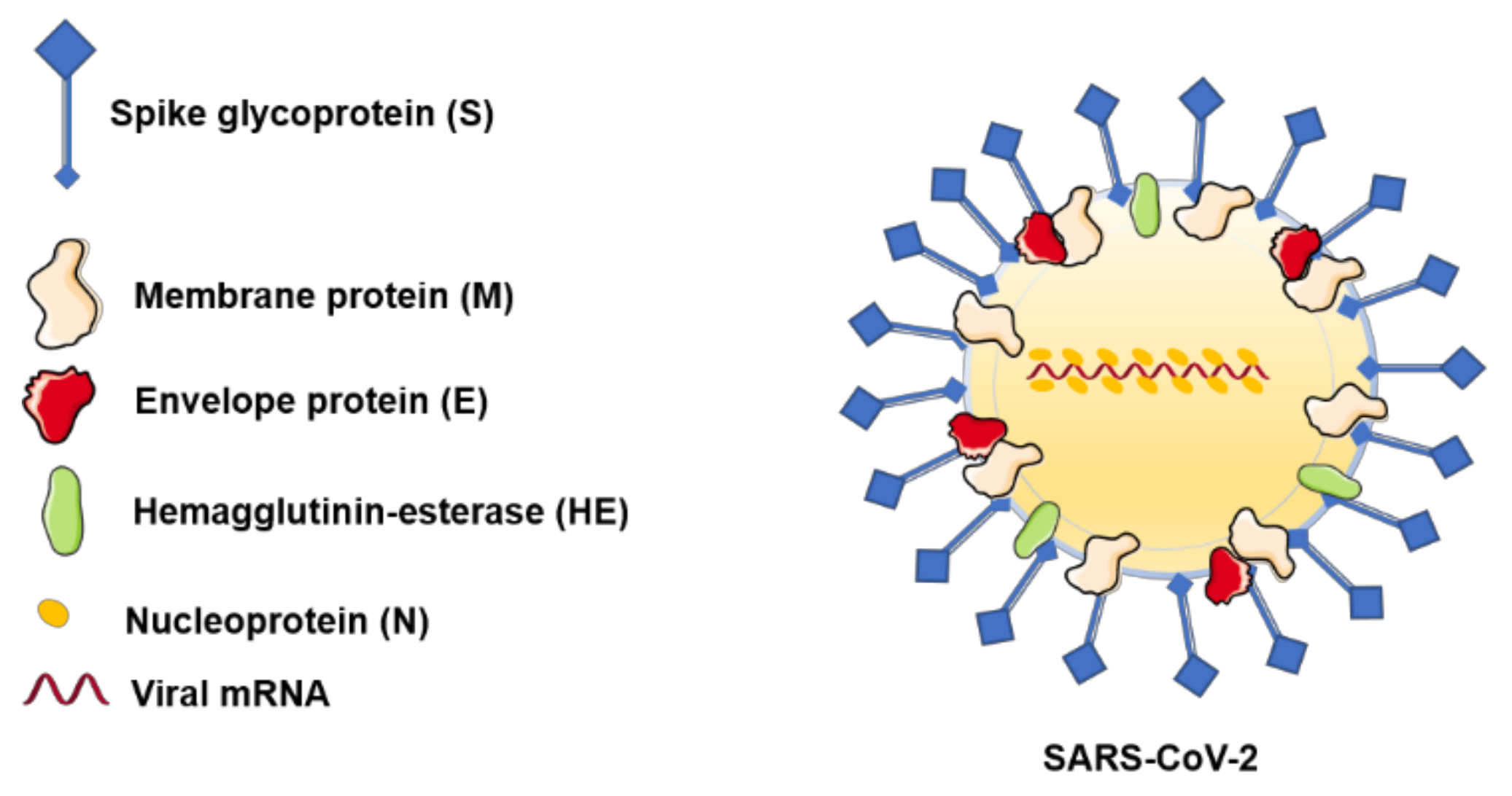
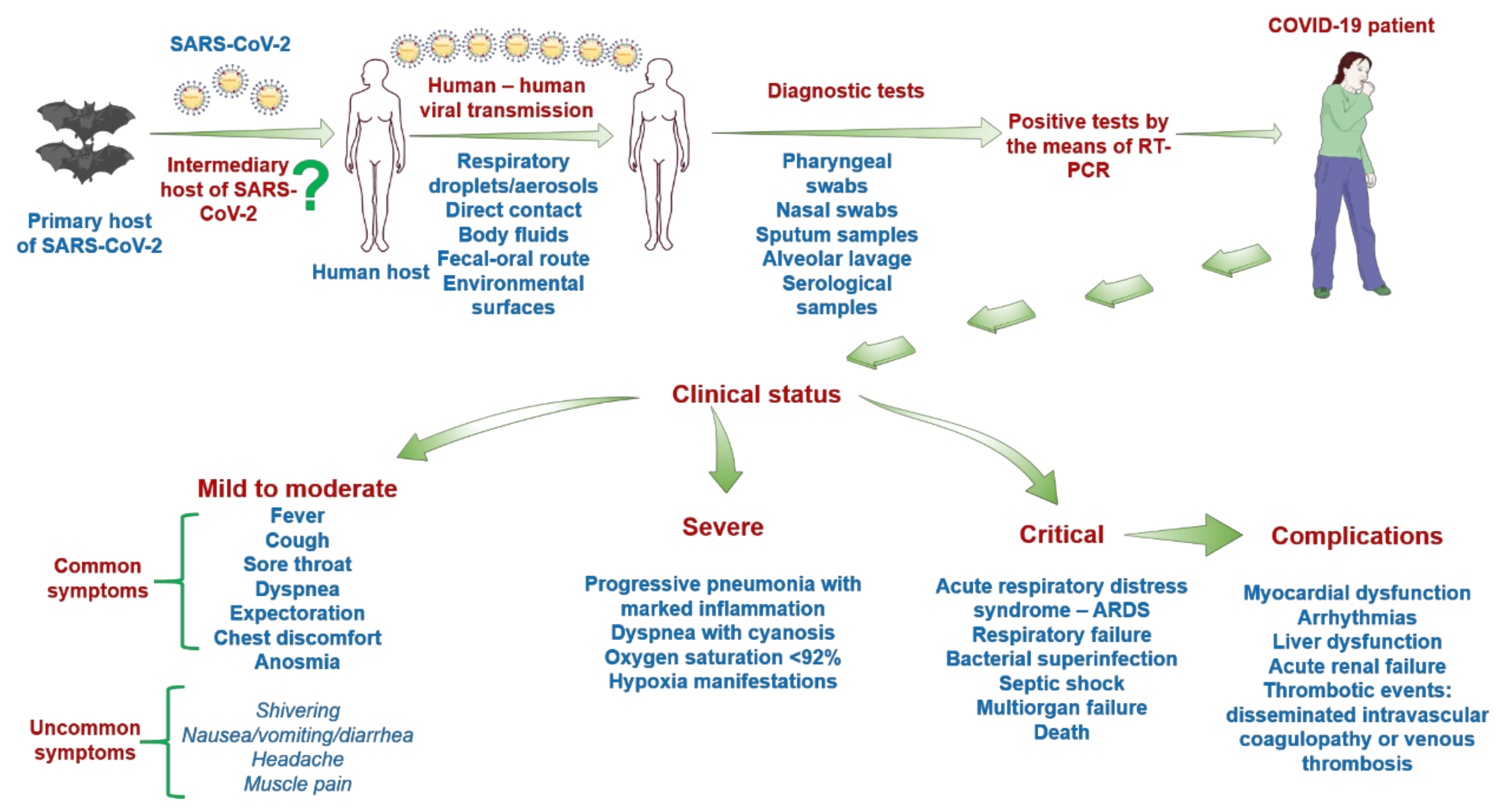
The outbreak determined by the newly emerged coronavirus, SARS-CoV-2, has initiated a roller effect both at medical and industrial levels, and its consequences are already seen in different areas. It has been more than five months since the first mention of this virus new potential, and if, at the beginning, there were more missing pieces, the latest data managed to almost complete the puzzle, as follows: (i) SARS-CoV-2 has a zoonotic origin (bats are considered the primary source); (ii) the genomic structure of the virus was elucidated, leading to the development of diagnostic tools and of potential novel innovative antivirals; (iii) human infection requires the binding of S spike glycoprotein to the human ACE2 receptor expressed by epithelial respiratory cells, vascular endothelial cells, cardiomyocytes, gastrointestinal cells, and hepatocytes, among others; (iv) human-to-human transmission occurs via respiratory droplets, direct contact, fecal–oral route, environmental transmission, and bodily fluids (key data for the infection spread repression); and (v) the clinical impact of SARS-CoV-2 infection (characteristic symptomatology, onset, clinical stages, complications) is mostly described, but the data in this area are constantly being updated.
The urge to find a therapy for SARS-CoV-2 infection determined the occurrence of multiple therapeutic alternatives, such as repurposed drugs, broad-spectrum antivirals (remdesivir, lopinavir/ritonavir, oseltamivir, and so on), antimalarial drugs (chloroquine and hydroxychloroquine), anti-inflammatory compounds (baricitinib, fedratinib, ruxolitinib, sunitinib), anticoagulants (low molecular weight heparin, unfractionated heparin), convalescent plasma, and novel potential antivirals (vaccines, anti-SARS-CoV-2 antibodies, natural compounds, and so on).
At present, remdesivir is considered the most promising therapy for SARS-CoV-2 infection being already included in the interim guidelines for COVID-19 treatment, based on the following considerations: proved to be an effective drug in severely and critically ill forms of COVID-19; shows a high human tolerance; the only side effect directly correlated to RDV is the hepatic injury (elevation of hepatic enzymes values); has a very low potential to induce drug–drug interactions; and the duration of treatment can be reduced at 5 days according to the latest clinical results. Although the mechanism of action of RDV is well defined, its toxicological profile needs further investigations in order to be fully established, so the administration of this compound should be performed under surveillance.
Chloroquine and hydroxychloroquine are recommended for compassionate use in mild-to-moderate (as single treatment option) and in severely and critically ill forms of COVID-19 (as comedication with remdesivir or lopinavir/ritonavir). These drugs present efficient oral absorption and distribution patterns, several drug–drug interactions were described (see detailed list at https://www.covid19-druginteractions.org/), and a thorough risk/benefit ratio analyze should be conducted before their administration mainly in combination with lopinavir/ritonavir (might increase the risk for prolongation of QT interval). Still, their clinical effectiveness as anti-COVID-19 agents is debatable at this point.
The use of lopinavir/ritonavir against SARS-CoV-2 infection determined controversial results; that is, according to some clinical studies, this combination had no significant antiviral effect, whereas in terms of adverse events and drug–drug interactions, the significance was reached (both lopinavir and ritonavir are CYP3A inhibitors and are subjected to multiple drug–drug interactions and adverse effects implicitly). Therefore, lopinavir/ritonavir is recommended as a second choice in the interim guidelines for remdesivir and its administration should be performed only under strict surveillance. Administration of anti-inflammatory agents (baricitinib, fedratinib, sunitinib, ruloxitinib, and so on) should be recommended as therapy for SARS-CoV-2 infection only when the laboratory results indicate a potential “cytokine storm” occurrence. The initiation of anticoagulant treatment should be performed based on laboratory results that indicate a potential risk for thrombotic events and coagulopathy disfunction; the administration of LMWHs is recommended in mild and moderate coagulation impairment, whereas in severe conditions, unfractionated heparin should be the first choice.
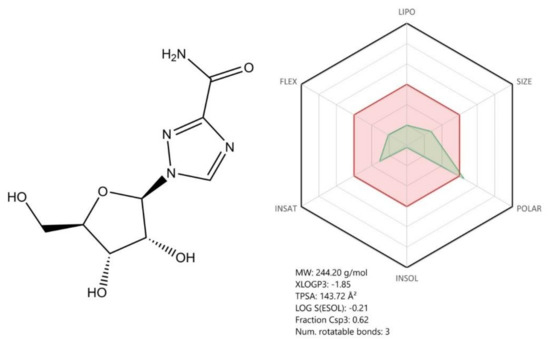
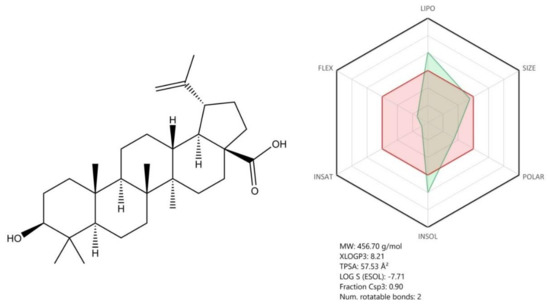
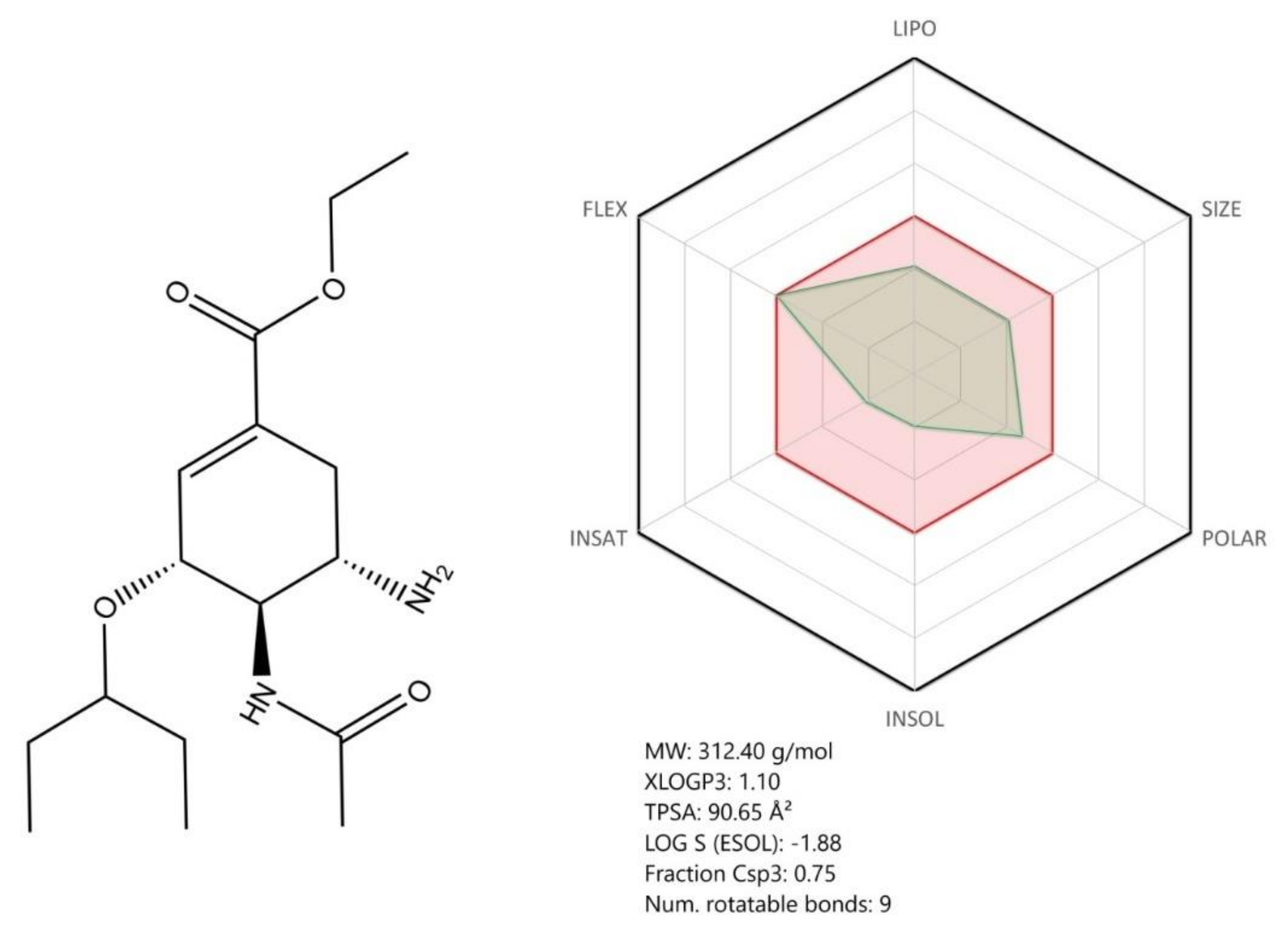
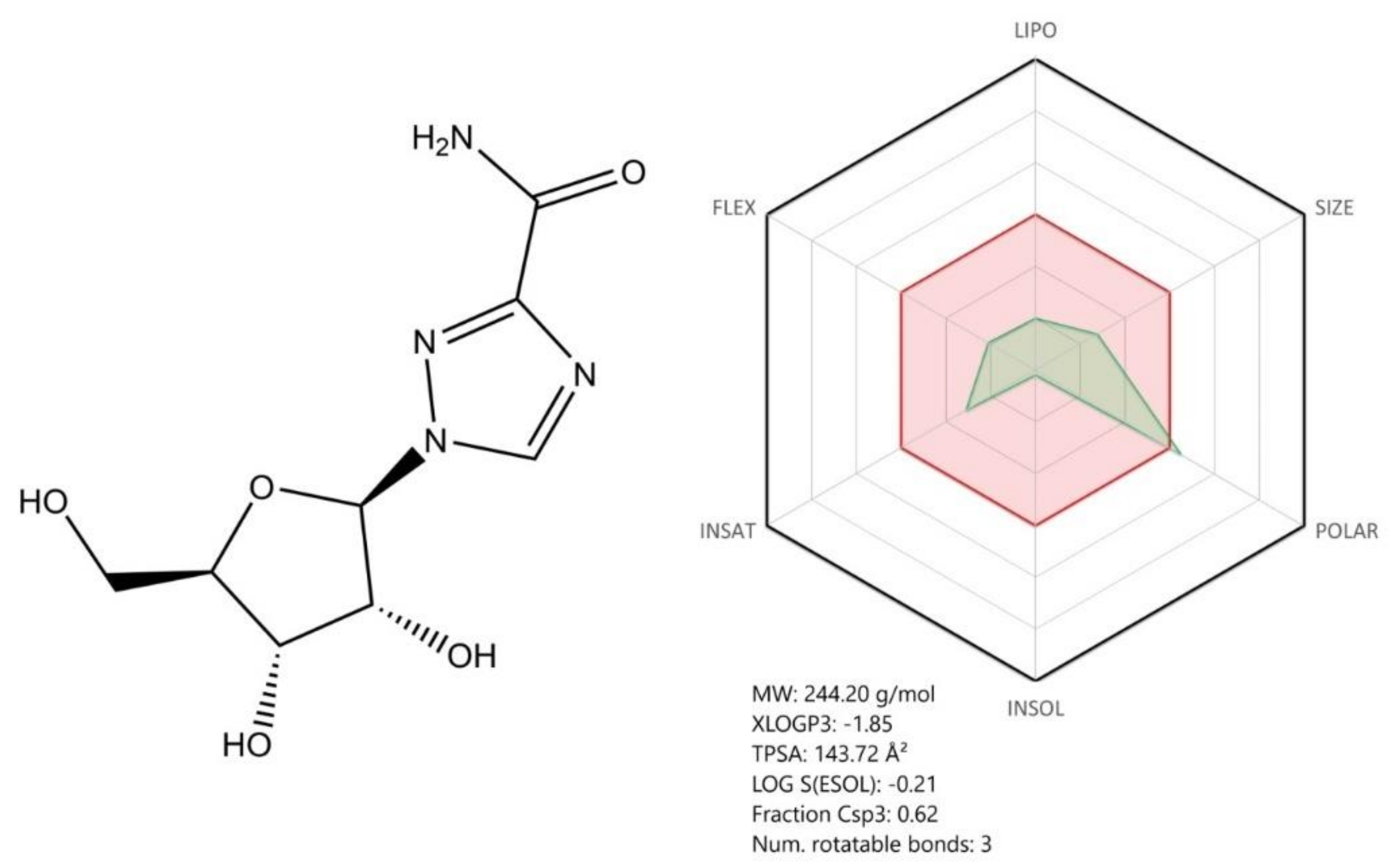
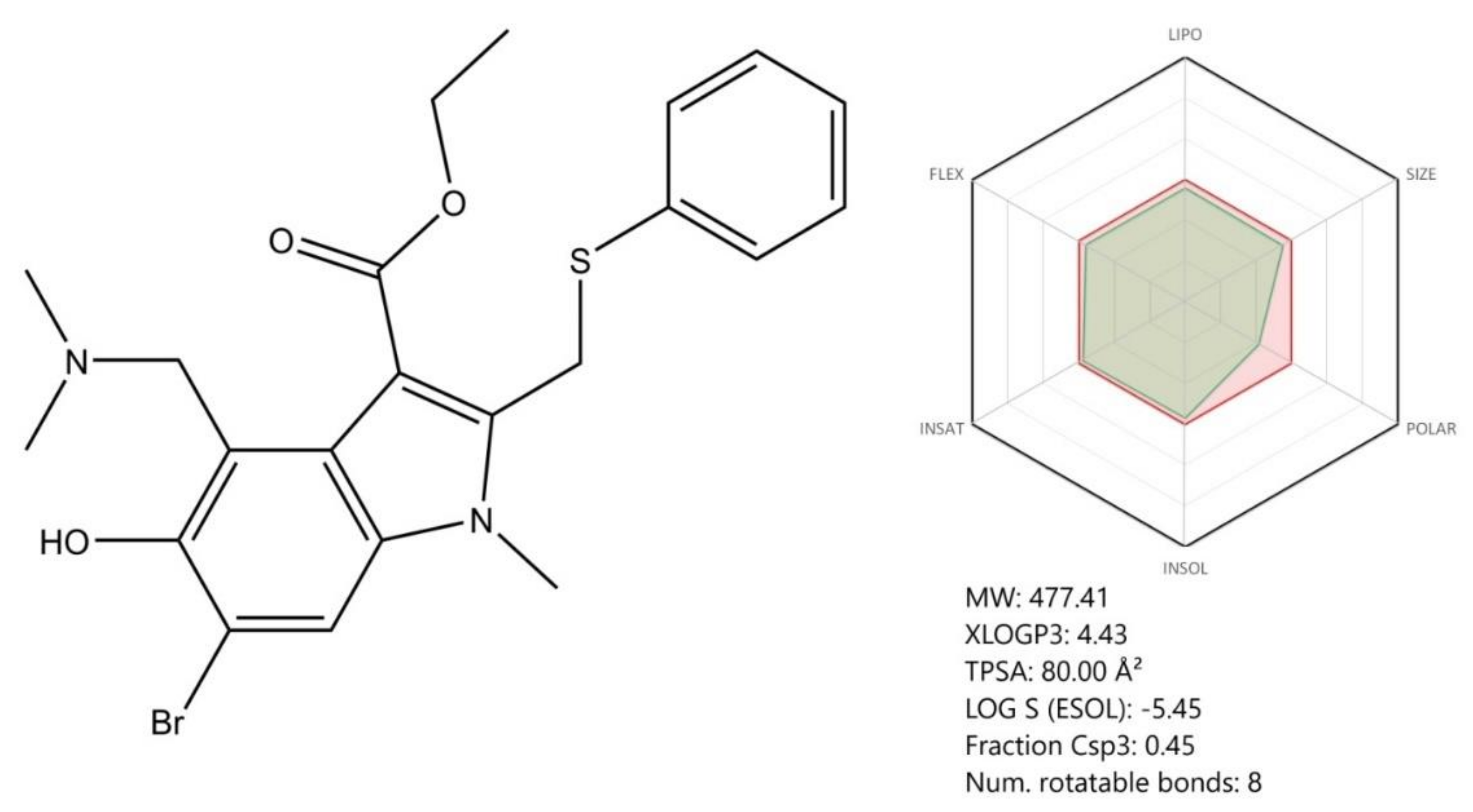
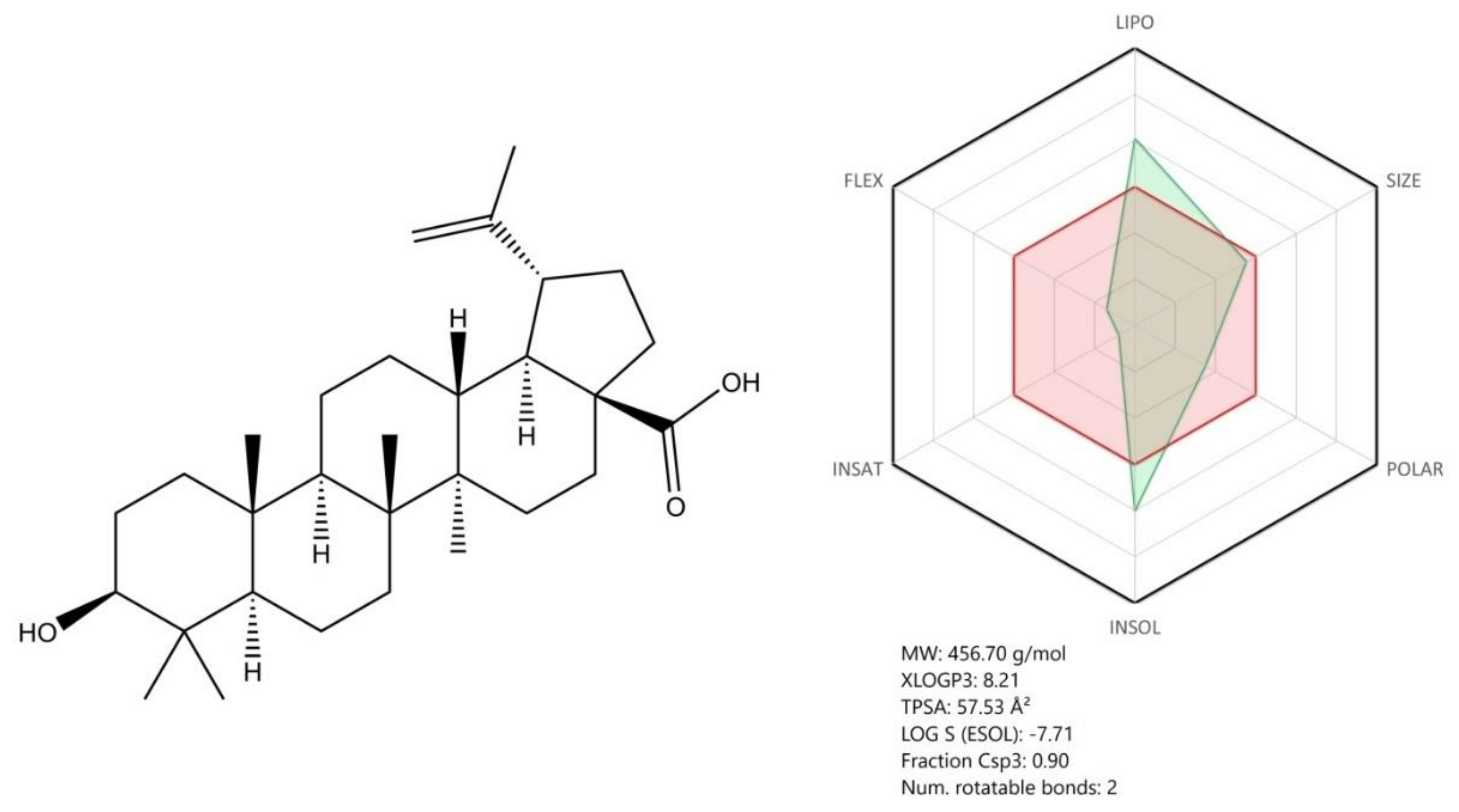
Other compounds described in our manuscript such as oseltamivir, ribavirin, arbidol, and natural compounds (betulinic acid) proved to have potent in vitro antiviral activity against SARS-CoV-2; still, further studies (preclinical and clinical) are required to confirm their effectiveness as anti-COVID-19 therapeutic agents. Convalescent plasma collected from COVID-19 patients represents a promising therapeutic alternative with immediate results; however, some limitations were noticed in terms of its obtaining and approval for use (the ongoing clinical trials results will confirm its effectiveness).
In defiance of the great efforts recorded, no drug was approved as a specific anti-SARS-CoV-2 treatment up to present. The considerable number of ongoing clinical trials (over 1300) that evaluate multiple potential antivirals represent the future perspectives concerning the elucidation of COVID-19 pathology and finding an appropriate treatment. The information regarding the progress recorded in the field of vaccine development is also optimistic and represents an option of selective and potent intervention.
Author Contributions
Conceptualization, C.A.D., V.L., and D.C.; Writing—Original draft preparation, C.A.D., D.C., M.M., I.M., V.L., I.P., and R.O.; Writing and critical revision for important intellectual content, C.A.D., C.S., A.M.T., and O.C.; Supervision, C.A.D. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Khan, S.; Siddique, R.; Shereen, M.A.; Ali, A.; Liu, J.; Bai, Q.; Bashir, N.; Xue, M. Emergence of a Novel Coronavirus, Severe Acute Respiratory Syndrome Coronavirus 2: Biology and Therapeutic Options. J. Clin. Microbiol. 2020, 58, e00187-20. [Google Scholar] [CrossRef]
- Sohrabi, C.; Alsafi, Z.; O’Neill, N.; Khan, M.; Kerwan, A.; Al-Jabir, A.; Iosifidis, C.; Agha, R. World Health Organization declares global emergency: A review of the 2019 novel coronavirus (COVID-19). Int. J. Surg. 2020, 76, 71–76. [Google Scholar] [CrossRef]
- Lai, C.C.; Liu, Y.H.; Wang, C.Y.; Wang, Y.H.; Hsueh, S.C.; Yen, M.Y.; Ko, W.C.; Hsueh, P.R. Asymptomatic carrier state, acute respiratory disease, and pneumonia due to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2): Facts and myths. J. Microbiol. Immunol. Infect. 2020. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization Home Page—WHO Director-General’s Opening Remarks at the Media Briefing on COVID-19—11 March 2020. Available online: https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020 (accessed on 25 April 2020).
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [PubMed]
- COVID-19 Map. Available online: https://news.google.com/covid19/map?hl=en-US&gl=US&ceid=US:en (accessed on 4 May 2020).
- ECDC. COVID-19 Situation Update Worldwide, as of 10 May 2020. Available online: https://www.ecdc.europa.eu/en/geographical-distribution-2019-ncov-cases (accessed on 11 May 2020).
- WHO. Home Page—Coronavirus Disease (COVID-19) Situation Report—105. Available online: https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200504-covid-19-sitrep-105.pdf?sfvrsn=4cdda8af_2 (accessed on 5 May 2020).
- Wu, D.; Wu, T.; Liu, Q.; Yang, Z. The SARS-CoV-2 Outbreak: What We Know. Int. J. Infect. Dis. 2020, 94, 44–48. [Google Scholar] [CrossRef]
- Zhang, Y.Z.; Holmes, E.C. A Genomic Perspective on the Origin and Emergence of SARS-CoV-2. Cell 2020, 181, 223–227. [Google Scholar] [CrossRef]
- Wang, C.; Horby, P.W.; Hayden, F.G.; Gao, G.F. A novel coronavirus outbreak of global health concern. Lancet 2020, 395, 470–473. [Google Scholar] [CrossRef]
- Shanmugaraj, B.; Siriwattananon, K.; Wangkanont, K.; Phoolcharoen, W. Perspectives on monoclonal antibody therapy as potential therapeutic intervention for Coronavirus disease-19 (COVID-19). Asian Pac. J. Allergy Immunol. 2020, 38, 10–18. [Google Scholar] [CrossRef]
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef]
- Liu, D.X.; Liang, J.Q.; Fung, T.S. Human Coronavirus-229E, -OC43, -NL63, and -HKU1. Ref. Modul. Life Sci. 2020. [Google Scholar] [CrossRef]
- Ashour, H.M.; Elkhatib, W.F.; Rahman, M.M.; Elshabrawy, H.A. Insights into the Recent 2019 Novel Coronavirus (SARS-CoV-2) in Light of Past Human Coronavirus Outbreaks. Pathogens 2020, 9, 186. [Google Scholar] [CrossRef] [PubMed]
- Kannan, S.; Shaik Syed Ali, P.; Sheeza, A.; Hemalatha, K. COVID-19 (Novel Coronavirus 2019)—Recent Trends. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 2006–2011. [Google Scholar] [CrossRef] [PubMed]
- Sun, P.; Lu, X.; Xu, C.; Sun, W.; Pan, B. Understanding of COVID-19 Based on Current Evidence. J. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- NCBI. GenBank. Available online: https://www.ncbi.nlm.nih.gov/genbank/sars-cov-2-seqs/ (accessed on 18 June 2020).
- Khailany, R.A.; Safdar, M.; Ozaslan, M. Genomic Characterization of a Novel SARS-CoV-2. Gene Rep. 2020, 19, 100682. [Google Scholar] [CrossRef]
- Penarrubia, A.L.; Ruiz, M.; Porco, R.; Rao, S.N.; Juanola-Falgarona, M.; Manissero, D.; López-Fontanals, M.; Pareja, J. Multiple Assays in a Real-Time RT-PCR SARS-CoV-2 Panel Can Mitigate the Risk of Loss of Sensitivity by New Genomic Variants during the COVID-19 Outbreak. Int. J. Infect. Dis. Off. Publ. Int. Soc. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.Y.; et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181, 894–904. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292. [Google Scholar] [CrossRef]
- Fu, Y.; Cheng, Y.; Wu, Y. Understanding SARS-CoV-2-Mediated Inflammatory Responses: From Mechanisms to Potential Therapeutic Tools. Virol. Sin. 2020. [Google Scholar] [CrossRef]
- Li, R.; Qiao, S.; Zhang, G. Analysis of Angiotensin-Converting Enzyme 2 (ACE2) From Different Species Sheds Some Light on Cross-Species Receptor Usage of a Novel Coronavirus 2019-nCoV. J. Infect. 2020, 80, 469–496. [Google Scholar] [CrossRef]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 Spike Receptor-Binding Domain Bound to the ACE2 Receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Ye, G.; Shi, K.; Wan, Y.; Luo, C.; Aihara, H.; Geng, Q.; Auerbach, A.; Li, F. Structural basis of receptor recognition by SARS-CoV-2. Nature 2020, 581, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.R.; Cao, Q.D.; Hong, Z.S.; Tan, Y.Y.; Chen, S.D.; Jin, H.J.; Tan, K.S.; Wang, D.Y.; Yan, Y. The Origin, Transmission and Clinical Therapies on Coronavirus Disease 2019 (COVID-19) Outbreak—An Update on the Status. Mil. Med. Res. 2020, 7, 11. [Google Scholar] [CrossRef]
- Neurath, M.F. Covid-19 and Immunomodulation in IBD. Gut 2020. [Google Scholar] [CrossRef]
- Shereen, M.A.; Khan, S.; Kazmi, A.; Bashir, N.; Siddique, R. COVID-19 infection: Origin, transmission, and characteristics of human coronaviruses. J. Adv. Res. 2020, 24, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.C.; Deng, Q.X.; Dai, S.X. Remdesivir for Severe Acute Respiratory Syndrome Coronavirus 2 Causing COVID-19: An Evaluation of the Evidence. Travel Med. Infect. Dis. 2020, 101647. [Google Scholar] [CrossRef]
- Nicola, M.; O’Neill, N.; Sohrabi, C.; Khan, M.; Agha, M.; Agha, R. Evidence based management guideline for the COVID-19 pandemic—Review article. Int. J. Surg. 2020, 77, 206–216. [Google Scholar] [CrossRef]
- Tian, Y.; Rong, L.; Nian, W.; He, Y. Review Article: Gastrointestinal Features in COVID-19 and the Possibility of Faecal Transmission. Aliment. Pharmacol. Ther. 2020, 51, 843–851. [Google Scholar] [CrossRef]
- Kluge, S.; Janssens, U.; Welte, T.; Weber-Carstens, S.; Marx, G.; Karagiannidis, C. German Recommendations for Critically Ill Patients with COVID-19. Med. Klin. Intensivmed. Notfmed. 2020. [Google Scholar] [CrossRef]
- Tian, X.; Li, C.; Huang, A.; Xia, S.; Lu, S.; Shi, Z.; Lu, L.; Jiang, S.; Yang, Z.; Wu, Y.; et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg. Microbes Infect. 2020, 9, 382–385. [Google Scholar] [CrossRef]
- Kuba, K.; Imai, Y.; Ohto-Nakanishi, T.; Penninger, J.M. Trilogy of ACE2: A Peptidase in the Renin-Angiotensin System, a SARS Receptor, and a Partner for Amino Acid Transporters. Pharmacol. Ther. 2010, 128, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Docea, A.O.; Tsatsakis, A.; Albulescu, D.; Cristea, O.; Zlatian, O.; Vinceti, M.; Moschos, S.A.; Tsoukalas, D.; Goumenou, M.; Drakoulis, N.; et al. A new threat from an old enemy: Re-emergence of coronavirus (Review). Int. J. Mol. Med. 2020, 45, 1631–1643. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.; Zheng, K.I.; Yan, Q.Q.; Rios, R.S.; Targher, G.; Byrne, C.D.; Poucke, S.V.; Liu, W.Y.; Zheng, M.H. COVID-19 and Liver Dysfunction: Current Insights and Emergent Therapeutic Strategies. J. Clin. Transl. Hepatol. 2020, 8, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Bikdeli, B.; Madhavan, M.V.; Jimenez, D.; Chuich, T.; Dreyfus, I.; Driggin, E.; Nigoghossian, C.; Ageno, W.; Madjid, M.; Guo, Y.; et al. COVID-19 and Thrombotic or Thromboembolic Disease: Implications for Prevention, Antithrombotic Therapy, and Follow-up. J. Am. Coll. Cardiol. 2020. [Google Scholar] [CrossRef]
- Meng, X.; Deng, Y.; Dai, Z.; Meng, Z. COVID-19 and Anosmia: A Review Based on Up-To-Date Knowledge. Am. J. Otolaryngol. 2020, 41, 102581. [Google Scholar] [CrossRef]
- Dawson, P.; Rabold, E.M.; Laws, R.L.; Conners, E.E.; Gharpure, R.; Yin, S.; Buono, S.A.; Dasu, T.; Bhattacharyya, S.; Westergaard, R.P.; et al. Loss of Taste and Smell as Distinguishing Symptoms of COVID-19. Clin. Infect. Dis. 2020, 21. [Google Scholar] [CrossRef]
- Li, J.; Fan, J.G. Characteristics and Mechanism of Liver Injury in 2019 Coronavirus Disease. J. Clin. Transl. Hepatol. 2020, 8, 13–17. [Google Scholar] [CrossRef]
- Hong, E.H.; Song, J.H.; Kang, K.B.; Sung, S.H.; Ko, H.J.; Yang, H. Anti-Influenza Activity of Betulinic Acid From Zizyphus Jujuba on Influenza A/PR/8 Virus. Biomol. Ther. 2015, 23, 345–349. [Google Scholar] [CrossRef]
- Kulkarni, R. Antibody-Dependent Enhancement of Viral Infections. Dyn. Immune Act. Viral Dis. 2019, 5, 9–41. [Google Scholar] [CrossRef]
- Davies, N.G.; Klepac, P.; Liu, Y.; Prem, K.; Jit, M.; CMMID COVID-19 Working Group; Eggo, R.M. Age-dependent Effects in the Transmission and Control of COVID-19 Epidemics. Nat. Med. 2020. [Google Scholar] [CrossRef]
- Xie, Z. Pay attention to SARS-CoV-2 infection in children. Pediatric Investig. 2020, 4, 1–4. [Google Scholar] [CrossRef]
- Liguoro, I.; Pilotto, C.; Bonanni, M.; Ferrari, M.E.; Pusiol, A.; Nocerino, A.; Vidal, E.; Cogo, P. SARS-COV-2 infection in children and newborns: A systematic review. Eur. J. Pediatr. 2020. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Zhou, Q.; Li, Y.; Garner, L.V.; Watkins, S.P.; Carter, L.J.; Smoot, J.; Gregg, A.C.; Daniels, A.D.; Jervey, S.; et al. Research and Development on Therapeutic Agents and Vaccines for COVID-19 and Related Human Coronavirus Diseases. ACS Cent. Sci. 2020, 6, 315–331. [Google Scholar] [CrossRef] [PubMed]
- WHO. Solidarity Clinical Trial for COVID-19 Treatments. Available online: https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---18-march-2020 (accessed on 25 April 2020).
- WHO. Public Statement for Collaboration on COVID-19 Vaccine Development. Available online: https://www.who.int/news-room/detail/13-04-2020-public-statement-for-collaboration-on-covid-19-vaccine-development (accessed on 25 April 2020).
- NIH U.S National Library of Medicine. Home—ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/results?cond=COVID-19 (accessed on 11 May 2020).
- WHO. WHO | Welcome to the WHO ICTRP. Available online: https://www.who.int/ictrp/en/ (accessed on 30 April 2020).
- Van Ierssel, S.; Dauby, N.; Bottieau, E.; Huits, R.; The Belgium Task Force. Interim Clinical Guidance for Adults with Suspected or Confirmed COVID-19 in Belgium 06 May 2020, Version 8. 2020.
- Van Ierssel, S.; Dauby, N.; Bottieau, E.; Huits, R.; The Belgium Task Force. Interim Clinical Guidance for Adults with Suspected or Confirmed COVID-19 in Belgium 06 May 2020, Version 5. 2020.
- Lombardy Section Italian Society Infectious and Tropical Diseases. Vademecum for the treatment of people with COVID-19. Edition 2.0, 13 March 2020. Infez. Med. 2020, 28, 143–152. [Google Scholar]
- Kupferschmidt, K.; Cohen, J. Race to Find COVID-19 Treatments Accelerates. Science 2020, 367, 1412–1413. [Google Scholar] [CrossRef]
- Gilead. Gilead Announces Results from Phase 3 Trial of Investigational Antiviral Remdesivir in Patients with Severe COVID-19. Available online: https://www.gilead.com/news-and-press/press-room/press-releases/2020/4/gilead-announces-results-from-phase-3-trial-of-investigational-antiviral-remdesivir-in-patients-with-severe-covid-19 (accessed on 30 April 2020).
- A NIH NIAID. NIH Clinical Trial Shows Remdesivir Accelerates Recovery from Advanced COVID-19 | NIH: National Institute of Allergy and Infectious Diseases. Available online: https://www.niaid.nih.gov/news-events/nih-clinical-trial-shows-remdesivir-accelerates-recovery-advanced-covid-19 (accessed on 30 April 2020).
- Agostini, M.L.; Andres, E.L.; Sims, A.C.; Graham, R.L.; Sheahan, T.P.; Lu, X.; Smith, E.C.; Case, J.B.; Feng, J.Y.; Jordan, R.; et al. Coronavirus Susceptibility to the Antiviral Remdesivir (GS-5734) Is Mediated by the Viral Polymerase and the Proofreading Exoribonuclease. mBio 2018, 9, e00221-18. [Google Scholar] [CrossRef]
- Ko, W.C.; Rolain, J.M.; Lee, N.Y.; Chen, P.L.; Huang, C.T.; Lee, P.I.; Hsueh, P.R. Arguments in Favour of Remdesivir for Treating SARS-CoV-2 Infections. Int. J. Antimicrob. Agents 2020, 55, 105933. [Google Scholar] [CrossRef]
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and Chloroquine Effectively Inhibit the Recently Emerged Novel Coronavirus (2019-nCoV) in Vitro. Cell Res. 2020, 30, 269–271. [Google Scholar] [CrossRef]
- Tchesnokov, E.P.; Feng, J.Y.; Porter, D.P.; Götte, M. Mechanism of Inhibition of Ebola Virus RNA-Dependent RNA Polymerase by Remdesivir. Viruses 2019, 11, 326. [Google Scholar] [CrossRef] [PubMed]
- Daina, A.; Michielin, O.; Zoete, V. wissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Zhang, C.; Huang, S.; Zheng, F.; Dai, Y. Controversial treatments: An updated understanding of the coronavirus disease 2019. J. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Amirian, E.S.; Levy, J.K. Current Knowledge about the Antivirals Remdesivir (GS-5734) and GS-441524 as Therapeutic Options for Coronaviruses. One Health 2020, 9, 100128. [Google Scholar] [CrossRef] [PubMed]
- Lo, M.K.; Jordan, R.; Arvey, A.; Sudhamsu, J.; Shrivastava-Ranjan, P.; Hotard, A.L.; Flint, M.; McMullan, L.K.; Siegel, D.; Clarke, M.O.; et al. GS-5734 and its parent nucleoside analog inhibit Filo-, Pneumo-, and Paramyxoviruses. Sci. Rep. 2017, 7, 43395. [Google Scholar] [CrossRef] [PubMed]
- Shannon, A.; Le, N.T.; Selisko, B.; Eydoux, C.; Alvarez, K.; Guillemot, J.C.; Decroly, E.; Peersen, O.; Ferron, F.; Canard, B. Remdesivir and SARS-CoV-2: Structural Requirements at Both nsp12 RdRp and nsp14 Exonuclease Active-Sites. Antiviral Res. 2020, 178, 104793. [Google Scholar] [CrossRef]
- Gordon, C.J.; Tchesnokov, E.P.; Woolner, E.; Perry, J.K.; Feng, J.Y.; Porter, D.P.; Götte, M. Remdesivir is a direct-acting antiviral that inhibits RNA-dependent RNA polymerase from severe acute respiratory syndrome coronavirus 2 with high potency. J. Biol. Chem. 2020, 95, 6785–6797. [Google Scholar] [CrossRef]
- Yin, W.; Mao, C.; Luan, X.; Shen, D.D.; Shen, Q.; Su, H.; Wang, X.; Zhou, F.; Zhao, W.; Gao, M.; et al. Structural Basis for Inhibition of the RNA-dependent RNA Polymerase From SARS-CoV-2 by Remdesivir. Science 2020. [Google Scholar] [CrossRef]
- Zhang, L.; Zhou, R. Binding mechanism of remdesivir to SARS-CoV-2 RNA dependent RNA polymerase. Published online ahead of print: 17 March 2020. [CrossRef]
- Grein, J.; Ohmagari, N.; Shin, D.; Diaz, G.; Asperges, E.; Castagna, A.; Feldt, T.; Green, G.; Green, M.L.; Lescure, F.X.; et al. Compassionate Use of Remdesivir for Patients With Severe Covid-19. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, D.; Du, G.; Du, R.; Zhao, J.; Jin, Y.; Fu, S.; Gao, L.; Cheng, Z.; Lu, Q.; et al. Remdesivir in adults with severe COVID-19: A randomised, double-blind, placebo-controlled, multicentre trial. Lancet 2020. [Google Scholar] [CrossRef]
- EMA. Conditions of Use, Conditions for Distribution and Patients Targeted and Conditions for Safety Monitoring Adressed to Member States for Remdesivir Available for Compassionate Use. Available online: https://www.ema.europa.eu/en/documents/other/conditions-use-conditions-distribution-patients-targeted-conditions-safety-monitoring-adressed_en-0.pdf (accessed on 21 November 2013).
- Gilead. Gilead Announces Results from Phase 3 Trial of Remdesivir in Patients with Moderate COVID-19. Available online: https://www.gilead.com/news-and-press/press-room/press-releases/2020/6/gilead-announces-results-from-phase-3-trial-of-remdesivir-in-patients-with-moderate-covid-19 (accessed on 19 June 2020).
- Gilead. Gilead Announces Approval of Veklury® (Remdesivir) in Japan for Patients with Severe COVID-19. Available online: https://www.gilead.com/news-and-press/press-room/press-releases/2020/5/gilead-announces-approval-of-veklury-remdesivir-in-japan-for-patients-with-severe-covid19 (accessed on 19 June 2020).
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; Hohmann, E.; Chu, H.Y.; Luetkemeyer, A.; Kline, S.; et al. Remdesivir for the Treatment of Covid-19—Preliminary Report. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- EMA. EMA Starts Rolling Review of Remdesivir for COVID-19. Available online: https://www.ema.europa.eu/en/news/ema-starts-rolling-review-remdesivir-covid-19 (accessed on 30 June 2020).
- U.S. Food and Drug Administration. Remdesivir EUA Letter of Authorization; Gilead Sciences, Inc.: Foster City, CA, USA, 2020.
- EMA. Summary on Compassionate Use Remdesivir Gilead. Available online: https://www.ema.europa.eu/en/documents/other/summary-compassionate-use-remdesivir-gilead_en.pdf (accessed on 25 April 2020).
- The University of Liverpool. COVID-19 Drug Interactions, 2020 Detailed Recommendations for Interactions with Experimental COVID-19 Therapies. Available online: https://www.covid19-druginteractions.org/ (accessed on 25 April 2020).
- Hu, T.Y.; Frieman, M.; Wolfram, J. Insights from Nanomedicine into Chloroquine Efficacy Against COVID-19. Nat. Nanotechnol. 2020, 15, 247–249. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.; Ye, F.; Zhang, M.; Cui, C.; Huang, B.; Niu, P.; Liu, X.; Zhao, L.; Dong, E.; Song, C.; et al. Optimized Dosing Design of Hydroxychloroquine for the Treatment of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Cao, R.; Xu, M.; Wang, X.; Zhang, H.; Hu, H.; Li, Y.; Hu, Z.; Zhong, W.; Wang, M. Hydroxychloroquine, a Less Toxic Derivative of Chloroquine, Is Effective in Inhibiting SARS-CoV-2 Infection in Vitro. Cell Discov. 2020, 6, 16. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.K.; Kumar, A.; Tiwari, G.; Kumar, R.; Misra, N. In Silico Investigations on the Potential Inhibitors for COVID-19 Protease. Published Online Ahead of Print: 23 March 2020. Available online: http://arxiv.org/abs/2003.10642 (accessed on 25 April 2020).
- Colson, P.; Rolain, J.M.; Lagier, J.C.; Brouqui, P.; Raoult, D. Chloroquine and hydroxychloroquine as available weapons to fight COVID-19. Int. J. Antimicrob. Agents 2020, 55, 105932. [Google Scholar] [CrossRef] [PubMed]
- Vincent, M.J.; Bergeron, E.; Benjannet, S.; Erickson, B.R.; Rollin, P.E.; Ksiazek, T.G.; Seidah, N.G.; Nichol, S.T. Chloroquine Is a Potent Inhibitor of SARS Coronavirus Infection and Spread. Virol. J. 2005, 2, 69. [Google Scholar] [CrossRef]
- Savarino, A.; Di Trani, L.; Donatelli, I.; Cauda, R.; Cassone, A. New Insights into the Antiviral Effects of Chloroquine. Lancet Infect. Dis. 2006, 6, 67–69. [Google Scholar] [CrossRef]
- Mingo, R.M.; Simmons, J.A.; Shoemaker, C.J.; Nelson, E.A.; Schornberg, K.L.; D’Souza, R.S.; Casanova, J.E.; White, J.M. Ebola Virus and Severe Acute Respiratory Syndrome Coronavirus Display Late Cell Entry Kinetics: Evidence That Transport to NPC1+ Endolysosomes Is a Rate-Defining Step. J. Virol. 2015, 89, 2931–2943. [Google Scholar] [CrossRef]
- Wu, C.; Liu, Y.; Yang, Y.; Zhang, P.; Zhong, W.; Wang, Y.; Wang, Q.; Xu, Y.; Li, M.; Li, X.; et al. Analysis of Therapeutic Targets for SARS-CoV-2 and Discovery of Potential Drugs by Computational Methods. Acta Pharm. Sin. B 2020. [Google Scholar] [CrossRef]
- PDR. Chloroquine Phosphate—Drud Summary. Available online: https://www.pdr.net/drug-summary/Chloroquine-Phosphate-chloroquine-phosphate-3418.2640 (accessed on 30 April 2020).
- Chowdhury, M.S.; Rathod, J.; Gernsheimer, J. A Rapid Systematic Review of Clinical Trials Utilizing Chloroquine and Hydroxychloroquine as a Treatment for COVID-19. Acad. Emerg. Med. 2020. [Google Scholar] [CrossRef]
- Chen, J.; Liu, D.; Liu, L.; Liu, P.; Xu, Q.; Xia, L.; Ling, Y.; Huang, D.; Song, S.; Zhang, D.; et al. A Pilot Study of Hydroxychloroquine in Treatment of Patients with Moderate COVID-19. Zhejiang Da Xue Xue Bao Yi Xue Ban 2020, 49, 215–219. [Google Scholar]
- Gautret, P.; Lagier, J.C.; Parola, P.; Hoang, V.T.; Meddeb, L.; Mailhe, M.; Doudier, B.; Courjon, J.; Giordanengo, V.; Vieira, V.E.; et al. Hydroxychloroquine and Azithromycin as a Treatment of COVID-19: Results of an Open-Label Non-Randomized Clinical Trial. Int. J. Antimicrob. Agents 2020, 105949. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Tian, Z.; Yang, X. Breakthrough: Chloroquine Phosphate Has Shown Apparent Efficacy in Treatment of COVID-19 Associated Pneumonia in Clinical Studies. Biosci. Trends 2020, 14, 72–73. [Google Scholar] [CrossRef]
- Zhou, D.; Dai, S.M.; Tong, Q. COVID-19: A Recommendation to Examine the Effect of Hydroxychloroquine in Preventing Infection and Progression. J. Antimicrob. Chemother. 2020. [Google Scholar] [CrossRef]
- Meo, S.A.; Klonoff, D.C.; Akram, J. Efficacy of Chloroquine and Hydroxychloroquine in the Treatment of COVID-19. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 4539–4547. [Google Scholar] [CrossRef] [PubMed]
- Gendelman, O.; Amital, H.; Bragazzi, N.L.; Watad, A.; Chodick, G. Continuous Hydroxychloroquine or Colchicine Therapy Does Not Prevent Infection with SARS-CoV-2: Insights from a Large Healthcare Database Analysis. Autoimmun. Rev. 2020, 102566. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Chen, Y.; Fan, X.; Wang, X.; Han, Q.; Liu, Z. Advances in the Use of Chloroquine and Hydroxychloroquine for the Treatment of COVID-19. Postgrad. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Shukla, A.M.; Archibald, L.K.; Shukla, A.W.; Mehta, H.J.; Cherabuddi, K. Chloroquine and Hydroxychloroquine in the Context of COVID-19. Drugs Context 2020, 9. [Google Scholar] [CrossRef]
- Van den Eynde, J.J. COVID-19: An Update about the Discovery Clinical Trial. Pharmaceuticals 2020, 13, 98. [Google Scholar] [CrossRef]
- Pastick, K.A.; Okafor, E.C.; Wang, F.; Lofgren, S.M.; Skipper, C.P.; Nicol, M.R.; Pullen, M.F.; Rajasingham, R.; McDonald, E.G.; Lee, T.C.; et al. Review: Hydroxychloroquine and Chloroquine for Treatment of SARS-CoV-2 (COVID-19). Open Forum Infect. Dis. 2020, 7. [Google Scholar] [CrossRef]
- Meyerowitz, E.A.; Vannier, A.; Friesen, M.; Schoenfeld, S.; Gelfand, J.A.; Callahan, M.V.; Kim, A.Y.; Reeves, P.M.; Poznansky, M.C. Rethinking the Role of Hydroxychloroquine in the Treatment of COVID-19. FASEB J. 2020, 34, 6027–6037. [Google Scholar] [CrossRef]
- Ruamviboonsuk, P.; Lai, T.; Chang, A.; Lai, C.C.; Mieler, W.F.; Lam, D.; Asia-Pacific Vitreo-Retina Society. Chloroquine and Hydroxychloroquine Retinal Toxicity Consideration in the Treatment of COVID-19. Asia Pac. J. Ophthalmol. 2020, 9, 85–87. [Google Scholar] [CrossRef]
- Van den Broek, M.; Möhlmann, J.E.; Abeln, B.; Liebregts, M.; van Dijk, V.F.; van de Garde, E. Chloroquine-induced QTc Prolongation in COVID-19 Patients. Neth. Heart J. 2020. [Google Scholar] [CrossRef] [PubMed]
- Szekely, Y.; Lichter, Y.; Shrkihe, B.A.; Bruck, H.; Oster, H.S.; Viskin, S. Chloroquine-induced torsades de pointes in a patient with coronavirus disease 2019. Heart Rhythm 2020, 30420–30423. [Google Scholar] [CrossRef] [PubMed]
- EMA. COVID-19: Chloroquine and Hydroxychloroquine Only to be Used in Clinical Trials or Emergency Use Programmes. Available online: https://www.ema.europa.eu/en/documents/press-release/covid-19-chloroquine-hydroxychloroquine-only-be-used-clinical-trials-emergency-use-programmes_en.pdf (accessed on 1 April 2020).
- WHO. Solidarity Clinical Trial for COVID-19 Treatments. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/global-research-on-novel-coronavirus-2019-ncov/solidarity-clinical-trial-for-covid-19-treatments (accessed on 19 June 2020).
- Muralidharan, N.; Sakthivel, R.; Velmurugan, D.; Gromiha, M.M. Computational Studies of Drug Repurposing and Synergism of Lopinavir, Oseltamivir and Ritonavir Binding with SARS-CoV-2 Protease against COVID-19. J. Biomol. Struct. Dyn. 2020. [Google Scholar] [CrossRef] [PubMed]
- Chu, C.M.; Cheng, V.C.; Hung, I.F.; Wong, M.M.; Chan, K.H.; Chan, K.S.; Kao, R.Y.; Poon, L.L.; Wong, C.L.; Guan, Y.; et al. Role of Lopinavir/Ritonavir in the Treatment of SARS: Initial Virological and Clinical Findings. Thorax 2004, 59, 252–256. [Google Scholar] [CrossRef]
- Lin, P.Y.; Chou, C.Y.; Chang, H.C.; Hsu, W.C.; Chang, G.G. Correlation between Dissociation and Catalysis of SARS-CoV Main Protease. Arch Biochem. Biophys. 2008, 472, 34–42. [Google Scholar] [CrossRef]
- Dayer, M.R.; Taleb-Gassabi, S.; Dayer, M.S. Lopinavir; a potent drug against coronavirus infection: Insight from molecular docking study. Arch. Clin. Infect. Dis. 2017, 12. [Google Scholar] [CrossRef]
- Tu, Y.F.; Chien, C.S.; Yarmishyn, A.A.; Lin, Y.Y.; Luo, Y.H.; Lin, Y.T.; Lai, W.Y.; Yang, D.M.; Chou, S.J.; Yang, Y.P.; et al. A Review of SARS-CoV-2 and the Ongoing Clinical Trials. Int. J. Mol. Sci. 2020, 21, 2657. [Google Scholar] [CrossRef]
- ESICM. Surviving Sepsis Campaign Rapid Guidelines on the Management of Critically Ill Adults with Coronavirus Disease 2019. Available online: https://www.esicm.org/ssc-covid19-guidelines/ (accessed on 1 May 2020).
- EMA. Kaletra, Product Sheet 2020. Available online: https://www.ema.europa.eu/en/documents/product-information/kaletra-epar-product-information_en.pdf (accessed on 25 April 2020).
- Sun, J.; Deng, X.; Chen, X.; Huang, J.; Huang, S.; Li, Y.; Feng, J.; Liu, J.; He, G. Incidence of Adverse Drug Reactions in COVID-19 Patients in China: An Active Monitoring Study by Hospital Pharmacovigilance System. Clin. Pharmacol. Ther. 2020. [Google Scholar] [CrossRef]
- IDSA. Infectious Diseases Society of America Guidelines on the Treatment and Management of Patients with COVID-19. Available online: https://www.idsociety.org/practice-guideline/covid-19-guideline-treatment-and-management/ (accessed on 25 April 2020).
- Gori, T.; Lelieveld, J.; Münzel, T. Perspective: Cardiovascular Disease and the Covid-19 Pandemic. Basic Res. Cardiol. 2020, 115, 32. [Google Scholar] [CrossRef]
- Bassetti, M.; Giacobbe, D.R.; Aliberti, S.; Barisione, E.; Centanni, S.; De Rosa, F.G.; Di Marco, F.; Gori, A.; Granata, G.; Mikulska, M.; et al. Balancing Evidence and Frontline Experience in the Early Phases of the COVID-19 Pandemic: Current Position of the Italian Society of Anti-infective Therapy (SITA) and the Italian Society of Pulmonology (SIP). Clin. Microbiol. Infect. 2020. [Google Scholar] [CrossRef] [PubMed]
- Naksuk, N.; Lazar, S.; Peeraphatdit, T.B. Cardiac Safety of Off-Label COVID-19 Drug Therapy: A Review and Proposed Monitoring Protocol. Eur. Heart J. Acute Cardiovasc. Care 2020. [Google Scholar] [CrossRef] [PubMed]
- Nitulescu, G.M.; Paunescu, H.; Moschos, S.A.; Petrakis, D.; Nitulescu, G.; Ion, G.N.D.; Spnadidos, D.A.; Nikolouzakis, T.K.; Drakoulis, N.; Tsatsakis, A. Comprehensive Analysis of Drugs to Treat SARS-CoV-2 Infection: Mechanistic Insights Into Current COVID-19 Therapies (Review). Int. J. Mol. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Lew, W.; Chen, X.; Kim, C.U. Discovery and Development of GS 4104 (Oseltamivir): An Orally Active Influenza Neuraminidase Inhibitor. Curr. Med. Chem. 2000, 7, 663–672. [Google Scholar] [CrossRef]
- Davies, B.E. Pharmacokinetics of Oseltamivir: An Oral Antiviral for the Treatment and Prophylaxis of Influenza in Diverse Populations. J. Antimicrob. Chemother. 2010, 65. [Google Scholar] [CrossRef]
- Thomas, E.; Ghany, M.G.; Liang, T.J. Review the application and mechanism of action of ribavirin in therapy of hepatitis C. Antivir. Chem. Chemother. 2012, 23, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Elfiky, A.A. Anti-HCV, nucleotide inhibitors, repurposing against COVID-19. Life Sci. 2020, 248, 117477. [Google Scholar] [CrossRef]
- Crotty, S.; Cameron, C.; Andino, R. Ribavirin’s Antiviral Mechanism of Action: Lethal Mutagenesis? J. Mol. Med. 2002, 80, 86–95. [Google Scholar] [CrossRef]
- Elfiky, A.A. Ribavirin, Remdesivir, Sofosbuvir, Galidesivir, and Tenofovir against SARS-CoV-2 RNA dependent RNA polymerase (RdRp): A molecular docking study. Life Sci. 2020, 253, 117592. [Google Scholar] [CrossRef]
- Mousavizadeh, L.; Ghasemi, S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J. Microbiol. Immunol. Infect. 2020. [Google Scholar] [CrossRef]
- Dong, L.; Hu, S.; Gao, J. Discovering drugs to treat coronavirus disease 2019 (COVID-19). Drug Discov. Ther. 2020, 14, 58–60. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.X.; Chen, X.P. Favipiravir: Pharmacokinetics and Concerns about Clinical Trials for 2019-NCoV Infection. Clin. Pharmacol. Ther. 2020. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Yang, M.; Liu, D.; Chen, J.; Shu, D.; Xia, J.; Liao, X.; Gu, Y.; Cai, Q.; Yang, Y.; et al. Experimental Treatment with Favipiravir for COVID-19: An Open-Label Control Study. Engineering 2020. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhang, Y.; Huang, J.; Yin, P.; Cheng, Z.; Wu, J.; Chen, S.; Zhang, Y.; Chen, B.; Lu, M.; et al. Favipiravir versus Arbidol for COVID-19: A Randomized Clinical Trial. medRxiv 2020. [Google Scholar] [CrossRef]
- Russian Ministry of Health. Government Registry of Medicines, Medicine Registration License ЛП-006225. Available online: https://grls.rosminzdrav.ru/Grls_View_v2.aspx?routingGuid=38ce634a-8cb4-43ee-9283-5406335095ee&t= (accessed on 18 June 2020).
- Khaerunnisa, S.; Kurniawan, H.; Awaluddin, R.; Suhartati, S. Potential Inhibitor of COVID-19 Main Protease (M pro) from Several Medicinal Plant Compounds by Molecular Docking Study. Preprints 2020. [Google Scholar] [CrossRef]
- Ríos, J.L.; Máñez, S. New Pharmacological Opportunities for Betulinic Acid. Planta Med. 2018, 84, 8–19. [Google Scholar] [CrossRef]
- Phillips, J.; Phillips, I.; Enya, B.; Zhao, H.; Nitta, T. Effect of Betulinic Acid and Its Ionic Derivatives on M-MuLV Replication. Biochem. Biophys. Res. Commun. 2018, 500, 365–369. [Google Scholar] [CrossRef]
- Wen, C.C.; Kuo, Y.H.; Jan, J.T.; Liang, P.H.; Wang, S.Y.; Liu, H.G.; Lee, C.K.; Chang, S.T.; Kuo, C.J.; Lee, S.S.; et al. Specific Plant Terpenoids and Lignoids Possess Potent Antiviral Activities Against Severe Acute Respiratory Syndrome Coronavirus. J. Med. Chem. 2007, 50, 4087–4095. [Google Scholar] [CrossRef]
- Zhang, D.H.; Wu, K.L.; Zhang, X.; Deng, S.Q.; Peng, B. In Silico Screening of Chinese Herbal Medicines With the Potential to Directly Inhibit 2019 Novel Coronavirus. J. Integr. Med. 2020, 18, 152–158. [Google Scholar] [CrossRef]
- Gupta, M.K.; Vemula, S.; Donde, R.; Gouda, G.; Behera, L.; Vadde, R. In-silico Approaches to Detect Inhibitors of the Human Severe Acute Respiratory Syndrome Coronavirus Envelope Protein Ion Channel. J. Biomol. Struct. Dyn. 2020. [Google Scholar] [CrossRef]
- Boopathirajan, P.M.K.; Vijayakumar, K. In-Silico Drug Discovery for Covid19 by Targeting Spike Glycoprotein of SARS COV 2 (Wuhan Corona Virus 2019 Outbreak) Against the Docking Analysis with Structure Predicted Human ‘ACE2-FC Region of IGG1’ Fusion Protein As a Protein Based Drug. IJCRT 2020, 8, 1667–1675. [Google Scholar]
- Cava, C.; Bertoli, G.; Castiglioni, I. In Silico Discovery of Candidate Drugs against Covid-19. Viruses 2020, 12, 404. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Q.; Yang, K.; Wang, W.; Jiang, L.; Song, J. Clinical Predictors of Mortality Due to COVID-19 Based on an Analysis of Data of 150 Patients from Wuhan, China. Intensive Care Med. 2020, 846–848. [Google Scholar] [CrossRef] [PubMed]
- La Rosée, P.; Horne, A.; Hines, M.; von Bahr Greenwood, T.; Machowicz, R.; Berliner, N.; Birndt, S.; Gil-Herrera, J.; Girschikofsky, M.; Jordan, M.B.; et al. Recommendations for the Management of Hemophagocytic Lymphohistiocytosis in Adults. Blood 2019, 133, 2465–2477. [Google Scholar] [CrossRef] [PubMed]
- Van der Ven, A.J.A.M.; Netea, M.G.; van der Meer, J.W.M.; de Mast, Q. Ebola Virus Disease Has Features of Hemophagocytic Lymphohistiocytosis Syndrome. Front. Med. 2015, 2, 4. [Google Scholar] [CrossRef] [PubMed]
- Huet, T.; Beaussier, H.; Voisin, O.; Jouveshomme, S.; Dauriat, G.; Lazareth, I.; Sacco, E.; Naccache, J.M.; Bézie, Y.; Laplanche, S.; et al. Anakinra for Severe Forms of COVID-19: A Cohort Study. Lancet Rheumatol. 2020. [Google Scholar] [CrossRef]
- King, A.; Vail, A.; O’Leary, C.; Hannan, C.; Brough, D.; Patel, H.; Galea, J.; Ogungbenro, K.; Wright, M.; Pathmanaban, O.; et al. Anakinra in COVID-19: Important Considerations for Clinical Trials. Lancet Rheumatol. 2020, 2, 20–22. [Google Scholar] [CrossRef]
- Calina, D.; Docea, A.O.; Petrakis, D.; Egorov, A.M.; Ishmukhametov, A.A.; Gabibov, A.G.; Shtilman, M.I.; Kostoff, R.; Carvalho, F.; Vinceti, M.; et al. Towards Effective COVID-19 Vaccines: Updates, Perspectives and Challenges (Review). Int. J. Mol. Med. 2020. [Google Scholar] [CrossRef]
- Bloch, E.M.; Shoham, S.; Casadevall, A.; Sachais, B.S.; Shaz, B.; Winters, J.L.; van Buskirk, C.; Grossman, B.J.; Joyner, M.; Henderson, J.P.; et al. Deployment of Convalescent Plasma for the Prevention and Treatment of COVID-19. J. Clin. Investig. 2020, 138745. [Google Scholar] [CrossRef]
- Casadevall, A.; Pirofski, L.A. The Convalescent Sera Option for Containing COVID-19. J. Clin. Investig. 2020, 130, 1545–1548. [Google Scholar] [CrossRef]
- U.S. Food and Drug Administration. Recommendations for Investigational COVID-19 Convalescent Plasma | FDA. Available online: https://www.fda.gov/vaccines-blood-biologics/investigational-new-drug-ind-or-device-exemption-ide-process-cber/recommendations-investigational-covid-19-convalescent-plasma (accessed on 4 May 2020).
- Alattar, R.; Ibrahim, T.; Shaar, S.H.; Abdalla, S.; Shukri, K.; Daghfal, J.N.; Khatib, M.Y.; Aboukamar, M.; Abukhattab, M.; Alsoub, H.A.; et al. Tocilizumab for the Treatment of Severe Coronavirus Disease 2019. J. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Jalkanen, J.; Hollmén, M.; Jalkanen, S. Interferon Beta-1a for COVID-19: Critical Importance of the Administration Route. Crit. Care 2020, 24, 335. [Google Scholar] [CrossRef] [PubMed]
- Siddiqi, H.K.; Mehra, M.R. COVID-19 Illness in Native and Immunosuppressed States: A Clinical-Therapeutic Staging Proposal. J. Heart Lung Transpl. 2020, 39, 405–407. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical Course and Risk Factors for Mortality of Adult Inpatients with COVID-19 in Wuhan, China: A Retrospective Cohort Study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Dastan, F.; Nadji, S.A.; Saffaei, A.; Marjani, M.; Moniri, A.; Jamaati, H.; Hashemian, S.M.; Baghaei, P.; Abedini, A.; Varahram, M.; et al. Subcutaneous Administration of Interferon Beta-1a for COVID-19: A Non-Controlled Prospective Trial. Int. Immunopharmacol. 2020, 85, 106688. [Google Scholar] [CrossRef]
- Hung, I.F.N.; Lung, K.C.; Tso, E.Y.K.; Liu, R.; Chung, T.W.H.; Chu, M.Y.; Ng, Y.Y.; Lo, J.; Chan, J.; Tam, A.R.; et al. Triple Combination of Interferon Beta-1b, Lopinavir-Ritonavir, and Ribavirin in the Treatment of Patients Admitted to Hospital with COVID-19: An Open-Label, Randomised, Phase 2 Trial. Lancet 2020, 395, 1695–1704. [Google Scholar] [CrossRef]
- Tang, N.; Bai, H.; Chen, X.; Gong, J.; Li, D.; Sun, Z. Anticoagulant Treatment Is Associated With Decreased Mortality in Severe Coronavirus Disease 2019 Patients with Coagulopathy. J. Thromb. Haemost. 2020, 18, 1094–1099. [Google Scholar] [CrossRef]
- Connors, J.M.; Levy, J.H. COVID-19 and its implications for thrombosis and anticoagulation. Blood 2020. [Google Scholar] [CrossRef]
- Song, J.C.; Wang, G.; Zhang, W.; Zhang, Y.; Li, W.Q.; Zhou, Z.; People’s Liberation Army Professional Committee of Critical Care Medicine, Chinese Society on Thrombosis and Haemostasis. Chinese Expert Consensus on Diagnosis and Treatment of Coagulation Dysfunction in COVID-19. Mil. Med. Res. 2020, 7, 19. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).