Nanoribbon-Based Electronic Detection of a Glioma-Associated Circular miRNA
Abstract
:1. Introduction
2. Materials and Methods
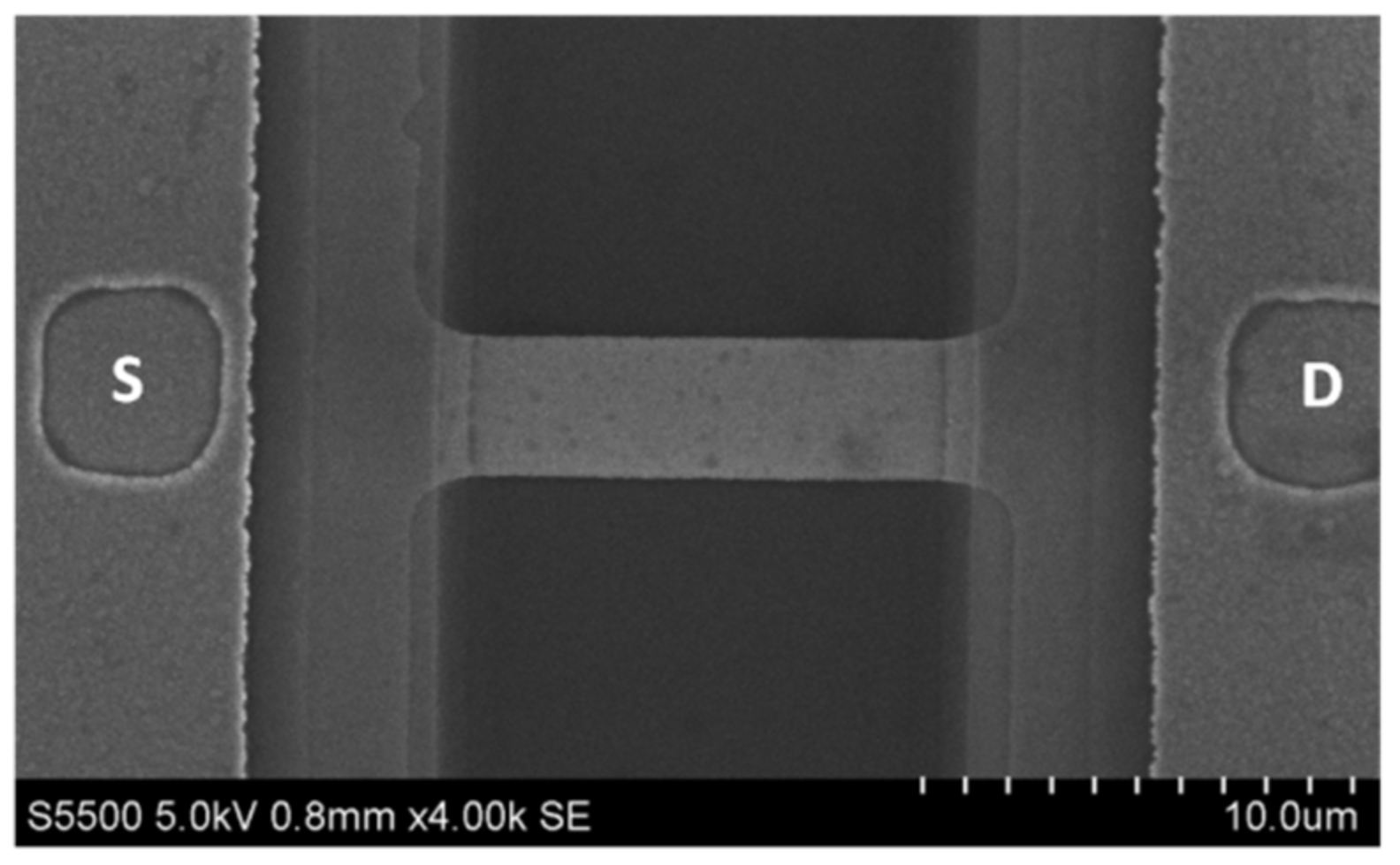
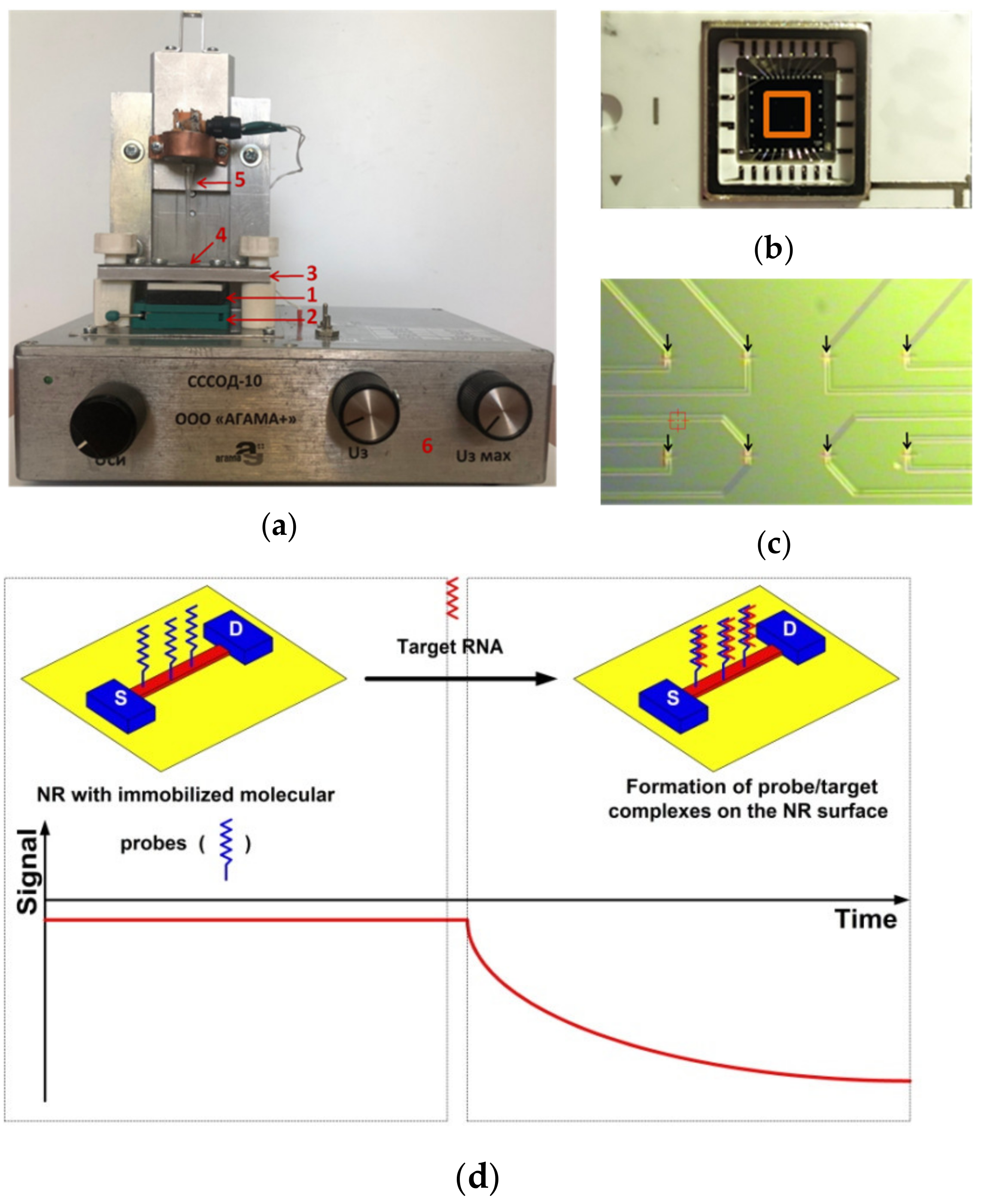
2.1. Nanoribbon Biosensor
2.2. Fabrication of the SOI-NR Chips
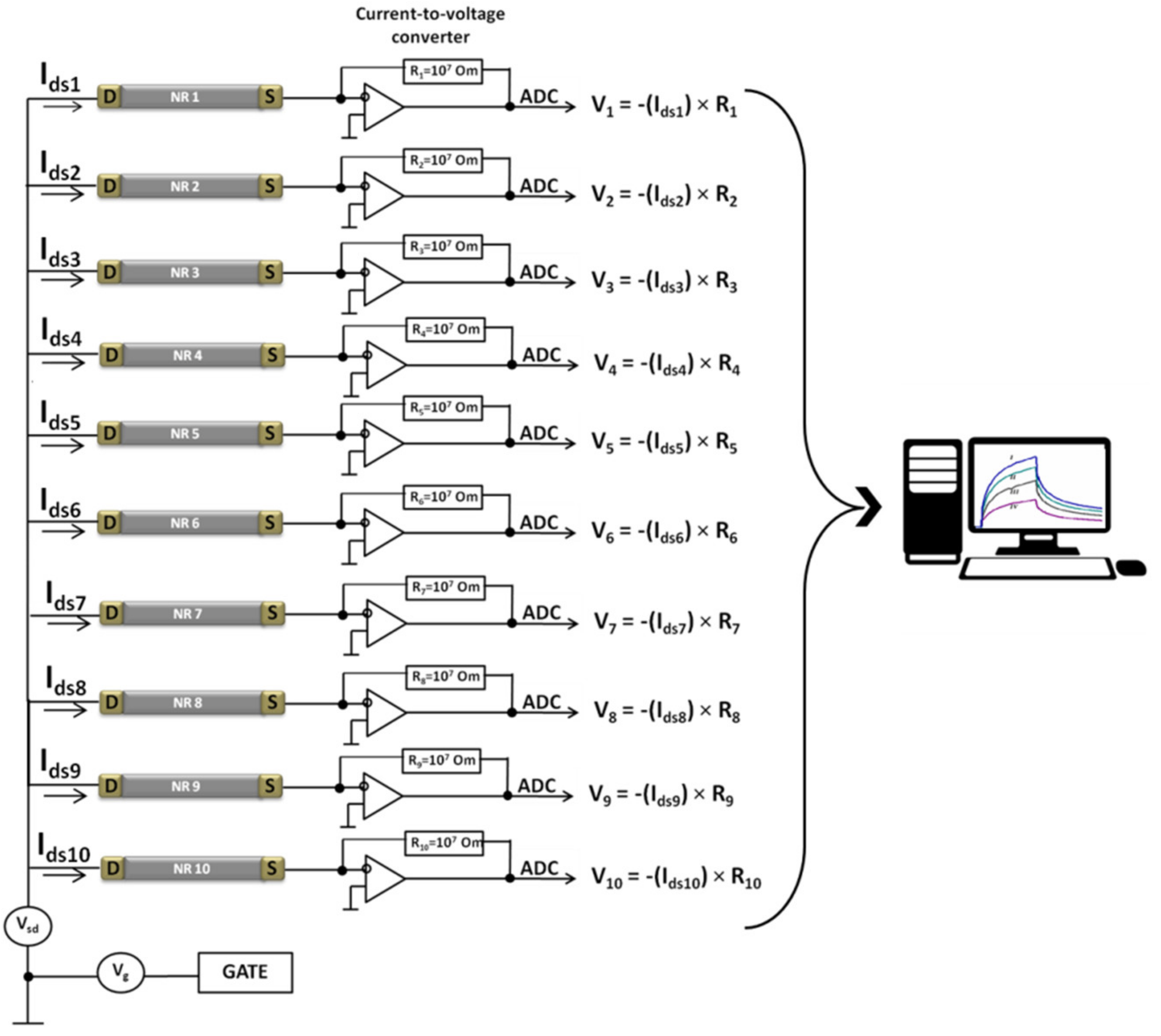
2.3. Electrical Measurements
2.4. Chemicals
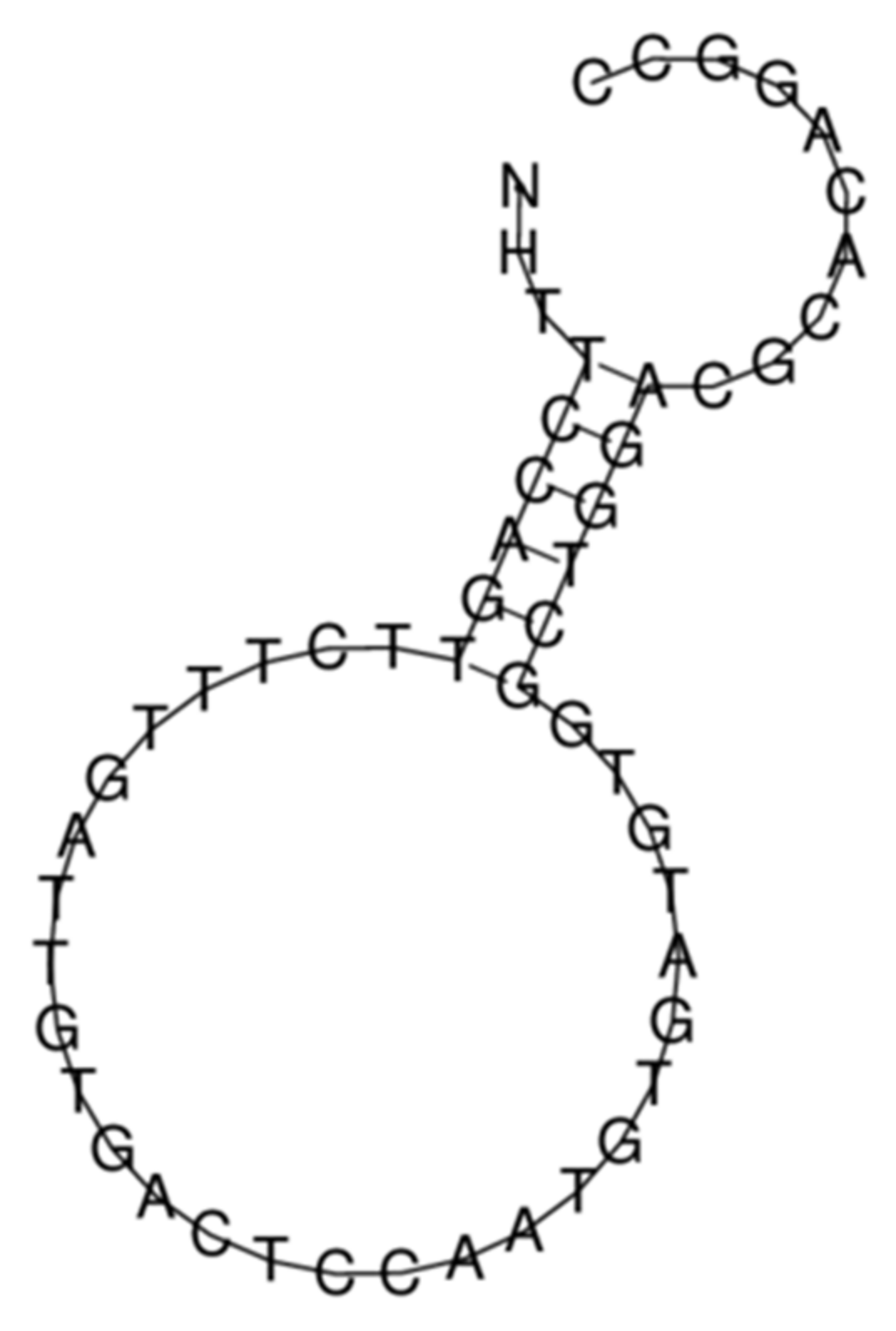
2.5. DNA Oligonucleotides and Circular RNA
2.6. Plasma Samples
2.7. Chemical Treatment of the SOI-NR Chip Surface
2.8. Covalent Immobilization of the oDNA Probes
2.9. Preparation of Solutions of Target oDNA in Purified Buffer
3. Results
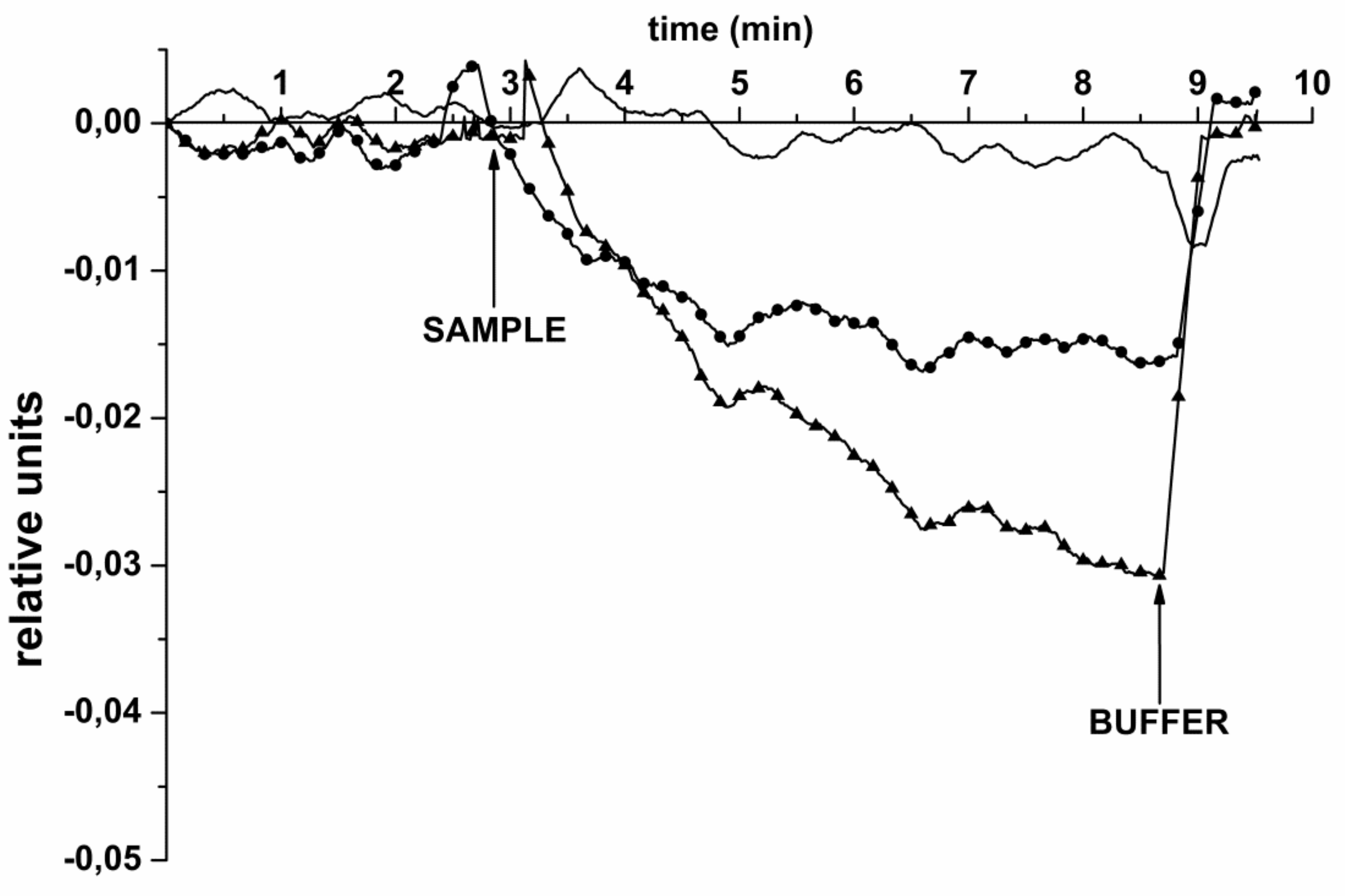
3.1. oDNA Detection in Buffer
3.2. Detection of circNFIX RNA, Isolated from Plasma
4. Discussion
- (1)
- Upon the registration of the biosensor signal in the experiments on the detection of oDNA in purified buffer solution at known target oDNA concentrations, the signal received from the working NR (whose surface is sensitized with immobilized molecular probes) should exceed that received from the control NR. The control NR is used in order to account for the non-specific adsorption, and the differential signal (i.e., the signal from the working NR minus the signal from the control NR) is used in the entire concentration range studied;
- (2)
- In the experiments with plasma samples, the differential signal (the signal from the working NR minus the signal from the control NR), registered after the addition of the sample of a glioma patient, must exceed each of the differential signals obtained upon the analysis of (a) the sample of a healthy volunteer, and (b) the sample of a patient suffering from a different type of disease (in our case, from prostatic hyperplasia).
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- McKinney, P.A. Brain tumours: Incidence, survival, and aetiology. J. Neurol. Neurosurg. Psychiatry 2004, 75, ii12–ii17. [Google Scholar] [CrossRef] [Green Version]
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ostrom, Q.; Gittleman, H.; Farah, P.; Ondracek, A.; Chen, Y.; Wolinsky, Y.; Stroup, N.E.; Kruchko, C.; Barnholtz-Sloan, J. CBTRUS Statistical Report: Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2006–2010. Neuro-Oncology 2013, 15, ii1–ii56. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koshy, M.; Villano, J.L.; Dolecek, T.A.; Howard, A.; Mahmood, U.; Chmura, S.J.; Weichselbaum, R.R.; McCarthy, B.J. Improved survival time trends for glioblastoma using the SEER 17 population-based registries. J. Neuro-Oncol. 2012, 107, 207–212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stupp, R.; Mason, W.P.; van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus Concomitant and Adjuvant Temozolomide for Glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef] [PubMed]
- Faguer, R.; Tanguy, J.-Y.; Rousseau, A.; Clavreul, A.; Menei, P. Early presentation of primary glioblastoma. Neurochirurgie 2014, 60, 188–193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ideguchi, M.; Kajiwara, K.; Goto, H.; Sugimoto, K.; Nomura, S.; Ikeda, E.; Suzuki, M. MRI findings and pathological features in early-stage glioblastoma. J. Neuro-Oncol. 2015, 123, 289–297. [Google Scholar] [CrossRef] [PubMed]
- Oyama, H.; Ando, Y.; Aoki, S.; Kito, A.; Maki, H.; Hattori, K.; Tanahashi, K. Glioblastoma Detected at the Initial Stage in its Developmental Process -Case Report-. Neurol. Med. Chir. 2010, 50, 414–417. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zachariah, M.A.; Oliveira-Costa, J.P.; Carter, B.S.; Stott, S.L.; Nahed, B.V. Blood-based biomarkers for the diagnosis and monitoring of gliomas. Neuro-Oncology 2018, 20, 1155–1161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adachi-Hayama, M.; Adachi, A.; Shinozaki, N.; Matsutani, T.; Hiwasa, T.; Takiguchi, M.; Saeki, N.; Iwadate, Y. Circulating anti-filamin C autoantibody as a potential serum biomarker for low-grade gliomas. BMC Cancer 2014, 14, 452. [Google Scholar] [CrossRef] [Green Version]
- Beermann, J.; Piccoli, M.-T.; Viereck, J.; Thum, T. Non-coding RNAs in Development and Disease: Background, Mechanisms, and Therapeutic Approaches. Physiol. Rev. 2016, 96, 1297–1325. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, J.; Xie, Q.; Xu, H.; Li, J.; Li, Y. Circular RNAs and cancer. Cancer Lett. 2017, 396, 138–144. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.U.; Agarwal, V.; Guo, H.; Bartel, D.P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014, 15, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zheng, Q.; Bao, C.; Li, S.; Guo, W.; Zhao, J.; Chen, D.; Gu, J.; He, X.; Huang, S. Circular RNA is enriched and stable in exosomes: A promising biomarker for cancer diagnosis. Cell Res. 2015, 25, 981–984. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, Q.; Ding, T.; Zhang, G.; Li, Z.; Zeng, L.; Zhu, Y.; Guo, J.; Hou, J.; Zhu, T.; Zheng, J.; et al. Circular RNA Expression Profiling Identifies Prostate Cancer- Specific circRNAs in Prostate Cancer. Cell. Physiol. Biochem. 2018, 50, 1903–1915. [Google Scholar] [CrossRef] [PubMed]
- Rynkeviciene, R.; Simiene, J.; Strainiene, E.; Stankevicius, V.; Usinskiene, J.; Kaubriene, E.M.; Meskinyte, I.; Cicenas, J.; Suziedelis, K. Non-Coding RNAs in Glioma. Cancers 2018, 11, 17. [Google Scholar] [CrossRef] [Green Version]
- Jinglei, L.; Kun, Z.; Nunu, H.; Nu, Z.; Liu, J.; Zhao, K.; Huang, N.; Zhang, N. Circular RNAs and human glioma. Cancer Biol. Med. 2019, 16, 11–23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, X.; Zhang, N.; Han, P.; Moon, B.-S.; Lai, R.K.; Wang, K.; Lu, W. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res. 2016, 44, e87. [Google Scholar] [CrossRef] [PubMed]
- Piper, M.; Gronostajski, R.; Messina, G. Nuclear Factor One X in Development and Disease. Trends Cell Biol. 2019, 29, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C.E.; Piper, M.; Plachez, C.; Yeh, Y.-T.; Baizer, J.S.; Osinski, J.M.; Litwack, E.D.; Richards, L.J.; Gronostajski, R.M. The transcription factor Nfix is essential for normal brain development. BMC Dev. Biol. 2008, 8, 52. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, H.; Zhang, Y.; Qi, L.; Ding, L.; Jiang, H.; Yu, H. NFIX Circular RNA Promotes Glioma Progression by Regulating miR-34a-5p via Notch Signaling Pathway. Front. Mol. Neurosci. 2018, 11, 225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Yang, L.; Chen, L.-L. The Biogenesis, Functions, and Challenges of Circular RNAs. Mol. Cell 2018, 71, 428–442. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szabo, L.; Salzman, J. Detecting circular RNAs: Bioinformatic and experimental challenges. Nat. Rev. Genet. 2016, 17, 679–692. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hayer, K.E.; Pizarro, A.D.; Lahens, N.F.; HogenEsch, J.B.; Grant, G. Benchmark analysis of algorithms for determining and quantifying full-length mRNA splice forms from RNA-seq data. Bioinformatics 2015, 31, 3938–3945. [Google Scholar] [CrossRef] [PubMed]
- Carrara, M.; Lum, J.; Cordero, F.; Beccuti, M.; Poidinger, M.; Donatelli, S.; Calogero, R.A.; Zolezzi, F. Alternative splicing detection workflow needs a careful combination of sample prep and bioinformatics analysis. BMC Bioinform. 2015, 16, S2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rissin, D.M.; Kan, C.W.; Campbell, T.G.; Howes, S.C.; Fournier, D.R.; Song, L.; Piech, T.; Patel, P.P.; Chang, L.; Rivnak, A.J.; et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nat. Biotechnol. 2010, 28, 595–599. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pleshakova, T.O.; Shumov, I.D.; Ivanov, Y.D.; Malsagova, K.; Kaysheva, A.L.; Archakov, A.I. AFM-based technologies as the way towards the reverse Avogadro number. Biochem. Suppl. Ser. B Biomed. Chem. 2015, 9, 244–257. [Google Scholar] [CrossRef]
- Malsagova, K.A.; Pleshakova, T.O.; Galiullin, R.A.; Shumov, I.D.; Kozlov, A.F.; Romanova, T.S.; Popov, V.P.; Glukhov, A.V.; Konev, V.A.; Archakov, A.I.; et al. Nanowire Aptamer-Sensitized Biosensor Chips with Gas Plasma-Treated Surface for the Detection of Hepatitis C Virus Core Antigen. Coatings 2020, 10, 753. [Google Scholar] [CrossRef]
- Ivanov, Y.D.; Pleshakova, T.; Malsagova, K.; Kozlov, A.; Kaysheva, A.; Shumov, I.; Galiullin, R.; Kurbatov, L.; Popov, V.; Naumova, O.; et al. Detection of marker miRNAs in plasma using SOI-NW biosensor. Sens. Actuators B Chem. 2018, 261, 566–571. [Google Scholar] [CrossRef]
- Li, J.; He, G.; Ueno, H.; Jia, C.; Noji, H.; Qi, C.; Guo, X. Direct real-time detection of single proteins using silicon nanowire-based electrical circuits. Nanoscale 2016, 8, 16172–16176. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Zhang, G.-J. Silicon nanowire-transistor biosensor for study of molecule-molecule interactions. Rev. Anal. Chem. 2014, 33. [Google Scholar] [CrossRef]
- Malsagova, K.; Pleshakova, T.; Popov, V.; Kupriyanov, I.; Galiullin, R.; Kozlov, A.; Shumov, I.; Kaysheva, A.; Tikhonenko, F.; Archakov, A.; et al. Optical Monitoring of the Production Quality of Si-Nanoribbon Chips Intended for the Detection of ASD-Associated Oligonucleotides. Micromachines 2021, 12, 147. [Google Scholar] [CrossRef] [PubMed]
- Malsagova, K.; Popov, V.; Kupriyanov, I.; Pleshakova, T.; Galiullin, R.; Kozlov, A.; Shumov, I.; Larionov, D.; Tikhonenko, F.; Kapustina, S.; et al. Raman Spectroscopy-Based Quality Control of “Silicon-On-Insulator” Nanowire Chips for the Detection of Brain Cancer-Associated MicroRNA in Plasma. Sensors 2021, 21, 1333. [Google Scholar] [CrossRef] [PubMed]
- Malsagova, K.; Pleshakova, T.; Galiullin, R.; Kozlov, A.; Shumov, I.; Popov, V.; Tikhonenko, F.; Glukhov, A.; Ziborov, V.; Petrov, O.; et al. Highly Sensitive Detection of CA 125 Protein with the Use of an n-Type Nanowire Biosensor. Biosensors 2020, 10, 210. [Google Scholar] [CrossRef] [PubMed]
- Malsagova, K.A.; Pleshakova, T.O.; Galiullin, R.A.; Kozlov, A.F.; Romanova, T.S.; Shumov, I.D.; Popov, V.P.; Tikhonenko, F.V.; Glukhov, A.V.; Smirnov, A.Y.; et al. SOI-Nanowire Biosensor for the Detection of Glioma-Associated miRNAs in Plasma. Chemosensors 2020, 8, 95. [Google Scholar] [CrossRef]
- Ivanov, Y.; Pleshakova, T.; Malsagova, K.; Kurbatov, L.; Popov, V.; Glukhov, A.; Smirnov, A.; Enikeev, D.; Potoldykova, N.; Alekseev, B.; et al. Detection of Marker miRNAs, Associated with Prostate Cancer, in Plasma Using SOI-NW Biosensor in Direct and Inversion Modes. Sensors 2019, 19, 5248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Malsagova, K.A.; Pleshakova, T.O.; Kozlov, A.F.; Shumov, I.D.; Ilnitskii, M.A.; Miakonkikh, A.V.; Popov, V.P.; Rudenko, K.V.; Glukhov, A.V.; Kupriyanov, I.N.; et al. Micro-Raman Spectroscopy for Monitoring of Deposition Quality of High-k Stack Protective Layer onto Nanowire FET Chips for Highly Sensitive miRNA Detection. Biosensors 2018, 8, 72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Doucey, M.-A.; Carrara, S. Nanowire Sensors in Cancer. Trends Biotechnol. 2019, 37, 86–99. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chiang, P.-L.; Chou, T.-C.; Wu, T.-H.; Li, C.-C.; Liao, C.-D.; Lin, J.-Y.; Tsai, M.-H.; Tsai, C.-C.; Sun, C.-J.; Wang, C.-H.; et al. Nanowire Transistor-Based Ultrasensitive Virus Detection with Reversible Surface Functionalization. Chem. Asian J. 2012, 7, 2073–2079. [Google Scholar] [CrossRef]
- Patolsky, F.; Zheng, G.; Hayden, O.; Lakadamyali, M.; Zhuang, X.; Lieber, C.M. Electrical detection of single viruses. Proc. Natl. Acad. Sci. USA 2004, 101, 14017–14022. [Google Scholar] [CrossRef] [Green Version]
- Elfström, N.; Juhasz, R.; Sychugov, I.; Engfeldt, T.; Karlström, A.E.; Linnros, J. Surface Charge Sensitivity of Silicon Nanowires: Size Dependence. Nano Lett. 2007, 7, 2608–2612. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, H.; Han, X.; Ou, X.; Lee, C.-S.; Zhang, X.; Lee, S.-T. Silicon nanowire based single-molecule SERS sensor. Nanoscale 2013, 5, 8172–8176. [Google Scholar] [CrossRef]
- Hahm, J.-I.; Lieber, C.M. Direct Ultrasensitive Electrical Detection of DNA and DNA Sequence Variations Using Nanowire Nanosensors. Nano Lett. 2004, 4, 51–54. [Google Scholar] [CrossRef]
- Zhang, P.; Gao, K.; Liang, Y.; Su, F.; Wang, F.; Li, Z. Ultrasensitive detection of circular RNA by accurate recognition of the specific junction site using stem-loop primer induced double exponential amplification. Talanta 2020, 217, 121021. [Google Scholar] [CrossRef] [PubMed]
- Boss, M.; Arenz, C. A Fast and Easy Method for Specific Detection of Circular RNA by Rolling-Circle Amplification. ChemBioChem 2020, 21, 793–796. [Google Scholar] [CrossRef] [Green Version]
- Jiao, J.; Gao, T.; Shi, H.; Sheng, A.; Xiang, Y.; Shu, Y.; Li, G. A method to directly assay circRNA in real samples. Chem. Commun. 2018, 54, 13451–13454. [Google Scholar] [CrossRef] [PubMed]
- Popov, V.P.; Antonova, A.I.; Frantsuzov, A.A.; Safronov, L.N.; Feofanov, G.N.; Naumova, O.V.; Kilanov, D.V. Properties of silicon-on-insulator structures and devices. Semiconductors 2001, 35, 1030–1037. [Google Scholar] [CrossRef]
- Ivanov, Y.D.; Tatur, V.Y.; Glukhov, A.V.; Ziborov, V.S. Nanowire biosensor for registration of biomolecules. Zametki Uchenogo 2021, 7, 56–61. [Google Scholar]


| Plasma Sample # | Age | Sex | Pathology | WHO Grade |
|---|---|---|---|---|
| 001 | 67 | M | Anaplastic oligodendroglioma | III |
| 002 | 42 | F | Anaplastic astrocytoma | III |
| 14 | 73 | M | Prostatic hyperplasia | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ivanov, Y.D.; Malsagova, K.A.; Popov, V.P.; Pleshakova, T.O.; Kozlov, A.F.; Galiullin, R.A.; Shumov, I.D.; Kapustina, S.I.; Tikhonenko, F.V.; Ziborov, V.S.; et al. Nanoribbon-Based Electronic Detection of a Glioma-Associated Circular miRNA. Biosensors 2021, 11, 237. https://doi.org/10.3390/bios11070237
Ivanov YD, Malsagova KA, Popov VP, Pleshakova TO, Kozlov AF, Galiullin RA, Shumov ID, Kapustina SI, Tikhonenko FV, Ziborov VS, et al. Nanoribbon-Based Electronic Detection of a Glioma-Associated Circular miRNA. Biosensors. 2021; 11(7):237. https://doi.org/10.3390/bios11070237
Chicago/Turabian StyleIvanov, Yuri D., Kristina A. Malsagova, Vladimir P. Popov, Tatyana O. Pleshakova, Andrey F. Kozlov, Rafael A. Galiullin, Ivan D. Shumov, Svetlana I. Kapustina, Fedor V. Tikhonenko, Vadim S. Ziborov, and et al. 2021. "Nanoribbon-Based Electronic Detection of a Glioma-Associated Circular miRNA" Biosensors 11, no. 7: 237. https://doi.org/10.3390/bios11070237
APA StyleIvanov, Y. D., Malsagova, K. A., Popov, V. P., Pleshakova, T. O., Kozlov, A. F., Galiullin, R. A., Shumov, I. D., Kapustina, S. I., Tikhonenko, F. V., Ziborov, V. S., Dolgoborodov, A. Y., Petrov, O. F., Gadzhieva, O. A., Bashiryan, B. A., Shimansky, V. N., Potoldykova, N. V., Enikeev, D. V., Usachev, D. Y., & Archakov, A. I. (2021). Nanoribbon-Based Electronic Detection of a Glioma-Associated Circular miRNA. Biosensors, 11(7), 237. https://doi.org/10.3390/bios11070237






