Efforts to Minimise the Bacterial Genome as a Free-Living Growing System
Abstract
Simple Summary
Abstract
1. Introduction
2. Genetic Requirement for Minimal Genome
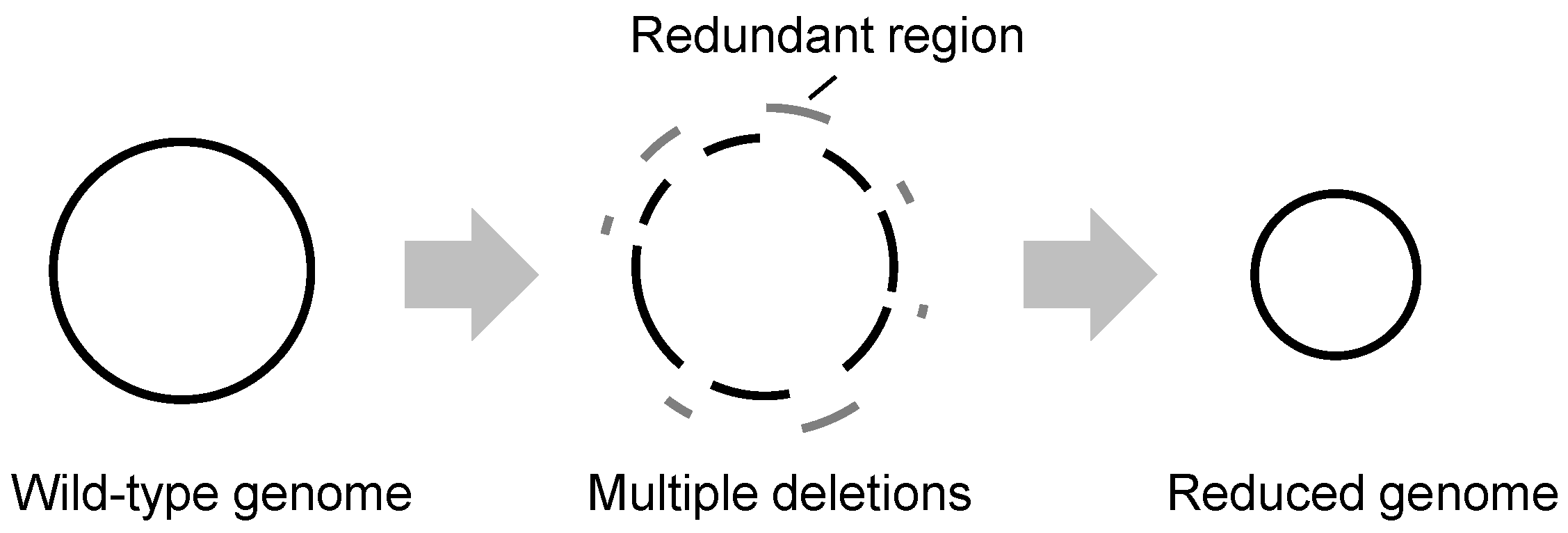
2.1. Genome Reduction
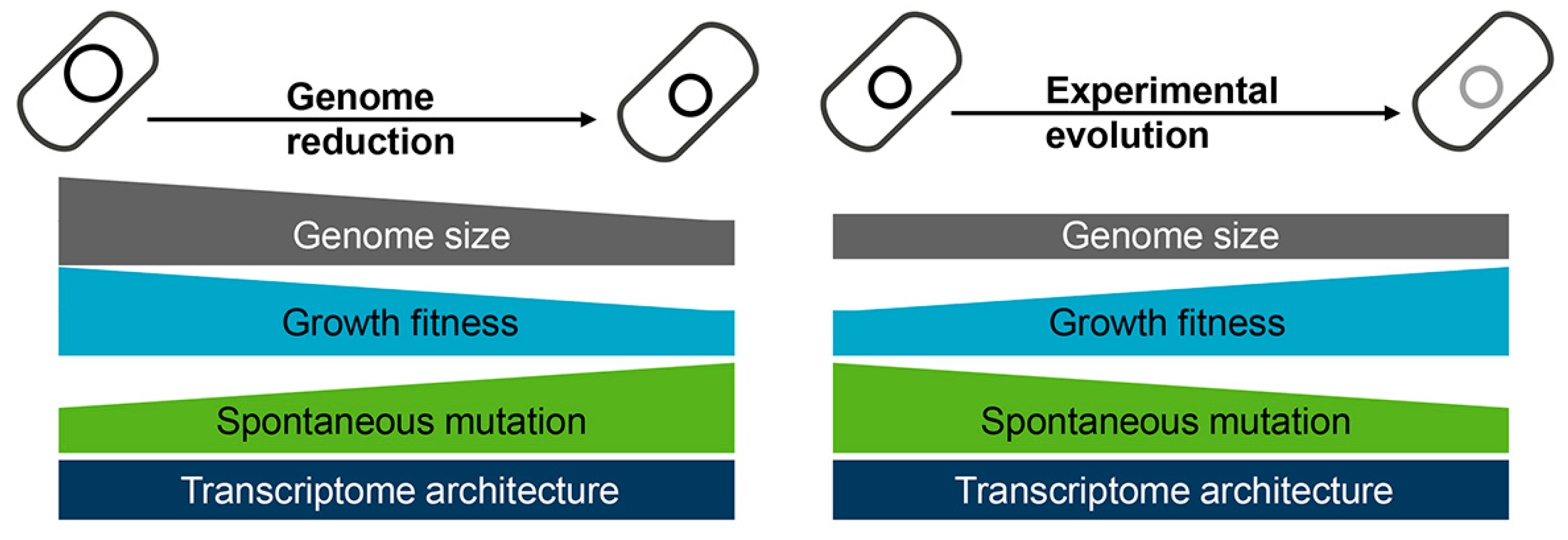
2.2. Evolutionary Approaches for Reduced Genome
3. Environmental Requirement for Minimal Genome
3.1. Culture Medium
3.2. Medium Optimization
4. Machine Learning-Based Minimal Genome Methods
4.1. Machine Learning-Assisted Medium Optimization
4.2. Active Learning
5. Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Campillo-Balderas, J.A.; Lazcano, A.; Becerra, A. Viral genome size distribution does not correlate with the antiquity of the host lineages. Front. Ecol. Evol. 2015, 3, 143. [Google Scholar] [CrossRef]
- Pellicer, J.; Fay, M.F.; Leitch, I.J. The largest eukaryotic genome of them all? Bot. J. Linn. Soc. 2010, 164, 10–15. [Google Scholar] [CrossRef]
- Leitch, I.J.; Pellicer, J.; Hidalgo, O.; Bennett, M.D. Plant DNA C-values Database (Release 7.1). New Phytol. 2019. [Google Scholar] [CrossRef]
- Rodríguez-Gijón, A.; Nuy, J.K.; Mehrshad, M.; Buck, M.; Schulz, F.; Woyke, T.; Garcia, S.L. A Genomic Perspective Across Earth’s Microbiomes Reveals That Genome Size in Archaea and Bacteria Is Linked to Ecosystem Type and Trophic Strategy. Front. Microbiol. 2021, 12, 761869. [Google Scholar] [CrossRef]
- Sezonov, G.; Joseleau-Petit, D.; D’Ari, R. Escherichia coli physiology in Luria-Bertani broth. J. Bacteriol. 2007, 189, 8746–8749. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 1983, 166, 557–580. [Google Scholar] [CrossRef]
- Blattner, F.R.; Plunkett, G., III; Bloch, C.A.; Perna, N.T.; Burland, V.; Riley, M.; Collado-Vides, J.; Glasner, J.D.; Rode, C.K.; Mayhew, G.F.; et al. The Complete Genome Sequence of Escherichia coli K-12. Science 1997, 277, 1453–1462. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Antonio, A.; Janga, S.C.; Thieffry, D. Functional organisation of Escherichia coli transcriptional regulatory network. J. Mol. Biol. 2008, 381, 238–247. [Google Scholar] [CrossRef]
- Fang, X.; Sastry, A.; Mih, N.; Kim, D.; Tan, J.; Yurkovich, J.T.; Lloyd, C.J.; Gao, Y.; Yang, L.; Palsson, B.O. Global transcriptional regulatory network for Escherichia coli robustly connects gene expression to transcription factor activities. Proc. Natl. Acad. Sci. USA 2017, 114, 10286–10291. [Google Scholar] [CrossRef]
- Fu, Y.; Jarboe, L.R.; Dickerson, J.A. Reconstructing genome-wide regulatory network of E. coli using transcriptome data and predicted transcription factor activities. BMC Bioinform. 2011, 12, 233. [Google Scholar] [CrossRef]
- Al-Aamri, A.; Kudlicki, A.S.; Maalouf, M.; Taha, K.; Homouz, D. Inferring Gene Regulatory Networks from RNA-seq Data Using Kernel Classification. Biology 2023, 12, 518. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Mahajan, S.; Jain, S. Feedbacks from the metabolic network to the genetic network reveal regulatory modules in E. coli and B. subtilis. PLoS ONE 2018, 13, e0203311. [Google Scholar] [CrossRef]
- Larsen, S.J.; Röttger, R.; Schmidt, H.H.H.W.; Baumbach, J. E. coli gene regulatory networks are inconsistent with gene expression data. Nucleic Acids Res. 2019, 47, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Hutchison, C.A., 3rd; Chuang, R.-Y.; Noskov, V.N.; Assad-Garcia, N.; Deerinck, T.J.; Ellisman, M.H.; Gill, J.; Kannan, K.; Karas, B.J.; Ma, L.; et al. Design and synthesis of a minimal bacterial genome. Science 2016, 351, aad6253. [Google Scholar] [CrossRef] [PubMed]
- Gibson, D.G.; Glass, J.I.; Lartigue, C.; Noskov, V.N.; Chuang, R.-Y.; Algire, M.A.; Benders, G.A.; Montague, M.G.; Ma, L.; Moodie, M.M.; et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 2010, 329, 52–56. [Google Scholar] [CrossRef]
- Gibson, D.G.; Benders, G.A.; Andrews-Pfannkoch, C.; Denisova, E.A.; Baden-Tillson, H.; Zaveri, J.; Stockwell, T.B.; Brownley, A.; Thomas, D.W.; Algire, M.A.; et al. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science 2008, 319, 1215–1220. [Google Scholar] [CrossRef]
- Posfai, G.; Kolisnychenko, V.; Bereczki, Z.; Blattner, F.R. Markerless gene replacement in Escherichia coli stimulated by a double-strand break in the chromosome. Nucleic Acids Res. 1999, 27, 4409–4415. [Google Scholar] [CrossRef]
- Kato, J.-I.; Hashimoto, M. Construction of long chromosomal deletion mutants of Escherichia coli and minimization of the genome. Methods Mol. Biol. 2008, 416, 279–293. [Google Scholar] [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Li, Y.; Lin, Z.; Huang, C.; Zhang, Y.; Wang, Z.; Tang, Y.-J.; Chen, T.; Zhao, X. Metabolic engineering of Escherichia coli using CRISPR–Cas9 meditated genome editing. Metab. Eng. 2015, 31, 13–21. [Google Scholar] [CrossRef]
- Ma, S.; Su, T.; Liu, J.; Lu, X.; Qi, Q. Reduction of the Bacterial Genome by Transposon-Mediated Random Deletion. ACS Synth. Biol. 2022, 11, 668–677. [Google Scholar] [CrossRef] [PubMed]
- LeBlanc, N.; Charles, T.C. Bacterial genome reductions: Tools, applications, and challenges. Front. Genome Ed. 2022, 4, 957289. [Google Scholar] [CrossRef] [PubMed]
- Karcagi, I.; Draskovits, G.; Umenhoffer, K.; Fekete, G.; Kovács, K.; Méhi, O.; Balikó, G.; Szappanos, B.; Györfy, Z.; Fehér, T.; et al. Indispensability of Horizontally Transferred Genes and Its Impact on Bacterial Genome Streamlining. Mol. Biol. Evol. 2016, 33, 1257–1269. [Google Scholar] [CrossRef]
- Posfai, G.; Plunkett, G., 3rd; Fehér, T.; Frisch, D.; Keil, G.M.; Umenhoffer, K.; Kolisnychenko, V.; Stahl, B.; Sharma, S.S.; de Arruda, M.; et al. Emergent properties of reduced-genome Escherichia coli. Science 2006, 312, 1044–1046. [Google Scholar] [CrossRef]
- Kato, J.; Hashimoto, M. Construction of consecutive deletions of the Escherichia coli chromosome. Mol. Syst. Biol. 2007, 3, 132. [Google Scholar] [CrossRef] [PubMed]
- Kotaka, Y.; Hashimoto, M.; Lee, K.-I.; Kato, J.-I. Mutations identified in engineered Escherichia coli with a reduced genome. Front. Microbiol. 2023, 14, 1189877. [Google Scholar] [CrossRef]
- Kurokawa, M.; Ying, B.-W. Experimental Challenges for Reduced Genomes: The Cell Model Escherichia coli. Microorganisms 2019, 8, 3. [Google Scholar] [CrossRef] [PubMed]
- Mori, H.; Mizoguchi, H.; Fujio, T. Escherichia coli minimum genome factory. Biotechnol. Appl. Biochem. 2007, 46, 157–167. [Google Scholar] [CrossRef]
- Mizoguchi, H.; Sawano, Y.; Kato, J.-I.; Mori, H. Superpositioning of Deletions Promotes Growth of Escherichia coli with a Reduced Genome. DNA Res. 2008, 15, 277–284. [Google Scholar] [CrossRef]
- Ran, H.; Wu, J.; Wu, D.; Duan, X. Enhanced Production of Recombinant Thermobifida fusca Isoamylase in Escherichia coli MDS42. Appl. Biochem. Biotechnol. 2016, 180, 464–476. [Google Scholar] [CrossRef]
- Lee, J.H.; Sung, B.H.; Kim, M.S.; Blattner, F.R.; Yoon, B.H.; Kim, J.H.; Kim, S.C. Metabolic engineering of a reduced-genome strain of Escherichia coli for L-threonine production. Microb. Cell Factories 2009, 8, 2. [Google Scholar] [CrossRef] [PubMed]
- Reuß, D.R.; Altenbuchner, J.; Mäder, U.; Rath, H.; Ischebeck, T.; Sappa, P.K.; Thürmer, A.; Guérin, C.; Nicolas, P.; Steil, L.; et al. Large-scale reduction of the Bacillus subtilis genome: Consequences for the transcriptional network, resource allocation, and metabolism. Genome Res. 2017, 27, 289–299. [Google Scholar] [CrossRef] [PubMed]
- Suárez, R.A.; Stülke, J.; van Dijl, J.M. Less Is More: Toward a Genome-Reduced Bacillus Cell Factory for “Difficult Proteins”. ACS Synth. Biol. 2019, 8, 99–108. [Google Scholar] [CrossRef]
- Zhang, F.; Huo, K.; Song, X.; Quan, Y.; Wang, S.; Zhang, Z.; Gao, W.; Yang, C. Engineering of a genome-reduced strain Bacillus amyloliquefaciens for enhancing surfactin production. Microb. Cell Factories 2020, 19, 223. [Google Scholar] [CrossRef]
- Kurokawa, M.; Seno, S.; Matsuda, H.; Ying, B.-W. Correlation between genome reduction and bacterial growth. DNA Res. Int. J. Rapid Publ. Rep. Genes Genomes 2016, 23, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Foster, P.L. Methods for Determining Spontaneous Mutation Rates. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2006; Volume 409, pp. 195–213. [Google Scholar]
- Nishimura, I.; Kurokawa, M.; Liu, L.; Ying, B.-W. Coordinated Changes in Mutation and Growth Rates Induced by Genome Reduction. MBio 2017, 8, e00676-17. [Google Scholar] [CrossRef] [PubMed]
- Lao, Z.; Matsui, Y.; Ijichi, S.; Ying, B.-W. Global coordination of the mutation and growth rates across the genetic and nutritional variety in Escherichia coli. Front. Microbiol. 2022, 13, 990969. [Google Scholar] [CrossRef] [PubMed]
- Giovannoni, S.J.; Tripp, H.J.; Givan, S.; Podar, M.; Vergin, K.L.; Baptista, D.; Bibbs, L.; Eads, J.; Richardson, T.H.; Noordewier, M.; et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science 2005, 309, 1242–1245. [Google Scholar] [CrossRef]
- Viklund, J.; Ettema, T.J.; Andersson, S.G.E. Independent Genome Reduction and Phylogenetic Reclassification of the Oceanic SAR11 Clade. Mol. Biol. Evol. 2011, 29, 599–615. [Google Scholar] [CrossRef]
- Bergh, B.V.D.; Swings, T.; Fauvart, M.; Michiels, J. Experimental Design, Population Dynamics, and Diversity in Microbial Experimental Evolution. Microbiol. Mol. Biol. Rev. 2018, 82, e00008-18. [Google Scholar] [CrossRef]
- Kawecki, T.J.; Lenski, R.E.; Ebert, D.; Hollis, B.; Olivieri, I.; Whitlock, M.C. Experimental evolution. Trends Ecol. Evol. 2012, 27, 547–560. [Google Scholar] [CrossRef] [PubMed]
- Dragosits, M.; Mattanovich, D. Adaptive laboratory evolution—Principles and applications for biotechnology. Microb. Cell Factories 2013, 12, 64. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Aida, H.; Kurokawa, M.; Chen, F.; Xia, Y.; Xu, J.; Li, K.; Ying, B.-W.; Yomo, T. Primordial mimicry induces morphological change in Escherichia coli. Commun. Biol. 2022, 5, 24. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, S.; Horinouchi, T.; Furusawa, C. Prediction of antibiotic resistance by gene expression profiles. Nat. Commun. 2014, 5, 5792. [Google Scholar] [CrossRef] [PubMed]
- Kishimoto, T.; Ying, B.-W.; Tsuru, S.; Iijima, L.; Suzuki, S.; Hashimoto, T.; Oyake, A.; Kobayashi, H.; Someya, Y.; Narisawa, D.; et al. Molecular Clock of Neutral Mutations in a Fitness-Increasing Evolutionary Process. PLoS Genet. 2015, 11, e1005392. [Google Scholar] [CrossRef]
- Toprak, E.; Veres, A.; Michel, J.-B.; Chait, R.; Hartl, D.L.; Kishony, R. Evolutionary paths to antibiotic resistance under dynamically sustained drug selection. Nat. Genet. 2011, 44, 101–105. [Google Scholar] [CrossRef]
- Lenski, R.E. Experimental evolution and the dynamics of adaptation and genome evolution in microbial populations. ISME J. 2017, 11, 2181–2194. [Google Scholar] [CrossRef]
- Kurokawa, M.; Nishimura, I.; Ying, B.-W. Experimental Evolution Expands the Breadth of Adaptation to an Environmental Gradient Correlated with Genome Reduction. Front. Microbiol. 2022, 13, 826894. [Google Scholar] [CrossRef]
- Csörgő, B.; Fehér, T.; Tímár, E.; Blattner, F.R.; Pósfai, G. Low-mutation-rate, reduced-genome Escherichia coli: An improved host for faithful maintenance of engineered genetic constructs. Microb. Cell Factories 2012, 11, 11. [Google Scholar] [CrossRef]
- Umenhoffer, K.; Fehér, T.; Balikó, G.; Ayaydin, F.; Pósfai, J.; Blattner, F.R.; Pósfai, G. Reduced evolvability of Escherichia coli MDS42, an IS-less cellular chassis for molecular and synthetic biology applications. Microb. Cell Factories 2010, 9, 38. [Google Scholar] [CrossRef]
- Choe, D.; Lee, J.H.; Yoo, M.; Hwang, S.; Sung, B.H.; Cho, S.; Palsson, B.; Kim, S.C.; Cho, B.-K. Adaptive laboratory evolution of a genome-reduced Escherichia coli. Nat. Commun. 2019, 10, 935. [Google Scholar] [CrossRef] [PubMed]
- Matsui, Y.; Nagai, M.; Ying, B.-W. Growth rate-associated transcriptome reorganization in response to genomic, environmental, and evolutionary interruptions. Front. Microbiol. 2023, 14, 1145673. [Google Scholar] [CrossRef] [PubMed]
- Gibson, G. The environmental contribution to gene expression profiles. Nat. Rev. Genet. 2008, 9, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Wornell, K.; Pardesi, B.; Lee, K.; Boycheva, S.; Roberton, A.M.; White, W.L. High-throughput Method for Novel Medium Development for Culture of Anaerobic Gut Bacteria. Curr. Protoc. 2022, 2, e463. [Google Scholar] [CrossRef]
- Lagier, J.-C.; Dubourg, G.; Million, M.; Cadoret, F.; Bilen, M.; Fenollar, F.; Levasseur, A.; Rolain, J.-M.; Fournier, P.-E.; Raoult, D. Culturing the human microbiota and culturomics. Nat. Rev. Microbiol. 2018, 16, 540–550. [Google Scholar] [CrossRef]
- Oberhardt, M.A.; Zarecki, R.; Gronow, S.; Lang, E.; Klenk, H.-P.; Gophna, U.; Ruppin, E. Harnessing the landscape of microbial culture media to predict new organism–media pairings. Nat. Commun. 2015, 6, 8493. [Google Scholar] [CrossRef]
- Neidhardt, F.C.; Bloch, P.L.; Smith, D.F. Culture medium for enterobacteria. J. Bacteriol. 1974, 119, 736–747. [Google Scholar] [CrossRef]
- Rowe, J.J.; Goldberg, I.D.; Amelunxen, R.E. Development of defined and minimal media for the growth of Bacillus stearothermophilus. J. Bacteriol. 1975, 124, 279–284. [Google Scholar] [CrossRef]
- Mishek, H.P.; Stock, S.A.; Florick, J.D.; Blomberg, W.R.; Franke, J.D. Development of a chemically-defined minimal medium for studies on growth and protein uptake of Gemmata obscuriglobus. J. Microbiol. Methods 2018, 145, 40–46. [Google Scholar] [CrossRef]
- Maser, A.; Peebo, K.; Vilu, R.; Nahku, R. Amino acids are key substrates to Escherichia coli BW25113 for achieving high specific growth rate. Res. Microbiol. 2020, 171, 185–193. [Google Scholar] [CrossRef]
- Ehrenberg, M.; Bremer, H.; Dennis, P.P. Medium-dependent control of the bacterial growth rate. Biochimie 2013, 95, 643–658. [Google Scholar] [CrossRef] [PubMed]
- Vijayakumari, S.J.; Sasidharannair, N.K.; Nambisan, B.; Mohandas, C. Optimization of media and temperature for enhanced antimicrobial production by bacteria associated with Rhabditis sp. Iran. J. Microbiol. 2013, 5, 136–141. [Google Scholar] [PubMed]
- Neuenschwander, S.M.; Ghai, R.; Pernthaler, J.; Salcher, M.M. Microdiversification in genome-streamlined ubiquitous freshwater Actinobacteria. ISME J. 2018, 12, 185–198. [Google Scholar] [CrossRef] [PubMed]
- Tong, M.; French, S.; El Zahed, S.S.; Ong, W.K.; Karp, P.D.; Brown, E.D. Gene Dispensability in Escherichia coli Grown in Thirty Different Carbon Environments. MBio 2020, 11, e02865-20. [Google Scholar] [CrossRef] [PubMed]
- Zampieri, M.; Hörl, M.; Hotz, F.; Müller, N.F.; Sauer, U. Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli. Nat. Commun. 2019, 10, 3354. [Google Scholar] [CrossRef]
- Paliy, O.; Gunasekera, T.S. Growth of E. coli BL21 in minimal media with different gluconeogenic carbon sources and salt contents. Appl. Microbiol. Biotechnol. 2007, 73, 1169–1172. [Google Scholar] [CrossRef]
- Wang, J.; Yan, D.; Dixon, R.; Wang, Y.-P. Deciphering the Principles of Bacterial Nitrogen Dietary Preferences: A Strategy for Nutrient Containment. MBio 2016, 7, e00792-16. [Google Scholar] [CrossRef]
- Smith, D.K.; Benedict, C.D.; Weinberg, E.D. Bacterial culture longevity: Control by inorganic phosphate and temperature. Appl. Microbiol. 1974, 27, 292–293. [Google Scholar] [CrossRef]
- Brown, D.R.; Barton, G.; Pan, Z.; Buck, M.; Wigneshweraraj, S. Nitrogen stress response and stringent response are coupled in Escherichia coli. Nat. Commun. 2014, 5, 4115. [Google Scholar] [CrossRef]
- Peterson, C.N.; Mandel, M.J.; Silhavy, T.J. Escherichia coli starvation diets: Essential nutrients weigh in distinctly. J. Bacteriol. 2005, 187, 7549–7553. [Google Scholar] [CrossRef]
- Brauer, M.J.; Yuan, J.; Bennett, B.D.; Lu, W.; Kimball, E.; Botstein, D.; Rabinowitz, J.D. Conservation of the metabolomic response to starvation across two divergent microbes. Proc. Natl. Acad. Sci. USA 2006, 103, 19302–19307. [Google Scholar] [CrossRef] [PubMed]
- Egli, T. On multiple-nutrient-limited growth of microorganisms, with special reference to dual limitation by carbon and nitrogen substrates. Antonie Van Leeuwenhoek 1991, 60, 225–234. [Google Scholar] [CrossRef]
- Rajpurohit, H.; Eiteman, M.A. Nutrient-Limited Operational Strategies for the Microbial Production of Biochemicals. Microorganisms 2022, 10, 2226. [Google Scholar] [CrossRef] [PubMed]
- Rauch, C.; Cherkaoui, M.; Egan, S.; Leigh, J. The bio-physics of condensation of divalent cations into the bacterial wall has implications for growth of Gram-positive bacteria. Biochim. Biophys. Acta Biomembr. 2017, 1859, 282–288. [Google Scholar] [CrossRef]
- Dai, X.; Zhu, M.; Warren, M.; Balakrishnan, R.; Okano, H.; Williamson, J.R.; Fredrick, K.; Hwa, T. Slowdown of Translational Elongation in Escherichia coli under Hyperosmotic Stress. MBio 2018, 9, e02375-17. [Google Scholar] [CrossRef]
- Christensen, D.G.; Orr, J.S.; Rao, C.V.; Wolfe, A.J. Increasing Growth Yield and Decreasing Acetylation in Escherichia coli by Optimizing the Carbon-to-Magnesium Ratio in Peptide-Based Media. Appl. Environ. Microbiol. 2017, 83, e03034-16. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, R.; Islas, L.; Obregon, A.-M.; Escalante, L.; Sanchez, S. Gentamicin formation in Micromonospora purpurea: Stimulatory effect of ammonium. J. Antibiot. 1995, 48, 479–483. [Google Scholar] [CrossRef] [PubMed]
- Adinarayana, K.; Ellaiah, P. Response surface optimization of the critical medium components for the production of alkaline protease by a newly isolated Bacillus sp. J. Pharm. Pharm. Sci. 2002, 5, 272–278. [Google Scholar] [PubMed]
- Chen, J.; Lan, X.; Jia, R.; Hu, L.; Wang, Y. Response Surface Methodology (RSM) Mediated Optimization of Medium Components for Mycelial Growth and Metabolites Production of Streptomyces alfalfae XN-04. Microorganisms 2022, 10, 1854. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Zheng, H.; Cheng, Y.; Song, S.; Zheng, Z.; Jia, S. Medium optimization for ε-poly-L-lysine production by Streptomyces diastatochromogenes using response surface methodology. Lett. Appl. Microbiol. 2018, 66, 124–131. [Google Scholar] [CrossRef]
- Plackett, R.L.; Burman, J.P. The design of optimum multifactorial experiments. Biometrika 1946, 33, 305–325. [Google Scholar] [CrossRef]
- Vaidya, R.; Vyas, P.; Chhatpar, H. Statistical optimization of medium components for the production of chitinase by Alcaligenes xylosoxydans. Enzym. Microb. Technol. 2003, 33, 92–96. [Google Scholar] [CrossRef]
- Shokri, D.; Emtiazi, G. Indole-3-acetic acid (IAA) production in symbiotic and non-symbiotic nitrogen-fixing bacteria and its optimization by Taguchi design. Curr. Microbiol. 2010, 61, 217–225. [Google Scholar] [CrossRef] [PubMed]
- Chenthamarakshan, A.; Parambayil, N.; Miziriya, N.; Soumya, P.S.; Lakshmi, M.S.K.; Ramgopal, A.; Dileep, A.; Nambisan, P. Optimization of laccase production from Marasmiellus palmivorus LA1 by Taguchi method of Design of experiments. BMC Biotechnol. 2017, 17, 12. [Google Scholar] [CrossRef]
- Senthilkumar, S.R.; AshokKumar, B.; Raj, K.C.; Gunasekaran, P. Optimization of medium composition for alkali-stable xylanase production by Aspergillus fischeri Fxn 1 in solid-state fermentation using central composite rotary design. Bioresour. Technol. 2005, 96, 1380–1386. [Google Scholar] [CrossRef]
- Książek, E.E.; Janczar-Smuga, M.; Pietkiewicz, J.J.; Walaszczyk, E. Optimization of Medium Constituents for the Production of Citric Acid from Waste Glycerol Using the Central Composite Rotatable Design of Experiments. Molecules 2023, 28, 3268. [Google Scholar] [CrossRef]
- Bren, A.; Park, J.O.; Towbin, B.D.; Dekel, E.; Rabinowitz, J.D.; Alon, U. Glucose becomes one of the worst carbon sources for E.coli on poor nitrogen sources due to suboptimal levels of cAMP. Sci. Rep. 2016, 6, 24834. [Google Scholar] [CrossRef]
- Azodi, C.B.; Tang, J.; Shiu, S.-H. Opening the Black Box: Interpretable Machine Learning for Geneticists. Trends Genet. 2020, 36, 442–455. [Google Scholar] [CrossRef]
- Schrider, D.R.; Kern, A.D. Supervised Machine Learning for Population Genetics: A New Paradigm. Trends Genet. 2018, 34, 301–312. [Google Scholar] [CrossRef]
- Hie, B.; Bryson, B.D.; Berger, B. Leveraging Uncertainty in Machine Learning Accelerates Biological Discovery and Design. Cell Syst. 2020, 11, 461–477.e9. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Vilchis, P.; De-La-Cruz-García, J.-S.; Ramirez-Arellano, A. Identification of Relevant Protein Interactions with Partial Knowledge: A Complex Network and Deep Learning Approach. Biology 2023, 12, 140. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.B.; Kim, W.J.; Kim, H.U.; Lee, S.Y. Machine learning applications in systems metabolic engineering. Curr. Opin. Biotechnol. 2020, 64, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Patra, P.; Disha, B.R.; Kundu, P.; Das, M.; Ghosh, A. Recent advances in machine learning applications in metabolic engineering. Biotechnol. Adv. 2023, 62, 108069. [Google Scholar] [CrossRef] [PubMed]
- Zhou, K.; Ng, W.; Cortés-Peña, Y.; Wang, X. Increasing metabolic pathway flux by using machine learning models. Curr. Opin. Biotechnol. 2020, 66, 179–185. [Google Scholar] [CrossRef]
- Ashino, K.; Sugano, K.; Amagasa, T.; Ying, B.-W. Predicting the decision making chemicals used for bacterial growth. Sci. Rep. 2019, 9, 7251. [Google Scholar] [CrossRef]
- Aida, H.; Hashizume, T.; Ashino, K.; Ying, B.-W. Machine learning-assisted discovery of growth decision elements by relating bacterial population dynamics to environmental diversity. Elife 2022, 11, e76846. [Google Scholar] [CrossRef]
- Aida, H.; Uchida, K.; Nagai, M.; Hashizume, T.; Masuo, S.; Takaya, N.; Ying, B.-W. Machine learning-assisted medium optimization revealed the discriminated strategies for improved production of the foreign and native metabolites. Comput. Struct. Biotechnol. J. 2023, 21, 2654–2663. [Google Scholar] [CrossRef]
- Hashizume, T.; Ozawa, Y.; Ying, B.-W. Employing active learning in the optimization of culture medium for mammalian cells. NPJ Syst. Biol. Appl. 2023, 9, 20. [Google Scholar] [CrossRef]
- Yoshida, K.; Watanabe, K.; Chiou, T.-Y.; Konishi, M. High throughput optimization of medium composition for Escherichia coli protein expression using deep learning and Bayesian optimization. J. Biosci. Bioeng. 2023, 135, 127–133. [Google Scholar] [CrossRef]
- Borkowski, O.; Koch, M.; Zettor, A.; Pandi, A.; Batista, A.C.; Soudier, P.; Faulon, J.-L. Large scale active-learning-guided exploration for in vitro protein production optimization. Nat. Commun. 2020, 11, 1872. [Google Scholar] [CrossRef]
- Bapat, P.M.; Wangikar, P.P. Optimization of rifamycin B fermentation in shake flasks via a machine-learning-based approach. Biotechnol. Bioeng. 2004, 86, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Cohn, D.A.; Ghahramani, Z.; Jordan, M.I. Active learning with statistical models. J. Artif. Intell. Res. 1996, 4, 129–145. [Google Scholar] [CrossRef]
- Lin, F.-R.; Shaw, M.J. Active training of backpropagation neural networks using the learning by experimentation methodology. Ann. Oper. Res. 1997, 75, 105–122. [Google Scholar] [CrossRef]
- Reker, D.; Schneider, G. Active-learning strategies in computer-assisted drug discovery. Drug Discov. Today 2015, 20, 458–465. [Google Scholar] [CrossRef]
- Osmanbeyoglu, H.U.; Wehner, J.A.; Carbonell, J.G.; Ganapathiraju, M.K. Active machine learning for transmembrane helix prediction. BMC Bioinform. 2010, 11, S58. [Google Scholar] [CrossRef]
- Wang, X.; Rai, N.; Pereira, B.M.P.; Eetemadi, A.; Tagkopoulos, I. Accelerated knowledge discovery from omics data by optimal experimental design. Nat. Commun. 2020, 11, 5026. [Google Scholar] [CrossRef]
- King, R.D.; Whelan, K.E.; Jones, F.M.; Reiser, P.G.K.; Bryant, C.H.; Muggleton, S.H.; Kell, D.B.; Oliver, S.G. Functional genomic hypothesis generation and experimentation by a robot scientist. Nature 2004, 427, 247–252. [Google Scholar] [CrossRef]
- Pál, C.; Papp, B.; Lercher, M.J.; Csermely, P.; Oliver, S.G.; Hurst, L.D. Chance and necessity in the evolution of minimal metabolic networks. Nature 2006, 440, 667–670. [Google Scholar] [CrossRef]





| Parent Genome | Strain Name | Genome Size (Mb) | Reduced Ratio | Growth Medium | Growth Fitness |
|---|---|---|---|---|---|
| W3110 (4.66 Mb) | MGF-01 (N28) | 3.6 | 22% | minimal | decreased |
| minimal, amino acids | decreased | ||||
| rich | decreased | ||||
| DGF-298 | 3.0 | 36% | rich | increased | |
| MG1655 (4.64 Mb) | MDS42 | 4.0 | 14% | minimal | equivalent |
| rich | equivalent | ||||
| minimal, amino acids | decreased | ||||
| MDS69 | 3.7 | 20% | rich | decreased | |
| Δ16 | 3.3 | 30% | minimal | decreased | |
| MS56 | 3.6 | 23% | minimal | increased, decreased | |
| rich | equivalent, decreased |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aida, H.; Ying, B.-W. Efforts to Minimise the Bacterial Genome as a Free-Living Growing System. Biology 2023, 12, 1170. https://doi.org/10.3390/biology12091170
Aida H, Ying B-W. Efforts to Minimise the Bacterial Genome as a Free-Living Growing System. Biology. 2023; 12(9):1170. https://doi.org/10.3390/biology12091170
Chicago/Turabian StyleAida, Honoka, and Bei-Wen Ying. 2023. "Efforts to Minimise the Bacterial Genome as a Free-Living Growing System" Biology 12, no. 9: 1170. https://doi.org/10.3390/biology12091170
APA StyleAida, H., & Ying, B.-W. (2023). Efforts to Minimise the Bacterial Genome as a Free-Living Growing System. Biology, 12(9), 1170. https://doi.org/10.3390/biology12091170








