Comparative Transcriptome Analysis of Two Chrysomycin-Producing Wild-Type and Mutant Strains of Streptomyces sp. 891
Abstract
:1. Introduction
2. Materials and Methods
2.1. Microbes and Cultivation
2.2. RNA Extraction
2.3. Illumina Sequencing, Assembly, and Annotation
2.4. Bioinformatics Analysis
3. Results
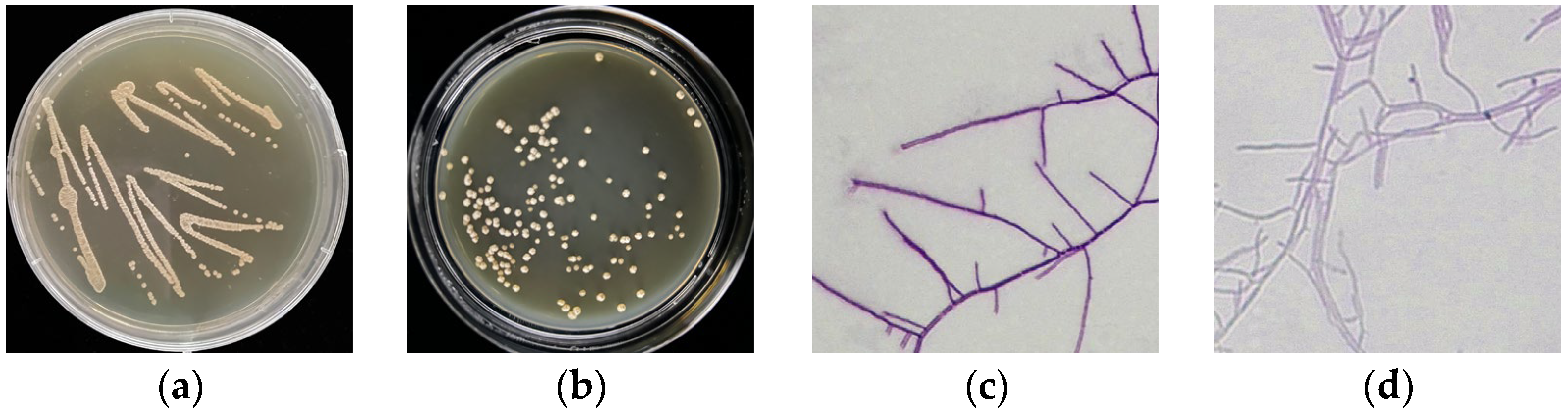
3.1. Morphological Features of Strains 891 and 891-B6
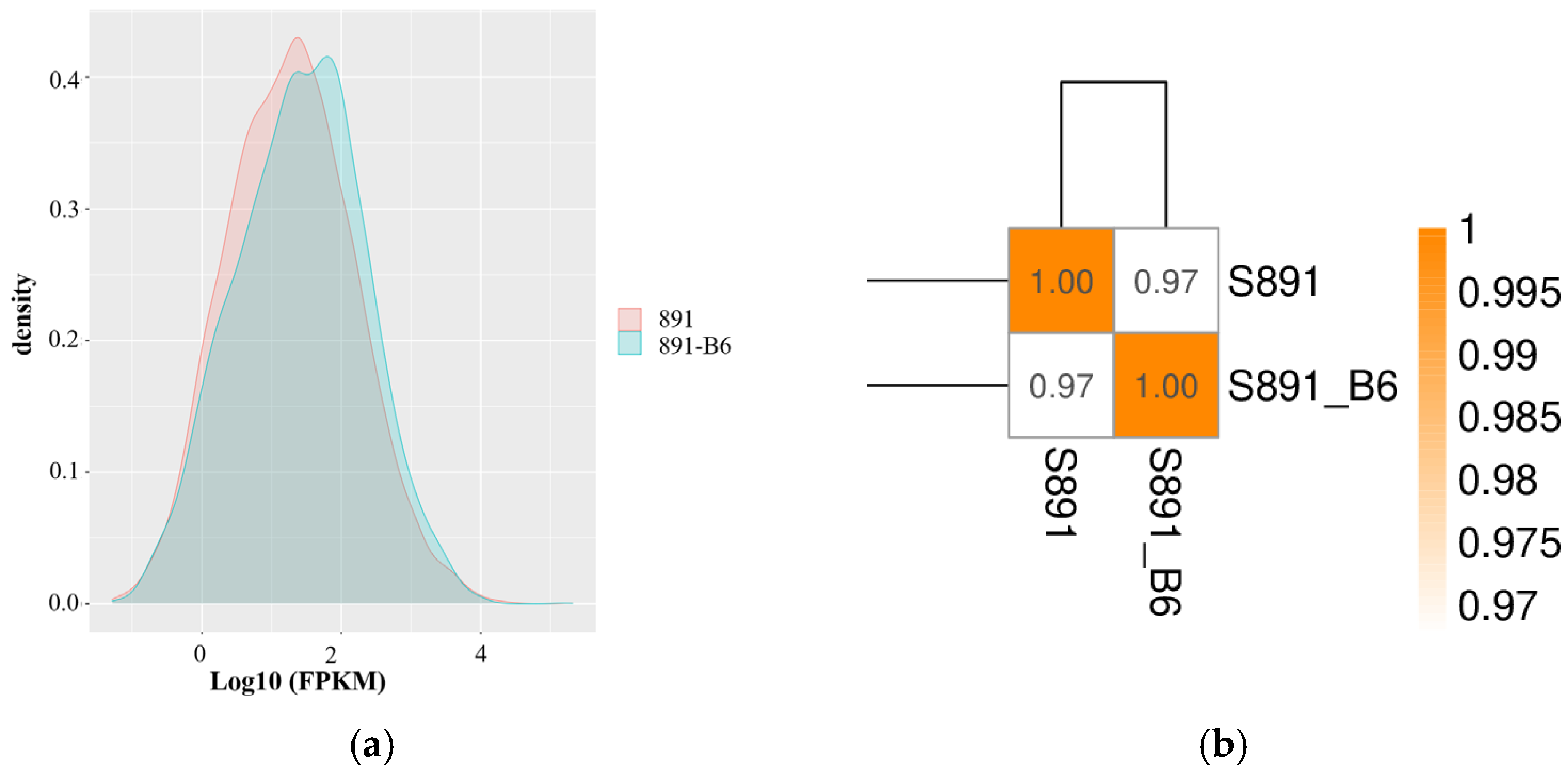
3.2. RNA-Seq Gene Expression Analysis
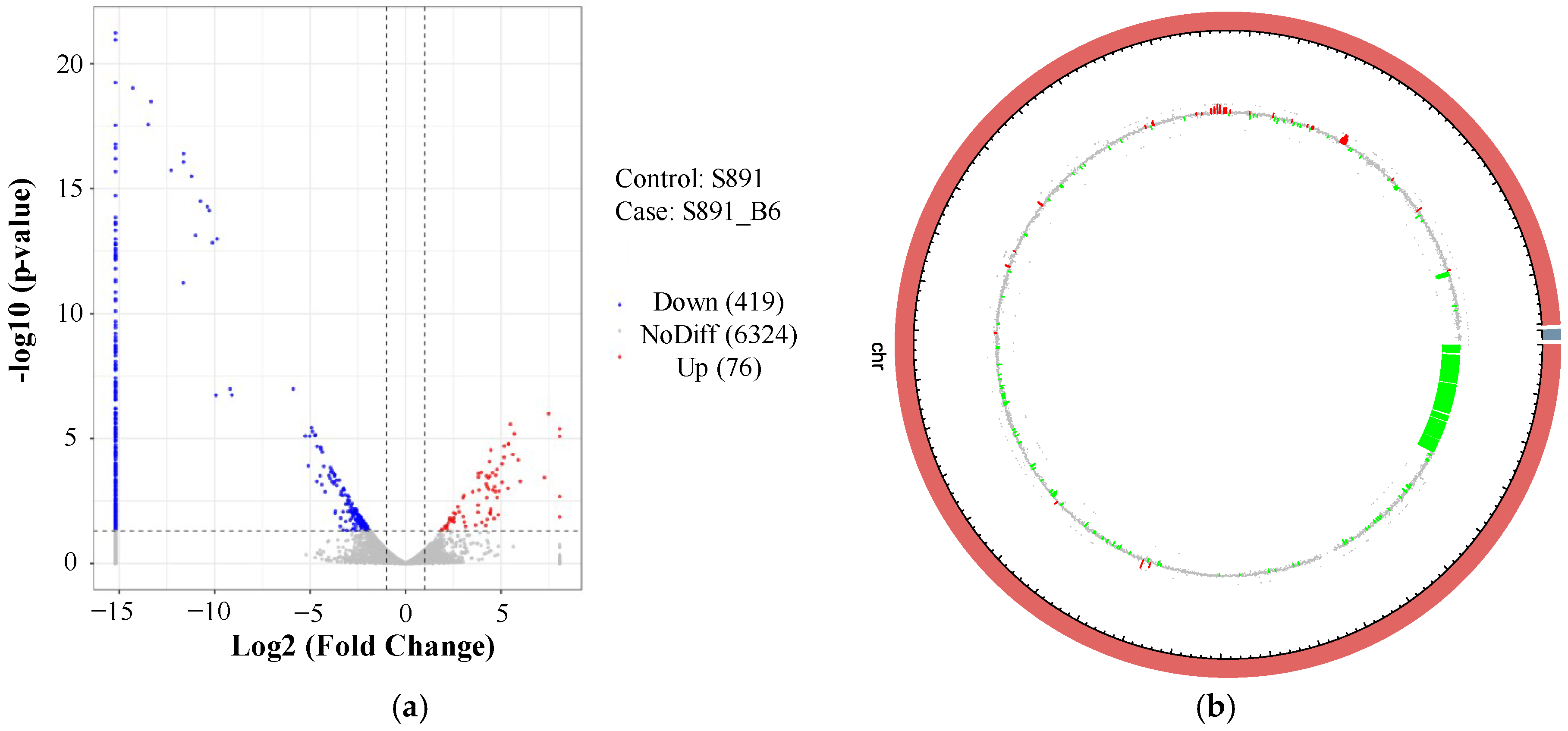
3.3. Analyses of DEGs and Identification of Regulated Genes
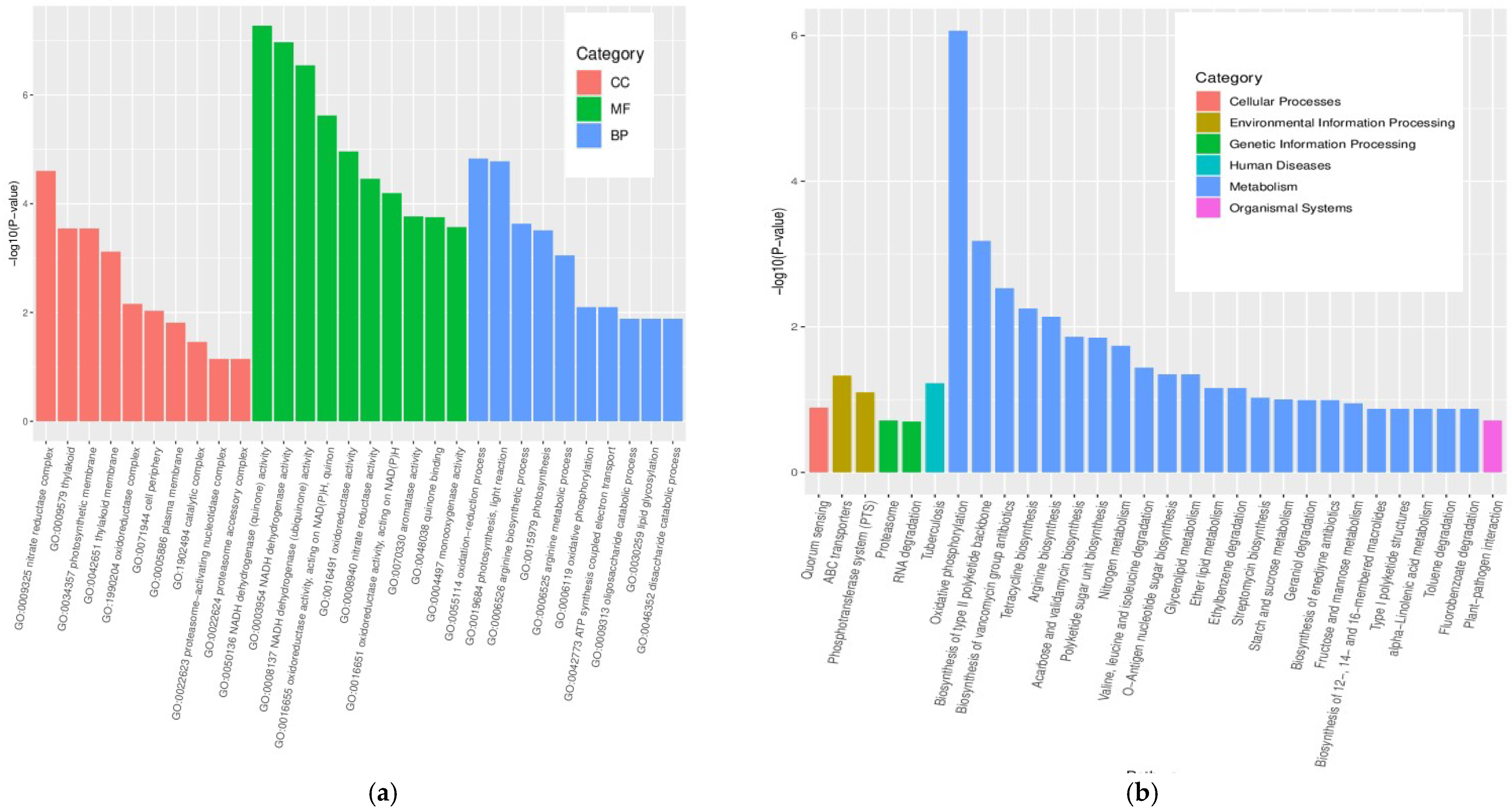
3.4. GO and KEGG Pathway Enrichment Analyses of DEGs
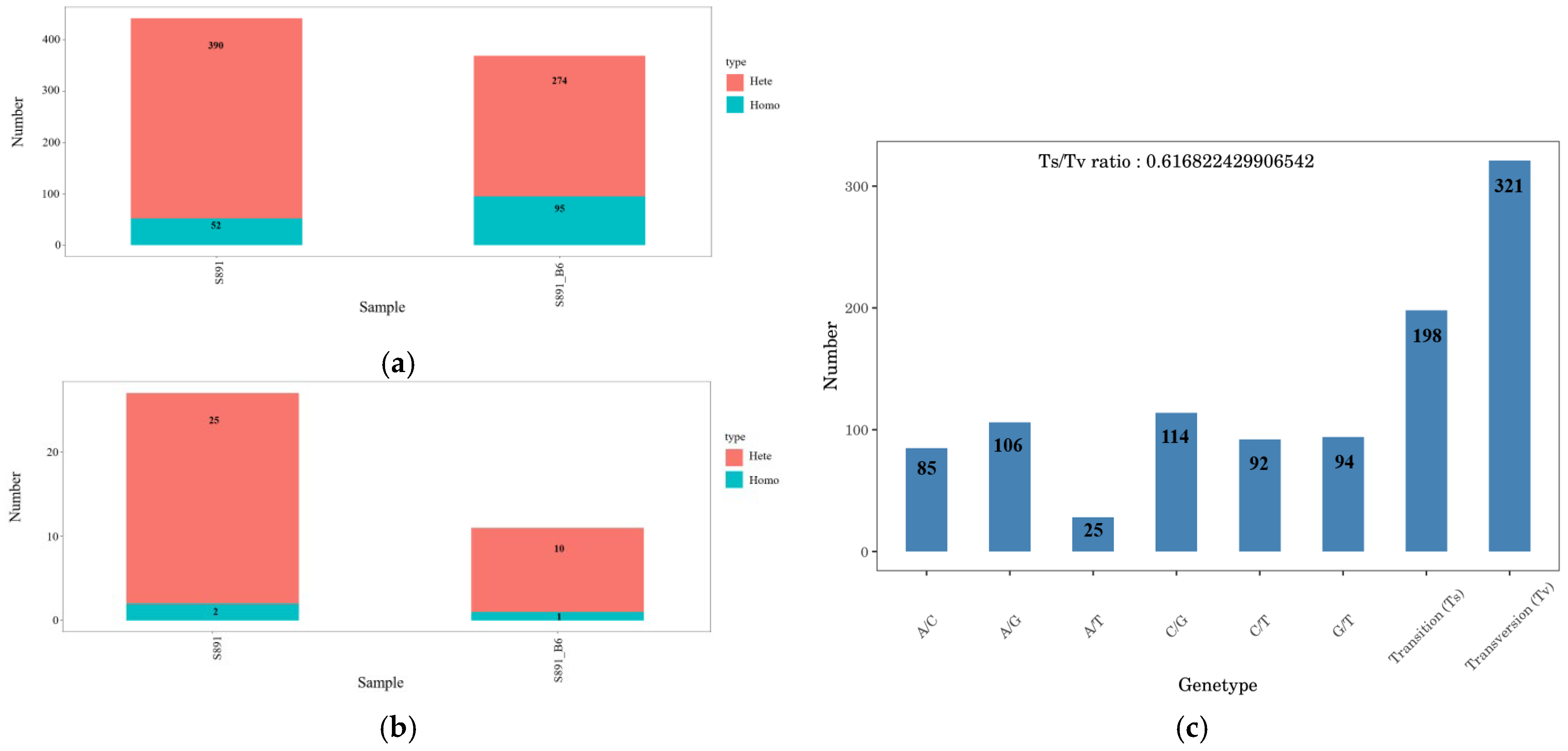
3.5. Analyses of SNPs and InDels
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Strelitz, F.; Flon, H.; Asheshov, I.N. Chrysomycin: A new antibiotic substance for bacterial viruses. J. Bacteriol. 1955, 69, 280–283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ni, H.J.; Lv, S.Y.; Sheng, Y.T.; Wang, H.; Chu, X.H.; Zhang, H.W. Optimization of fermentation conditions and medium compositions for the production of chrysomycin a by a marine-derived strain Streptomyces sp. 891. Prep. Biochem. Biotechnol. 2021, 51, 998–1003. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.K.; Pathania, A.S.; Parshad, R.; Raina, C.; Ali, A.; Gupta, A.P.; Kushwaha, M.; Aravinda, S.; Bhushan, S.; Bharate, S.B.; et al. Chrysomycins A-C, antileukemic naphthocoumarins from Streptomyces sporoverrucosus. RSC Adv. 2013, 3, 21046–21053. [Google Scholar] [CrossRef]
- Matson, J.A.; Rose, W.C.; Bush, J.A.; Myllymaki, R.; Bradner, W.T.; Doyle, T.W. Antitumor activity of chrysomycins M and V. J. Antibiot. 1989, 42, 1446–1448. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wei, T.T.; Byrne, K.M.; Warnick-Pickle, D.; Greenstein, M. Studies on the mechanism of action of gilvocarcin V and chrysomycin A. J. Antibiot. 1982, 35, 545–548. [Google Scholar] [CrossRef] [Green Version]
- Muralikrishnan, B.; Dan, V.M.; Vinodh, J.S.; Jamsheena, V.; Ramachandran, R.; Thomas, S.; Dastager, S.G.; Kumar, K.S.; Lankalapalli, R.S.; Kumar, R.A.; et al. Anti-microbial activity of chrysomycin A produced by Streptomyces sp. against Mycobacterium tuberculosis. RSC Adv. 2017, 7, 36335–36339. [Google Scholar] [CrossRef] [Green Version]
- Liu, H.; Cai, Y.; Chu, Y.; Yu, X.; Song, F.; Wang, H.; Zhang, H.; Sun, X. Formulation of chrysomycin A cream for the treatment of skin infections. Molecules 2022, 27, 4613. [Google Scholar] [CrossRef]
- Wu, F.; Zhang, J.; Song, F.; Wang, S.; Guo, H.; Wei, Q.; Dai, H.; Chen, X.; Xia, X.; Liu, X.; et al. Chrysomycin A derivatives for the treatment of multi-drug-resistant tuberculosis. ACS Cent. Sci. 2020, 6, 928–938. [Google Scholar] [CrossRef]
- Wada, S.-I.; Sawa, R.; Iwanami, F.; Nagayoshi, M.; Kubota, Y.; Iijima, K.; Hayashi, C.; Shibuya, Y.; Hatano, M.; Igarashi, M.; et al. Structures and biological activities of novel 4’-acetylated analogs of chrysomycins A and B. J. Antibiot. 2017, 70, 1078–1082. [Google Scholar] [CrossRef]
- Weiss, U.; Yoshihira, K.; Highet, R.J.; White, R.J.; Wei, T.T. The chemistry of the antibiotics chrysomycin A and B. Antitumor activity of chrysomycin A. J. Antibiot. 1982, 35, 1194–1201. [Google Scholar] [CrossRef]
- Liu, D.-N.; Liu, M.; Zhang, S.-S.; Shang, Y.-F.; Song, F.-H.; Zhang, H.-W.; Du, G.-H.; Wang, Y.-H. Chrysomycin A inhibits the proliferation, migration and invasion of U251 and U87-MG glioblastoma cells to exert its anti-cancer effects. Molecules 2022, 27, 6148. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Zhang, S.S.; Liu, D.N.; Yang, Y.L.; Wang, Y.H.; Du, G.H. Chrysomycin A attenuates Neuroinflammation by down-regulating NLRP3/Cleaved caspase-1 signaling pathway in LPS-stimulated mice and BV2 cells. Int. J. Mol. Sci. 2021, 22, 6799. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Tang, Y.; Liu, Y.; Pei, X.; Huang, Z.; Song, F.; Zhang, H. Comprehensive genomic analysis of marine strain Streptomyces sp. 891, an excellent producer of chrysomycin A with therapeutic potential. Mar. Drugs 2022, 20, 287. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Hu, Z.; Li, S.; Yang, H.; Li, S.; Lv, C.; Zaynab, M.; Cheng, C.H.K.; Chen, H.; Yang, X. Comparative Transcriptomic Analysis Uncovers Genes Responsible for the DHA Enhancement in the Mutant Aurantiochytrium sp. Microorganisms 2020, 8, 529. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, X.; Xie, Y.; Li, C.; Donovan, D.M.; Gehring, A.; Irwin, P.; He, Y. Comparative transcriptome analysis reveals differentially expressed genes related to antimicrobial properties of lysostaphin in Staphylococcus aureus. Antibiotics 2022, 11, 125. [Google Scholar] [CrossRef]
- Tekarslan-Sahin, S.H.; Alkim, C.; Sezgin, T. Physiological and transcriptomic analysis of a salt-resistant Saccharomyces cerevisiae mutant obtained by evolutionary engineering. Bosn J. Basic Med. Sci. 2018, 18, 55–65. [Google Scholar] [CrossRef] [Green Version]
- Cardoso, M.H.; Almeida, K.C.; Candido, E.S.; Fernandes, G.D.R.; Dias, S.C.; Alencar, S.A.; Franco, O.L. Comparative transcriptome analyses of magainin I-susceptible and -resistant Escherichia coli strains. Microbiol.-SGM 2018, 164, 1383–1393. [Google Scholar] [CrossRef]
- Zhang, H.; Lv, S.; Tang, Y.; Zhu, W.; Wang, H. Streptomycete Mutant and Application. CN Patent 113278545 A, 20 August 2021. [Google Scholar]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Smith, A.D.; Yan, X.; Chen, C.; Dawson, H.D.; Bhagwat, A.A. Understanding the host-adapted state of Citrobacter rodentium by transcriptomic analysis. Arch. Microbiol. 2016, 198, 353–362. [Google Scholar] [CrossRef]
- Li, P.E.; Lo, C.C.; Anderson, J.J.; Davenport, K.W.; Bishop-Lilly, K.A.; Xu, Y.; Ahmed, S.; Feng, S.; Mokashi, V.P.; Chain, P.S. Enabling the democratization of the genomics revolution with a fully integrated web-based bioinformatics platform. Nucleic Acids Res. 2017, 45, 67–80. [Google Scholar] [CrossRef]
- Blin, K.; Kim, H.U.; Medema, M.H.; Weber, T. Recent development of antiSMASH and other computational approaches to mine secondary metabolite biosynthetic gene clusters. Brief. Bioinform. 2019, 20, 1103–1113. [Google Scholar] [CrossRef] [PubMed]
- Kharel, M.K.; Nybo, S.E.; Shepherd, M.D.; Rohr, J. Cloning and characterization of the ravidomycin and chrysomycin biosynthetic gene clusters. ChemBioChem 2010, 11, 523–532. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huntley, R.P.; Harris, M.A.; Alam-Faruque, Y.; Blake, J.A.; Carbon, S.; Dietze, H.; Dimmer, E.C.; Foulger, R.E.; Hill, D.P.; Khodiyar, V.K.; et al. A method for increasing expressivity of gene ontology annotations using a compositional approach. BMC Bioinf. 2014, 15, 155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murray, C.J.L.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef] [PubMed]
- Vestergaard, M.; Frees, D.; Ingmer, H. Antibiotic Resistance and the MRSA Problem. Microbiol. Spectr. 2019, 7, GPP3-0057-2018. [Google Scholar] [CrossRef] [PubMed]

| Gene ID | Gene Name | LogFC | p-Value | Proposed Function |
|---|---|---|---|---|
| gene 5694 | chryE | 3.79 | 0.00 | putative TDP-glucose-4,6-dehydratase |
| gene 5695 | chryD | 4.27 | 0.00 | putative glucose-1-phosphate thymidylyltransferase |
| gene 5696 | chryXa | 3.00 | 0.00 | unknow |
| gene 5697 | chryGT | 4.72 | 0.00 | putative C-glycosyltransferase |
| gene 5698 | chryMT | 4.90 | 0.00 | putative O-methyltransferase |
| gene 5699 | chryOII | 5.61 | 0.00 | putative anthrone monooxygenase |
| gene 5700 | chryRM | 5.40 | 0.00 | putative C-methyltransferase |
| gene 5701 | chryRM | 4.86 | 0.01 | putative C-methyltransferase |
| gene 5702 | chryU | - | 0.00 | putative NDP-hexose 4-ketoreductase |
| gene 5703 | chryCMT | 5.40 | 0.00 | putative C-methyltransferase |
| gene 5704 | chryF | 4.41 | 0.00 | putative polyketide ketoreductase |
| gene 5705 | chryV | 4.71 | 0.00 | hypothetical protein |
| gene 5706 | chryH | 4.44 | 0.01 | putative oxidoreductase/NADH-dependent FMN reductase |
| gene 5707 | chryOI | 5.69 | 0.00 | putative FAD-dependent monooxygenase |
| gene 5708 | chryG | 6.01 | 0.00 | putative cyclase |
| gene 5710 | chryK | 5.16 | 0.00 | putative bifunctional cyclase/dehydratase |
| gene 5711 | chryOIV | 5.06 | 0.00 | putative FAD-dependent monooxygenase |
| gene 5712 | chryA | 4.33 | 0.00 | putative ketoacyl synthase (KSα) |
| gene 5713 | chryB | 8.07 | 0.00 | putative chain length factor (KSβ) |
| gene 5714 | chryC | - | 0.01 | putative acyl carrier protein |
| gene 5715 | chryOIII | 5.06 | 0.00 | putative P450 monooxygenase |
| gene 5717 | chryP | 4.23 | 0.00 | putative malonyl-CoA carrier protein transacylase |
| gene 5718 | chryQ | 5.37 | 0.00 | putative propionyl-CoA carrier protein transacylase |
| gene 5723 | chryX1 | 4.63 | 0.02 | putative spore-associated protein precursor |
| gene 5724 | - | 5.49 | 0.00 | unknown |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, W.; Pei, X.; Chen, X.; Wu, Y.; Song, F.; Zhang, H. Comparative Transcriptome Analysis of Two Chrysomycin-Producing Wild-Type and Mutant Strains of Streptomyces sp. 891. Metabolites 2022, 12, 1170. https://doi.org/10.3390/metabo12121170
Zhu W, Pei X, Chen X, Wu Y, Song F, Zhang H. Comparative Transcriptome Analysis of Two Chrysomycin-Producing Wild-Type and Mutant Strains of Streptomyces sp. 891. Metabolites. 2022; 12(12):1170. https://doi.org/10.3390/metabo12121170
Chicago/Turabian StyleZhu, Wangjie, Xinwei Pei, Xiaoyu Chen, You Wu, Fuhang Song, and Huawei Zhang. 2022. "Comparative Transcriptome Analysis of Two Chrysomycin-Producing Wild-Type and Mutant Strains of Streptomyces sp. 891" Metabolites 12, no. 12: 1170. https://doi.org/10.3390/metabo12121170






