Integrated Analysis of lncRNAs and mRNAs Reveals Complex Gene Network Mediated by lncRNAs and Regulatory Function of MuLRR-RLK-AS in Response to Phytoplasma Infection in Mulberry
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. Construction of cDNA Libraries and High-Throughput Sequencing
2.3. Identification of lncRNAs and Target Prediction
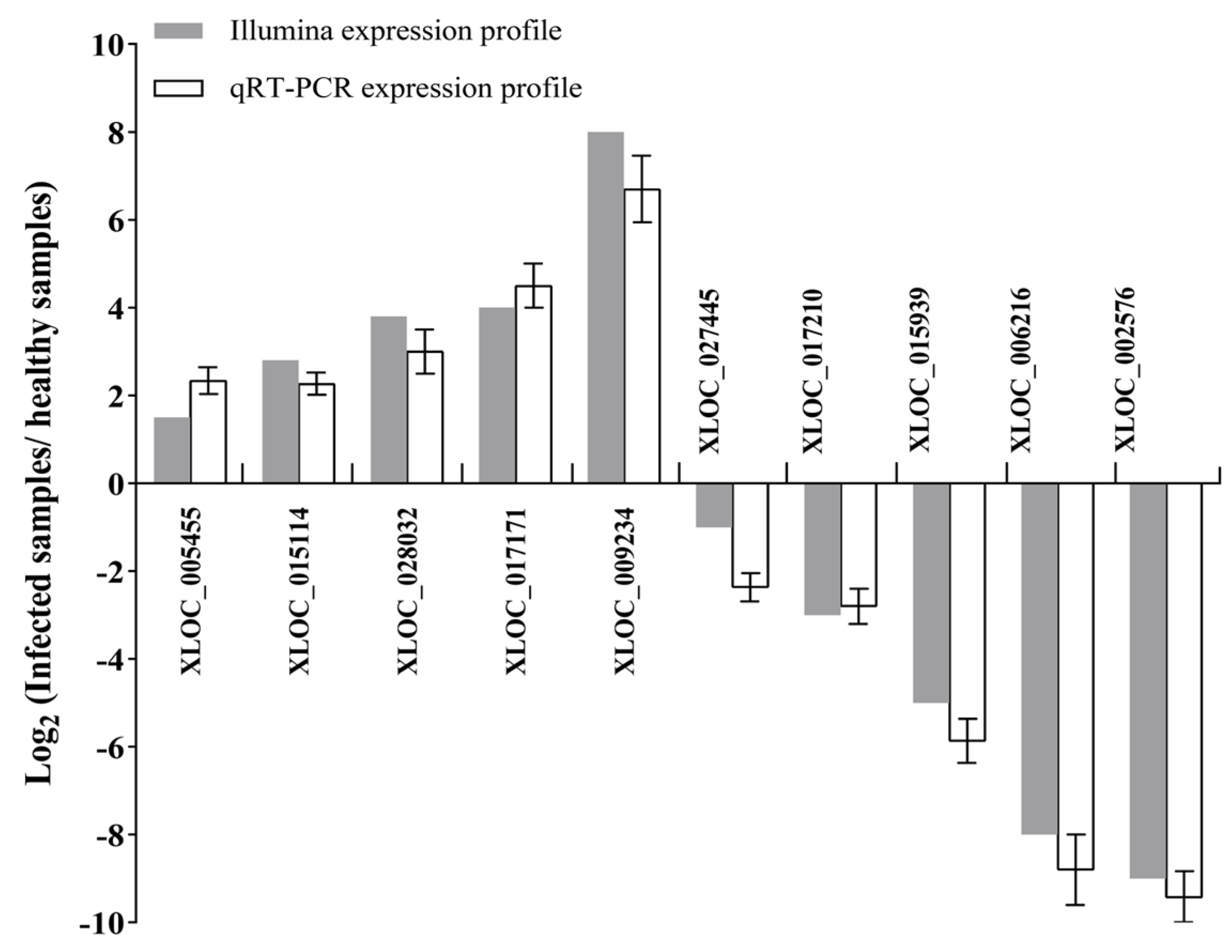
2.4. Gene Expression Analysis and qRT-PCR Validation
2.5. Gene Ontology Analysis
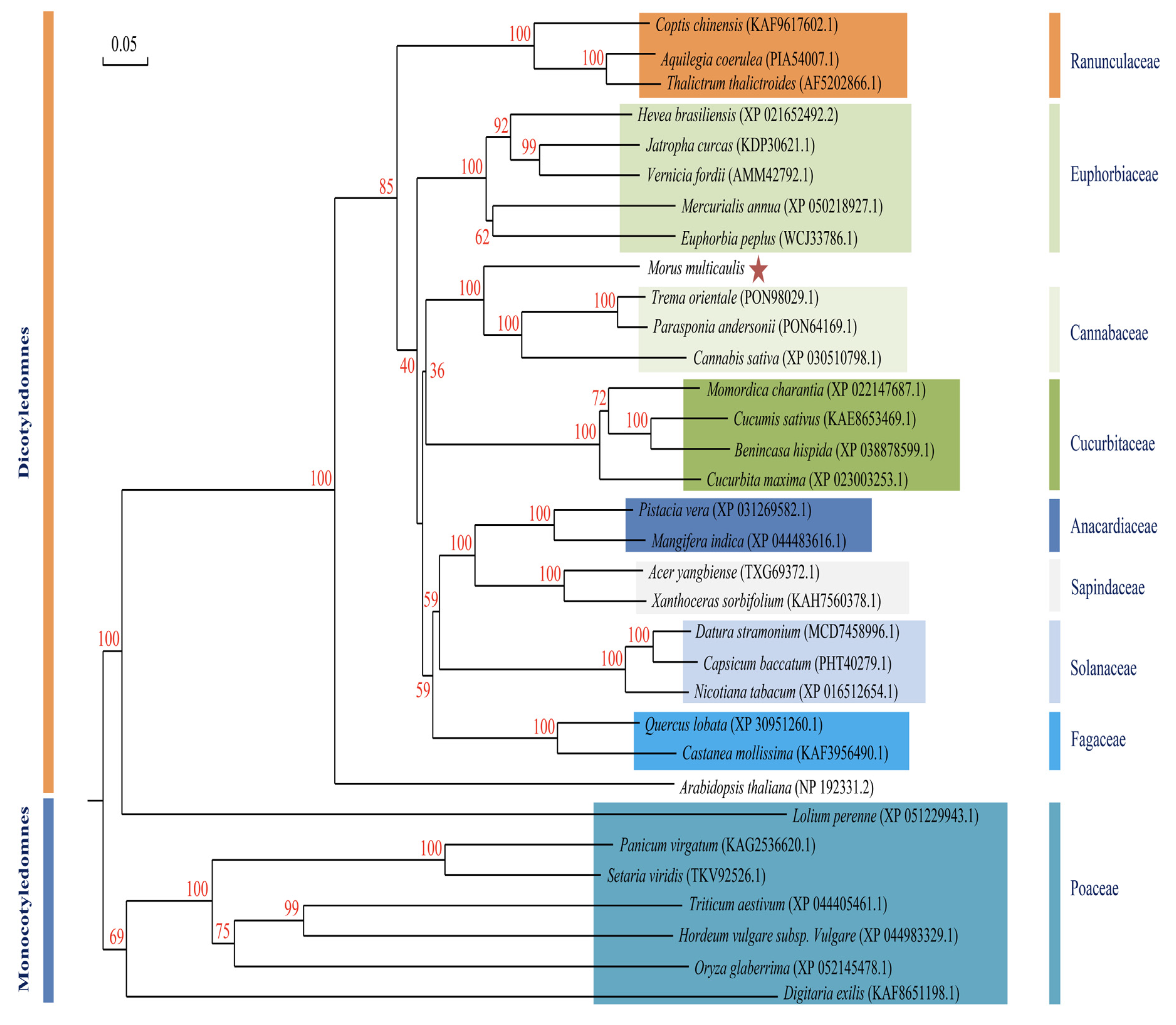
2.6. Gene Cloning and Phylogenetic Analysis
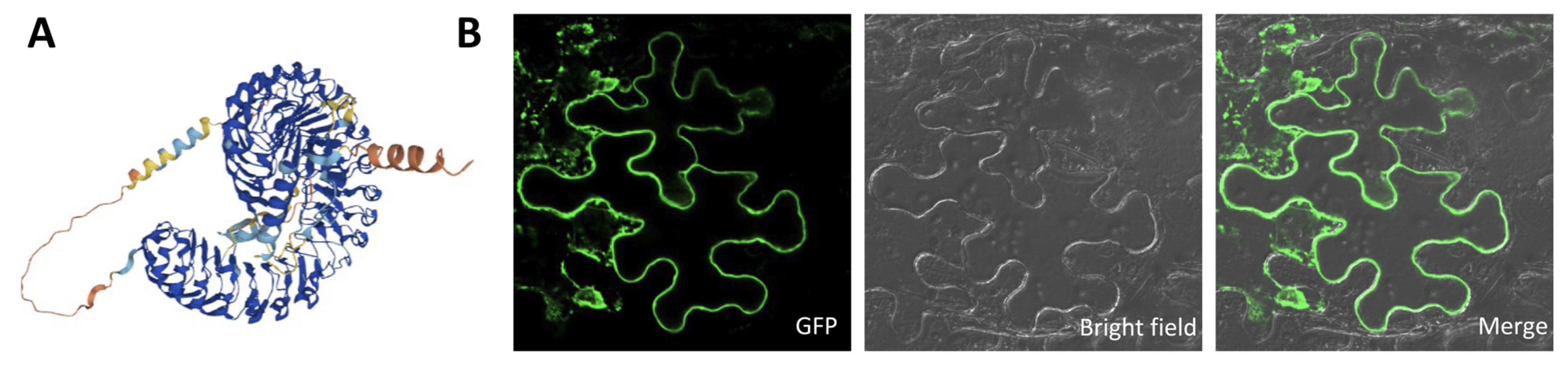
2.7. Subcellular Localization
2.8. Promoter Activity Analysis
2.9. Production of Transgenic Arabidopsis
2.10. Plant Treatment
2.11. Detection of Colony-Forming Units
2.12. Statistic Analysis
3. Results
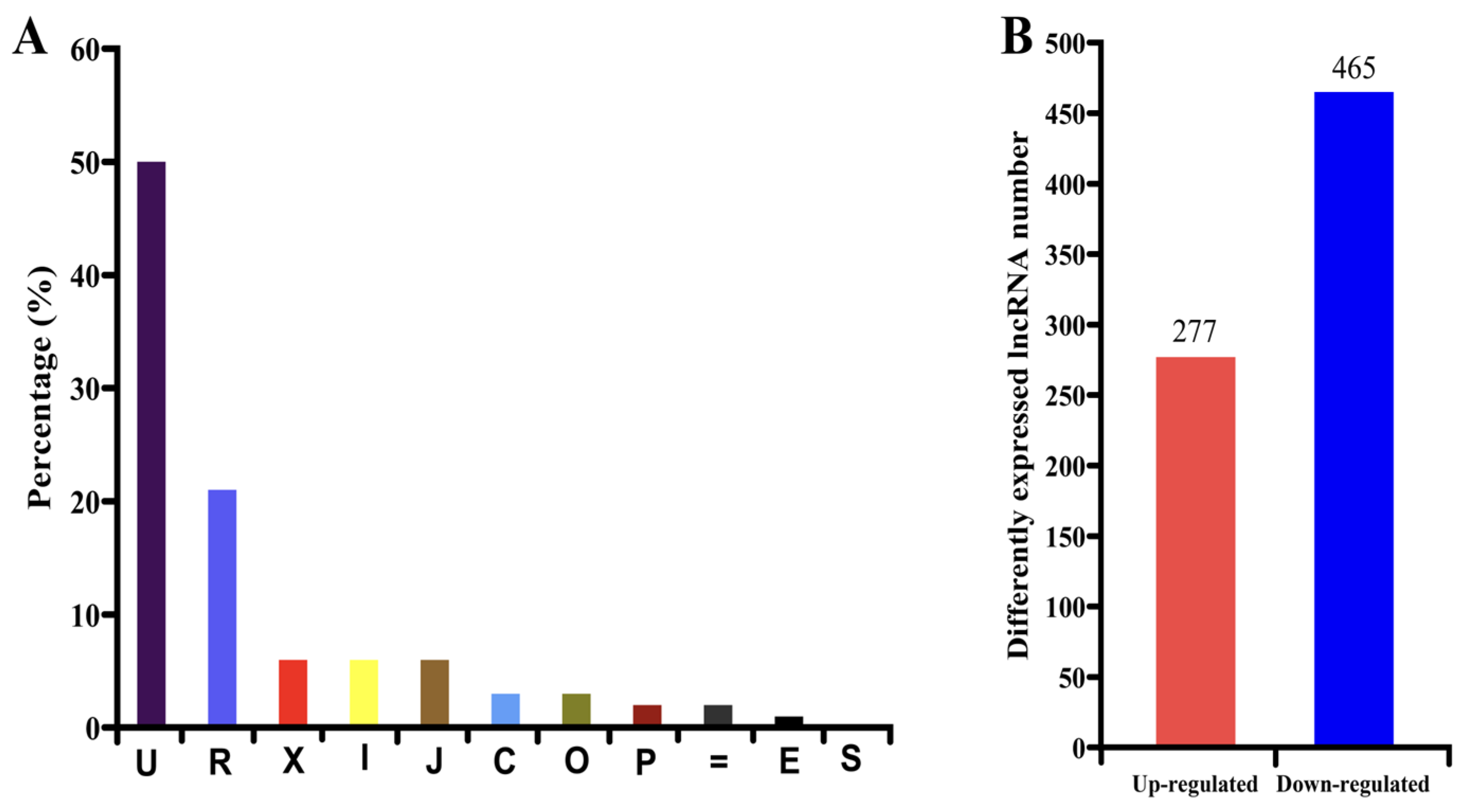
3.1. High-Throughput Sequencing and DEG Analysis
3.2. Identification and Characterization of lncRNAs
3.3. Target Prediction of the Differentially Expressed lncRNAs (DELs)
3.4. Characterization of MuLRR-RLK-AS Trans-Target Gene MuLRR-RLK
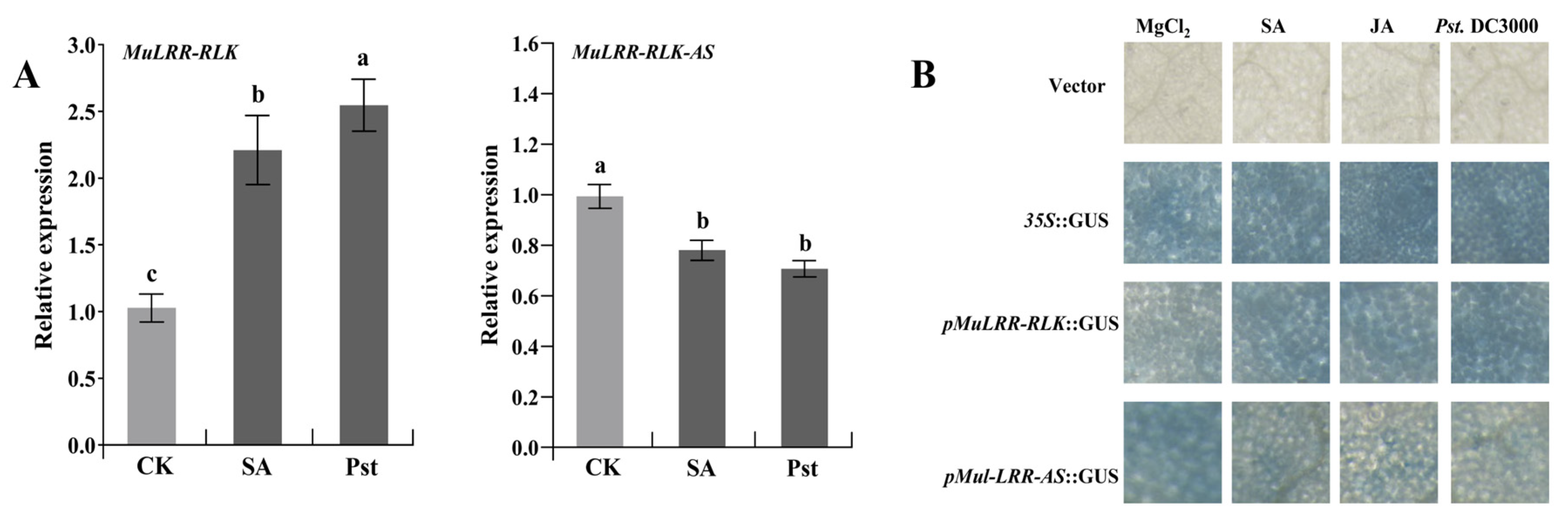
3.5. Expression Patterns of MuLRR-RLK and MuLRR-RLK-AS
3.6. MuLRR-RLK-AS Represses the Expression of MuLRR-RLK
3.7. MuLRR-RLK-AS Is a Negative Regulator of Plant Disease Resistance
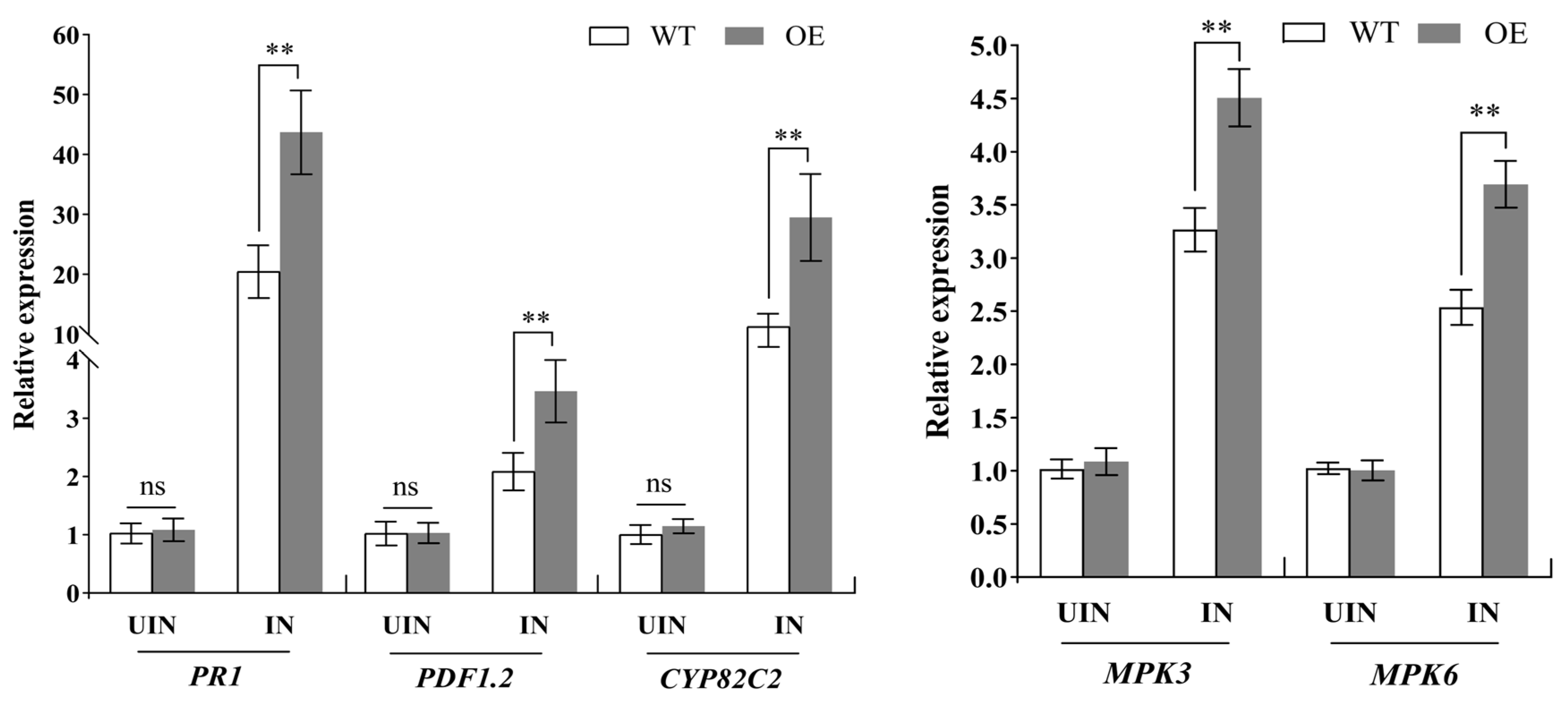
3.8. Ectopic Expression of MuLRR-RLK Affects the Expression of Defense-Related Genes
4. Discussion
4.1. LncRNAs Are Involved in Regulating Gene Expression in Both Cis- and Trans-Manners in Response to Phytoplasma Infection
4.2. MuLRR-RLK-AS Plays an Important Role in Regulating MuLRR-RLK Gene Expression in Response to Phytoplasma Infection
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Dhanyalakshmi, K.H.; Nataraja, K.N. Mulberry (Morus spp.) has the features to treat as a potential perennial model system. Plant Signal Behav. 2018, 13, e1491267. [Google Scholar] [CrossRef]
- Yuan, Q.; Zhao, L. The mulberry (Morus alba L.) Fruit-a review of characteristic components and health benefits. J. Agric. Food Chem. 2017, 65, 10383–10394. [Google Scholar] [CrossRef] [PubMed]
- He, N.; Zhang, C.; Qi, X.; Zhao, S.; Tao, Y.; Yang, G.; Lee, T.H.; Wang, X.; Cai, Q.; Li, D.; et al. Draft genome sequence of the mulberry tree. Morus Notabilis. Nat. Commun. 2013, 4, 2445. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Dong, X.; Xu, Y.; Dong, Q.; Wang, Y.; Gai, Y.; Ji, X. Transcriptome and DNA methylome reveal insights into phytoplasma infection responses in mulberry (Morus multicaulis Perr.). Front. Plant Sci. 2021, 12, 697702. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Wei, W.; Saiki, T.; Kawakita, H.; Watanabe, K.; Sato, M. Distribution patterns of mulberry dwarf phytoplasma in reproductive organs, winter buds, and roots of mulberry trees. J. Gen. Plant Pathol. 2004, 70, 168–173. [Google Scholar] [CrossRef]
- Luo, L.; Zhang, X.; Meng, F.; Wang, Y.; Zhou, Y.; Liu, J. Draft genome sequences resources of mulberry dwarf phytoplasma strain MDGZ-01 associated with mulberry yellow dwarf (MYD) diseases. Plant Dis. 2022, 106, 2239–2242. [Google Scholar] [CrossRef] [PubMed]
- Gai, Y.P.; Yuan, S.S.; Liu, Z.Y.; Zhao, H.N.; Liu, Q.; Qin, R.L.; Fang, L.J.; Ji, X.L. Integrated phloem sap mRNA and protein expression analysis reveals phytoplasma-infection responses in mulberry. Mol. Cell Proteom. 2018, 17, 1702–1719. [Google Scholar] [CrossRef]
- Gai, Y.P.; Zhao, H.N.; Zhao, Y.N.; Zhu, B.S.; Yuan, S.S.; Li, S.; Guo, F.Y.; Ji, X.L. MiRNA-seq-based profiles of miRNAs in mulberry phloem sap provide insight into the pathogenic mechanisms of mulberry yellow dwarf disease. Sci. Rep. 2018, 8, 812. [Google Scholar] [CrossRef]
- Gai, Y.P.; Han, X.J.; Li, Y.Q.; Yuan, C.Z.; Mo, Y.Y.; Guo, F.Y.; Liu, Q.X.; Ji, X.L. Metabolomic analysis reveals the potential metabolites and pathogenesis involved in mulberry yellow dwarf disease. Plant Cell Environ. 2014, 37, 1474–1490. [Google Scholar] [CrossRef] [PubMed]
- Gai, Y.P.; Li, Y.Q.; Guo, F.Y.; Yuan, C.Z.; Mo, Y.Y.; Zhang, H.L.; Wang, H.; Ji, X.L. Analysis of phytoplasma-responsive sRNAs provide insight into the pathogenic mechanisms of mulberry yellow dwarf disease. Sci. Rep. 2014, 4, 5378. [Google Scholar] [CrossRef]
- Song, L.; Fang, Y.; Chen, L.; Wang, J.; Chen, X. Role of non-coding RNAs in plant immunity. Plant Commun. 2021, 2, 100180. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.C.; Zhang, J.; Zhang, D.Y.; Nan, Y.H.; Ge, S.; Guo, Y.L. Identification of long noncoding natural antisense transcripts (lncNATs) correlated with drought stress response in wild rice (Oryza nivara). BMC Genom. 2021, 22, 424. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Liu, J.; Cheng, J.; Sun, Q.; Zhang, Y.; Liu, J.; Li, H.; Zhang, Z.; Wang, P.; Cai, C.; et al. lncRNA7 and lncRNA2 modulate cell wall defense genes to regulate cotton resistance to Verticillium wilt. Plant Physiol. 2022, 189, 264–284. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Jiang, N.; Meng, J.; Yang, G.; Liu, W.; Zhou, X.; Ma, N.; Hou, X.; Luan, Y. LncRNA33732-respiratory burst oxidase module associated with WRKY1 in tomato- Phytophthora infestans interactions. Plant J. 2019, 97, 933–946. [Google Scholar] [CrossRef]
- Jiang, N.; Cui, J.; Hou, X.; Yang, G.; Xiao, Y.; Han, L.; Meng, J.; Luan, Y. Sl-lncRNA15492 interacts with Sl-miR482a and affects Solanum lycopersicum immunity against Phytophthora infestans. Plant J. 2020, 103, 1561–1574. [Google Scholar] [CrossRef]
- Hong, Y.; Zhang, Y.; Cui, J.; Meng, J.; Chen, Y.; Zhang, C.; Yang, J.; Luan, Y. The lncRNA39896-miR166b-HDZs module affects tomato resistance to Phytophthora infestans. J. Integr. Plant Biol. 2022, 64, 1979–1993. [Google Scholar] [CrossRef]
- Liu, G.; Liu, F.; Wang, Y.; Liu, X. A novel long noncoding RNA CIL1 enhances cold stress tolerance in Arabidopsis. Plant Sci. 2022, 323, 111370. [Google Scholar] [CrossRef]
- Liu, N.; Xu, Y.; Li, Q.; Cao, Y.; Yang, D.; Liu, S.; Wang, X.; Mi, Y.; Liu, Y.; Ding, C.; et al. A lncRNA fine-tunes salicylic acid biosynthesis to balance plant immunity and growth. Cell Host Microbe. 2022, 30, 1124–1138.e8. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.F.; Zhang, Y.C.; Sun, Y.M.; Yu, Y.; Lei, M.Q.; Yang, Y.W.; Lian, J.P.; Feng, Y.Z.; Zhang, Z.; Yang, L.; et al. The parent-of-origin lncRNA MISSEN regulates rice endosperm development. Nat. Commun. 2021, 12, 6525. [Google Scholar] [CrossRef] [PubMed]
- Ransohoff, J.D.; Wei, Y.; Khavari, P.A. The functions and unique features of long intergenic non-coding RNA. Nat. Rev. Mol. Cell Biol. 2018, 19, 143–157. [Google Scholar] [CrossRef]
- Wang, Y.; Deng, X.W.; Zhu, D. From molecular basics to agronomic benefits: Insights into noncoding RNA-mediated gene regulation in plants. J. Integr. Plant Biol. 2022, 64, 2290–2308. [Google Scholar] [CrossRef] [PubMed]
- Quinn, J.; Chang, H. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 2016, 17, 47–62. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Liang, S.; Liu, S.; Meng, Z.; Wang, J.; Liang, S. Sequence pre-training-based graph neural network for predicting lncRNA-miRNA associations. Brief. Bioinform. 2023, 24, bbad317. [Google Scholar] [CrossRef] [PubMed]
- Ai, G.; Li, T.; Zhu, H.; Dong, X.; Fu, X.; Xia, C.; Pan, W.; Jing, M.; Shen, D.; Xia, A.; et al. BPL3 binds the long non-coding RNA nalncFL7 to suppress FORKED-LIKE7 and modulate HAI1-mediated MPK3/6 dephosphorylation in plant immunity. Plant Cell 2023, 35, 598–616. [Google Scholar] [CrossRef]
- Li, C.; Lai, X.; Yu, X.; Xiong, Z.; Chen, J.; Lang, X.; Feng, H.; Wan, X.; Liu, K. Plant long noncoding RNAs: Recent progress in understanding their roles in growth, development, and stress responses. Biochem. Biophys. Res. Commun. 2023, 671, 270–277. [Google Scholar] [CrossRef]
- Wang, J.Y.; Yu, W.G.; Yang, Y.W.; Li, X.; Chen, T.Z.; Liu, T.L.; Ma, N.; Yang, X.; Liu, R.Y.; Zhang, B.L. Genome-wide analysis of tomato long non-coding RNAs and identification as endogenous target mimic for microRNA in response to TYLCV infection. Sci. Rep. 2015, 5, 16946. [Google Scholar] [CrossRef]
- Zhou, Y.; Cho, W.K.; Byun, H.S.; Chavan, V.; Kil, E.J.; Lee, S.; Hong, S.W. Genome wide identification of long non-coding RNAs in tomato plants irradiated by neutrons followed by infection with tomato yellow leaf curl virus. PeerJ 2019, 7, e6286. [Google Scholar] [CrossRef]
- Zhang, T.; Liang, Q.; Li, C.; Fu, S.; Kundu, J.K.; Zhou, X.; Wu, J. Transcriptome analysis of rice reveals the lncRNA-mRNA regulatory network in response to rice black-streaked dwarf virus infection. Viruses 2020, 12, 951. [Google Scholar] [CrossRef]
- Gao, R.; Liu, P.; Irwanto, N.; Loh, R.; Wong, S.M. Upregulation of LINC-AP2 is negatively correlated with AP2 gene expression with Turnip crinkle virus infection in Arabidopsis thaliana. Plant Cell Rep. 2016, 35, 2257–2267. [Google Scholar] [CrossRef]
- Kang, S.H.; Sun, Y.D.; Atallah, O.O.; Huguet-Tapia, J.C.; Noble, J.D.; Folimonova, S.Y. A long non-coding RNA of Citrus tristeza virus: Role in the virus interplay with the host immunity. Viruses 2019, 11, 436. [Google Scholar] [CrossRef]
- Zhu, Q.H.; Stephen, S.; Taylor, J.; Helliwell, C.A.; Wang, M.B. Long noncoding RNAs responsive to Fusarium oxysporum infection in Arabidopsis thaliana. New Phytol. 2014, 201, 574–584. [Google Scholar] [CrossRef]
- Wang, N.; Cao, P.; Xia, W.; Fang, L.; Yu, H. Identification and characterization of long non-coding RNAs in response to early infection by Melampsoralarici-populina using genome-wide high-throughput RNA sequencing. Tree Genet. Genomes. 2017, 13, 34. [Google Scholar] [CrossRef]
- Yao, Z.; Chen, Q.; Chen, D.; Zhan, L.; Zeng, K.; Gu, A.; Zhou, J.; Zhang, Y.; Zhu, Y.; Gao, W.; et al. The susceptibility of sea-island cotton recombinant inbred lines to Fusarium oxysporum f. sp. vasinfectum infection is characterized by altered expression of long noncoding RNAs. Sci. Rep. 2019, 9, 2894. [Google Scholar] [CrossRef]
- Jiang, N.; Cui, J.; Shi, Y.; Yang, G.; Zhou, X.; Hou, X.; Meng, J.; Luan, Y. Tomato lncRNA23468 functions as a competing endogenous RNA to modulate NBS-LRR genes by decoying miR482b in the tomato-Phytophthora infestans interaction. Hortic. Res. 2019, 6, 28. [Google Scholar] [CrossRef]
- Zhao, X.; Sun, X.; Chen, Y.; Wu, H.; Liu, Y.; Jiang, Y.; Xie, F.; Chen, Y. Mining of long non-coding RNAs with target genes in response to rust based on full-length transcriptome in Kentucky bluegrass. Front. Plant Sci. 2023, 14, 1158035. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Liu, Y.; Li, L.; Li, D.; Zhang, Q.; Guo, Y.; Wang, S.; Zhong, C.; Huang, H. Whole transcriptome sequencing of Pseudomonas syringae pv. actinidiae-infected kiwifruit plants reveals species specific interaction between long non-coding RNA and coding genes. Sci. Rep. 2017, 7, 4910. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Zhou, Y.F.; Feng, Y.Z.; He, H.; Lian, J.P.; Yang, Y.W.; Lei, M.Q.; Zhang, Y.C.; Chen, Y.Q. Transcriptional landscape of pathogen-responsive lncRNAs in rice unveils the role of ALEX1 in jasmonate pathway and disease resistance. Plant Biotechnol. J. 2020, 18, 679–690. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.S.; Sun, H.X.; Park, B.S.; Huang, C.H.; Yeh, S.D.; Jung, C.; Chua, N.H. ELF18-INDUCED LONG-NONCODING RNA associates with mediator to enhance expression of innate immune response genes in Arabidopsis. Plant Cell 2017, 29, 1024–1038. [Google Scholar] [CrossRef] [PubMed]
- Rosli, H.G.; Sirvent, E.; Bekier, F.N.; Ramos, R.N.; Pombo, M.A. Genome-wide analysis uncovers tomato leaf lncRNAs transcriptionally active upon Pseudomonas syringae pv. tomato challenge. Sci. Rep. 2021, 11, 24523. [Google Scholar] [CrossRef]
- Wang, J.; Song, L.; Jiao, Q.; Yang, S.; Gao, R.; Lu, X.; Zhou, G. Comparative genome analysis of jujube witches’-broom Phytoplasma, an obligate pathogen that causes jujube witches’-broom disease. BMC Genom. 2018, 19, 689. [Google Scholar] [CrossRef] [PubMed]
- Tokuda, R.; Iwabuchi, N.; Kitazawa, Y.; Nijo, T.; Suzuki, M.; Maejima, K.; Oshima, K.; Namba, S.; Yamaji, Y. Potential mobile units drive the horizontal transfer of phytoplasma effector phyllogen genes. Front. Genet. 2023, 14, 1132432. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.B.; Fan, G.Q.; Zhai, X.Q.; Dong, Y.P. Genome-wide analysis of lncRNAs in Paulownia tomentosa infected with phytoplasmas. Acta Physiol. Plant. 2018, 40, 49. [Google Scholar] [CrossRef]
- Fan, G.; Cao, Y.; Wang, Z. Regulation of long noncoding RNAs responsive to phytoplasma infection in Paulownia tomentosa. Int. J. Genom. 2018, 2018, 3174352. [Google Scholar] [CrossRef] [PubMed]
- Deng, P.; Liu, S.; Nie, X.; Weining, S.; Wu, L. Conservation analysis of long non-coding RNAs in plants. Sci. China Life Sci. 2018, 61, 190–198. [Google Scholar] [CrossRef]
- Ji, X.; Gai, Y.; Zheng, C.; Mu, Z. Comparative proteomic analysis provides new insights into mulberry dwarf responses in mulberry (Morus alba L.). Proteomics 2009, 9, 5328–5339. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Mazziotta, C.; Cervellera, C.F.; Badiale, G.; Vitali, I.; Touzé, A.; Tognon, M.; Martini, F.; Rotondo, J.C. Distinct retinoic gene signatures discriminate Merkel cell polyomavirus-positive from -negative Merkel cell carcinoma cells. J. Med. Virol. 2023, 95, e28949. [Google Scholar] [CrossRef]
- Tomczak, A.; Mortensen, J.M.; Winnenburg, R.; Liu, C.; Alessi, D.T.; Swamy, V.; Vallania, F.; Lofgren, S.; Haynes, W.; Shah, N.H.; et al. Interpretation of biological experiments changes with evolution of the Gene Ontology and its annotations. Sci. Rep. 2018, 8, 5115. [Google Scholar] [CrossRef]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. GUS fusions: Beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [CrossRef]
- Harrison, S.J.; Mott, E.K.; Parsley, K.; Aspinall, S.; Gray, J.C.; Cottage, A. A rapid and robust method of identifying transformed Arabidopsis thaliana seedlings following floral dip transformation. Plant Methods 2006, 2, 19. [Google Scholar] [CrossRef]
- Patra, G.K.; Gupta, D.; Rout, G.R.; Panda, S.K. Role of long non coding RNA in plants under abiotic and biotic stresses. Plant Physiol. Biochem. 2023, 194, 96–110. [Google Scholar] [CrossRef]
- Sun, Q.; Hao, Q.; Prasanth, K.V. Nuclear long noncoding RNAs: Key regulators of gene expression. Trends Genet. 2018, 34, 142–157. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Luo, X.; Sun, F.; Hu, J.; Zha, X.; Su, W.; Yang, J. Overexpressing lncRNA LAIR increases grain yield and regulates neighbouring gene cluster expression in rice. Nat. Commun. 2018, 9, 3516. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Long, X.; Gao, M.; Zhao, Y.; Guan, X. Global identification of natural antisense transcripts in Gossypium hirsutum and Gossypium barbadense under chilling stress. iScience 2023, 26, 107362. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Li, J.; Lian, B.; Gu, H.; Li, Y.; Qi, Y. Global identification of Arabidopsis lncRNAs reveals the regulation of MAF4 by a natural antisense RNA. Nat. Commun. 2018, 9, 5056. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Marchese, F.P.; Raimondi, I.; Huarte, M. The multidimensional mechanisms of long noncoding RNA function. Genome Biol. 2017, 18, 206. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.; Mattick, J. Structure and function of long noncoding RNAs in epigenetic regulation. Nat. Struct. Mol. Biol. 2013, 20, 300–307. [Google Scholar] [CrossRef]
- Yan, J.; Su, P.; Meng, X.; Liu, P. Phylogeny of the plant receptor-like kinase (RLK) gene family and expression analysis of wheat RLK genes in response to biotic and abiotic stresses. BMC Genom. 2023, 24, 224. [Google Scholar] [CrossRef]
- Wang, L.; Xu, G.; Li, L.; Ruan, M.; Bennion, A.; Wang, G.L.; Li, R.; Qu, S. The OsBDR1-MPK3 module negatively regulates blast resistance by suppressing the jasmonate signaling and terpenoid biosynthesis pathway. Proc. Natl. Acad. Sci. USA 2023, 120, e2211102120. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Li, L.; Wang, P.; Zhang, S.; Wu, J. Genome-wide characterization, evolution, and expression analysis of the leucine-rich repeat receptor-like protein kinase (LRR-RLK) gene family in Rosaceae genomes. BMC Genom. 2017, 18, 763. [Google Scholar] [CrossRef]
- Wang, Z.; Cheng, J.; Fan, A.; Zhao, J.; Yu, Z.; Li, Y.; Zhang, H.; Xiao, J.; Muhammad, F.; Wang, H.; et al. LecRK-V, an L-type lectin receptor kinase in Haynaldia villosa, plays positive role in resistance to wheat powdery mildew. Plant Biotechnol. J. 2018, 16, 50–62. [Google Scholar] [CrossRef]
- Hu, L.; Ye, M.; Kuai, P.; Ye, M.; Erb, M.; Lou, Y. OsLRR-RLK1, an early responsive leucine-rich repeat receptor-like kinase, initiates rice defense responses against a chewing herbivore. New Phytol. 2018, 219, 1097–1111. [Google Scholar] [CrossRef]
- Liu, L.; Liu, J.; Xu, N. Ligand recognition and signal transduction by lectin receptor-like kinases in plant immunity. Front. Plant Sci. 2023, 14, 1201805. [Google Scholar] [CrossRef] [PubMed]
- Dermastia, M. Plant hormones in phytoplasma infected plants. Front. Plant Sci. 2019, 10, 477. [Google Scholar] [CrossRef] [PubMed]





| LncRNAs | Up/Down Stream | Co-Localization Differentially Expressed Target Genes | |||||||
|---|---|---|---|---|---|---|---|---|---|
| ID | log2FoldChange (IL/HL) | p Value | Up/Down | ID | log2FoldChange (IL/HL) | p Value | Up/Down | Description | |
| XLOC_000009 | −1.659904691 | 2.01 × 10−5 | Down | Upstream_2k | XP_010086539.1 | −6.285402219 | 9.71 × 10−11 | Down | Uncharacterized protein LOC21384651 |
| XLOC_003582 | −1.357748564 | 4.19 × 10−63 | Down | Downstream_2k | XP_010089960.1 | 1.622764533 | 2.69 × 10−30 | Up | NADP-dependent glyceraldehyde-3-phosphate dehydrogenase |
| XLOC_004104 | 2.30176161 | 0 | Up | Upstream_2k | XP_010090441.1 | −1.498805857 | 8.26 × 10−9 | Down | NAC domain containing protein 50 isoform X1 |
| XLOC_004314 | −11.62763875 | 4.50 × 10−161 | Down | Downstream_2k | XP_010090692.1 | 5.044394119 | 5.83 × 10−8 | Up | Uncharacterized protein |
| XLOC_005352 | −1.359639818 | 7.41 × 10−74 | Down | Downstream_2k | XP_010091710.1 | −1.75161237 | 0.00017364 | Down | Hypothetical protein |
| XLOC_005353 | −2.142379817 | 1.19 × 10−54 | Down | Downstream_2k | XP_010091710.1 | −1.75161237 | 0.00017364 | Down | Hypothetical protein |
| XLOC_006216 | −8.087982542 | 0.000355214 | Down | Downstream_2k | XP_010092498.1 | 11.06173466 | 0 | Up | Hypothetical protein |
| XLOC_006461 | −2.220725888 | 3.24 × 10−11 | Down | Upstream_2k | XP_010092702.1 | −2.115477217 | 8.81 × 10−8 | Down | Pentatricopeptide repeat-containing protein |
| XLOC_006690 | −1.496801279 | 2.07 × 10−6 | Down | Downstream_2k | XP_010092905.1 | 1.325507985 | 2.24 × 10−35 | Up | Common plant regulatory factor 1 |
| XLOC_006961 | −1.32577553 | 4.79 × 10−5 | Down | Upstream_2k | XP_010093164.1 | −2.191426071 | 1.03 × 10−19 | Down | Hypothetical protein |
| XLOC_007096 | −1.048521438 | 2.67 × 10−8 | Down | Downstream_2k | XP_010093303.1 | −1.415037499 | 1.18 × 10−5 | Down | ATP-dependent zinc metalloprotease FtsH |
| XLOC_008373 | 2.902444746 | 5.16 × 10−6 | Up | Upstream_2k | XP_010094597.1 | 1.378394431 | 4.77 × 10−91 | Up | 4-coumarate--CoA ligase-like 5 |
| XLOC_008398 | −2.45656083 | 0.474176 | Down | Upstream_2k | XP_010094597.1 | 1.378394431 | 4.77 × 10−91 | Up | 4-coumarate--CoA ligase-like 5 |
| XLOC_016866 | −1.108741537 | 3.09 × 10−6 | Down | Downstream_2k | XP_010094598.1 | −1.708344916 | 7.10 × 10−19 | Down | Protein ROOT PRIMORDIUM DEFECTIVE 1 |
| XLOC_009663 | −2.043818661 | 1.86 × 10−23 | Down | Upstream_2k | XP_010095745.1 | −2.172404286 | 9.14 × 10−119 | Down | Pyruvate kinase isozyme A |
| XLOC_011123 | −1.699511675 | 1.81 × 10−14 | Down | Downstream_2k | XP_010097263.1 | −1.570240995 | 7.69 × 10−21 | Down | Hypothetical protein |
| XLOC_011929 | 1.009110152 | 2.46 × 10−50 | Up | Downstream_2k | XP_010097937.1 | −1.653442239 | 6.36 × 10−61 | Down | Retrovirus-related Pol polyprotein from transposon TNT 1–94 |
| XLOC_011968 | −3.450165156 | 0.000183881 | Down | Downstream_2k | XP_010097994.1 | −6.087462841 | 0.00017296 | Down | GDT1-like protein 2 |
| XLOC_012270 | −1.271836651 | 1.07 × 10−23 | Down | Upstream_2k | XP_010098256.1 | −1.115122616 | 1.46 × 10−36 | Down | DEAD-box ATP-dependent RNA helicase ISE2 |
| XLOC_012337 | −2.438980184 | 0.474176 | Down | Upstream_2k | XP_010098340.1 | −2.850856561 | 0.00017039 | Down | Zinc finger CCCH domain-containing protein 58 |
| XLOC_012366 | −2.450477186 | 0.0266538 | Down | Upstream_2k | XP_010098340.1 | −2.850856561 | 0.00017039 | Down | Zinc finger CCCH domain-containing protein 58 |
| XLOC_014320 | −2.086989256 | 5.76 × 10−12 | Down | Downstream_2k | XP_010100223.1 | −1.047305715 | 1.35 × 10−7 | Down | Ubiquitin carboxyl-terminal hydrolase 10 |
| XLOC_014820 | −1.871361676 | 6.68 × 10−98 | Down | Downstream_2k | XP_010100733.1 | −4.037089319 | 0.00018388 | Down | Hypothetical protein |
| XLOC_015257 | −2.853283109 | 0.474176 | Down | Upstream_2k | XP_010101095.1 | −1.461049897 | 2.36 × 10−7 | Down | Histone-lysine N-methyltransferase |
| XLOC_015724 | 12.61184784 | 4.13 × 10−69 | Up | Downstream_2k | XP_010101540.1 | −2.722049907 | 0 | Down | GDSL esterase/lipase |
| XLOC_017908 | −1.9200952 | 1.66 × 10−142 | Down | Upstream_2k | XP_010103622.1 | −1.972293071 | 6.04 × 10−21 | Down | Dihydroorotate dehydrogenase (quinone) |
| XLOC_018202 | −1.743622263 | 5.76 × 10−133 | Down | Upstream_2k | XP_010103913.1 | −1.545434137 | 3.59 × 10−9 | Down | Hypothetical protein |
| XLOC_018332 | 2.51571606 | 0.000105589 | Up | Downstream_2k | XP_010104046.1 | −1.906890596 | 9.50 × 10−8 | Down | Hypothetical protein |
| XLOC_018333 | 1.508625478 | 2.35 × 10−15 | Up | Downstream_2k | XP_010104046.1 | −1.906890596 | 9.50 × 10−8 | Down | Hypothetical protein |
| XLOC_018334 | 1.136596235 | 1.14 × 10−12 | Up | Downstream_2k | XP_010104046.1 | −1.906890596 | 9.50 × 10−8 | Down | Hypothetical protein |
| XLOC_018352 | 1.274845329 | 8.73 × 10−16 | Up | Upstream_2k | XP_010104047.1 | −1.465663572 | 4.78 × 10−11 | Down | Glucuronoxylan 4-O-methyltransferase 1 |
| XLOC_018600 | 1.232238957 | 0.000128139 | Up | Downstream_2k | XP_010104288.1 | −1.080959321 | 2.26 × 10−9 | Down | Uncharacterized protein LOC21388859 |
| XLOC_018675 | −1.476521201 | 4.78 × 10−16 | Down | Upstream_2k | XP_010104373.1 | −1.862496476 | 0.00022881 | Down | Two-component response regulator-like protein |
| XLOC_018680 | −2.179955771 | 2.14 × 10−6 | Down | Upstream_2k | XP_010104373.1 | −1.862496476 | 0.00022881 | Down | Two-component response regulator-like protein |
| XLOC_018692 | −2.139247136 | 2.33 × 10−37 | Down | Upstream_2k | XP_010104373.1 | −1.862496476 | 0.00022881 | Down | Two-component response regulator-like protein |
| XLOC_018696 | 12.2276009 | 3.01 × 10−28 | Up | Upstream_2k | XP_010104373.1 | −1.862496476 | 0.00022881 | Down | Two-component response regulator-like protein |
| XLOC_019166 | −1.120989021 | 2.10 × 10−94 | Down | Downstream_2k | XP_010104915.1 | −1.100273908 | 1.37 × 10−28 | Down | Uncharacterized protein LOC21406865 |
| XLOC_019555 | −2.58203706 | 0.0595904 | Down | Downstream_2k | XP_010105201.1 | −5.087462841 | 2.00 × 10−5 | Down | Sn1-specific diacylglycerol lipase beta |
| XLOC_019810 | 14.10733152 | 1.15 × 10−47 | Up | Downstream_2k | XP_010105413.1 | −1.70571466 | 2.40 × 10−31 | Down | Putative GDP-L-fucose synthase 2 |
| XLOC_020110 | −1.030050297 | 2.09 × 10−24 | Down | Upstream_2k | XP_010105705.1 | −1.650322233 | 8.17 × 10−45 | Down | Guanylate kinase 2 |
| XLOC_020112 | 1.243959167 | 7.39 × 10−25 | Up | Upstream_2k | XP_010105705.1 | −1.650322233 | 8.17 × 10−45 | Down | Guanylate kinase 2 |
| XLOC_020267 | −1.009344657 | 8.55 × 10−6 | Down | Downstream_2k | XP_010105833.1 | −2.321928095 | 1.57 × 10−10 | Down | Pre-mRNA-processing protein 40A isoform X2 |
| XLOC_020549 | −3.468308658 | 2.93 × 10−10 | Down | Upstream_2k | XP_010106108.1 | −7.6794801 | 8.38 × 10−20 | Down | Hypothetical protein |
| XLOC_020799 | −3.174801049 | 8.27 × 10−9 | Down | Upstream_2k | XP_010106332.1 | 4.558967292 | 1.88 × 10−156 | Up | Hypothetical protein |
| XLOC_020823 | −1.290572462 | 9.68 × 10−10 | Down | Upstream_2k | XP_010106349.1 | −3.019590728 | 1.52 × 10−41 | Down | Hypothetical protein |
| XLOC_020815 | −1.550204751 | 4.40 × 10−5 | Down | Downstream_2k | XP_010106354.1 | 2.674712213 | 1.36 × 10−243 | Up | G-type lectin S-receptor-like serine/threonine-protein kinase |
| XLOC_021165 | −1.220913717 | 3.65 × 10−8 | Down | Upstream_2k | XP_010106679.1 | 1.85708704 | 4.53 × 10−17 | Up | Inorganic pyrophosphatase 1 |
| XLOC_021166 | −2.999715655 | 2.63 × 10−30 | Down | Upstream_2k | XP_010106679.1 | 1.85708704 | 4.53 × 10−17 | Up | Inorganic pyrophosphatase 1 |
| XLOC_021967 | −1.459141927 | 3.89 × 10−9 | Down | Downstream_2k | XP_010107456.1 | −3.378511623 | 2.13 × 10−13 | Down | uncharacterized protein LOC21385704 isoform X1 |
| XLOC_022178 | −1.074733339 | 1.69 × 10−54 | Down | Downstream_2k | XP_010107670.1 | 1.48112669 | 2.62 × 10−11 | Up | probable indole-3-acetic acid-amido synthetase GH3.6 |
| XLOC_023091 | 2.645694109 | 1.02 × 10−6 | Up | Downstream_2k | XP_010108540.1 | −2.264499815 | 3.96 × 10−15 | Down | hypothetical protein |
| XLOC_024046 | −2.458590043 | 0.230884 | Down | Downstream_2k | XP_010109450.1 | −1.638260727 | 1.48 × 10−60 | Down | mitochondrial carrier protein MTM1 isoform X1 |
| XLOC_024072 | −1.363720387 | 1.03 × 10−15 | Down | Downstream_2k | XP_010109466.1 | −1.469695084 | 8.90 × 10−40 | Down | hypothetical protein |
| XLOC_024250 | 1.511943985 | 4.91 × 10−15 | Up | Downstream_2k | XP_010109645.1 | 1.030214613 | 9.23 × 10−51 | Up | transcription factor DIVARICATA |
| XLOC_024279 | −2.968724068 | 0.474176 | Down | Upstream_2k | XP_010109661.1 | −2.164282855 | 1.62 × 10−187 | Down | pentatricopeptide repeat-containing protein |
| XLOC_024407 | −1.218449856 | 1.04 × 10−22 | Down | Upstream_2k | XP_010109786.1 | −1.513069582 | 3.18 × 10−11 | Down | uncharacterized protein At4g10930 |
| XLOC_025437 | −2.561579471 | 0.474176 | Down | Downstream_2k | XP_010110803.1 | 1.12963528 | 2.27 × 10−5 | Up | RING finger protein B |
| XLOC_025452 | −2.474769468 | 0.474176 | Down | Upstream_2k | XP_010110804.1 | 5.720122084 | 8.40 × 10−129 | Up | leucoanthocyanidin reductase |
| XLOC_025453 | 3.77129058 | 6.73 × 10−9 | Up | Upstream_2k | XP_010110804.1 | 5.720122084 | 8.40 × 10−129 | Up | leucoanthocyanidin reductase |
| XLOC_025463 | 1.448311774 | 5.88 × 10−9 | Up | Upstream_2k | XP_010110804.1 | 5.720122084 | 8.40 × 10−129 | Up | leucoanthocyanidin reductase |
| XLOC_025605 | 1.268119766 | 1.27 × 10−140 | Up | Downstream_2k | XP_010110915.1 | −1.776621795 | 1.26 × 10−42 | Down | hypothetical protein |
| XLOC_025845 | 1.086054103 | 0.000228706 | Up | Upstream_2k | XP_010111144.1 | −5.584962501 | 2.31 × 10−6 | Down | hypothetical protein |
| XLOC_025906 | 1.313631149 | 7.94 × 10−25 | Up | Upstream_2k | XP_010111198.1 | −1.252187024 | 3.51 × 10−8 | Down | putative E3 ubiquitin-protein ligase RING1a isoform X2 |
| XLOC_025907 | 2.290116081 | 5.36 × 10−23 | Up | Upstream_2k | XP_010111198.1 | −1.252187024 | 3.51 × 10−8 | Down | putative E3 ubiquitin-protein ligase RING1a isoform X2 |
| XLOC_025909 | −1.440184589 | 9.82 × 10−6 | Up | Upstream_2k | XP_010111198.1 | −1.252187024 | 3.51 × 10−8 | Down | putative E3 ubiquitin-protein ligase RING1a isoform X2 |
| XLOC_026031 | −1.440184589 | 9.82 × 10−6 | Down | Downstream_2k | XP_010111442.1 | 1.015266757 | 0.0001083 | Up | hypothetical protein |
| XLOC_026652 | −2.355017768 | 0.474176 | Down | Upstream_2k | XP_010111922.1 | −2.30256277 | 1.38 × 10−15 | Down | uncharacterized protein LOC21393124 |
| XLOC_026790 | −1.123960181 | 9.54 × 10−5 | Down | Upstream_2k | XP_010112073.1 | −2.692490965 | 1.52 × 10−20 | Down | ATP-dependent Clp protease ATP-binding subunit ClpX |
| LncRNAs | Target Genes | |||||||
|---|---|---|---|---|---|---|---|---|
| ID | log2FoldChange (B/J) | p Value | Up/Down | ID | log2FoldChange (BB/JJ) | p Value | Up/Down | Descrition |
| XLOC_001590 | 3.837441902 | 0.000471504 | Up | XP_010088049.1 | 2.605901838 | 6.15 × 10−43 | Up | Chaperone protein DnaJ |
| XLOC_003017 | 4.009664269 | 0.012449373 | Up | XP_010089382.1 | −1.15417093 | 2.37 × 10−8 | Down | PAP-specific phosphatase HAL2-like protein |
| XLOC_003046 | −2.433070931 | 0.014546012 | Down | XP_010089404.1 | −2.06667104 | 3.31 × 10−9 | Down | Hypothetical protein L484_013795 |
| XLOC_003503 | −1.599653771 | 1.22743 × 10−11 | Down | XP_010089852.1 | −7.033423002 | 2.00 × 10−5 | Down | Subtilisin-like protease |
| XLOC_004551 | −1.779844507 | 0.034037713 | Down | XP_010090841.1 | −1.385653692 | 4.59 × 10−6 | Down | Putative polygalacturonase |
| XLOC_004954 | 2.379663284 | 7.51328 × 10-−5 | Up | XP_010091231.1 | 2.011383996 | 0 | Up | Hypothetical protein L484_005255 ] |
| XLOC_005131 | −2.31198078 | 0.003175704 | Down | XP_010091411.1 | −2.021479727 | 7.08 × 10−11 | Down | Chlorophyll a-b binding protein 16 |
| XLOC_005342 | −8.732455766 | 2.79104 × 10−56 | Down | XP_010091645.1 | 2.479780264 | 5.94 × 10−26 | Up | UDP-glycosyltransferase 89A2 |
| XLOC_006046 | −1.595761698 | 2.7063 × 10−13 | Down | XP_010092313.1 | −2.662965013 | 1.18 × 10−40 | Down | Putative galacturonosyltransferase 12 |
| XLOC_006462 | −2.022357412 | 4.62933 × 10−36 | Down | XP_010092676.1 | −1.69538649 | 2.14 × 10−20 | Down | Hypothetical protein L484_019750 |
| XLOC_009619 | −1.020506348 | 6.48115 × 10−7 | Down | XP_010095699.1 | −1.156119202 | 2.20 × 10−7 | Down | Putative endo-1,4-beta-xylanase C |
| XLOC_010285 | −1.479609008 | 0.00017518 | Down | XP_010096371.1 | 5.169925001 | 0.00017525 | Up | Myb-related protein B |
| XLOC_011021 | −2.356597415 | 1.18979 × 10−10 | Down | XP_010097040.1 | −2.938599455 | 1.17 × 10−8 | Down | Hypothetical protein L484_003871 |
| XLOC_011087 | −1.189537055 | 1.92884 × 10−8 | Down | XP_010097097.1 | −1.310381065 | 0 | Down | Hypothetical protein L484_019536 |
| XLOC_011132 | −1.325768866 | 3.97956 × 10−11 | Down | XP_010097155.1 | −1.830074999 | 7.78 × 10−6 | Down | ATPase 9, plasma membrane-type |
| XLOC_011724 | −1.892272481 | 7.40323 × 10−13 | Down | XP_010097661.1 | −1.523561956 | 4.05 × 10−14 | Down | Phytosulfokine receptor 2 |
| XLOC_012537 | −1.437284073 | 0.002462778 | Down | XP_010098515.1 | −1.659924558 | 4.93 × 10−27 | Down | Hypothetical protein L484_025954 |
| XLOC_012821 | −1.545482034 | 1.99649 × 10−14 | Down | XP_010098738.1 | −1.74723393 | 2.02 × 10−8 | Down | Pectinesterase 3 |
| XLOC_012923 | 1.151448857 | 5.66295 × 10−6 | Up | XP_010098833.1 | −1.27364808 | 3.01 × 10−6 | Down | UDP-glycosyltransferase 73C3 |
| XLOC_013301 | −1.135788866 | 0.000554496 | Down | XP_010099215.1 | −1.963474124 | 3.86 × 10−34 | Down | Hypothetical protein L484_010155 |
| XLOC_013697 | −2.587472054 | 9.67889 × 10−17 | Down | XP_010099597.1 | −1.17290886 | 5.38 × 10−6 | Down | Histone H3-like centromeric protein |
| XLOC_015378 | −1.534366505 | 0.000274527 | Down | XP_010101197.1 | −1.021061616 | 0.000108539 | Down | Hypothetical protein L484_015001 |
| XLOC_016174 | −1.659968292 | 5.73067 × 10−20 | Down | XP_010101961.1 | −2.408084739 | 2.01 × 10−5 | Down | Hypothetical protein L484_011978 |
| XLOC_016448 | 9.017824501 | 0.008623184 | Up | XP_010102287.1 | 5.984893108 | 1.89 × 10−19 | Up | Hypothetical protein L484_024569 |
| XLOC_017317 | −2.24788776 | 1.18979 × 10−10 | Down | XP_010103054.1 | −2.815174427 | 7.64 × 10−92 | Down | Hypothetical protein L484_001885 |
| XLOC_017564 | −1.780652222 | 5.73067 × 10−20 | Down | XP_010103289.1 | 4.930737338 | 3.35 × 10−92 | Up | 3’-hydroxy-N-methyl-(S)-coclaurine 4’-O-methyltransferase |
| XLOC_018767 | −2.14733932 | 8.58013 × 10−35 | Down | XP_010104430.1 | −4.087462841 | 0.000183881 | Down | Hypothetical protein L484_016029 |
| XLOC_019411 | −1.597933878 | 0.006303773 | Down | XP_010105051.1 | −2.297266041 | 3.97 × 10−15 | Down | Hypothetical protein L484_001492 |
| XLOC_020739 | −2.772398916 | 2.36945 × 10−8 | Down | XP_010106301.1 | −3.795859283 | 3.06 × 10−10 | Down | Magnesium-transporting ATPase, P-type 1 |
| XLOC_020789 | 1.043080361 | 4.3553 × 10−7 | Up | XP_010106332.1 | 4.558967292 | 1.88 × 10−156 | Up | Hypothetical protein L484_004731 |
| XLOC_021217 | −1.307207494 | 8.68908 × 10−7 | Down | XP_010106725.1 | −1.634715536 | 5.88 × 10−5 | Down | Myb-related protein Myb4 |
| XLOC_021596 | −2.443238328 | 0.003175704 | Down | XP_010107118.1 | −1.093574115 | 9.06 × 10−5 | Down | NAC domain-containing protein 1 |
| XLOC_022461 | −1.911411449 | 0.031545424 | Down | XP_010107850.1 | −2.275735285 | 7.51 × 10−53 | Down | Hypothetical protein L484_027437 |
| XLOC_022591 | 2.284500804 | 8.77802 × 10−9 | Up | XP_010108041.1 | 1.83518913 | 1.25 × 10−6 | Up | Exocyst complex component 7 |
| XLOC_022840 | 1.471119577 | 0.008783891 | Up | XP_010108284.1 | −6.906890596 | 3.53 × 10−19 | Down | Hypothetical protein L484_007137 |
| XLOC_022990 | −9.988102379 | 7.94406 × 10−5 | Down | XP_010108424.1 | 2.03562391 | 8.43 × 10−11 | Up | Nudix hydrolase 15 |
| XLOC_025085 | 9.747083561 | 6.51127 × 10−23 | Up | XP_010110394.1 | 1.094718657 | 0.000123589 | Up | Hypothetical protein L484_022797 |
| XLOC_025632 | −2.675688053 | 7.45831 × 10−12 | Down | XP_010110923.1 | −1.80937409 | 2.02 × 10−73 | Down | Omega-hydroxypalmitate O-feruloyl transferase |
| XLOC_026011 | −2.847603572 | 7.57567 × 10−8 | Down | XP_010111400.1 | 1.961110987 | 1.28 × 10−25 | Up | Multiple C2 and transmembrane domain-containing protein 2 |
| XLOC_026715 | −2.99257368 | 2.63073 × 10−53 | Down | XP_010111977.1 | −3.89077093 | 2.45 × 10−17 | Down | Putative peptide/nitrate transporter |
| XLOC_026802 | −2.726806211 | 4.61238 × 10−40 | Down | XP_010112045.1 | −1.807354922 | 1.62 × 10−8 | Down | Endoglucanase 9 |
| XLOC_027017 | −1.271269054 | 5.03723 × 10−10 | Down | XP_010112258.1 | −1.332575339 | 0.000230944 | Down | MADS-box transcription factor 27 |
| XLOC_027018 | −2.185948672 | 1.18979 × 10−10 | Down | XP_010112258.1 | −1.332575339 | 0.000230944 | Down | MADS-box transcription factor 27 |
| XLOC_027445 | −1.081848858 | 0.033465549 | Down | XP_010112683.1 | 4.024384159 | 5.29 × 10−30 | Up | Putative LRR receptor-like serine/threonine-protein kinase |
| Site Name | Number | Sequence | Function |
|---|---|---|---|
| ABRE | 2 | ACGTG | Cis-acting element involved in abscisic acid responsiveness |
| ACE | 1 | GACACGTATG | Cis-acting element involved in light responsiveness |
| ARE | 1 | AAACCA | Cis-acting regulatory element essential for anaerobic induction |
| ATCT-motif | 1 | AATCTAATCC | Part of a conserved DNA module involved in light responsiveness |
| AuxRR-core | 1 | GGTCCAT | Cis-acting regulatory element involved in auxin responsiveness |
| Box 4 | 1 | ATTAAT | Part of a conserved DNA module involved in light responsiveness |
| CAAT-box | 31 | CCAAT | Common cis-acting element in promoter and enhancer regions |
| CAT-box | 1 | GCCACT | Cis-acting regulatory element related to meristem expression |
| GARE-motif | 1 | TCTGTTG | Gibberellin-responsive element |
| GATA-motif | 1 | GATAGGA | Part of a light responsive element |
| G-Box | 3 | TACGTG | Cis-acting regulatory element involved in light responsiveness |
| MBS | 1 | CGGTCA | MYB binding site involved in drought-inducibility |
| P-box | 1 | CCTTTTG | Gibberellin-responsive element |
| TATA-box | 13 | TATA | Core promoter element of approximately −30 transcription starts |
| TATC-box | 1 | TATCCCA | Cis-acting element involved in gibberellin-responsiveness |
| TCA-element | 1 | CCATCTTTTT | Cis-acting element involved in salicylic acid responsiveness |
| TCT-motif | 1 | TCTTAC | Part of a light responsive element |
| TGA-element | 1 | AACGAC | Auxin-responsive element |
| Site Name | Number | Sequence | Function |
|---|---|---|---|
| ABRE | 2 | ACGTG | Cis-acting element involved in abscisic acid responsiveness |
| ARE | 2 | AAACCA | Cis-acting regulatory element essential for anaerobic induction |
| Box 4 | 1 | ATTAAT | Part of a conserved DNA module involved in light responsiveness |
| CAAT-box | 46 | CCAAT | Common cis-acting element in promoter and enhancer regions |
| CAT-box | 1 | GCCACT | Cis-acting regulatory element related to meristem expression |
| CGTCA-motif | 2 | CGTCA | Cis-acting regulatory element involved in meja-responsiveness |
| G-Box | 3 | TACGTG | Cis-acting regulatory element involved in light responsiveness |
| GCN4_motif | 1 | TGAGTCA | Cis-regulatory element involved in endosperm expression |
| GT1-motif | 1 | GGTTAA | Light responsive element |
| LTR | 1 | CCGAAA | Cis-acting element involved in low-temperature responsiveness |
| MBS | 1 | CGGTCA | MYB binding site involved in drought-inducibility |
| P-box | 1 | CCTTTTG | Gibberellin-responsive element |
| O2-site | 2 | GATGA(C/T)(A/G)TG(A/G) | Cis-acting regulatory element involved in zein metabolism regulation |
| TATA-box | 46 | TATA | Core promoter element of approximately −30 transcription starts |
| TC-rich repeats | 1 | GTTTTCTTAC | Cis-acting element involved in defense and stress responsiveness |
| TCA-element | 1 | CCATCTTTTT | Cis-acting element involved in salicylic acid responsiveness |
| TCT-motif | 1 | TCTTAC | Part of a light responsive element |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Z.; Liu, C.; Zhao, T.; Yang, L.; Shang, Q.; Wang, G.; Liu, Z.; Gai, Y.; Ji, X. Integrated Analysis of lncRNAs and mRNAs Reveals Complex Gene Network Mediated by lncRNAs and Regulatory Function of MuLRR-RLK-AS in Response to Phytoplasma Infection in Mulberry. Biomolecules 2024, 14, 308. https://doi.org/10.3390/biom14030308
Liu Z, Liu C, Zhao T, Yang L, Shang Q, Wang G, Liu Z, Gai Y, Ji X. Integrated Analysis of lncRNAs and mRNAs Reveals Complex Gene Network Mediated by lncRNAs and Regulatory Function of MuLRR-RLK-AS in Response to Phytoplasma Infection in Mulberry. Biomolecules. 2024; 14(3):308. https://doi.org/10.3390/biom14030308
Chicago/Turabian StyleLiu, Zixuan, Chaorui Liu, Teng Zhao, Lulu Yang, Qiqi Shang, Gefan Wang, Zhaoyang Liu, Yingping Gai, and Xianling Ji. 2024. "Integrated Analysis of lncRNAs and mRNAs Reveals Complex Gene Network Mediated by lncRNAs and Regulatory Function of MuLRR-RLK-AS in Response to Phytoplasma Infection in Mulberry" Biomolecules 14, no. 3: 308. https://doi.org/10.3390/biom14030308






