Exploring the Molecular Mechanism of Comorbidity of Type 2 Diabetes Mellitus and Colorectal Cancer: Insights from Bulk Omics and Single-Cell Sequencing Validation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Download and Pre-Processing of Bulk RNA-Seq Data
2.2. Weighted Gene Co-Expression Network Analysis (WGCNA)
2.3. Identification of Differentially Expressed Genes (DEGs)
2.4. Functional Enrichment Analysis
2.5. Immune Landscape Assessment and Immunotherapy Response Prediction
2.6. Integration and Processing of scRNA-Seq Data
2.7. Pseudotime Analysis
2.8. Connectivity Map (CMap) Drug Identification
2.9. Molecular Docking Analysis
2.10. Statistical Analysis
3. Results
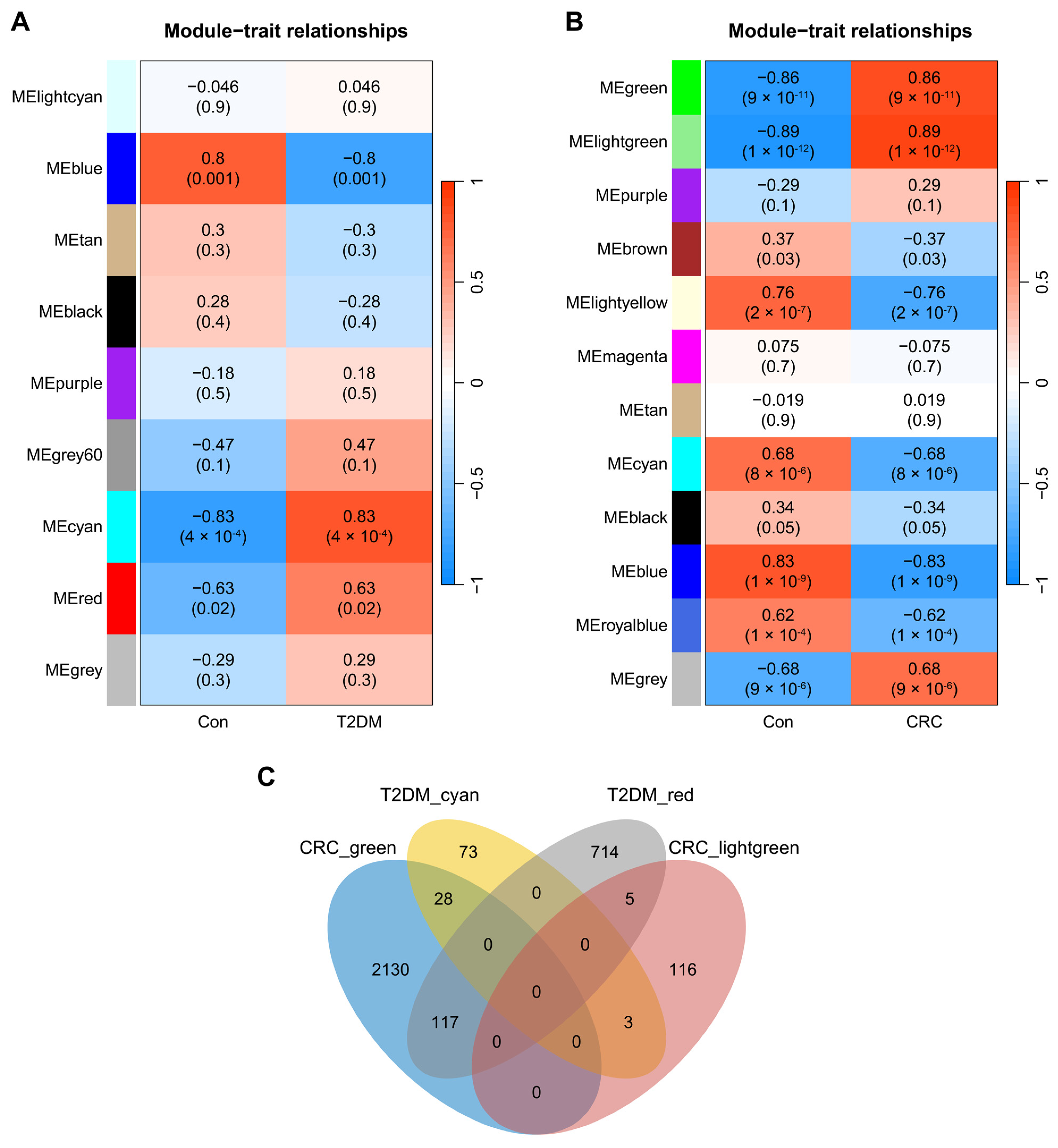
3.1. Co-Expression Modules in T2DM and CRC
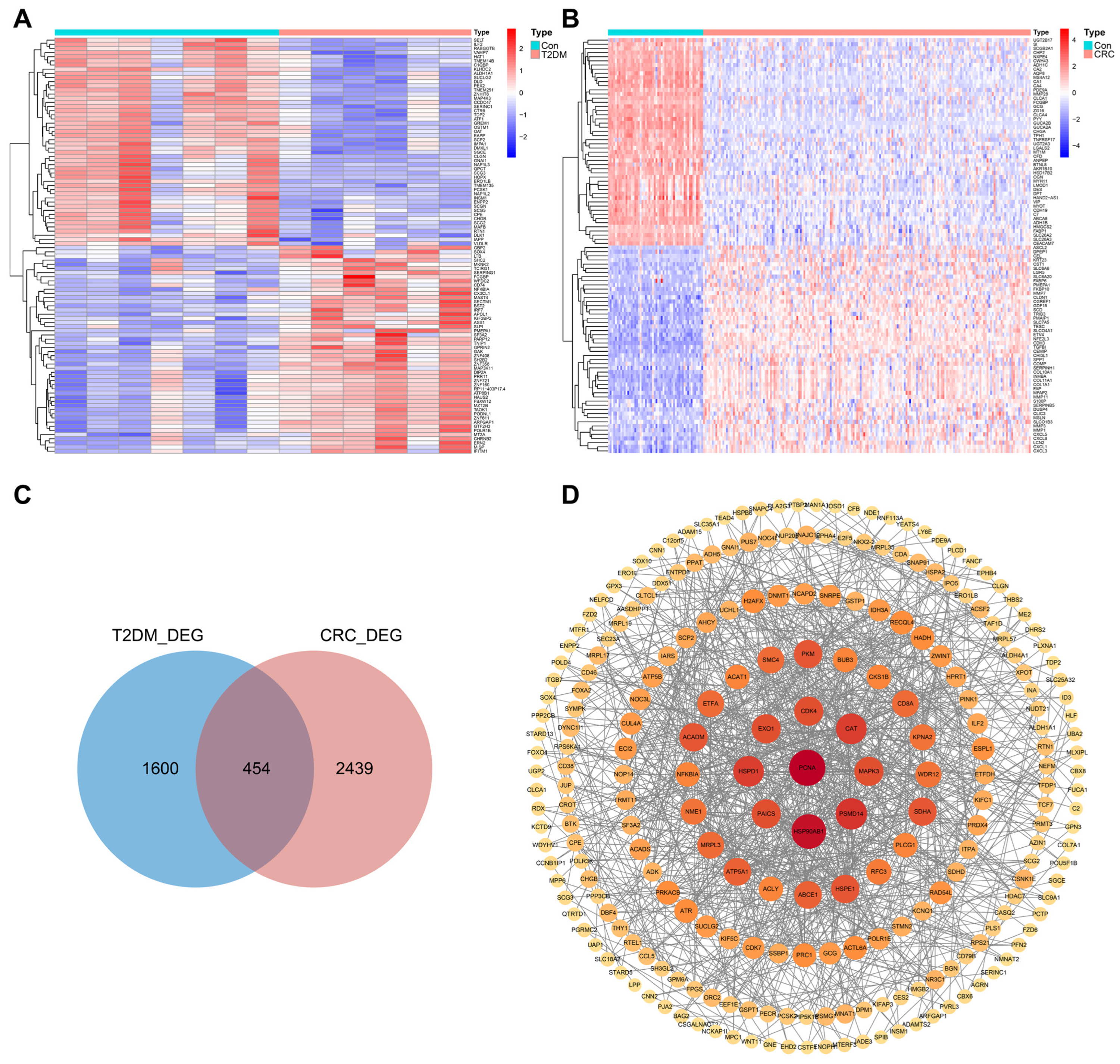
3.2. Identification of DEGs in T2DM and CRC
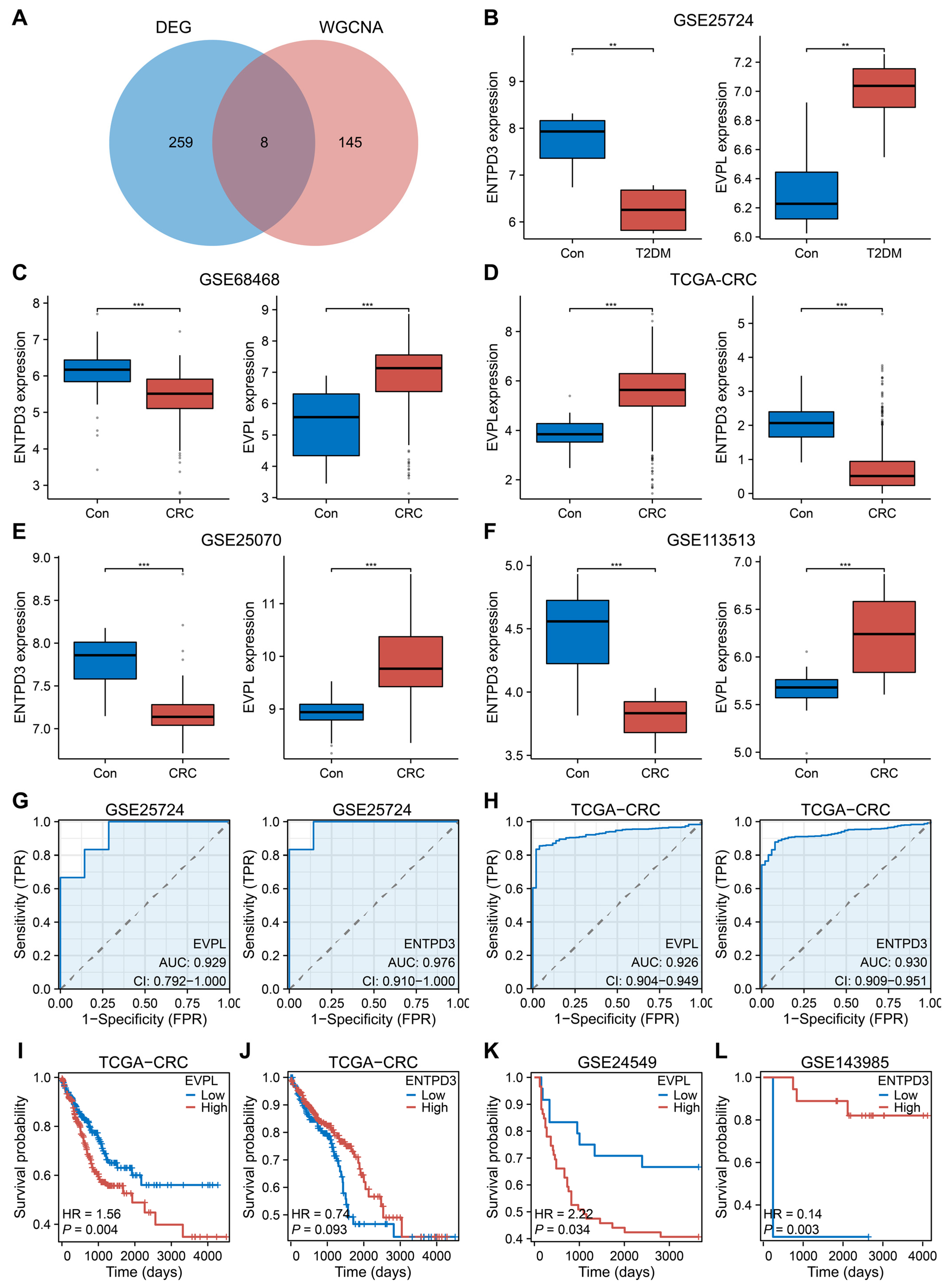
3.3. EVPL and ENTPD3 Are the Optimal Shared Genes
3.4. Biological Processes Associated with EVPL and ENTPD3
3.5. Abnormal Expression of EVPL and ENTPD3 Predicts Altered Immune Landscape in T2DM and CRC
3.6. EVPL and ENTPD3 Are Predictive Biomarkers for Cancer Immunotherapy
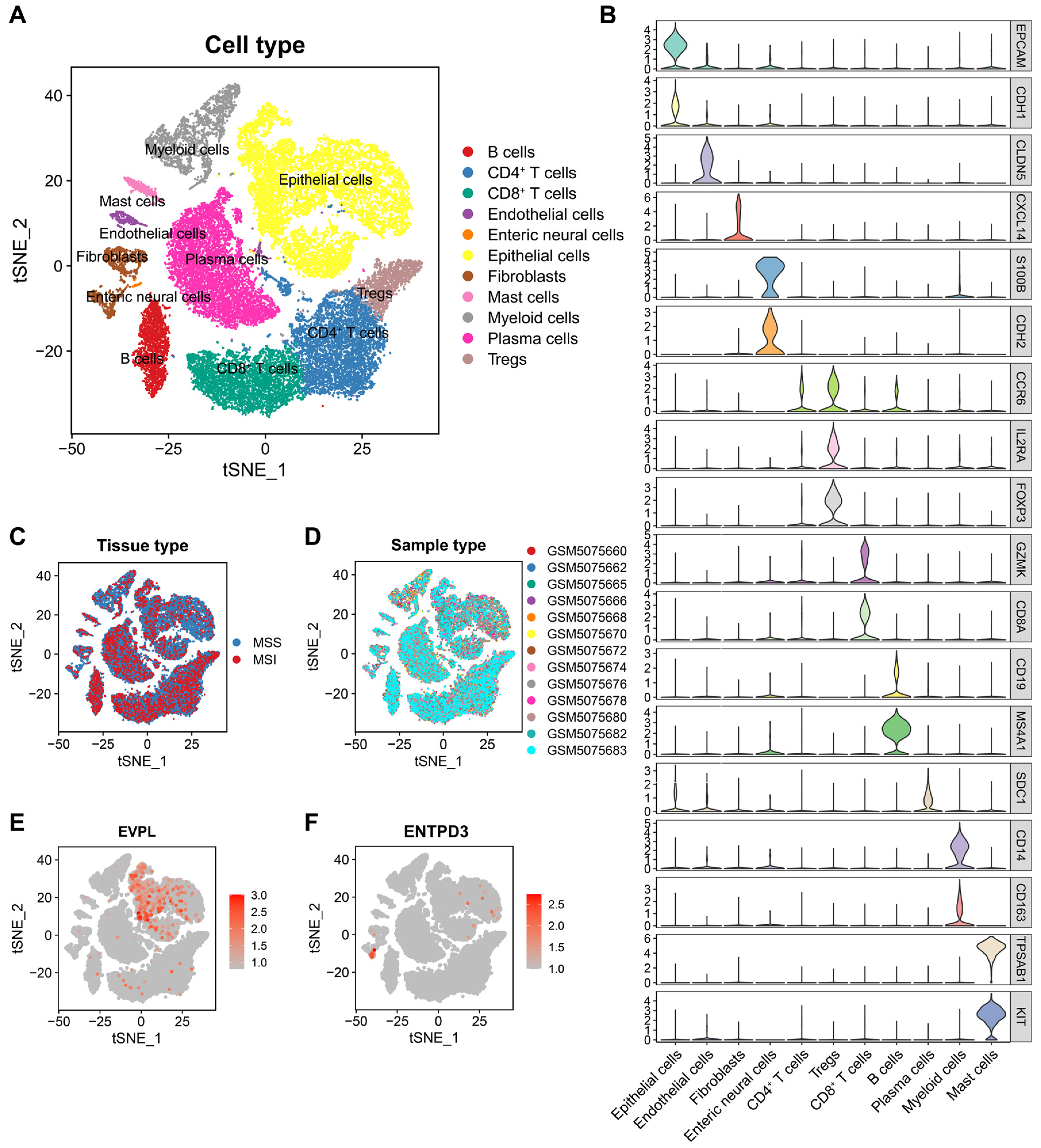
3.7. EVPL and ENTPD3 Are Expressed in Different Cell Subpopulations in CRC
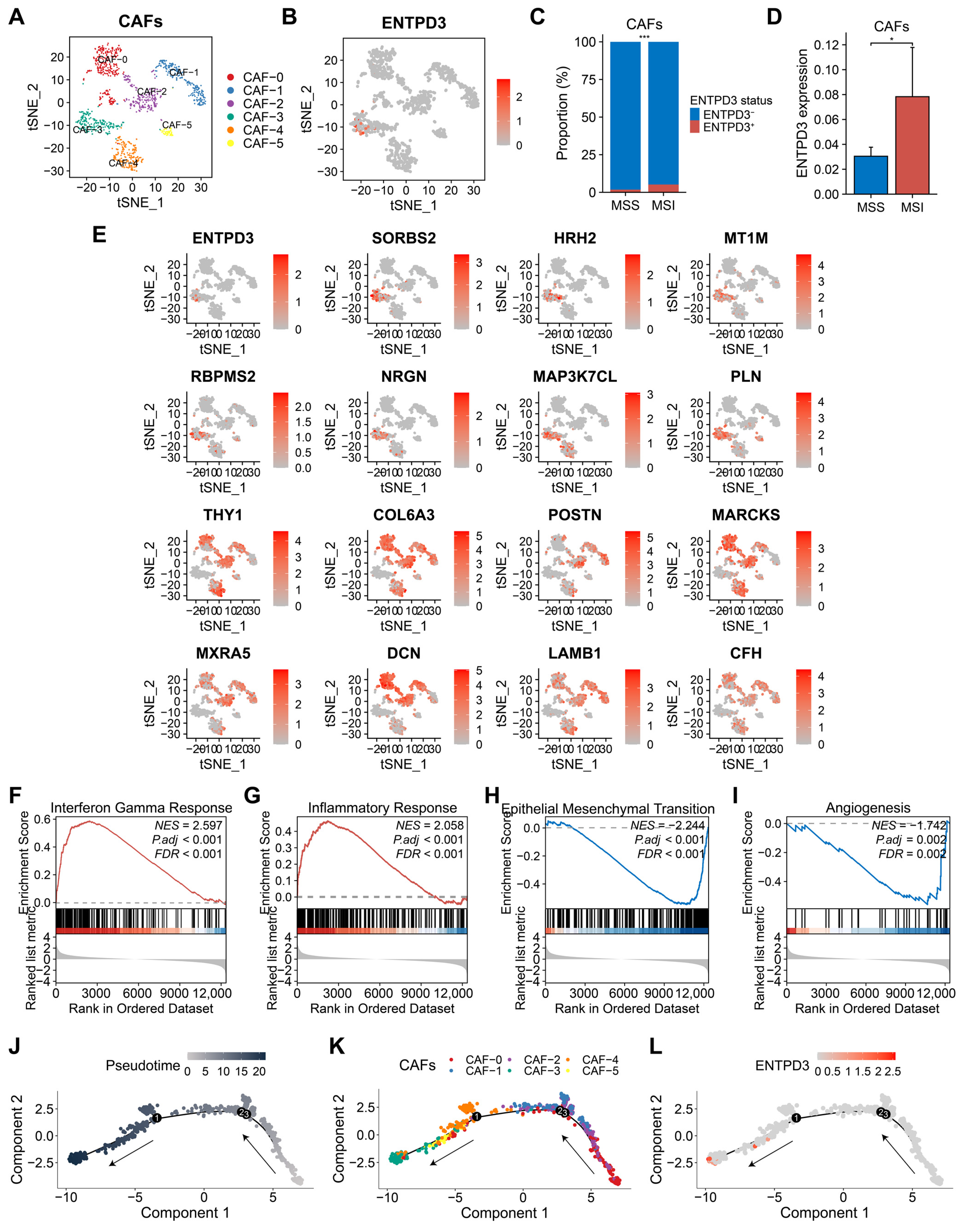
3.8. ENTPD3+ CAFs Exhibit High Heterogeneity and Tumor-Suppressive Features Distinct from Classical CAFs
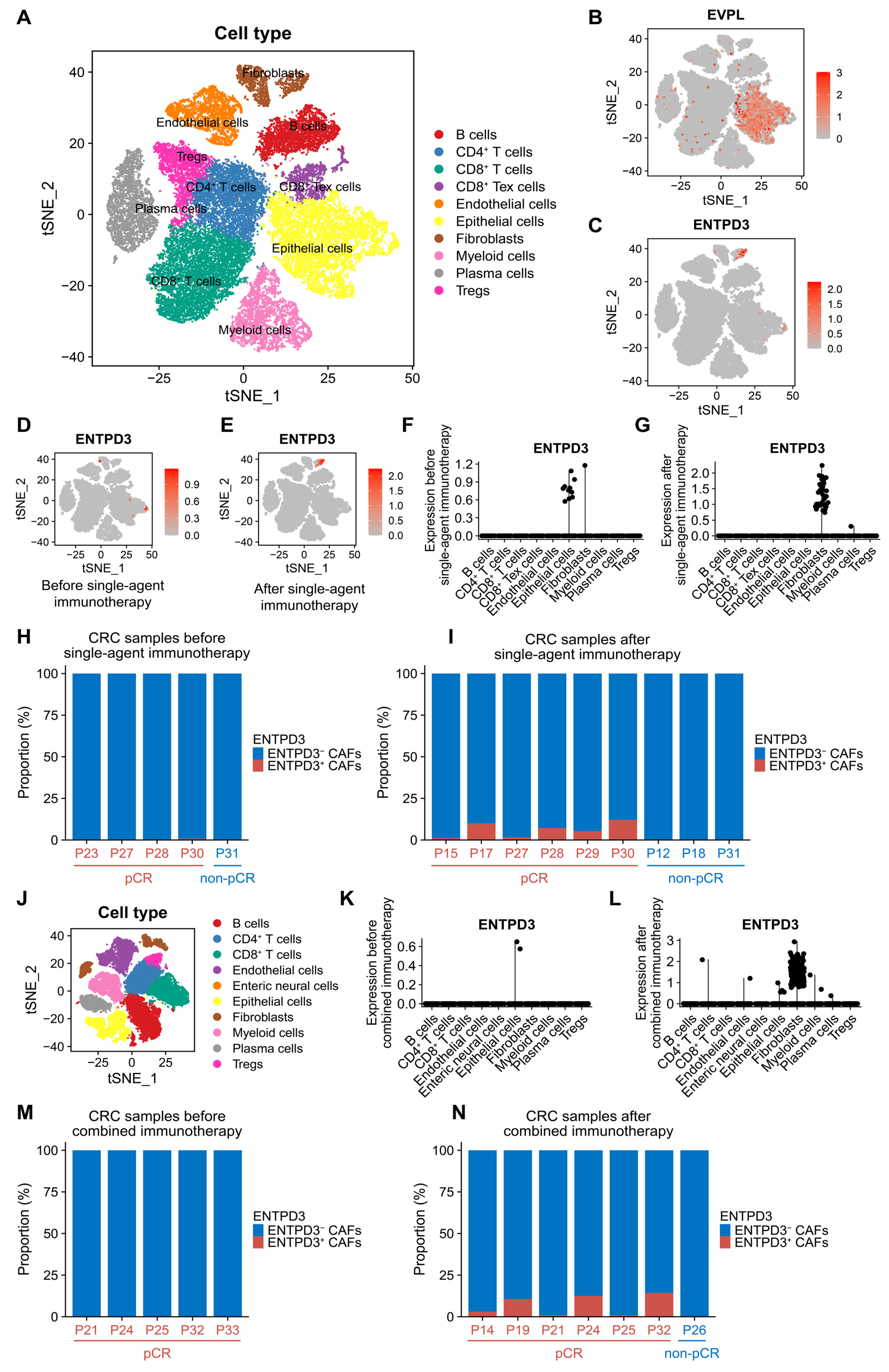
3.9. ENTPD3+ CAFs Are Extraordinarily Sensitive and Specific in Response to Real Effective Immunotherapy
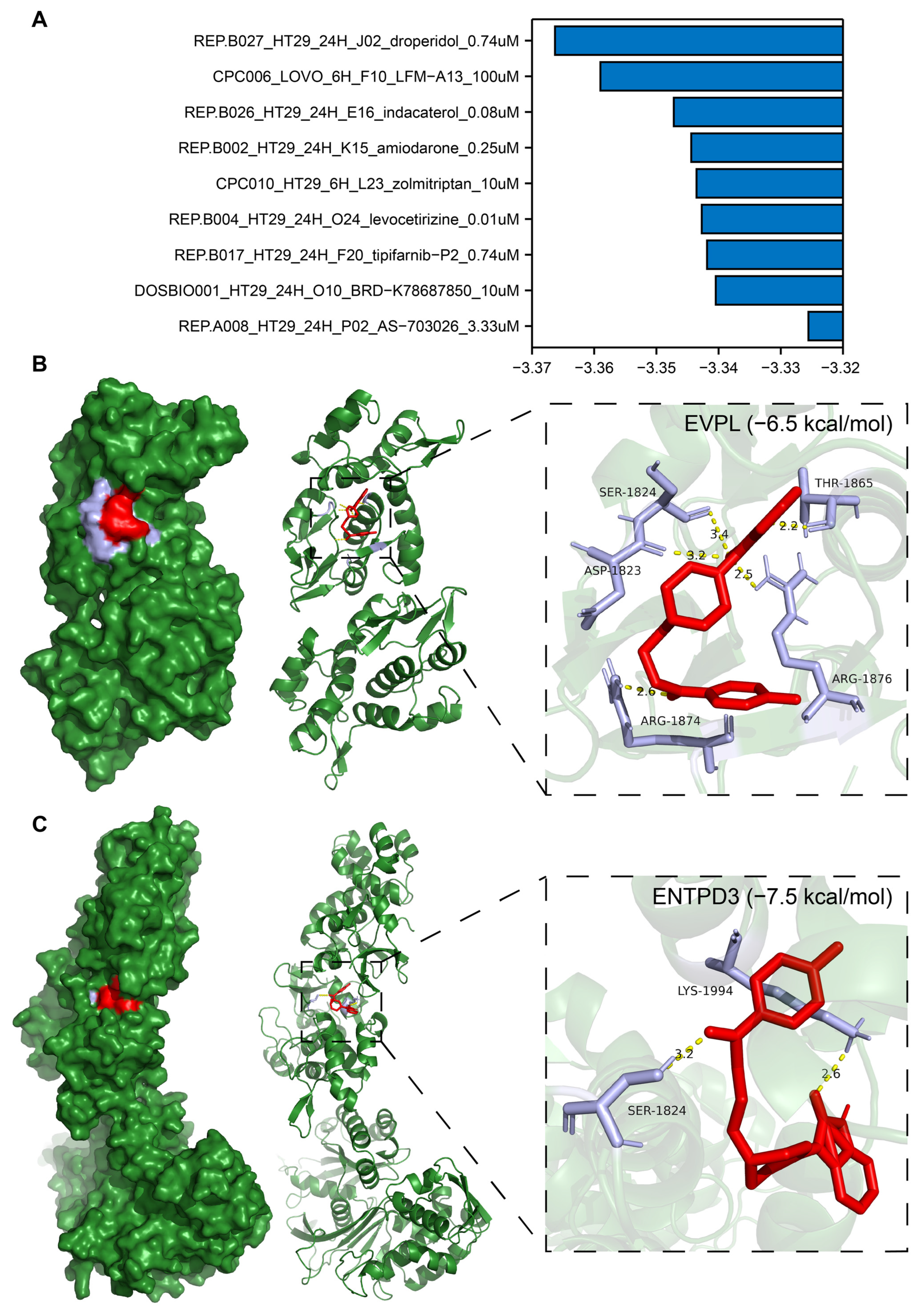
3.10. Identification of Potential Drugs Targeting EVPL and ENTPD3
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Grady, W.M.; Carethers, J.M. Genomic and epigenetic instability in colorectal cancer pathogenesis. Gastroenterology 2008, 135, 1079–1099. [Google Scholar] [CrossRef] [PubMed]
- Unger, R.H.; Orci, L. Paracrinology of islets and the paracrinopathy of diabetes. Proc. Natl. Acad. Sci. USA 2010, 107, 16009–16012. [Google Scholar] [CrossRef]
- Sun, H.; Saeedi, P.; Karuranga, S.; Pinkepank, M.; Ogurtsova, K.; Duncan, B.B.; Stein, C.; Basit, A.; Chan, J.C.N.; Mbanya, J.C.; et al. IDF Diabetes Atlas: Global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diabetes Res. Clin. Pract. 2022, 183, 109119. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.A.; Carstensen, B.; Witte, D.; Bowker, S.L.; Lipscombe, L.; Renehan, A.G.; Consortium, D.C.R. Diabetes and cancer (1): Evaluating the temporal relationship between type 2 diabetes and cancer incidence. Diabetologia 2012, 55, 1607–1618. [Google Scholar] [CrossRef] [PubMed]
- Peeters, P.J.H.L.; Bazelier, M.T.; Leufkens, H.G.M.; de Vries, F.; De Bruin, M.L. The risk of colorectal cancer in patients with type 2 diabetes: Associations with treatment stage and obesity. Diabetes Care 2015, 38, 495–502. [Google Scholar] [CrossRef] [PubMed]
- Renehan, A.G.; Yeh, H.C.; Johnson, J.A.; Wild, S.H.; Gale, E.A.M.; Moller, H.; Consortium, D.C.R. Diabetes and cancer (2): Evaluating the impact of diabetes on mortality in patients with cancer. Diabetologia 2012, 55, 1619–1632. [Google Scholar] [CrossRef]
- Dehal, A.N.; Newton, C.C.; Jacobs, E.J. Impact of diabetes mellitus and insulin use on survival after colorectal cancer diagnosis: The Cancer Prevention Study-II Nutrition Cohort. J. Clin. Oncol. 2012, 30, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Ling, S.P.; Zaccardi, F.; Issa, E.; Davies, M.J.; Khunti, K.; Brown, K. Inequalities in cancer mortality trends in people with type 2 diabetes: 20 year population-based study in England. Diabetologia 2023, 66, 657–673. [Google Scholar] [CrossRef] [PubMed]
- Rojas, A.; González, I.; Morales, E.; Pérez-Castro, R.; Romero, J.; Figueroa, H. Diabetes and cancer: Looking at the multiligand/RAGE axis. World J. Diabetes 2011, 2, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Giovannucci, E. Insulin, insulin-like growth factors and colon cancer: A review of the evidence. J. Nutr. 2001, 131, 3109S–3120S. [Google Scholar] [CrossRef]
- Othman, E.M.; Leyh, A.; Stopper, H. Insulin mediated DNA damage in mammalian colon cells and human lymphocytes in vitro. Mutat Res. 2013, 745–746, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.S.; Liu, X.W.; Nagel, J.M.; Chamberland, J.P.; Diakopoulos, K.N.; Brinkoetter, M.T.; Hatziapostolou, M.; Wu, Y.; Robson, S.C.; Iliopoulos, D.; et al. Salutary effects of adiponectin on colon cancer: In vivo and in vitro studies in mice. Gut 2013, 62, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Jia, W.; Xie, G.; Jia, W. Bile acid-microbiota crosstalk in gastrointestinal inflammation and carcinogenesis. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 111–128. [Google Scholar] [CrossRef] [PubMed]
- Piatkiewicz, P.; Milek, T.; Bernat-Karpinska, M.; Ohams, M.; Czech, A.; Ciostek, P. The dysfunction of NK cells in patients with type 2 diabetes and colon cancer. Arch. Immunol. Ther. Exp. 2013, 61, 245–253. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.; Yuan, C.; Morze, J.; Fu, R.; Wang, K.; Wang, L.; Sun, F.; Ji, J.S.; Giovannucci, E.L.; Song, M. New onset of type 2 diabetes after colorectal cancer diagnosis: Results from three prospective US cohort studies, systematic review, and meta-analysis. EBioMedicine 2022, 86, 104345. [Google Scholar] [CrossRef]
- Yang, Y.-X.; Hennessy, S.; Lewis, J.D. Insulin therapy and colorectal cancer risk among type 2 diabetes mellitus patients. Gastroenterology 2004, 127, 1044–1050. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.J.; Zheng, Z.J.; Kan, H.D.; Song, Y.Q.; Cui, W.; Zhao, G.M.; Kip, K.E. Reduced risk of colorectal cancer with metformin therapy in patients with type 2 diabetes: A meta-analysis. Diabetes Care 2011, 34, 2323–2328. [Google Scholar] [CrossRef] [PubMed]
- Colmers, I.N.; Bowker, S.L.; Johnson, J.A. Thiazolidinedione use and cancer incidence in type 2 diabetes: A systematic review and meta-analysis. Diabetes Metab. 2012, 38, 475–484. [Google Scholar] [CrossRef] [PubMed]
- Dimou, N.; Kim, A.E.; Flanagan, O.; Murphy, N.; Diez-Obrero, V.; Shcherbina, A.; Aglago, E.K.; Bouras, E.; Campbell, P.T.; Casey, G.; et al. Probing the diabetes and colorectal cancer relationship using gene–environment interaction analyses. Br. J. Cancer. 2023, 129, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.Q.; Li, D.C.; Gao, W.X.; Zhao, W.; Jin, L.J.; Chen, P.; Liu, H.; Zhao, Y.J.; Dong, G.L. Identification of the shared gene signature and biological mechanism between type 2 diabetes and colorectal cancer. Front. Genet. 2023, 14, 1202849. [Google Scholar] [CrossRef] [PubMed]
- Li, H.P.; Courtois, E.T.; Sengupta, D.; Tan, Y.L.; Chen, K.H.; Goh, J.J.L.; Kong, S.L.; Chua, C.; Hon, L.K.; Tan, W.S.; et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 2017, 49, 708–718. [Google Scholar] [CrossRef] [PubMed]
- Barrett, T.; Suzek, T.O.; Troup, D.B.; Wilhite, S.E.; Ngau, W.C.; Ledoux, P.; Rudnev, D.; Lash, A.E.; Fujibuchi, W.; Edgar, R. NCBI GEO: Mining millions of expression profiles--database and tools. Nucleic Acids Res. 2005, 33, D562–D566. [Google Scholar] [CrossRef] [PubMed]
- The Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Gautier, L.; Cope, L.; Bolstad, B.M.; Irizarry, R.A. Affy—Analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 2004, 20, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.M.; Du, P.; Huber, W.; Kibbe, W.A. Model-based variance-stabilizing transformation for Illumina microarray data. Nucleic Acids Res. 2008, 36, e11. [Google Scholar] [CrossRef] [PubMed]
- Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3, Article3. [Google Scholar] [CrossRef]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.C.; Wang, L.G.; Han, Y.Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Liberzon, A.; Subramanian, A.; Pinchback, R.; Thorvaldsdóttir, H.; Tamayo, P.; Mesirov, J.P. Molecular signatures database (MSigDB) 3.0. Bioinformatics 2011, 27, 1739–1740. [Google Scholar] [CrossRef] [PubMed]
- Barbie, D.A.; Tamayo, P.; Boehm, J.S.; Kim, S.Y.; Moody, S.E.; Dunn, I.F.; Schinzel, A.C.; Sandy, P.; Meylan, E.; Scholl, C.; et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 2009, 462, 108–112. [Google Scholar] [CrossRef] [PubMed]
- Hänzelmann, S.; Castelo, R.; Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013, 14, 7. [Google Scholar] [CrossRef] [PubMed]
- Bindea, G.; Mlecnik, B.; Tosolini, M.; Kirilovsky, A.; Waldner, M.; Obenauf, A.C.; Angell, H.; Fredriksen, T.; Lafontaine, L.; Berger, A.; et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 2013, 39, 782–795. [Google Scholar] [CrossRef] [PubMed]
- Yoshihara, K.; Shahmoradgoli, M.; Martínez, E.; Vegesna, R.; Kim, H.; Torres-Garcia, W.; Treviño, V.; Shen, H.; Laird, P.W.; Levine, D.A.; et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 2013, 4, 2612. [Google Scholar] [CrossRef] [PubMed]
- Gide, T.N.; Quek, C.; Menzies, A.M.; Tasker, A.T.; Shang, P.; Holst, J.; Madore, J.; Lim, S.Y.; Velickovic, R.; Wongchenko, M.; et al. Distinct Immune Cell Populations Define Response to Anti-PD-1 Monotherapy and Anti-PD-1/Anti-CTLA-4 Combined Therapy. Cancer Cell. 2019, 35, 238. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.T.; Cristescu, R.; Bass, A.J.; Kim, K.M.; Odegaard, J.I.; Kim, K.; Liu, X.Q.; Sher, X.W.; Jung, H.; Lee, M.; et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat. Med. 2018, 24, 1449. [Google Scholar] [CrossRef] [PubMed]
- Mariathasan, S.; Turley, S.J.; Nickles, D.; Castiglioni, A.; Yuen, K.; Wang, Y.L.; Kadel, E.E.; Koeppen, H.; Astarita, J.L.; Cubas, R.; et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 2018, 554, 544–548. [Google Scholar] [CrossRef] [PubMed]
- Braun, D.A.; Hou, Y.; Bakouny, Z.; Ficial, M.; Angelo, M.S.; Forman, J.; Ross-Macdonald, P.; Berger, A.C.; Jegede, O.A.; Elagina, L.; et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat. Med. 2020, 26, 909. [Google Scholar] [CrossRef]
- Hao, Y.H.; Hao, S.; Andersen-Nissen, E.; Mauck, W.M.; Zheng, S.W.; Butler, A.; Lee, M.J.; Wilk, A.J.; Darby, C.; Zager, M.; et al. Integrated analysis of multimodal single-cell data. Cell 2021, 184, 3573–3587. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, P.G.; Muñoz-Aguirre, M.; Reverter, F.; Godinho, C.P.S.; Sousa, A.; Amadoz, A.; Sodaei, R.; Hidalgo, M.R.; Pervouchine, D.; Carbonell-Caballero, J.; et al. The effects of death and post-mortem cold ischemia on human tissue transcriptomes. Nat. Commun. 2018, 9, 490. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Cacchiarelli, D.; Grimsby, J.; Pokharel, P.; Li, S.Q.; Morse, M.; Lennon, N.J.; Livak, K.J.; Mikkelsen, T.S.; Rinn, J.L. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 2014, 32, 381–386. [Google Scholar] [CrossRef]
- Lamb, J.; Crawford, E.D.; Peck, D.; Modell, J.W.; Blat, I.C.; Wrobel, M.J.; Lerner, J.; Brunet, J.P.; Subramanian, A.; Ross, K.N.; et al. The Connectivity Map: Using gene-expression signatures to connect small molecules, genes, and disease. Science 2006, 313, 1929–1935. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Narayan, R.; Corsello, S.M.; Peck, D.D.; Natoli, T.E.; Lu, X.D.; Gould, J.; Davis, J.F.; Tubelli, A.A.; Asiedu, J.K.; et al. A Next Generation Connectivity Map: L1000 Platform and the First 1,000,000 Profiles. Cell 2017, 171, 1437–1452.e17. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.L.; Xiao, J.W.; Suzek, T.O.; Zhang, J.; Wang, J.Y.; Bryant, S.H. PubChem: A public information system for analyzing bioactivities of small molecules. Nucleic Acids Res. 2009, 37, W623–W633. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Gaillard, T. Evaluation of AutoDock and AutoDock Vina on the CASF-2013 Benchmark. J. Chem. Inf. Model 2018, 58, 1697–1706. [Google Scholar] [CrossRef] [PubMed]
- Robin, X.; Turck, N.; Hainard, A.; Tiberti, N.; Lisacek, F.; Sanchez, J.C.; Müller, M. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 2011, 12, 77. [Google Scholar] [CrossRef] [PubMed]
- Dinse, G.E.; Lagakos, S.W. Nonparametric estimation of lifetime and disease onset distributions from incomplete observations. Biometrics 1982, 38, 921–932. [Google Scholar] [CrossRef] [PubMed]
- Wareham, N.J.; Pfister, R. Diabetes: Glycated hemoglobin is a marker of diabetes and CVD risk. Nat. Rev. Cardiol. 2010, 7, 367–368. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Song, E. Turning foes to friends: Targeting cancer-associated fibroblasts. Nat. Rev. Drug Discov. 2019, 18, 99–115. [Google Scholar] [CrossRef] [PubMed]
- Biffi, G.; Tuveson, D.A. Diversity and Biology of Cancer-Associated Fibroblasts. Physiol. Rev. 2021, 101, 147–176. [Google Scholar] [CrossRef] [PubMed]
- Marble, A. Diabetes and Cancer. N. Engl. J. Med. 1934, 211, 339–349. [Google Scholar] [CrossRef]
- Ruhrberg, C.; Williamson, J.A.; Sheer, D.; Watt, F.M. Chromosomal Localisation of the Human Envoplakin Gene (EVPL) to the Region of the Tylosis Oesophageal Cancer Gene (TOCG) on 17q25. Genomics 1996, 37, 381–385. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Chen, T.C.; Zhao, J.Y.; Peng, Y.; Chen, X.X.; Tu, P.; Zhu, X.J.; Liu, Z.; Wang, M.Y. Extremities of the N-terminus of envoplakin and C-terminus of its linker subdomain are major epitopes of paraneoplastic pemphigus. J. Dermatol. Sci. 2016, 84, 24–29. [Google Scholar] [CrossRef] [PubMed]
- Iwaya, T.; Maesawa, C.; Kimura, T.; Ogasawara, S.; Ikeda, K.; Kimura, Y.; Noda, Y.; Ishida, K.; Sato, N.; Saito, K.; et al. Infrequent mutation of the human envoplakin gene is closely linked to the tylosis oesophageal cancer locus in sporadic oesophageal squamous cell carcinomas. Oncol. Rep. 2005, 13, 703–707. [Google Scholar] [CrossRef] [PubMed]
- Eltzschig, H.K.; Sitkovsky, M.V.; Robson, S.C. Purinergic signaling during inflammation. N. Engl. J. Med. 2012, 367, 2322–2333. [Google Scholar] [CrossRef] [PubMed]
- Robson, S.C.; Sévigny, J.; Zimmermann, H. The E-NTPDase family of ectonucleotidases: Structure function relationships and pathophysiological significance. Purinergic Signal. 2006, 2, 409–430. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, H.; Zebisch, M.; Sträter, N. Cellular function and molecular structure of ecto-nucleotidases. Purinergic Signal. 2012, 8, 437–502. [Google Scholar] [CrossRef] [PubMed]
- Belcher, S.M.; Zsarnovszky, A.; Crawford, P.A.; Hemani, H.; Spurling, L.; Kirley, T.L. Immunolocalization of ecto-nucleoside triphosphate diphosphohydrolase 3 in rat brain: Implications for modulation of multiple homeostatic systems including feeding and sleep–wake behaviors. Neuroscience 2006, 137, 1331–1346. [Google Scholar] [CrossRef] [PubMed]
- Lavoie, E.G.; Fausther, M.; Kauffenstein, G.; Kukulski, F.; Künzli, B.M.; Friess, H.; Sévigny, J. Identification of the ectonucleotidases expressed in mouse, rat, and human Langerhans islets: Potential role of NTPDase3 in insulin secretion. Am. J. Physiol. Endocrinol. Metab. 2010, 299, E647–E656. [Google Scholar] [CrossRef] [PubMed]
- Lavoie, E.G.; Gulbransen, B.D.; Martín-Satué, M.; Aliagas, E.; Sharkey, K.A.; Sévigny, J. Ectonucleotidases in the digestive system: Focus on NTPDase3 localization. Am. J. Physiol. Gastrointest. Liver Physiol. 2011, 300, G608–G620. [Google Scholar] [CrossRef] [PubMed]
- Zhong, A.H.; Gordon Jiang, Z.; Cummings, R.D.; Robson, S.C. Various N-glycoforms differentially upregulate E-NTPDase activity of the NTPDase3/CD39L3 ecto-enzymatic domain. Purinergic. Signal. 2017, 13, 601–609. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Esparcia, P.; Hernández-Ortega, K.; Ansoleaga, B.; Carmona, M.; Ferrer, I. Purine metabolism gene deregulation in Parkinson’s disease. Neuropathol. Appl. Neurobiol. 2015, 41, 926–940. [Google Scholar] [CrossRef]
- Ansoleaga, B.; Jové, M.; Schlüter, A.; Garcia-Esparcia, P.; Moreno, J.; Pujol, A.; Pamplona, R.; Portero-Otín, M.; Ferrer, I. Deregulation of purine metabolism in Alzheimer’s disease. Neurobiol. Aging. 2015, 36, 68–80. [Google Scholar] [CrossRef] [PubMed]
- Feldbrügge, L.; Moss, A.C.; Yee, E.U.; Csizmadia, E.; Mitsuhashi, S.; Longhi, M.S.; Sandhu, B.; Stephan, H.; Wu, Y.; Cheifetz, A.S.; et al. Expression of Ecto-nucleoside Triphosphate Diphosphohydrolases-2 and -3 in the Enteric Nervous System Affects Inflammation in Experimental Colitis and Crohn’s Disease. J. Crohns Colitis 2017, 11, 1113–1123. [Google Scholar] [CrossRef] [PubMed]
- Saunders, D.C.; Brissova, M.; Phillips, N.; Shrestha, S.; Walker, J.T.; Aramandla, R.; Poffenberger, G.; Flaherty, D.K.; Weller, K.P.; Pelletier, J.; et al. Ectonucleoside Triphosphate Diphosphohydrolase-3 Antibody Targets Adult Human Pancreatic β Cells for In Vitro and In Vivo Analysis. Cell Metab. 2019, 29, 745–754. [Google Scholar] [CrossRef] [PubMed]
- Docherty, F.M.; Riemondy, K.A.; Castro-Gutierrez, R.; Dwulet, J.M.; Shilleh, A.H.; Hansen, M.S.; Williams, S.P.M.; Armitage, L.H.; Santostefano, K.E.; Wallet, M.A.; et al. ENTPD3 Marks Mature Stem Cell-Derived β-Cells Formed by Self-Aggregation In Vitro. Diabetes 2021, 70, 2554–2567. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Qi, Y.; Chen, M.; Wang, Z.; Zeng, D.; Xiao, Y.; Li, S.; Lin, H.; Wei, X.; Zhang, G. GATA Binding Protein 3 Boosts Extracellular ATP Hydrolysis and Inhibits Metastasis of Breast Cancer by Up-regulating Ectonucleoside Triphosphate Diphosphohydrolase 3. Int. J. Biol. Sci. 2019, 15, 2522–2537. [Google Scholar] [CrossRef] [PubMed]
- Enjyoji, K.; Kotani, K.; Thukral, C.; Blumel, B.; Sun, X.F.; Wu, Y.; Imai, M.; Friedman, D.; Csizmadia, E.; Bleibel, W.; et al. Deletion of cd39/entpd1 results in hepatic insulin resistance. Diabetes 2008, 57, 2311–2320. [Google Scholar] [CrossRef] [PubMed]
- Brooks-Worrell, B.; Narla, R.; Palmer, J.P. Biomarkers and immune-modulating therapies for Type 2 diabetes. Trends Immunol. 2012, 33, 546–553. [Google Scholar] [CrossRef] [PubMed]
- SantaCruz-Calvo, S.; Bharath, L.; Pugh, G.; SantaCruz-Calvo, L.; Lenin, R.R.; Lutshumba, J.; Liu, R.; Bachstetter, A.D.; Zhu, B.; Nikolajczyk, B.S. Adaptive immune cells shape obesity-associated type 2 diabetes mellitus and less prominent comorbidities. Nat. Rev. Endocrinol. 2022, 18, 23–42. [Google Scholar] [CrossRef] [PubMed]
- Catenacci, D.V.T.; Junttila, M.R.; Karrison, T.; Bahary, N.; Horiba, M.N.; Nattam, S.R.; Marsh, R.; Wallace, J.; Kozloff, M.; Rajdev, L.; et al. Randomized Phase Ib/II Study of Gemcitabine Plus Placebo or Vismodegib, a Hedgehog Pathway Inhibitor, in Patients With Metastatic Pancreatic Cancer. J. Clin. Oncol. 2015, 33, 4284–4292. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, R.K.; McDonough, S.; Philip, P.A.; Hingorani, S.R.; Lacy, J.; Kortmansky, J.S.; Thumar, J.; Chiorean, E.G.; Shields, A.F.; Behl, D.; et al. Phase IB/II Randomized Study of FOLFIRINOX Plus Pegylated Recombinant Human Hyaluronidase Versus FOLFIRINOX Alone in Patients With Metastatic Pancreatic Adenocarcinoma: SWOG S1313. J Clin Oncol. 2019, 37, 1062–1069. [Google Scholar] [CrossRef] [PubMed]
- Overman, M.J.; McDermott, R.; Leach, J.L.; Lonardi, S.; Lenz, H.-J.; Morse, M.A.; Desai, J.; Hill, A.; Axelson, M.; Moss, R.A.; et al. Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142): An open-label, multicentre, phase 2 study. Lancet Oncol. 2017, 18, 1182–1191. [Google Scholar] [CrossRef] [PubMed]
- Apfel, C.C.; Korttila, K.; Abdalla, M.; Kerger, H.; Turan, A.; Vedder, I.; Zernak, C.; Danner, K.; Jokela, R.; Pocock, S.J.; et al. A factorial trial of six interventions for the prevention of postoperative nausea and vomiting. N. Engl. J. Med. 2004, 350, 2441–2451. [Google Scholar] [CrossRef] [PubMed]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, Y.; Yang, L.; Wu, H.; Xu, H.; Peng, J.; Wang, Y.; Zhou, F. Exploring the Molecular Mechanism of Comorbidity of Type 2 Diabetes Mellitus and Colorectal Cancer: Insights from Bulk Omics and Single-Cell Sequencing Validation. Biomolecules 2024, 14, 693. https://doi.org/10.3390/biom14060693
Luo Y, Yang L, Wu H, Xu H, Peng J, Wang Y, Zhou F. Exploring the Molecular Mechanism of Comorbidity of Type 2 Diabetes Mellitus and Colorectal Cancer: Insights from Bulk Omics and Single-Cell Sequencing Validation. Biomolecules. 2024; 14(6):693. https://doi.org/10.3390/biom14060693
Chicago/Turabian StyleLuo, Yongge, Lei Yang, Han Wu, Hui Xu, Jin Peng, You Wang, and Fuxiang Zhou. 2024. "Exploring the Molecular Mechanism of Comorbidity of Type 2 Diabetes Mellitus and Colorectal Cancer: Insights from Bulk Omics and Single-Cell Sequencing Validation" Biomolecules 14, no. 6: 693. https://doi.org/10.3390/biom14060693




