Heterologous Expression of Either Human or Soya Bean Ferritins in Budding Yeast Reveals Common Functions Protecting Against Oxidative Agents and Counteracting Double-Strand Break Accumulation
Abstract
1. Introduction
2. Materials and Methods
2.1. Yeast Strains and Plasmids
2.2. Media and Growth Conditions
2.3. Vacuole and Staining
2.4. Cell Survival
2.5. Protein Extraction and Immunoblot Analyses
2.6. Fluorescence Microscopy
2.7. Statistical Analysis
3. Results
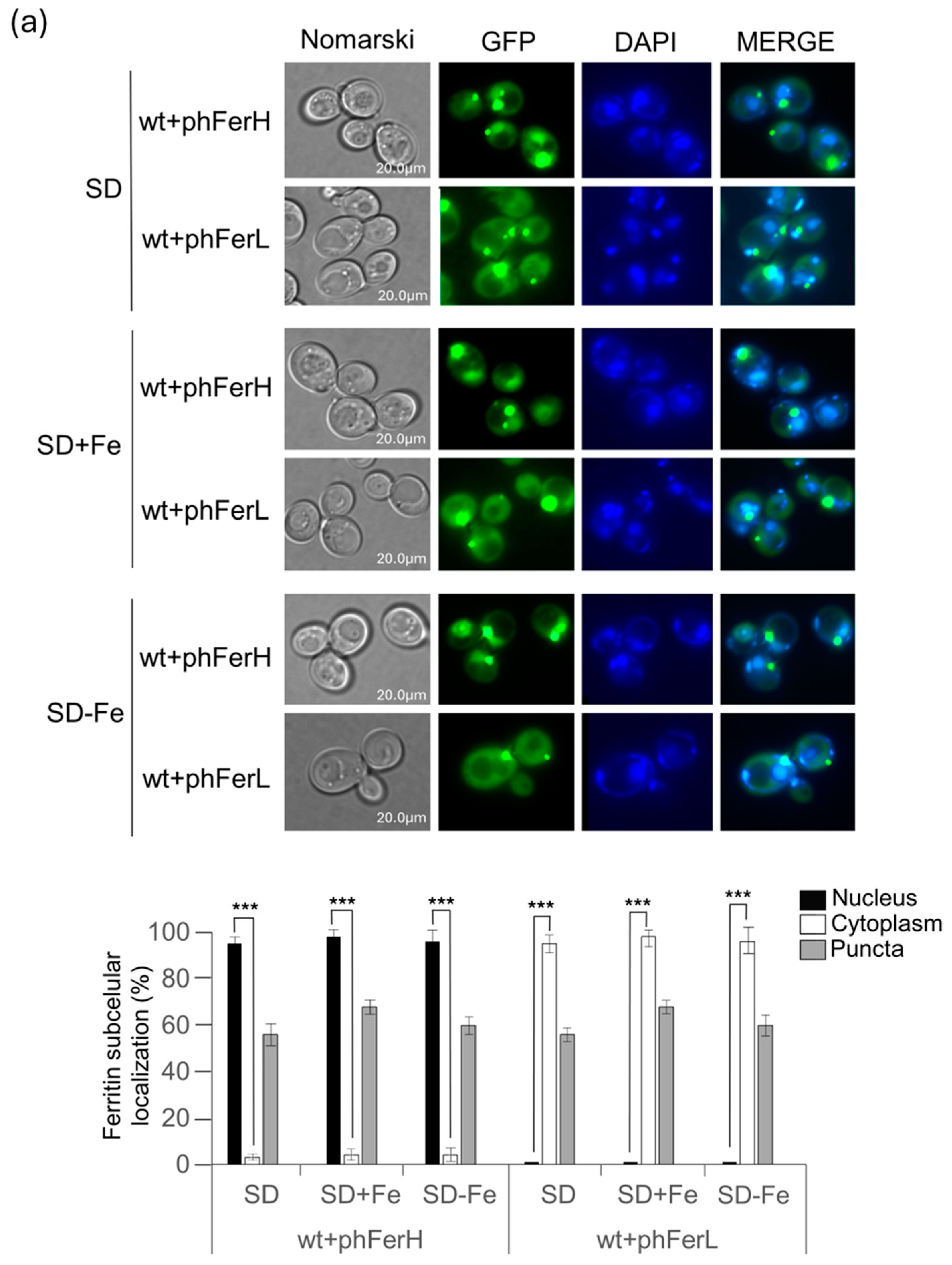
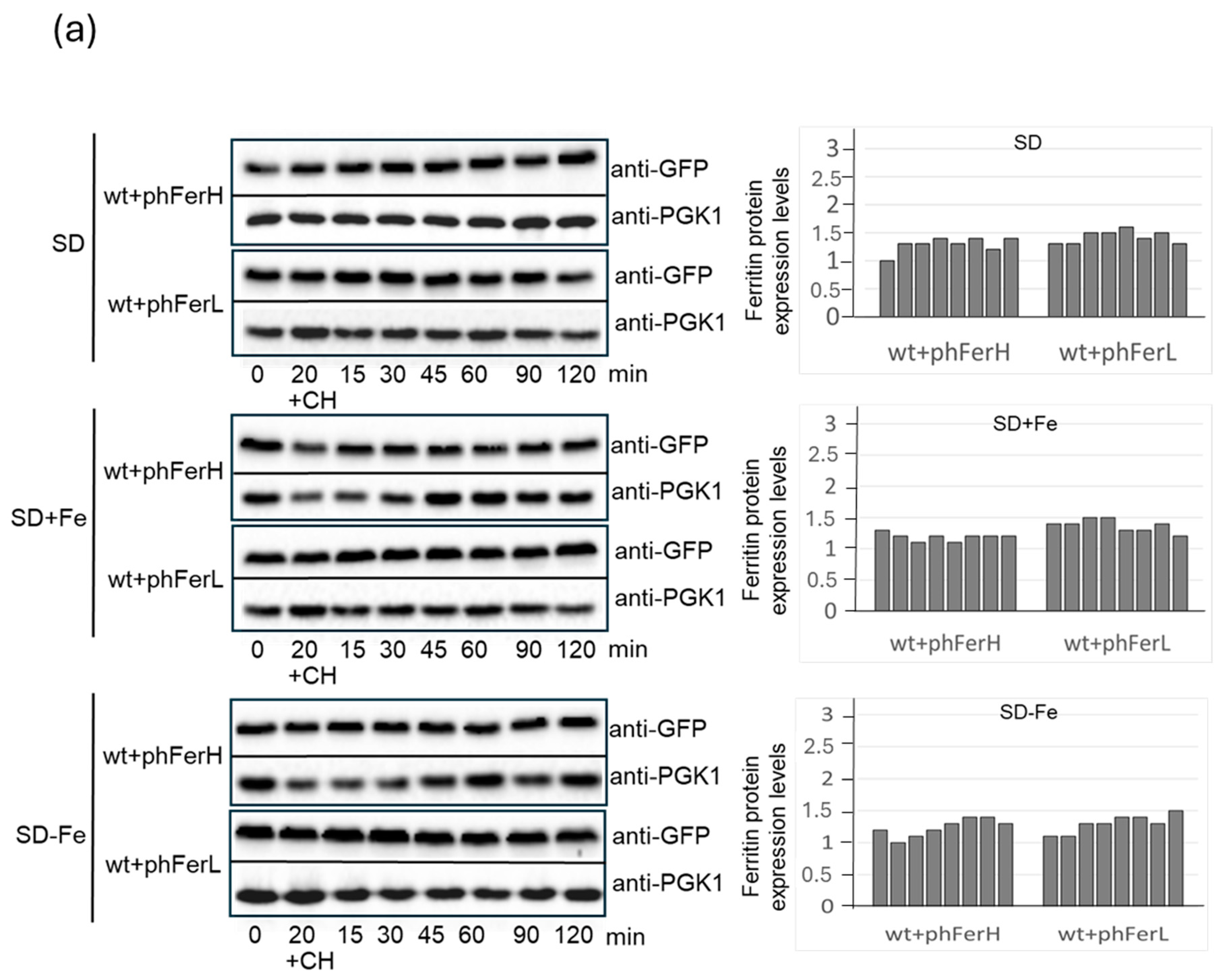
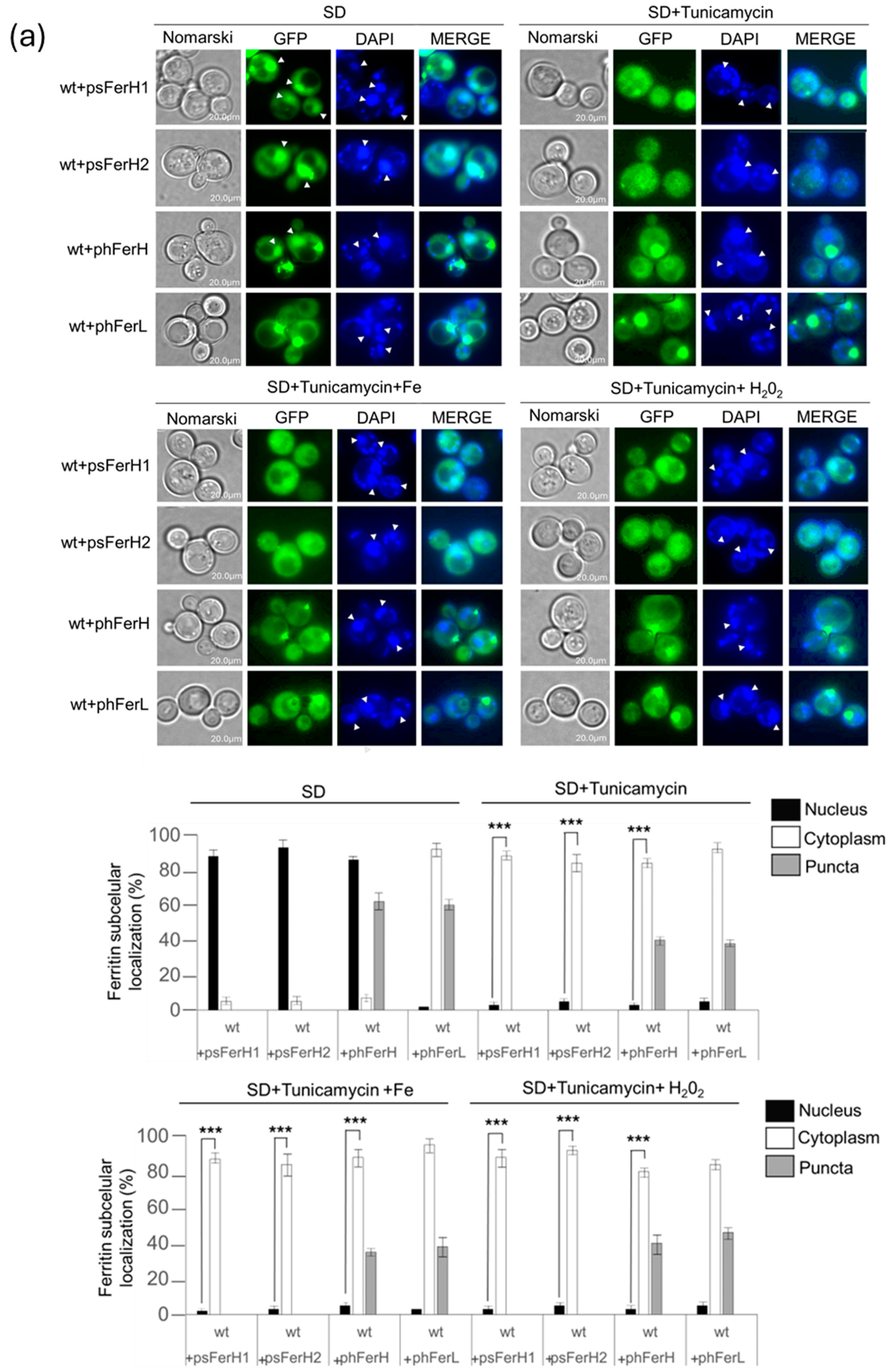
3.1. Expression of Human (H and L) and Soya Bean Ferritins (H1 and H2) in Budding Yeast Reveals New Information Concerning In Vivo Localization and Protein Stability
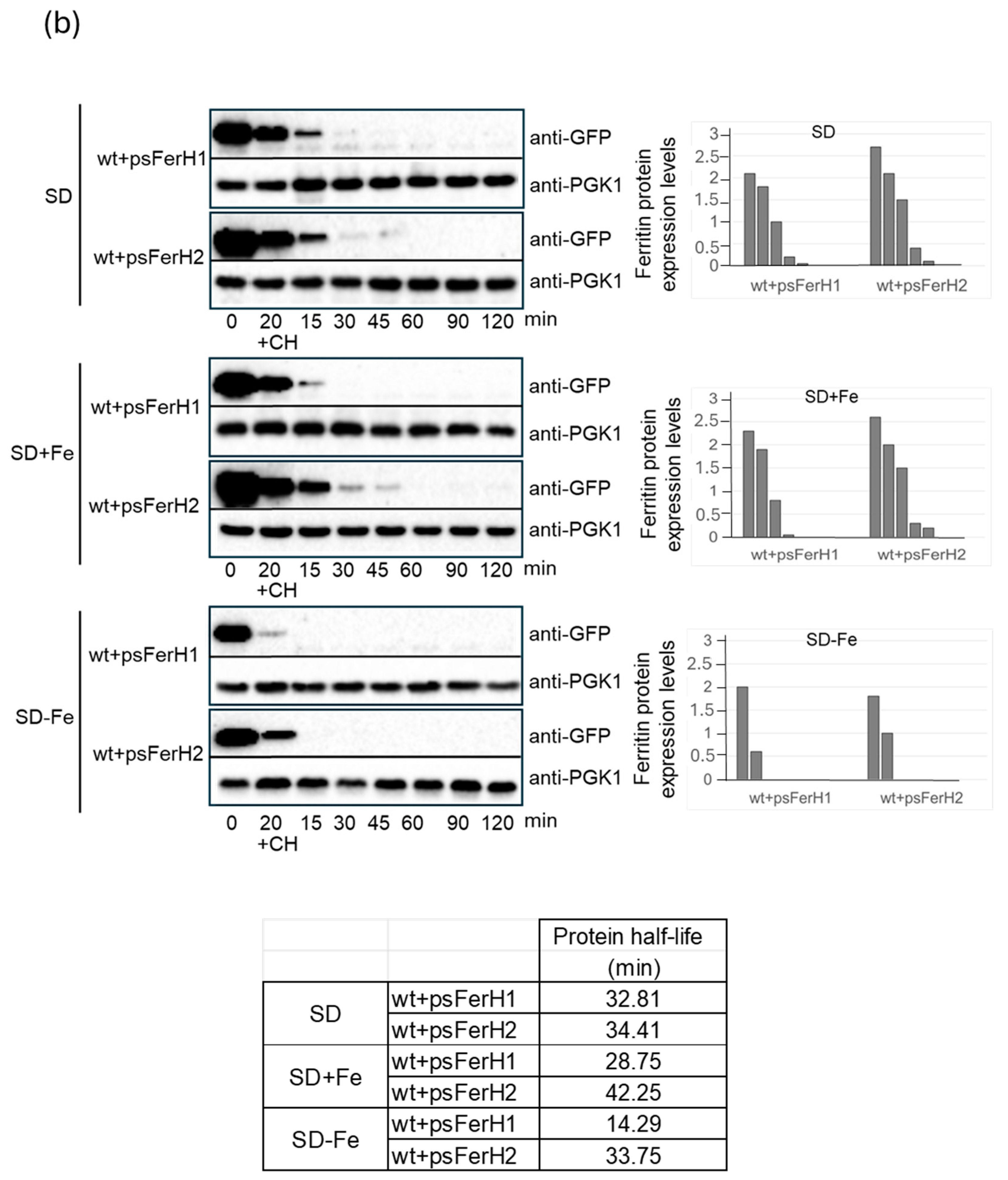
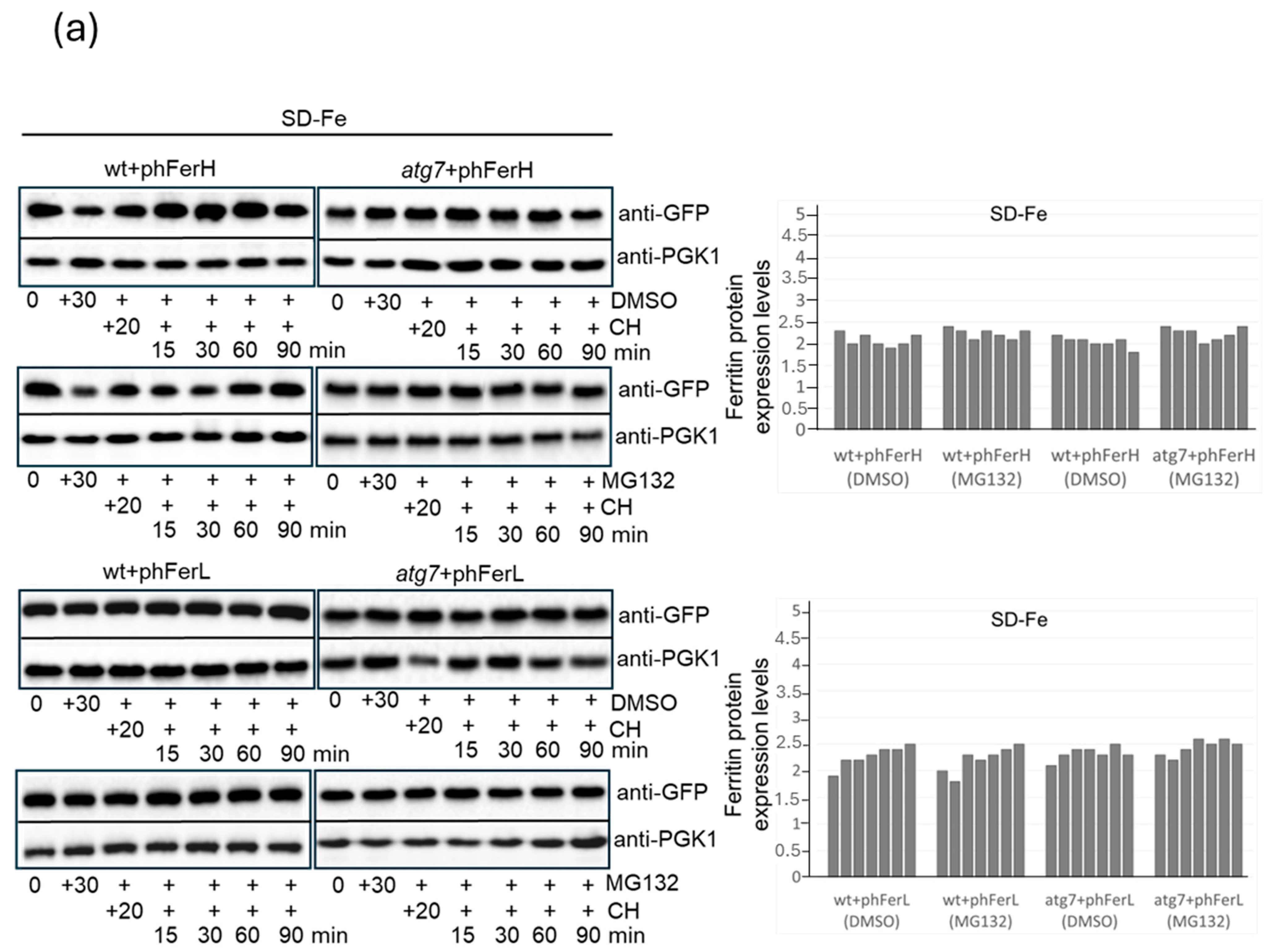
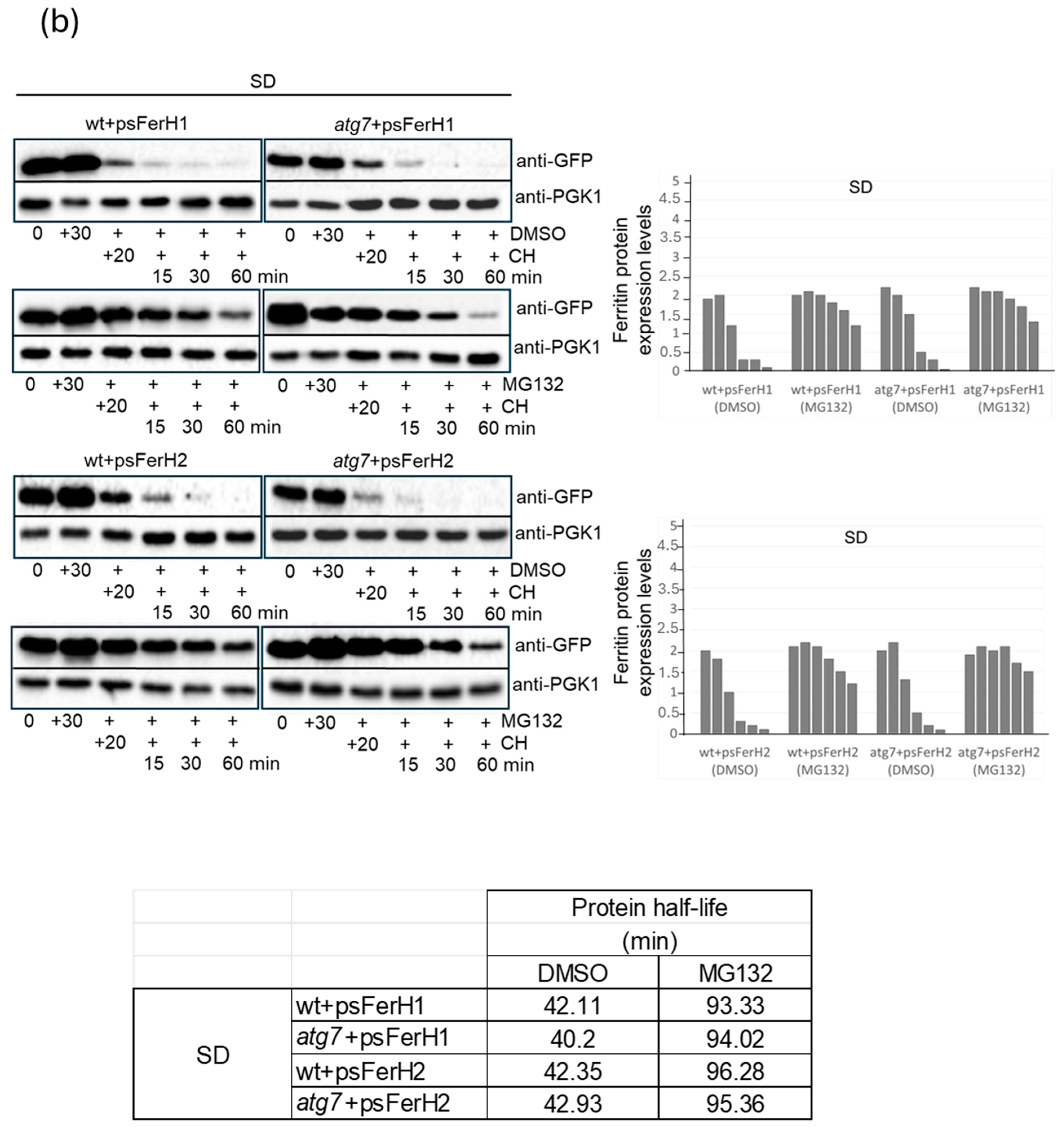
3.2. In Conditions of Iron Scarcity, Both H1 and H2 Soya Bean Ferritins Are Degraded in Proteasome to Liberate Iron, and Only H2 Ferritin Is Also Degraded Through a Mechanism Dependent on Bulk Autophagy
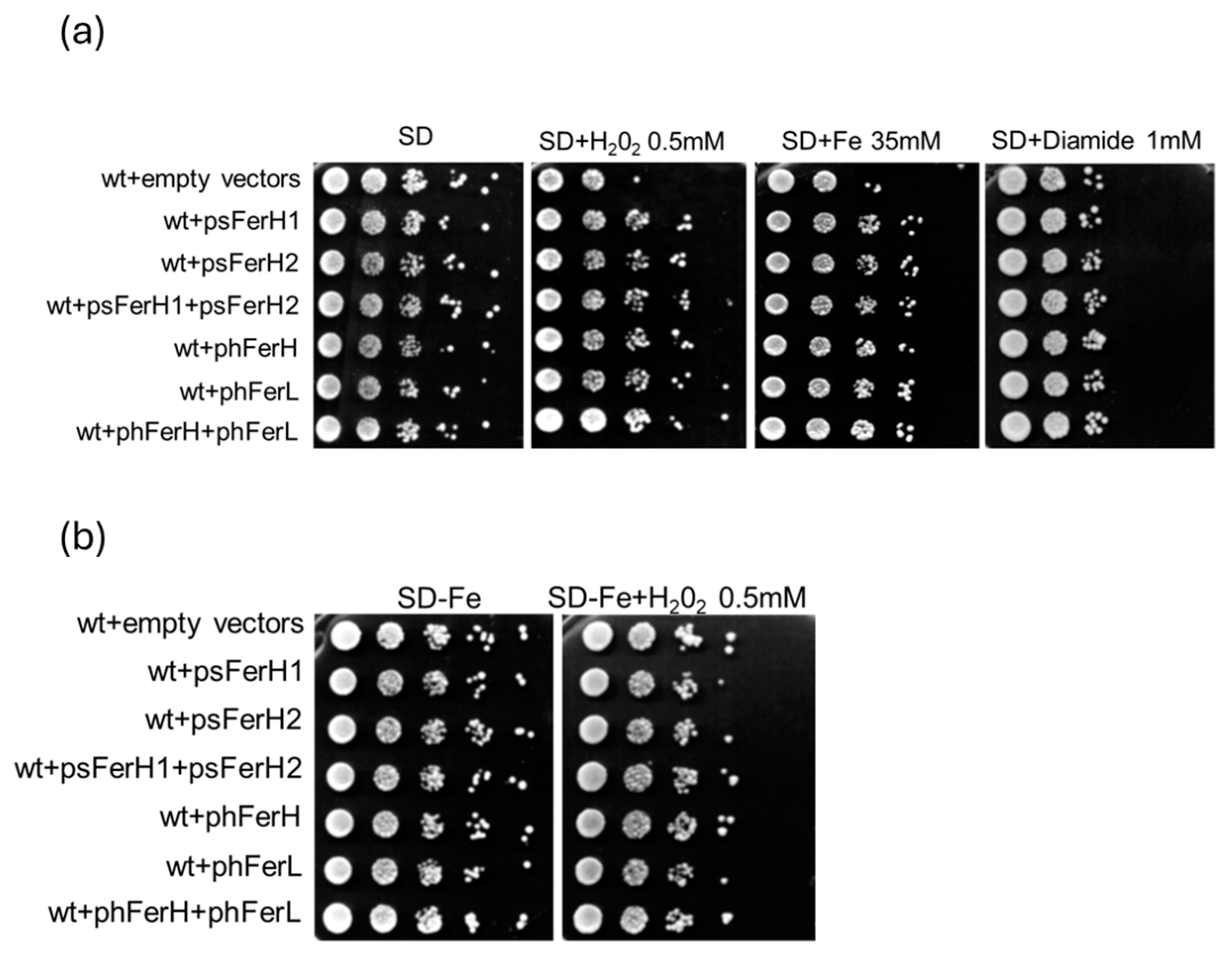
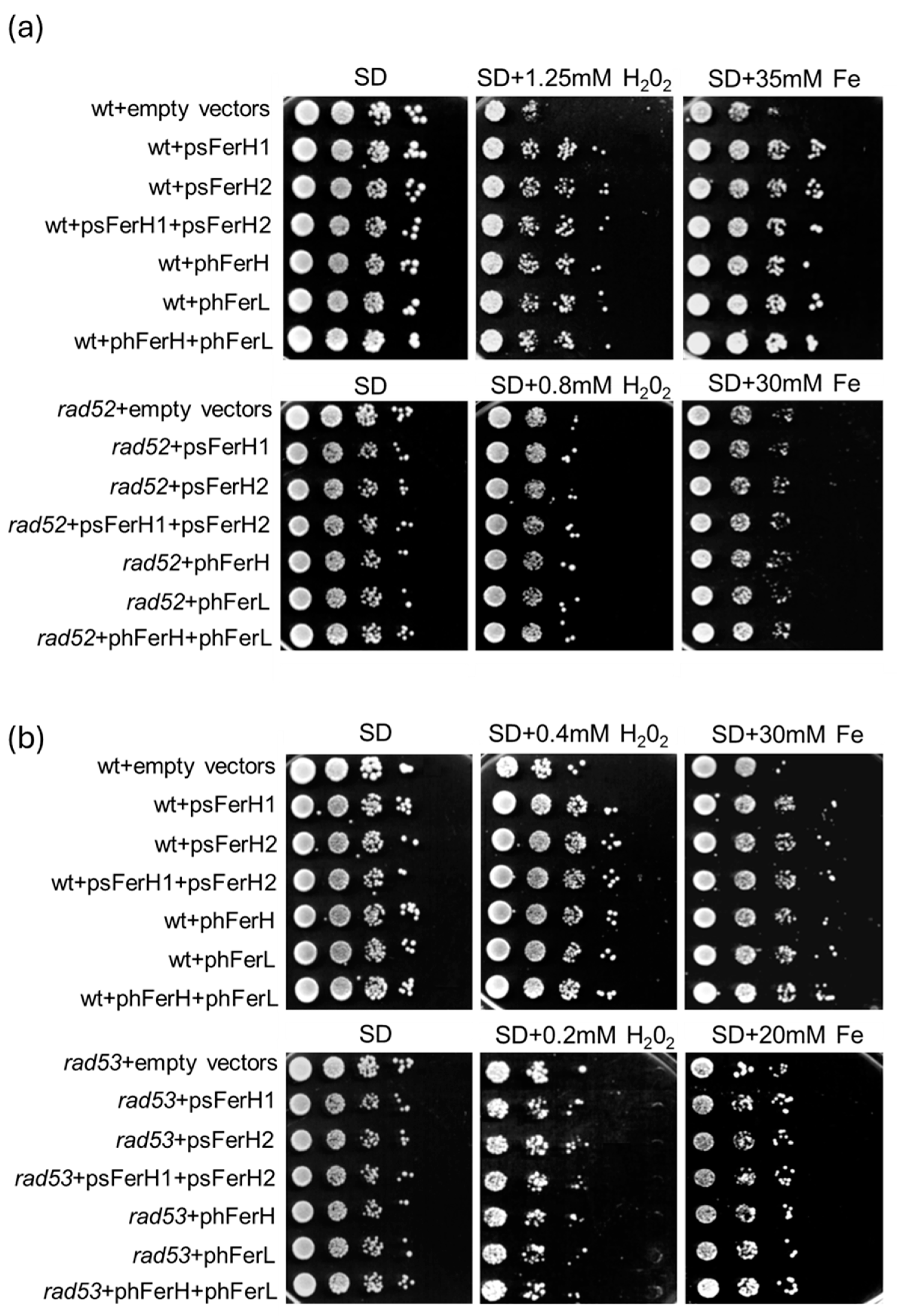
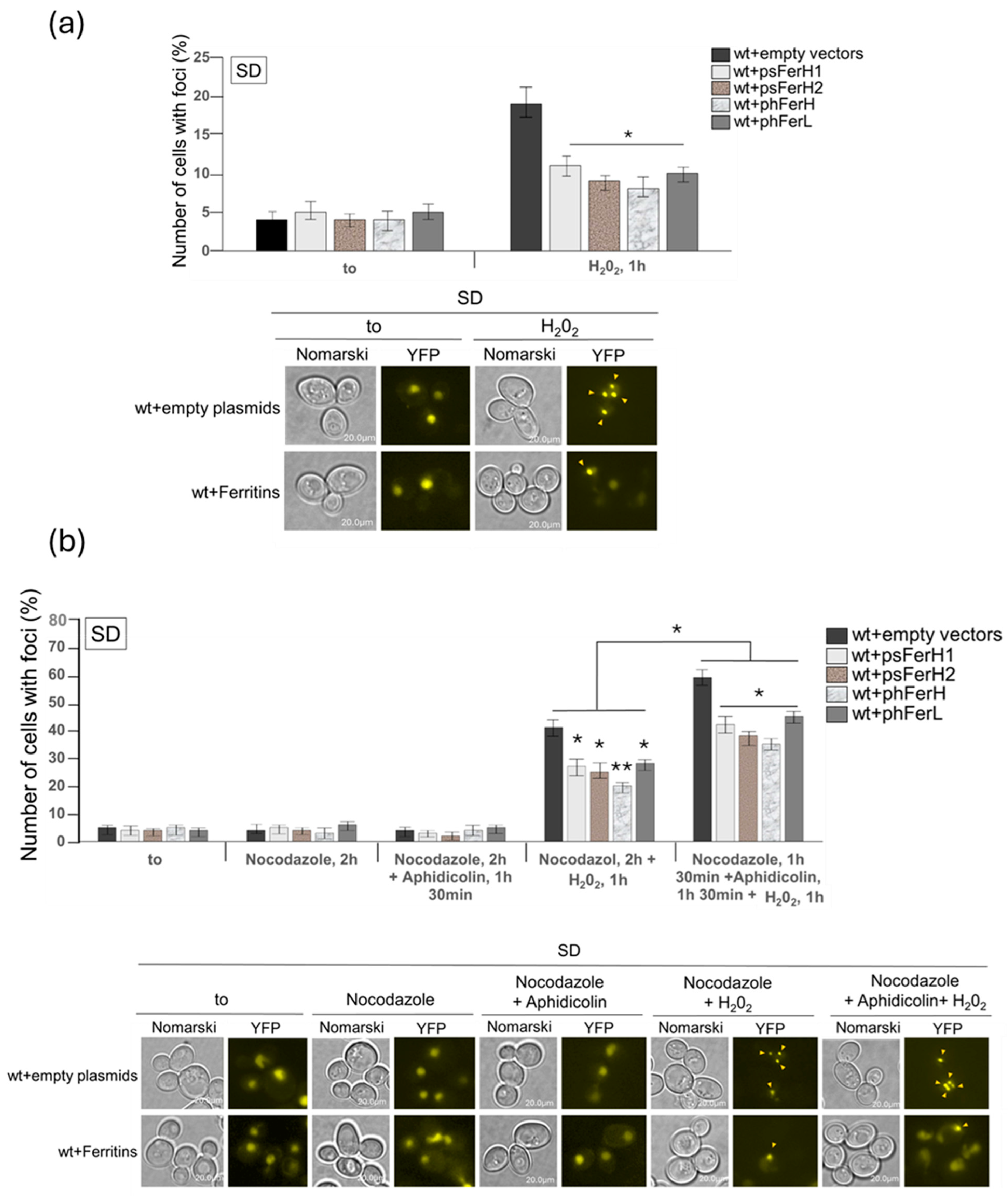
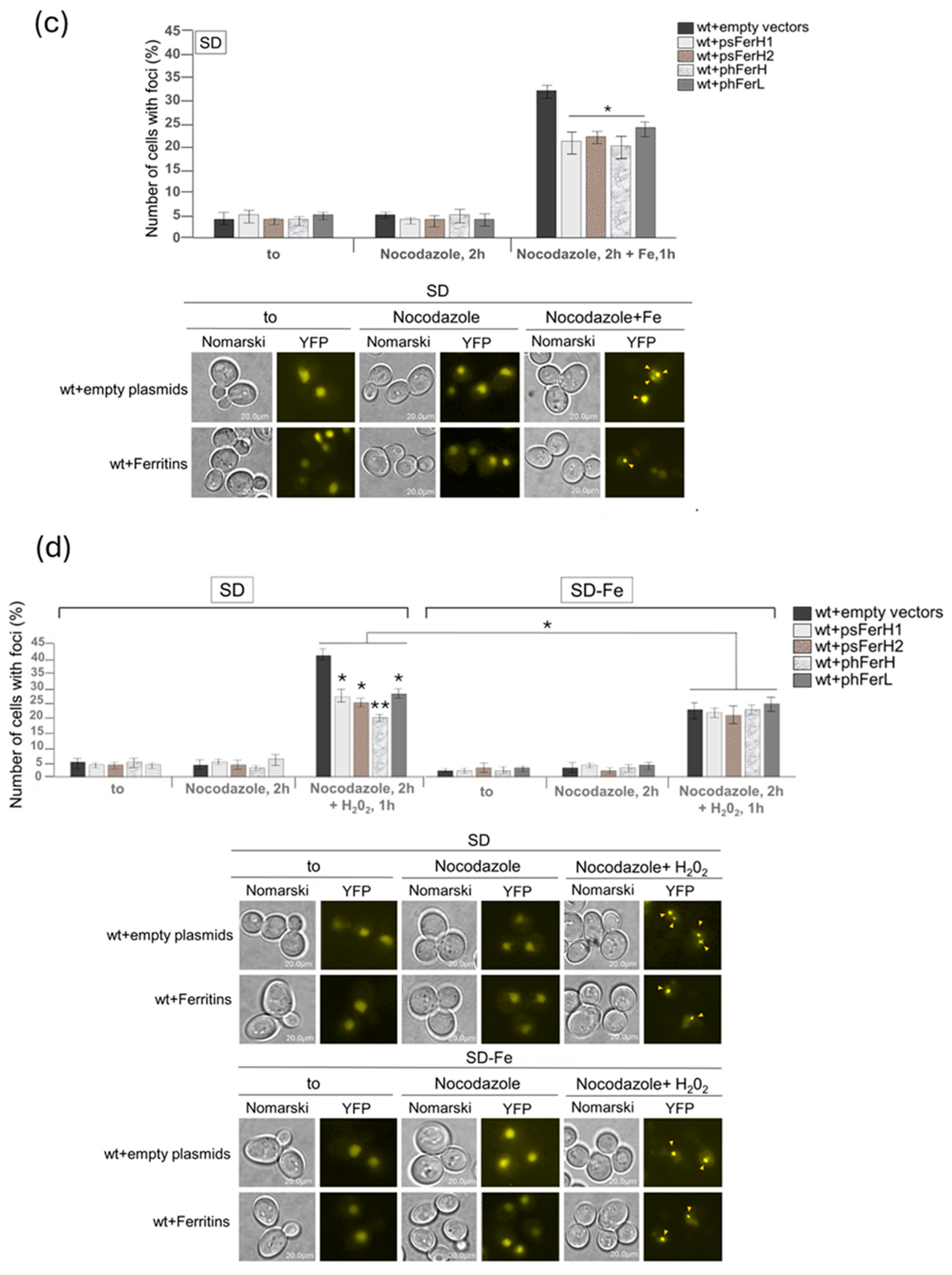
3.3. Both Human and Soya Bean Ferritins Protect Yeast Cells Against the Oxidative Effects Caused by H2O2 and Reduce the Impact of DSB in the DNA
3.4. Glycosylation Is Important in the Cellular Defense Against the Oxidative Effect Caused by ROS Dependent on Iron Carried Out by Both Human and Soya Bean Ferritins in the Budding Yeast Model
4. Discussion
5. Conclusions
- (a)
- DNA damage protection against the oxidative effects provoked by iron as a consequence of Fenton reaction. In this study we demonstrate that the four ferritin chains reduce the formation of DSB.
- (b)
- Reduction of oxidative damage and cell lethality caused by H2O2 in a manner totally dependent on iron.
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ramos, L.; Romero, A.M.; Martínez, M.T.; Puig, S. Iron Regulatory Mechanisms in Saccharomyces cerevisiae. Front. Microbiol. 2020, 9, 582830. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ward, R.J.; Zucca, F.A.; Duyn, J.H.; Crichton, R.R.; Zecca, L. The role of iron in brain ageing and neurodegenerative disorders. Lancet Neurol. 2014, 13, 1045–1060. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Angelova, D.; Iron, B.D. Aging Neurodegeneration. Metals 2015, 5, 2070–2092. [Google Scholar] [CrossRef]
- Biasiotto, G.; Di Lorenzo, D.; Archetti, S.; Zanella, I. Iron and Neurodegeneration: Is Ferritinophagy the Link? Mol. Neurobiol. 2016, 53, 5542–5574. [Google Scholar] [CrossRef] [PubMed]
- Koppenol, W.H.; Hider, R.H. Iron and redox cycling. Do’s and don’ts. Free Radic. Biol. Med. 2019, 133, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Rowe, L.A.; Degtyareva, N.; Doetsch, P.W. DNA damage-induced reactive oxygen species (ROS) stress response in Saccharomyces cerevisiae. Free Radic. Biol. Med. 2008, 45, 1167–1177. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Srinivas, U.S.; Tan, B.W.Q.; Vellayappan, B.A.; Jeyasekharan, A.D. ROS and the DNA damage response in cancer. Redox Biol. 2019, 25, 101084. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zimmermann, M.B.; Hurrell, R.F. Nutritional iron deficiency. Lancet 2007, 370, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Arosio, P.; Ingrassia, R.; Cavadini, P. Ferritins: A family of molecules for iron storage, antioxidation and more. Biochim. Biophys. Acta 2009, 1790, 589–599. [Google Scholar] [CrossRef] [PubMed]
- Arosio, P.; Levi, S. Ferritin, iron homeostasis, and oxidative damage. Free Radic. Biol. Med. 2002, 33, 457–463. [Google Scholar] [CrossRef] [PubMed]
- Arosio, P.; Elia, L.; Poli, M. Ferritin, cellular iron storage and regulation. IUBMB Life 2017, 69, 414–422. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Theil, E.C. Ferritins: Dynamic management of biological iron and oxygen chemistry. Acc. Chem. Res. 2005, 38, 167–175. [Google Scholar] [CrossRef] [PubMed]
- Waldo, G.S.; Wright, E.; Whang, Z.H.; Briat, J.F.; Theil, E.C.; Sayers, D.E. Formation of the ferritin iron mineral occurs in plastids. Plant Physiol. 1995, 109, 797–802. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Laulhere, J.P.; Laboure, A.M.; Briat, J.F. Mechanism of the transition from plant ferritin to phytosiderin. J. Biol. Chem. 1989, 264, 3629–3635. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Hu, X.; Zhao, G. Two different H-type subunits from pea seed (Pisum sativum) ferritin that are responsible for fast Fe (II) oxidation. Biochimie 2009, 91, 230–239. [Google Scholar] [CrossRef] [PubMed]
- Lönnerdal, B. Soybean ferritin: Implications for iron status of vegetarians. Am. J. Clin. Nutr. 2009, 89, 1680S–1685S. [Google Scholar] [CrossRef] [PubMed]
- Lv, C.; Zhao, G.; Lönnerdal, B. Bioavailability of iron from plant and animal ferritins. J. Nutr. Biochem. 2015, 26, 532–540. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, X.; Hong, J.; Tang, A.; Liu, Y.; Xie, N.; Nie, G.; Yan, X.; Liang, M. Biochemistry of mammalian ferritins in the regulation of cellular iron homeostasis and oxidative responses. Sci. China Life Sci. 2021, 64, 352–362. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Yu, X.; Xie, J.; Xu, H. New Insights into the Role of Ferritin in Iron Homeostasis and Neurodegenerative Diseases. Mol. Neurobiol. 2021, 58, 2812–2823. [Google Scholar] [CrossRef] [PubMed]
- Surguladze, N.; Thompson, K.M.; Beard, J.L.; Connor, J.R.; Fried, M.G. Interactions and reactions of ferritin with DNA. J. Biol. Chem. 2004, 279, 14694–14702. [Google Scholar] [CrossRef] [PubMed]
- Surguladze, N.; Patton, S.; Cozzi, A.; Fried, M.G.; Connor, J.R. Characterization of nuclear ferritin and mechanism of translocation. Biochem. J. 2005, 388 Pt 3, 731–740. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Cai, C.; Ching, A.; Lagace, C.; Linsenmayer, T. Nuclear ferritin-mediated protection of corneal epithelial cells from oxidative damage to DNA. Dev. Dyn. 2008, 237, 2676–2683. [Google Scholar] [CrossRef] [PubMed]
- Grissi, C.; Taverna Porro, M.; Perona, M.; Atia, M.; Negrin, L.; Moreno, M.S.; Sacanell, J.; Olivera, M.S.; Del Grosso, M.; Durán, H.; et al. Superparamagnetic iron oxide nanoparticles induce persistent large foci of DNA damage in human melanoma cells post-irradiation. Radiat. Environ. Biophys. 2023, 62, 357–369. [Google Scholar] [CrossRef] [PubMed]
- Lisby, M.; Mortensen, U.H.; Rothstein, R. Colocalization of multiple DNA double-strand breaks at a single Rad52 repair centre. Nat. Cell Biol. 2003, 5, 572–577. [Google Scholar] [CrossRef] [PubMed]
- Mortensen, U.H.; Lisby, M.; Rothstein, R. Rad52. Curr. Biol. 2009, 19, R676–R677. [Google Scholar] [CrossRef] [PubMed]
- Pujol, N.; Belli, G.; Herrero, E.; Nogues, A.; de la Torre Ruiz, M.A. Glutaredoxins Grx3 and Grx4 regulate nuclear localisation of Aft1 and the oxidative stress response in Saccharomyces cerevisiae. J. Cell Sci. 2006, 119, 4554–4564. [Google Scholar] [CrossRef] [PubMed]
- Petkova, M.I.; Pujol, N.; de la Torre Ruiz, M.A. Mtl1 O-mannosylation mediated by both Pmt1 and Pmt2 is important for cell survival under oxidative conditions and TOR blockade. Fungal Genet. Biol. 2012, 49, 903–914. [Google Scholar] [CrossRef] [PubMed]
- Pujol, N.; Alfonso, A.; Puig, S.; de la Torre Ruiz, M.A. Both human and soya bean ferritins highly improve the accumulation of bioavailable iron and contribute to extend the chronological life in budding yeast. Microb. Biotechnol. 2022, 15, 1525–1541. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Murillo, M.; Cabello, M.J.; Clemente, M.; Monje, F.; Prado, F. Defective histone supply causes condensing dependent chromatin alterations, SAC activation and chromosome decatenation impairment. Nucleic Acids Res. 2014, 42, 12469–12482, Erratum in Nucleic Acids Res. 2016, 44, 3479–3480. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zhao, X.; Muller, E.G.; Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 1998, 2, 329–340. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, C.; Michaelis, S.; Mitchell, A. Methods in Yeast Genetics; Cold Spring Harbor Laboratory Press Ltd.: New York, NY, USA, 1994; pp. 107–121. ISBN 9780879694517. Available online: https://books.google.es/books?id=u4MMAQAAMAAJ (accessed on 23 February 2015).
- Montella, S.; Pujol, N.; Mechoud, M.A.; de la Torre Ruiz, M.A. Bulk autophagy induction and life extension is achieved when iron is the only limited nutrient in Saccharomyces cerevisiae. Biochem. J. 2021, 478, 811–837. [Google Scholar] [CrossRef] [PubMed]
- Pujol, N.; Petkova, M.I.; Serrano, L.; de la Torre-Ruiz, M.A. The MAP kinase Slt2 is involved in vacuolar function and actin remodeling in Saccharomyces cerevisiae mutants affected by endogenous oxidative stress. Appl. Environ. Microbiol. 2013, 79, 6459–6471. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Pujol, N.; Torre Ruiz, M.A. Physical interaction between the MAPK Slt2 of the PKC1-MAPK pathway and Grx3/Grx4 glutaredoxins is required for the oxidative stress response in budding yeast. Free Radic. Biol. Med. 2017, 103, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Leroy, C.; Mann, C.; Marsolier, M.C. Silent repair accounts for cell cycle specificity in the signaling of oxidative DNA lesions. EMBO J. 2001, 20, 2896–2906. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- De Domenico, I.; Vaughn, M.B.; Li, L.; Bagley, D.; Musci, G.; Ward, D.M.; Kaplan, J. Ferroportin-mediated mobilization of ferritin iron precedes ferritin degradation by the proteasome. EMBO J. 2006, 25, 5396–5404. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dowdle, W.E.; Nyfeler, B.; Nagel, J.; Elling, R.A.; Liu, S.; Triantafellow, E.; Menon, S.; Wang, Z.; Honda, A.; Pardee, G.; et al. Selective VPS34 inhibitor blocks autophagy and uncovers a role for NCOA4 in ferritin degradation and iron homeostasis in vivo. Nat. Cell Biol. 2014, 16, 1069–1079. [Google Scholar] [CrossRef] [PubMed]
- Ravet, K.; Touraine, B.; Kim, S.A.; Cellier, F.; Thomine, S.; Guerinot, M.L.; Briat, J.F.; Gaymard, F. Post-translational regulation of AtFER2 ferritin in response to intracellular iron trafficking during fruit development in Arabidopsis. Mol. Plant 2009, 2, 1095–1106. [Google Scholar] [CrossRef] [PubMed]
- Gryzik, M.; Srivastava, A.; Longhi, G.; Bertuzzi, M.; Gianoncelli, A.; Carmona, F.; Poli, M.; Arosio, P. Expression and characterization of the ferritin binding domain of Nuclear Receptor Coactivator-4 (NCOA4). Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 2710–2716. [Google Scholar] [CrossRef] [PubMed]
- Perfecto, A.; Rodriguez, I.; Rodriguez, J.; Sharp, P.; Balk, J.; Fairweather, S. Pea Ferritin Stability under Gastric pH Conditions Determines the Mechanism of Iron Uptake in Caco-2 Cells. J. Nutr. 2018, 148, 1229–1235, Erratum in J. Nutr. 2019, 149, 542. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kim, M.; Rho, Y.; Jin, K.S.; Ahn, B.; Jung, S.; Kim, H.; Ree, M. pH-dependent structures of ferritin and apoferritin in solution: Disassembly and reassembly. Biomacromolecules 2011, 12, 1629–1640. [Google Scholar] [CrossRef] [PubMed]
- Zang, J.; Chen, H.; Zhao, G.; Wang, F.; Ren, F. Ferritin cage for encapsulation and delivery of bioactive nutrients: From structure, property to applications. Crit. Rev. Food Sci. Nutr. 2017, 57, 3673–3683. [Google Scholar] [CrossRef] [PubMed]
- Masuda, T.; Goto, F.; Yoshihara, T. A novel plant ferritin subunit from soybean that is related to a mechanism in iron release. J. Biol. Chem. 2001, 276, 19575–19579. [Google Scholar] [CrossRef] [PubMed]
- Plays, M.; Müller, S.; Rodriguez, R. Chemistry and biology of ferritin. Metallomics 2021, 12, mfab021. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ahmad, S.; Moriconi, F.; Naz, N. Ferritin L and Ferritin H are differentially located within hepatic and extra hepatic organs under physiological and acute phase conditions. Int. J. Clin. Exp. Pathol. 2013, 6, 622–629. [Google Scholar] [PubMed] [PubMed Central]
- Chiou, B.; Connor, J.R. Emerging and Dynamic Biomedical Uses of Ferritin. Pharmaceuticals 2018, 11, 124. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Cozzi, A.; Santambrogio, P.; Privitera, D.; Broccoli, V.; Rotundo, L.I.; Garavaglia, B.; Benz, R.; Altamura, S.; Goede, J.S.; Muckenthaler, M.U.; et al. Human L-ferritin deficiency is characterized by idiopathic generalized seizures and atypical restless leg syndrome. J. Exp. Med. 2013, 10, 1779–1791. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Li, W.; Garringer, H.J.; Goodwin, C.B.; Richine, B.; Acton, A.; VanDuyn, N.; Muhoberac, B.B.; Irimia Dominguez, J.; Chan, R.J.; Peacock, M.; et al. Systemic and cerebral iron homeostasis in ferritin knock-out mice. PLoS ONE 2015, 10, e0117435. [Google Scholar] [CrossRef] [PubMed]
- Briat, J.F.; Duc, C.; Ravet, K.; Gaymard, F. Ferritins and iron storage in plants. Biochim. Biophys. Acta 2010, 1800, 806–814. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Pan, X.; Xu, Y.; Li, Y.; Xu, N.; Huang, Z.; Ye, J.; Gao, D.; Guo, M. Agrobacterium tumefaciens ferritins play an important role in full virulence through regulating iron homeostasis and oxidative stress survival. Mol. Plant Pathol. 2020, 21, 1167–1178. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Deák, M.; Horváth, G.V.; Davletova, S.; Török, K.; Sass, L.; Vass, I.; Barna, B.; Király, Z.; Dudits, D. Plants ectopically expressing the iron-binding protein, ferritin, are tolerant to oxidative damage and pathogens. Nat. Biotechnol. 1999, 17, 192–196, Erratum in Nat. Biotechnol. 1999, 17, 393. [Google Scholar] [CrossRef] [PubMed]
- Stein, R.J.; Ricachenevsky, F.K.; Fett, J.P. Differential regulation of the two rice ferritin genes (OsFER1 and OsFER2). Plant Sci. 2009, 177, 563–569. [Google Scholar] [CrossRef]
- Newcomb, T.G.; Loeb, L.A. DNA Repair in Prokaryotes and Lower Eukaryotes; Nickoloff, J.A., Hoekstra, M.F., Eds.; Humana: Totawa, NJ, USA, 1998; pp. 65–84. [Google Scholar]
- Driessens, N.; Versteyhe, S.; Ghaddhab, C.; Burniat, A.; De Deken, X.; Van Sande, J.; Dumont, J.E.; Miot, F.; Corvilain, B. Hydrogen peroxide induces DNA single- and double-strand breaks in thyroid cells and is therefore a potential mutagen for this organ. Endocr. Relat. Cancer 2009, 16, 845–856. [Google Scholar] [CrossRef] [PubMed]
- Ragu, S.; Faye, G.; Iraqui, I.; Masurel-Heneman, A.; Kolodner, R.D.; Huang, M.E. Oxygen metabolism and reactive oxygen species cause chromosomal rearrangements and cell death. Proc. Natl. Acad. Sci. USA 2007, 104, 9747–9752. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Groesser, T.; Chang, H.; Fontenay, G.; Chen, J.; Costes, S.V.; Helen Barcellos-Hoff, M.; Parvin, B.; Rydberg, B. Persistence of γ-H2AX and 53BP1 foci in proliferating and non-proliferating human mammary epithelial cells after exposure to γ-rays or iron ions. Int. J. Radiat. Biol. 2011, 87, 696–710. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Akatsuka, S.; Nagai, H.; Chew, S.H.; Ohara, H.; Okazaki, Y.; Yamashita, Y.; Yoshikawa, Y.; Yasui, H.; Ikuta, K.; et al. Iron overload signature in chrysotile-induced malignant mesothelioma. J. Pathol. 2012, 228, 366–377. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Akatsuka, S.; Motooka, Y.; Kong, Y.; Zheng, H.; Mashimo, T.; Imaoka, T.; Toyokuni, S. BRCA1 haploinsufficiency impairs iron metabolism to promote chrysotile-induced mesothelioma via ferroptosis resistance. Cancer Sci. 2023, 14, 1423–1436. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Liao, X.; Lv, C.; Zhang, X.; Masuda, T.; Li, M.; Zhao, G. A novel strategy of natural plant ferritin to protect DNA from oxidative damage during iron oxidation. Free Radic. Biol. Med. 2012, 53, 375–382. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Yang, S.; Liao, X.; Deng, J.; Zhao, G. The interaction of DNA with phytoferritin during iron oxidation. Food Chem. 2014, 153, 292–297. [Google Scholar] [CrossRef] [PubMed]
- Theil, E.C. The ferritin family of iron storage proteins. Adv. Enzymol. Relat. Areas Mol. Biol. 1990, 63, 421–449. [Google Scholar] [CrossRef] [PubMed]
- Yanatori, I.; Nishina, S.; Kishi, F.; Hino, K. Newly uncovered biochemical and functional aspects of ferritin. FASEB J. 2023, 37, e23095. [Google Scholar] [CrossRef] [PubMed]
- Santana, N.; Gikandi, A.; Mancias, J.D. The Role of NCOA4-Mediated Ferritinophagy in Ferroptosis. Adv. Exp. Med. Biol. 2021, 1301, 41–57. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Foissac, X.; Carss, A.; Gatehouse, A.M.; Gatehouse, J.A. Ferritin acts as the most abundant binding protein for snowdrop lectin in the midgut of rice brown planthoppers (Nilaparvata lugens). Insect Biochem. Mol. Biol. 2000, 30, 297–305. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Hevi, S.; Chuck, S.L. Regulated secretion of glycosylated human ferritin from hepatocytes. Blood 2004, 103, 2369–2376. [Google Scholar] [CrossRef] [PubMed]
- Yanatori, I.; Kishi, F.; Toyokuni, S. New iron export pathways acting via holo-ferritin secretion. Arch. Biochem. Biophys. 2023, 746, 109737. [Google Scholar] [CrossRef] [PubMed]
- Bolun, S.; Beibei, T.; Gaoshang, L.; Changjie, Y.; Panxue, Z.; Chao, L.; Wenge, Y. Effect of Glycosylation on the Physicochemical Properties, Structure and Iron Bioavailability of Ferritin Extracted from Tegillarca granosa. J. Ocean Univ. China (Ocean. Coast. Sea Res.) 2023, 22, 1068–1078. [Google Scholar] [CrossRef]
- MacKenzie, E.L.; Iwasaki, K.; Tsuji, Y. Intracellular iron transport and storage: From molecular mechanisms to health implications. Antioxid. Redox Signal. 2008, 10, 997–1030. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pujol Carrión, N.; de la Torre-Ruiz, M.Á. Heterologous Expression of Either Human or Soya Bean Ferritins in Budding Yeast Reveals Common Functions Protecting Against Oxidative Agents and Counteracting Double-Strand Break Accumulation. Biomolecules 2025, 15, 447. https://doi.org/10.3390/biom15030447
Pujol Carrión N, de la Torre-Ruiz MÁ. Heterologous Expression of Either Human or Soya Bean Ferritins in Budding Yeast Reveals Common Functions Protecting Against Oxidative Agents and Counteracting Double-Strand Break Accumulation. Biomolecules. 2025; 15(3):447. https://doi.org/10.3390/biom15030447
Chicago/Turabian StylePujol Carrión, Nuria, and Maria Ángeles de la Torre-Ruiz. 2025. "Heterologous Expression of Either Human or Soya Bean Ferritins in Budding Yeast Reveals Common Functions Protecting Against Oxidative Agents and Counteracting Double-Strand Break Accumulation" Biomolecules 15, no. 3: 447. https://doi.org/10.3390/biom15030447
APA StylePujol Carrión, N., & de la Torre-Ruiz, M. Á. (2025). Heterologous Expression of Either Human or Soya Bean Ferritins in Budding Yeast Reveals Common Functions Protecting Against Oxidative Agents and Counteracting Double-Strand Break Accumulation. Biomolecules, 15(3), 447. https://doi.org/10.3390/biom15030447





