Genome-Wide Identification and Expression of Chitinase Class I Genes in Garlic (Allium sativum L.) Cultivars Resistant and Susceptible to Fusarium proliferatum
Abstract
:1. Introduction
2. Results
2.1. In Silico Genome-Wide Identification of Class I Chitinase Genes in A. sativum cv. Ershuizao
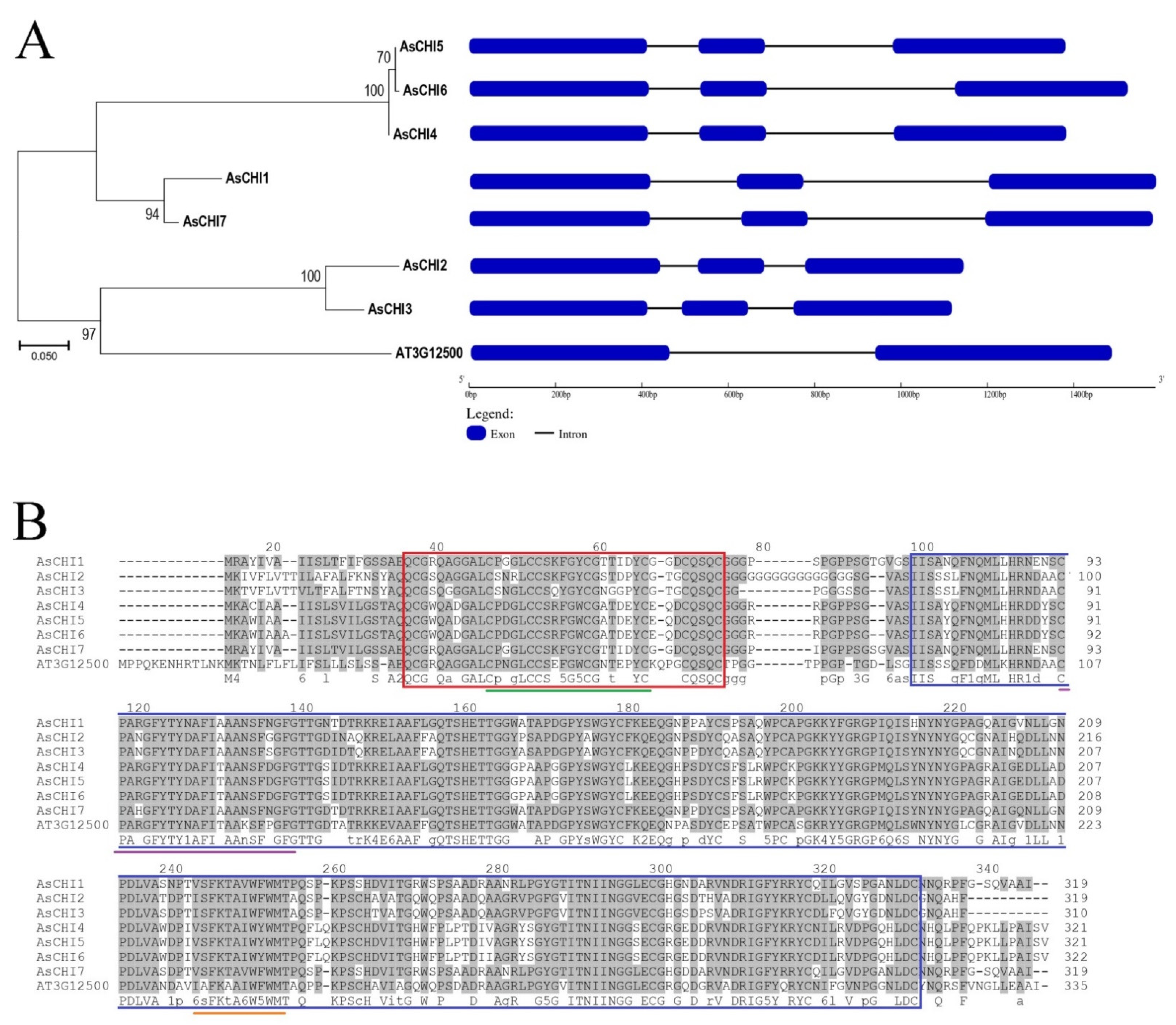
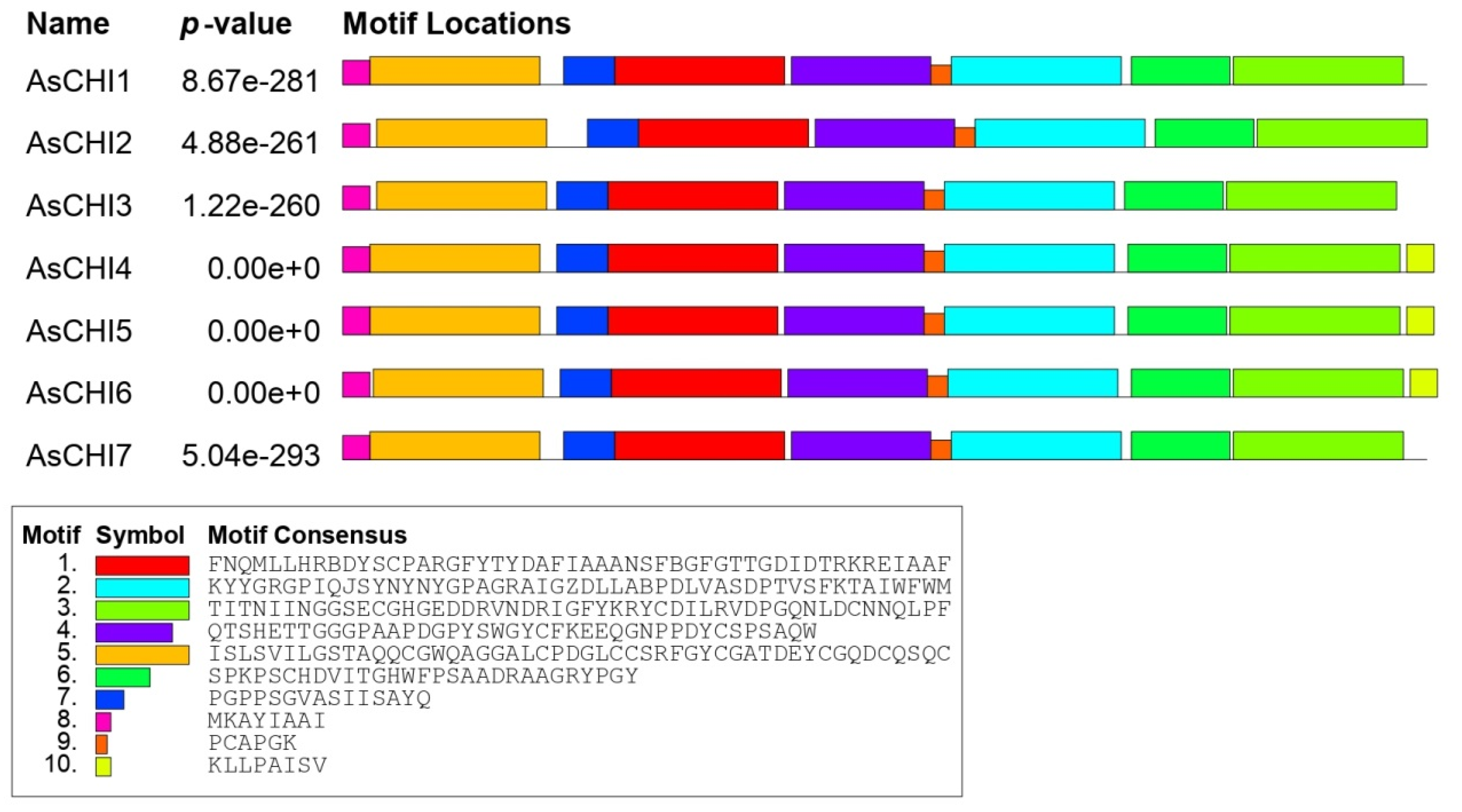
2.2. Structural and Phylogenetic Analyses of Class I Chitinases Predicted in Garlic
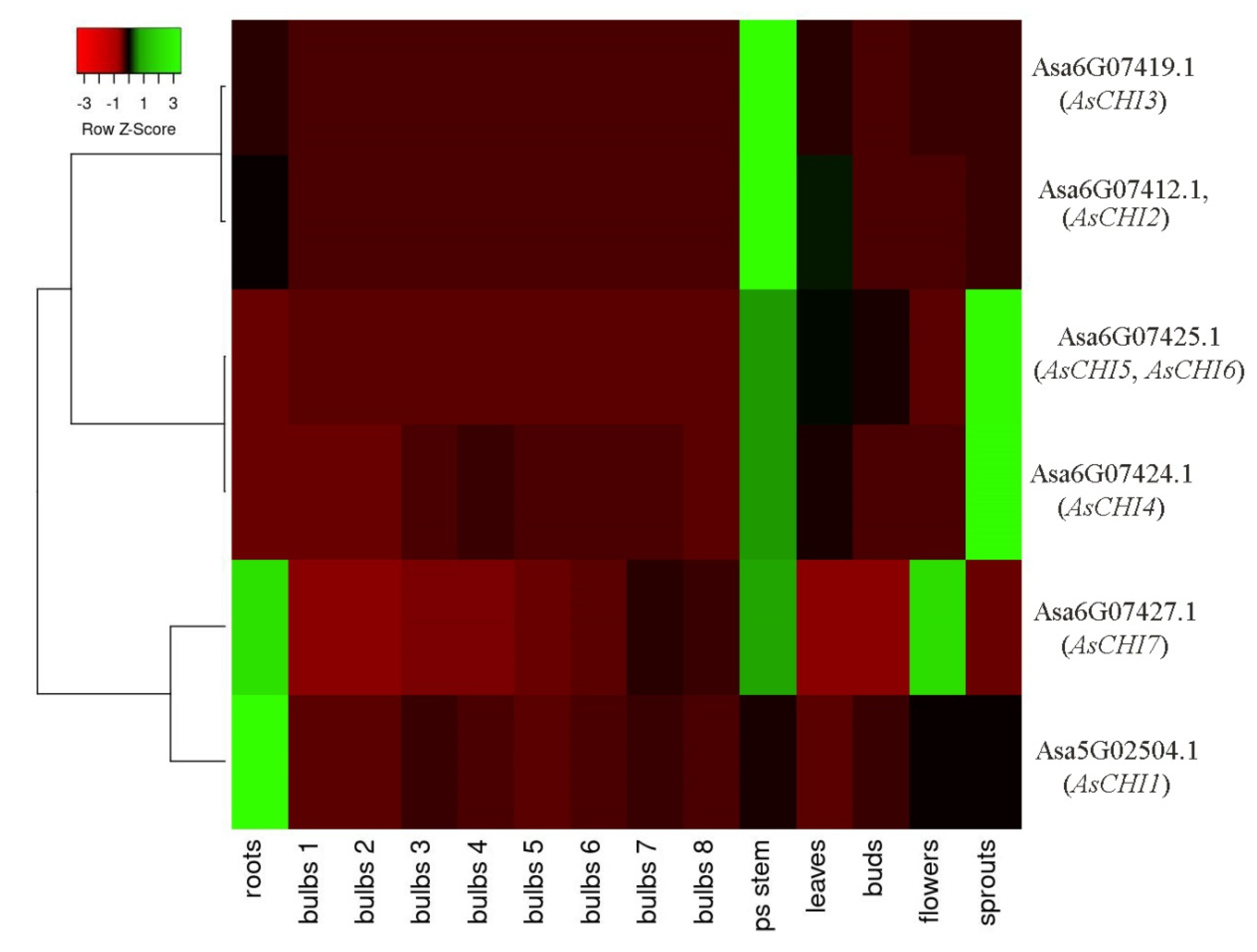
2.3. In Silico Analysis of the Expression of A. sativum Class I Chitinases
2.4. Promoter Analysis of Garlic Class I Chitinase Genes
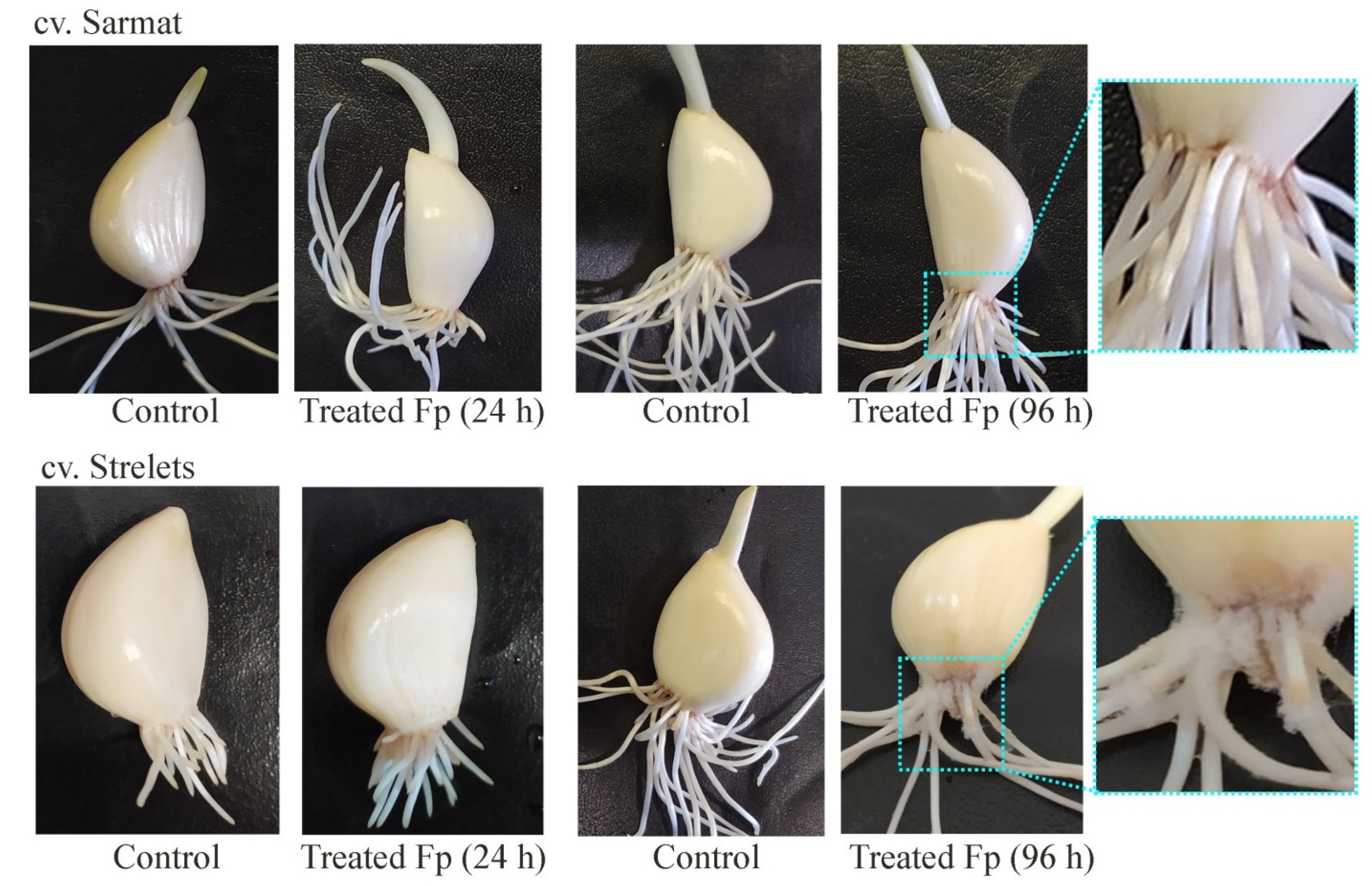
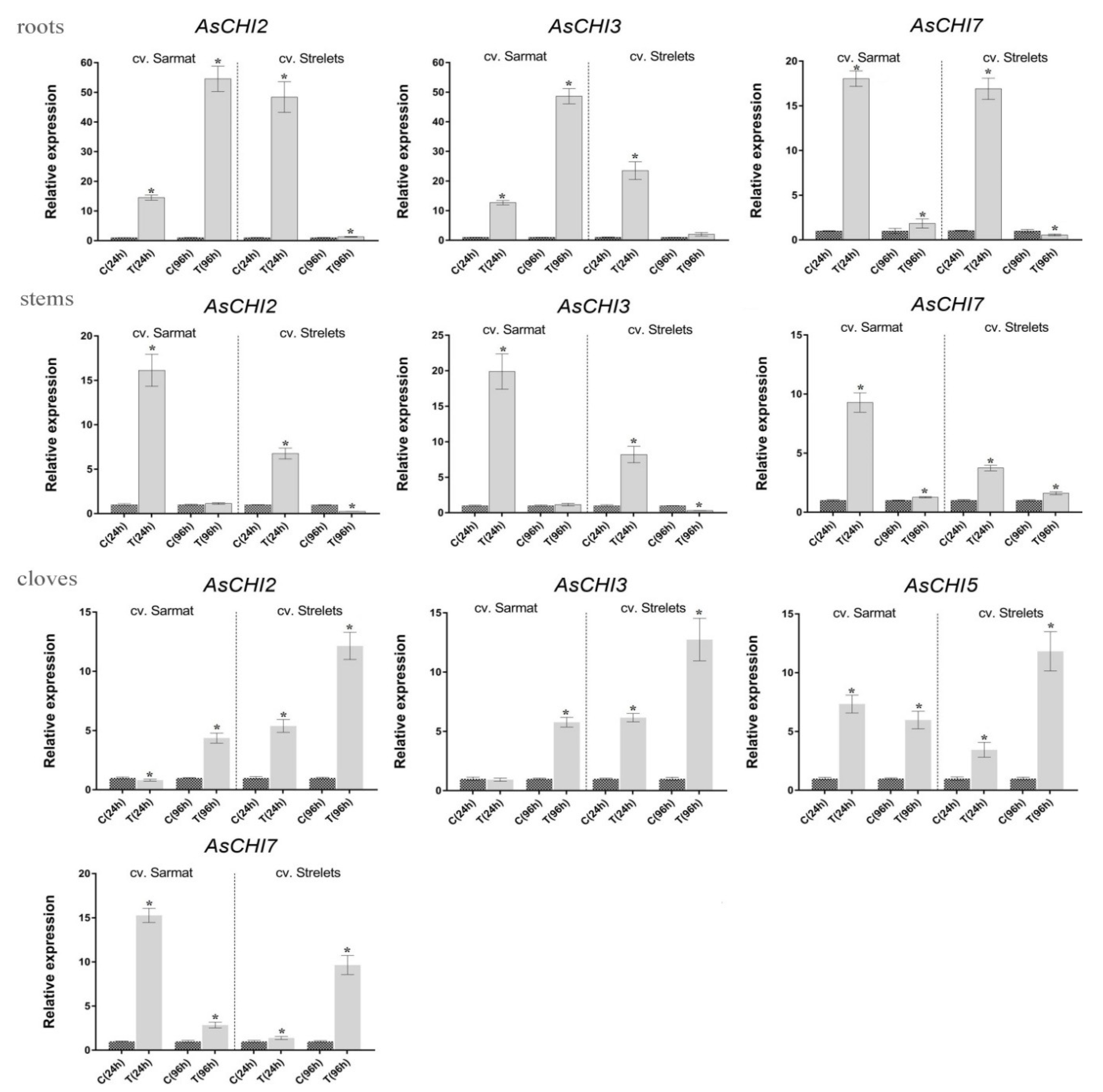
2.5. Class I Chitinase Gene Expression in cv. Sarmat and Strelets Infected with F. proliferatum
2.6. Expression and Characterization of AsCHI CDSs in A. sativum cv. Sarmat and Strelets
2.7. Analysis of Promoters in Class I Chitinase Genes Differentially Expressed in cv. Sarmat and Strelets after F. proliferatum Infection
3. Discussion
4. Materials and Methods
4.1. In Silico Identification and Structural Characterization of A. sativum Class I Chitinase Genes
4.2. In Silico mRNA Expression Analysis
4.3. Gene Identification
4.4. Plants, Fungi, and Fusarium Infection Assay
4.5. RNA Extraction and Quantitative Real-Time Reverse Transcription PCR (qRT-PCR)
4.6. Promoter and 5′-UTR Identification and Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mnayer, D.; Fabiano-Tixier, A.S.; Petitcolas, E.; Hamieh, T.; Nehme, N.; Ferrant, C.; Fernandez, X.; Chemat, F. Chemical composition, antibacterial and antioxidant activities of six essentials oils from the Alliaceae Family. Molecules 2014, 19, 20034–20053. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Snowdon, A. A Color Atlas of Post-Harvest Diseases of Fruits and Vegetables; Wolfe Scientific Ltd.: London, UK, 1990. [Google Scholar]
- Akhter, A.; Hage-Ahmed, K.; Soja, G.; Steinkellner, S. Potential of Fusarium wilt-inducing chlamydospores, in vitro behaviour in root exudates and physiology of tomato in biochar and compost amended soil. Plant Soil 2016, 406, 425–440. [Google Scholar] [CrossRef] [Green Version]
- Gálvez, L.; Urbaniak, M.; Waśkiewicz, A.; Stępień, Ł.; Palmero, D. Fusarium proliferatum—Causal agent of garlic bulb rot in Spain: Genetic variability and mycotoxin production. Food Microbiol. 2017, 67, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Chand, S.K.; Nanda, S.; Mishra, R.; Joshi, R.K. Multiple garlic (Allium sativum L.) microRNAs regulate the immunity against the basal rot fungus Fusarium oxysporum f. sp. Cepae. Plant Sci. 2017, 257, 9–21. [Google Scholar] [CrossRef]
- Pavlou, G.C.; Vakalounakis, D.J.; Ligoxigakis, E.K. Control of root and stem rot of cucumber, caused by Fusarium oxysporum f. sp. radicis-cucumerinum, by grafting onto resistant rootstocks. Plant Dis. 2002, 86, 379–382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Z.; Wang, T.; He, C.; Cheng, K.; Zeng, R.; Song, Y. Control of Panama disease of banana by intercropping with Chinese chive (Allium tuberosum Rottler): Cultivar differences. BMC Plant Biol. 2020, 20, 432. [Google Scholar] [CrossRef]
- Zhang, H.; Mallik, A.; Zeng, R.S. Control of Panama disease of banana by rotating and intercropping with Chinese chive (Allium tuberosum Rottler): Role of plant volatiles. J. Chem. Ecol. 2013, 39, 243–252. [Google Scholar] [CrossRef]
- Xu, N.; Wei, M.; Wang, C.; Shi, W.; Tian, F.M.; Wang, X. Composition of Welsh onion (Allium fistulosum L.) root exudates and their allelopathy on cucumber sprouts and Fusarium oxysporum f. sp. cucumerinum. Allelop. J. 2013, 32, 243–256. [Google Scholar]
- Zuo, G.W.; Li, C.Y.; Li, B.; Wei, Y.R.; Hu, C.H.; Yang, Q.S.; Yang, J.; Sheng, O.; Kuang, R.B.; Deng, G.M.; et al. The toxic mechanism and bioactive components of Chinese leek root exudates acting against Fusarium oxysporum f. sp. cubense, tropical race 4. Eur. J. Plant Pathol. 2015, 143, 447–460. [Google Scholar] [CrossRef]
- Mylona, K.; Garcia-Cela, E.; Sulyok, M.; Medina, A.; Magan, N. Influence of Two Garlic-Derived Compounds, Propyl Propane Thiosulfonate (PTS) and Propyl Propane Thiosulfinate (PTSO), on Growth and Mycotoxin Production by Fusarium Species In Vitro and in Stored Cereals. Toxins 2019, 11, 495. [Google Scholar] [CrossRef] [Green Version]
- Abdelrahman, M.; El-Sayed, M.; Sato, S.; Hirakawa, H.; Ito, S.I.; Tanaka, K.; Mine, Y.; Sugiyama, N.; Suzuki, Y.; Yamauchi, N.; et al. RNA-sequencing-based transcriptome and biochemical analyses of steroidal saponin pathway in a complete set of Allium fistulosum-A. cepa monosomic addition lines. PLoS ONE 2017, 12, e0181784. [Google Scholar] [CrossRef] [Green Version]
- Nishioka, T.; Marian, M.; Kobayashi, I.; Kobayashi, Y.; Yamamoto, K.; Tamaki, H.; Suga, H.; Shimizu, M. Microbial basis of Fusarium wilt suppression by Allium cultivation. Sci. Rep. 2019, 9, 1715. [Google Scholar] [CrossRef]
- Chand, S.K.; Nanda, S.; Joshi, R.K. Regulation of miR394 in Response to Fusarium oxysporum f. sp. cepae (FOC) Infection in Garlic (Allium sativum L.). Front. Plant Sci. 2016, 7, 258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.; Wang, H.; Zhu, W.; Li, W.; Wang, F. Transcriptome Analysis Reveals the Effects of Chinese Chive (Allium tuberosum R.) Extract on Fusarium oxysporum f. sp. radicis-lycopersici Spore Germination. Cur. Microbiol. 2020, 77, 855–864. [Google Scholar] [CrossRef] [PubMed]
- Gow, N.A.R.; Latge, J.P.; Munro, C.A. The Fungal Cell Wall: Structure, Biosynthesis, and Function. Microbiol. Spect. 2017, 5. [Google Scholar] [CrossRef] [Green Version]
- Arakane, Y.; Taira, T.; Ohnuma, T.; Fukamizo, T. Chitin-related enzymes in agro-biosciences. Curr. Drug Targets 2012, 13, 442–470. [Google Scholar] [CrossRef] [PubMed]
- Desaki, Y.; Miyata, K.; Suzuki, M.; Shibuya, N.; Kaku, H. Plant immunity and symbiosis signaling mediated by LysM receptors. Inn. Immun. 2018, 24, 92–100. [Google Scholar] [CrossRef]
- Volk, H.; Marton, K.; Flajšman, M.; Radišek, S.; Tian, H.; Hein, I.; Podlipnik, Č.; Thomma, B.P.H.J.; Košmelj, K.; Javornik, B.; et al. Chitin-binding protein of Verticillium nonalfalfae disguises fungus from plant chitinases and suppresses chitin-triggered host immunity. Mol. Plant Microb. Interact. 2019, 32, 1378–1390. [Google Scholar] [CrossRef] [Green Version]
- Van Loon, L.C.; Rep, M.; Pieterse, C.M.J. Significance of inducible defense-related proteins in infected plants. Ann. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ali, S.; Ganai, B.A.; Kamili, A.N.; Bhat, A.A.; Mir, Z.A.; Bhat, J.A.; Tyagi, A.; Islam, S.T.; Mushtaq, M.; Yadav, P.; et al. Pathogenesis-related proteins and peptides as promising tools for engineering plants with multiple stress tolerance. Microbiol. Res. 2018, 212–213, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Udaya Prakash, N.A.; Jayanthi, M.; Sabarinathan, R.; Kangueane, P.; Mathew, L.; Sekar, K. Evolution, homology conservation, and identification of unique sequence signatures in GH19 family chitinases. J. Mol. Evol. 2010, 70, 466–478. [Google Scholar] [CrossRef]
- Neuhaus, J.M.; Fritig, B.; Linthorst, H.J.M.; Meins, F.; Mikkelsen, J.D.; Ryals, J. A revised nomenclature for chitinase genes. Plant Mol. Biol. Rep. 1996, 14, 102–104. [Google Scholar] [CrossRef]
- Takenaka, Y.; Nakano, S.; Tamoi, M.; Sakuda, S.; Fukamizo, T. Chitinase gene expression in response to environmental stresses in Arabidopsis thaliana: Chitinase inhibitor allosamidin enhances stress tolerance. Biosci. Biotechnol. Biochem. 2009, 73, 1066–1071. [Google Scholar] [CrossRef] [Green Version]
- Yang, H.; Zhang, T.; Masuda, T.; Lv, C.; Sun, L.; Qu, G.; Zhao, G. Chitinase III in pomegranate seeds (Punica granatum Linn.): A high-capacity calcium-binding protein in amyloplasts. Plant J. 2011, 68, 765–776. [Google Scholar] [CrossRef]
- Ohnuma, T.; Numata, T.; Osawa, T.; Mizuhara, M.; Vårum, K.M.; Fukamizo, T. Crystal structure and mode of action of a class V chitinase from Nicotiana tabacum. Plant Mol. Biol. 2011, 75, 291–304. [Google Scholar] [CrossRef]
- Ohnuma, T.; Numata, T.; Osawa, T.; Mizuhara, M.; Lampela, O.; Juffer, A.H.; Skriver, K.; Fukamizo, T. A class V chitinase from Arabidopsis thaliana: Gene responses, enzymatic properties, and crystallographic analysis. Planta 2011, 234, 123–137. [Google Scholar] [CrossRef] [PubMed]
- Ohnuma, T.; Umemoto, N.; Kondo, K.; Numata, T.; Fukamizo, T. Complete subsite mapping of a “loopful” GH19 chitinase from rye seeds based on its crystal structure. FEBS Lett. 2013, 587, 2691–2697. [Google Scholar] [CrossRef] [Green Version]
- Oliveira, S.T.; Azevedo, M.I.G.; Cunha, R.M.S.; Silva, C.F.B.; Muniz, C.R.; Monteiro-Júnior, J.E.; Carneiro, R.F.; Nagano, C.S.; Girão, M.S.; Freitas, C.D.T.; et al. Structural and functional features of a class VI chitinase from cashew (Anacardium occidentale L.) with antifungal properties. Phytochemistry 2020, 180, 112527. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Caballero, S.; Cano-Sánchez, P.; Mares-Mejía, I.; Díaz-Sánchez, A.G.; Macías-Rubalcava, M.L.; Hermoso, J.A.; Rodríguez-Romero, A. Comparative study of two GH19 chitinase-like proteins from Hevea brasiliensis, one exhibiting a novel carbohydrate-binding domain. FEBS J. 2014, 281, 4535–4554. [Google Scholar] [CrossRef] [PubMed]
- Balu, K.E.; Ramya, K.S.; Radha, A.; Krishnasamy, G. Structure of intact chitinase with hevein domain from the plant Simarouba glauca, known for its traditional anti-inflammatory efficacy. Int. J. Biol. Macromol. 2020, 161, 1381–1392. [Google Scholar] [CrossRef] [PubMed]
- Broekaert, W.F.; Mariën, W.; Terras, F.R.; De Bolle, M.F.; Proost, P.; Van Damme, J.; Dillen, L.; Claeys, M.; Rees, S.B.; Vanderleyden, J.; et al. Antimicrobial peptides from Amaranthus caudatus seeds with sequence homology to the cysteine/glycine-rich domain of chitin-binding proteins. Biochemistry 1992, 31, 4308–4314. [Google Scholar] [CrossRef]
- Taira, T.; Toma, N.; Ishihara, M. Purification, characterization, and antifungal activity of chitinases from pineapple (Ananas comosus) leaf. Biosci. Biotechnol. Biochem. 2005, 69, 189–196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cletus, J.; Balasubramanian, V.; Vashisht, D.; Sakthivel, N. Transgenic expression of plant chitinases to enhance disease resistance. Biotechnol. Lett. 2013, 35, 1719–1732. [Google Scholar] [CrossRef]
- Tobias, P.A.; Christie, N.; Naidoo, S.; Guest, D.I.; Külheim, C. Identification of the Eucalyptus grandis chitinase gene family and expression characterization under different biotic stress challenges. Tree Physiol. 2017, 37, 565–582. [Google Scholar] [CrossRef]
- Durechova, D.; Jopcik, M.; Rajninec, M.; Moravcikova, J.; Libantova, J. Expression of Drosera rotundifolia Chitinase in Transgenic Tobacco Plants Enhanced Their Antifungal Potential. Mol. Biotechnol. 2019, 61, 916–928. [Google Scholar] [CrossRef]
- Mohapatra, R.K.; Nanda, S. In silico analysis of onion chitinases using transcriptome data. Bioinformation 2018, 14, 440–445. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; Zhu, S.; Li, N.; Cheng, Y.; Zhao, J.; Qiao, X.; Lu, L.; Liu, S.; Wang, Y.; Liu, C.; et al. A Chromosome-Level Genome Assembly of Garlic (Allium sativum) Provides Insights into Genome Evolution and Allicin Biosynthesis. Mol. Plant 2020, 13, 1328–1339. [Google Scholar] [CrossRef]
- Huet, J.; Azarkan, M.; Looze, Y.; Villeret, V.; Wintjens, R. Crystallization and preliminary X-ray analysis of a family 19 glycosyl hydrolase from Carica papaya latex. Acta Crystallogr. Sect. F Struct. Biol. Crystal. Commun. 2008, 64, 371–374. [Google Scholar] [CrossRef] [PubMed]
- Misra, B. Molecular Evolution and Functional Divergence of Chitinase Gene Family in Hevea brasiliensis Genome. Winnower 2015, 8, e144125.54243. [Google Scholar] [CrossRef]
- Hauenschild, F.; Favre, A.; Schnitzler, J.; Michalak, I.; Freiberg, M.; Muellner-Riehl, A.N. Spatio-temporal evolution of Allium L. in the Qinghai-Tibet-Plateau region: Immigration and in situ radiation. Plant Diver. 2017, 39, 167–179. [Google Scholar] [CrossRef] [PubMed]
- Khandagale, K.; Krishna, R.; Roylawar, P.; Ade, A.B.; Benke, A.; Shinde, B.; Singh, M.; Gawande, S.J.; Rai, A. Omics approaches in Allium research: Progress and way ahead. PeerJ 2020, 8, e9824. [Google Scholar] [CrossRef]
- Taylor, A.; Vagany, V.; Barbara, D.J.; Thomas, B.; Pink, D.A.C.; Jones, J.E.; Clarkson, J.P. Identification of differential resistance to six Fusarium oxysporum f. sp. cepae isolates in commercial onion cultivars through the development of a rapid seedling assay. Plant Pathol. 2013, 62, 103–111. [Google Scholar] [CrossRef]
- Chen, J.; Piao, Y.; Liu, Y.; Li, X.; Piao, Z. Genome-wide identification and expression analysis of chitinase gene family in Brassica rapa reveals its role in clubroot resistance. Plant Sci. 2018, 270, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Jeffares, D.C.; Penkett, C.J.; Bähler, J. Rapidly regulated genes are intron poor. Trends Genet. 2008, 24, 375–378. [Google Scholar] [CrossRef]
- Zhou, F.; Guo, Y.; Qiu, L.J. Genome-wide identification and evolutionary analysis of leucine-rich repeat receptor-like protein kinase genes in soybean. BMC Plant Biol. 2016, 16, 58. [Google Scholar] [CrossRef] [Green Version]
- Passarinho, P.A.; de Vries, S.C. Arabidopsis chitinases: A genomic survey. Arabidopsis Book 2002, 1, e0023. [Google Scholar] [CrossRef] [Green Version]
- de Gerhardt, L.B.A.; Sachetto-Martins, G.; Contarini, M.G.; Sandroni, M.; de Ferreira, R.P.; de Lima, V.M.; Cordeiro, M.C.; de Oliveira, D.E.; Margis-Pinheiro, M. Arabidopsis thaliana class IV chitinase is early induced during the interaction with Xanthomonas campestris. FEBS Lett. 1997, 419, 69–75. [Google Scholar] [CrossRef] [Green Version]
- Rahman, T.A.; Oirdi, M.E.; Gonzalez-Lamothe, R.; Bouarab, K. Necrotrophic pathogens use the salicylic acid signaling pathway to promote disease development in tomato. Mol. Plant-Microbe Interact. 2012, 25, 1584–1593. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Veluthakkal, R.; Dasgupta, M.G. Isolation and characterization of pathogen defence-related class I chitinase from the actinorhizal tree Casuarina equisetifolia. For. Pathol. 2012, 42, 467–480. [Google Scholar] [CrossRef]
- Grover, A. Plant Chitinases: Genetic Diversity and Physiological Roles. Crit. Rev. Plant Sci. 2012, 31, 57–73. [Google Scholar] [CrossRef]
- Bartholomew, E.S.; Black, K.; Feng, Z.; Liu, W.; Shan, N.; Zhang, X.; Wu, L.; Bailey, L.; Zhu, N.; Qi, C.; et al. Comprehensive Analysis of the Chitinase Gene Family in Cucumber (Cucumis sativus L.): From Gene Identification and Evolution to Expression in Response to Fusarium oxysporum. Int. J. Mol. Sci. 2019, 20, 5309. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, Y.; Jia, S.; Wang, C.; Wang, F.; Wang, F.; Zhao, K. BjMYB1, a transcription factor implicated in plant defence through activating BjCHI1 chitinase expression by binding to a W-box-like element. J. Exp. Bot. 2016, 67, 4647–4658. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, Y.; Zan, X.; Wu, X.; Yao, L.; Chen, Y.; Jia, S.; Zhao, K. Identification of Fungus-Responsive Cis–Acting Element in the Promoter of Brassica Juncea Chitinase Gene, BjCHI1. Plant Sci. 2014, 215–216, 190–198. [Google Scholar] [CrossRef] [PubMed]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Kracher, B.; Ziegler, J.; Birkenbihl, R.P.; Somssich, I.E. Negative regulation of ABA signaling by WRKY33 is critical for Arabidopsis immunity towards Botrytis cinerea 2100. Elife 2015, 4, e07295. [Google Scholar] [CrossRef] [PubMed]
- Du, M.; Zhai, Q.; Deng, L.; Li, S.; Li, H.; Yan, L.; Huang, Z.; Wang, B.; Jiang, H.; Huang, T.; et al. Closely related NAC transcription factors of tomato differentially regulate stomatal closure and reopening during pathogen attack. Plant Cell 2014, 26, 3167–3184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, J.; Duan, G.; Li, C.; Liu, L.; Han, G.; Zhang, Y.; Wang, C. The Crosstalks Between Jasmonic Acid and Other Plant Hormone Signaling Highlight the Involvement of Jasmonic Acid as a Core Component in Plant Response to Biotic and Abiotic Stresses. Front. Plant Sci. 2019, 10, 1349. [Google Scholar] [CrossRef] [Green Version]
- Zhang, C.; Huang, M.; Sang, X.; Li, P.; Ling, Y.; Zhao, F.; Du, D.; Li, Y.; Yang, Z.; He, G. Association between sheath blight resistance and chitinase activity in transgenic rice plants expressing McCHIT1 from bitter melon. Transgenic Res. 2019, 28, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yang, J.; Li, H.; Niu, L.; Xing, G.; Zhang, Y.; Xu, W.; Zhao, Q.; Li, Q.; Dong, Y. Overexpression of the chitinase gene CmCH1 from Coniothyrium minitans renders enhanced resistance to Sclerotinia sclerotiorum in soybean. Transgenic Res. 2020, 29, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0. molecular biology and evolution. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An Upgraded Gene Feature Visualization Server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar] [PubMed]
- Choi, Y.; Sims, G.E.; Murphy, S.; Miller, J.R.; Chan, A.P. Predicting the functional eect of amino acid substitutions and indels. PLoS ONE 2012, 7, e46688. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Babicki, S.; Arndt, D.; Marcu, A.; Liang, Y.; Grant, J.R.; Maciejewski, A.; Wishart, D.S. Heatmapper: Web-enabled heat mapping for all. Nucl. Acids Res. 2016, 44, W147–W153. [Google Scholar] [CrossRef]
- Puchooa, D. A simple, rapid and efficient method for the extraction of genomic DNA from lychee (Litchi chinensis Sonn.). Afr. J. Biotechnol. 2004, 3, 253–255. [Google Scholar]
- Leyronas, C.; Chrétien, P.L.; Troulet, C.; Duffaud, M.; Villeneuve, F.; Morris, C.E.; Hunyadi, H. First report of Fusarium proliferatum causing garlic clove rot in France. Plant Dis. 2018, 102, 2658. [Google Scholar] [CrossRef]
- Sugui, J.A.; Deising, H.B. Isolation of infection-specific sequence tags expressed during early stages of maize anthracnose disease development. Mol. Plant Pathol. 2002, 3, 197–203. [Google Scholar] [CrossRef]
- Liu., M.; Wu, Z.; Jiang, F. Selection and validation of garlic reference genes for quantitative real-time PCR normalization. Plant Cell Tissue Organ Cul. 2015, 122, 435–444. [Google Scholar] [CrossRef]
- Schwinn, K.E.; Ngo, H.; Kenel, F.; Brummell, D.A.; Albert, N.W.; McCallum, J.A.; Pither-Joyce, M.; Crowhurst, R.N.; Eady, C.; Davies, K.M. The onion (Allium cepa L.) R2R3-MYB gene MYB1 regulates anthocyanin biosynthesis. Front. Plant Sci. 2016, 7, 1865. [Google Scholar] [CrossRef] [Green Version]
- Lescot, M. PlantCARE, a Database of Plant Cis-Acting Regulatory Elements and a Portal to Tools for in Silico Analysis of Promoter Sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]



| Gene | Genomic Location (Strand) | Transcript ID in RNA-Seq Data [37] | Size (bp) | Number of Exons | CDS (bp) |
|---|---|---|---|---|---|
| AsCHI1 | ch05: 744404486–744406074 (−) | Asa5G02504.1 | 1589 | 3 | 960 |
| AsCHI2 | ch06: 936506601–963507740 (+) | Asa6G07412.1 | 1140 | 3 | 960 |
| AsCHI3 | ch06: 937609134–937610251 (+) | Asa6G07419.1 | 1118 | 3 | 933 |
| AsCHI4 | ch06: 938113646–938115026 (−) | Asa6G07424.1 | 1381 | 3 | 966 |
| AsCHI5 | ch06: 938164705–938166085 (−) | Asa6G07425.1 * | 1381 | 3 | 966 |
| AsCHI6 | ch06: 938218393–938219916 (−) | 1524 | 3 | 969 | |
| AsCHI7 | ch06: 938640561–938642142 (−) | Asa6G07427.1 | 1582 | 3 | 960 |
| Protein Symbol | Size (aa) | MW (kDa) | pI | Location of Specific Regions (aa) | Aliphatic Index | GRAVY | Functional Annotation in Gene Ontology Categories | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Signal Peptide | CBD1 | GH19 Domain | Cellular Component | Biological Process | Molecular Function | ||||||
| AsCHI1 | 319 | 33.72 | 8.47 | 1–19 | 21–58 | 75–306 | 57.62 | −0.304 | Secretory pathway | chitin catabolic process (GO:0006032) cell wall macromolecule catabolic process (GO:0016998) defense response to fungus (GO:0050832) carbohydrate metabolic process (GO:0005975) | chitinase activity (GO:0004568) chitin binding (GO:0008061) |
| AsCHI2 | 319 | 33.43 | 7.39 | 1–21 | 23–60 | 82–313 | 52.73 | −0.275 | Secretory pathway | chitin catabolic process (GO:0006032) cell wall macromolecule catabolic process (GO:0016998) polysaccharide catabolic process (GO:0000272) defense response to fungus (GO:0050832) | chitinase activity (GO:0004568) chitin binding (GO:0008061) |
| AsCHI3 | 310 | 32.74 | 5.40 | 1–21 | 23–60 | 73–304 | 54.26 | −0.246 | Secretory pathway | chitin catabolic process (GO:0006032) cell wall macromolecule catabolic process (GO:0016998) polysaccharide catabolic process (GO:0000272) defense response to fungus (GO:0050832) | chitinase activity (GO:0004568) chitin binding (GO:0008061) |
| AsCHI4 | 321 | 35.17 | 6.05 | 1–19 | 21–58 | 73–305 | 62.99 | −0.346 | Secretory pathway | chitin catabolic process (GO:0006032) cell wall macromolecule catabolic process (GO:0016998) polysaccharide catabolic process (GO:0000272) defense response (GO:0006952) response to fungus (GO:0009620) | chitinase activity (GO:0004568) chitin binding (GO:0008061) |
| AsCHI5 | 321 | 35.26 | 5.88 | 1–19 | 21–58 | 73–305 | 62.99 | −0.357 | Secretory pathway | chitin catabolic process (GO:0006032) cell wall macromolecule catabolic process (GO:0016998) defense response (GO:0006952) response to fungus (GO:0009620) carbohydrate metabolic process (GO:0005975) | chitinase activity (GO:0004568) chitin binding (GO:0008061) |
| AsCHI6 | 322 | 35.34 | 5.88 | 1–20 | 22–59 | 74–306 | 63.42 | −0.349 | Secretory pathway | chitin catabolic process (GO:0006032) cell wall macromolecule catabolic process (GO:0016998) defense response (GO:0006952) response to fungus (GO:0009620) carbohydrate metabolic process (GO:0005975) | chitinase activity (GO:0004568) chitin binding (GO:0008061) |
| AsCHI7 | 319 | 33.79 | 6.19 | 1–19 | 21–58 | 75–306 | 60.06 | −0.325 | Secretory pathway | chitin catabolic process (GO:0006032) cell wall macromolecule catabolic process (GO:0016998) defense response to fungus (GO:0050832) carbohydrate metabolic process (GO:0005975) | chitinase activity (GO:0004568) chitin binding (GO:0008061) |
| Functional Description | Annotation | Motif | Number of Elements Found in Gene Promoters | ||||||
|---|---|---|---|---|---|---|---|---|---|
| AsCHI1 | AsCHI2 | AsCHI3 | AsCHI4 | AsCHI5 | AsCHI6 | AsCHI7 | |||
| Hormone responsive | Cis-acting elements, involved in abscisic acid response | ABRE | 6 | 1 | 2 | 1 | |||
| ABRE3a | 4 | ||||||||
| ABRE4 | 4 | ||||||||
| CARE | 1 | 1 | |||||||
| Cis-acting elements involved in auxin response | AUXRR-core | 1 | 1 | 1 | |||||
| TGA-box | 1 | 1 | |||||||
| TGA-element | 1 | 1 | 5 | ||||||
| Cis-acting element involved in MeJA response | CGTCA-motif | 1 | 1 | 5 | 2 | 7 | 2 | 1 | |
| Cis-acting element involved in salicylic acid response | TCA-element | 2 | 1 | 1 | |||||
| Cis-acting element involved in gibberellin response | TATC-box | 1 | |||||||
| P-box | 1 | 1 | |||||||
| GARE-motif | 1 | 2 | 1 | 1 | |||||
| Ethylene-responsive element | ERE | 1 | 2 | 9 | 1 | 2 | 2 | 2 | |
| Stress responsive | Cis-acting element essential for anaerobic induction | ARE | 4 | 3 | 1 | 8 | 2 | 3 | 2 |
| Dehydration-responsive element | DRE1 | 2 | |||||||
| DRE core | 1 | 1 | 1 | ||||||
| Cis-acting element involved in low temperature response | LTR | 1 | 1 | 3 | 1 | ||||
| MYB-binding site involved in drought response | MBS | 1 | 1 | ||||||
| Stress-responsive element | STRE | 4 | 1 | 2 | 4 | 5 | 1 | 1 | |
| Salt and heavy metal stress response element | F-box | 1 | 1 | ||||||
| Maximal elicitor-mediated activation | AT-rich sequence | 2 | |||||||
| Cis-acting element involved in defense and stress responses | TC-rich repeats | 1 | 1 | ||||||
| Fungal elicitor and wound responses | W-box | 1 | 2 | 1 | 2 | 1 | |||
| Wound-responsive element | Wun-motif | 2 | 1 | 2 | 2 | ||||
| Wound and pathogen response | WRE3 | 1 | 1 | 1 | |||||
| Gene | cv. Sarmat | cv. Strelets | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NCBI ID | Number of SNPs | Gene (bp) | CDS (bp) | Protein (aa) | NCBI ID | Number of SNPs | Gene (bp) | CDS (bp) | Protein (aa) | |
| AsCHI2 | MW770892 | 4 | 1141 | 960 | 319 | MW770893 | 4 | 1141 | 960 | 319 |
| AsCHI3 | MW770894 | 7 | 1118 | 933 | 310 | MW770895 | 7 | 1118 | 933 | 310 |
| AsCHI5 | MW770896 | 2 | 1397 | 966 | 321 | MW770897 | 0 | 1397 | 966 | 321 |
| AsCHI7 | MW770898 | 6 | 1579 | 960 | 319 | MW770899 | 6 | 1579 | 960 | 319 |
| Genes/Gene Groups | Primer Sequences (5′→3′) | Amplicon Size (bp) | Application |
|---|---|---|---|
| AsCHI1 AsCHI7 | ATAAAAGYGGTGGTACATTGC GTACATAAAACTCATRTGCGWA | ~1100 | Gene amplification and sequencing |
| AsCHI2 | GTAGATRCAGTCCTRCTGCT ATATCATATGACGACTTCGC | ~1100 | |
| AsCHI3 | GTAGATRCAGTCCTRCTGCT ATTGCACATGTATCATATGAGG | ~1100 | |
| AsCHI4 AsCHI5 AsCHI6 | TAAAAGGAGAGGTACGCAC GTAATTATTGCAAGCATCGTAA | ~1100 | |
| AsCHI2 | CTTTCCAGAAACCTGTGACT TGCAGCTGCTATGAAGGCA | ~1000 | Regulatory region amplification and sequencing |
| AsCHI3 | GTAAATGAGCATGGGTAAGTTG GTATTGGCTGCAGCATAGC | ~1000 | |
| AsCHI7 | AGCACCACCAGCTTGTCTA ATGAGAACCGCGTTGATCGT | ~1000 | |
| AsCHI1 AsCHI7 | GTACCACTGGGGATACCGAT CCCCATGAATATGGTCCATCG | 114 | qRT-PCR |
| AsCHI2 | GGAACCACTGGAGACATCAATG GCCTTGTTCTTGCTTGAAGCAG | 140 | |
| AsCHI3 | GGAACCACTGGAGACATCGATA GCCTTGTTCTTGCTTGAAGCAG | 140 | |
| AsCHI4 AsCHI5 AsCHI6 | GGTACCACCGGGAGTATTGAC ACCCCATGAATATGGTCCACCT | 116 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Filyushin, M.A.; Anisimova, O.K.; Kochieva, E.Z.; Shchennikova, A.V. Genome-Wide Identification and Expression of Chitinase Class I Genes in Garlic (Allium sativum L.) Cultivars Resistant and Susceptible to Fusarium proliferatum. Plants 2021, 10, 720. https://doi.org/10.3390/plants10040720
Filyushin MA, Anisimova OK, Kochieva EZ, Shchennikova AV. Genome-Wide Identification and Expression of Chitinase Class I Genes in Garlic (Allium sativum L.) Cultivars Resistant and Susceptible to Fusarium proliferatum. Plants. 2021; 10(4):720. https://doi.org/10.3390/plants10040720
Chicago/Turabian StyleFilyushin, Mikhail A., Olga K. Anisimova, Elena Z. Kochieva, and Anna V. Shchennikova. 2021. "Genome-Wide Identification and Expression of Chitinase Class I Genes in Garlic (Allium sativum L.) Cultivars Resistant and Susceptible to Fusarium proliferatum" Plants 10, no. 4: 720. https://doi.org/10.3390/plants10040720







