Regulation of Glycosylphosphatidylinositol-Anchored Protein (GPI-AP) Expression by F-Box/LRR-Repeat (FBXL) Protein in Wheat (Triticum aestivum L.)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. Gene Cloning and Gene Expression Analysis
2.3. Yeast Two-Hybrid (Y2H) Assay and Library Screening
2.4. Co-Transformation and β-Galactosidase (β-Gal) Activity Assay
2.5. Subcellular Localization Analysis and Bimolecular Fluorescence Complementation (BiFC) Assay
2.6. MG132 Treatment and Western Blot Analysis
3. Results
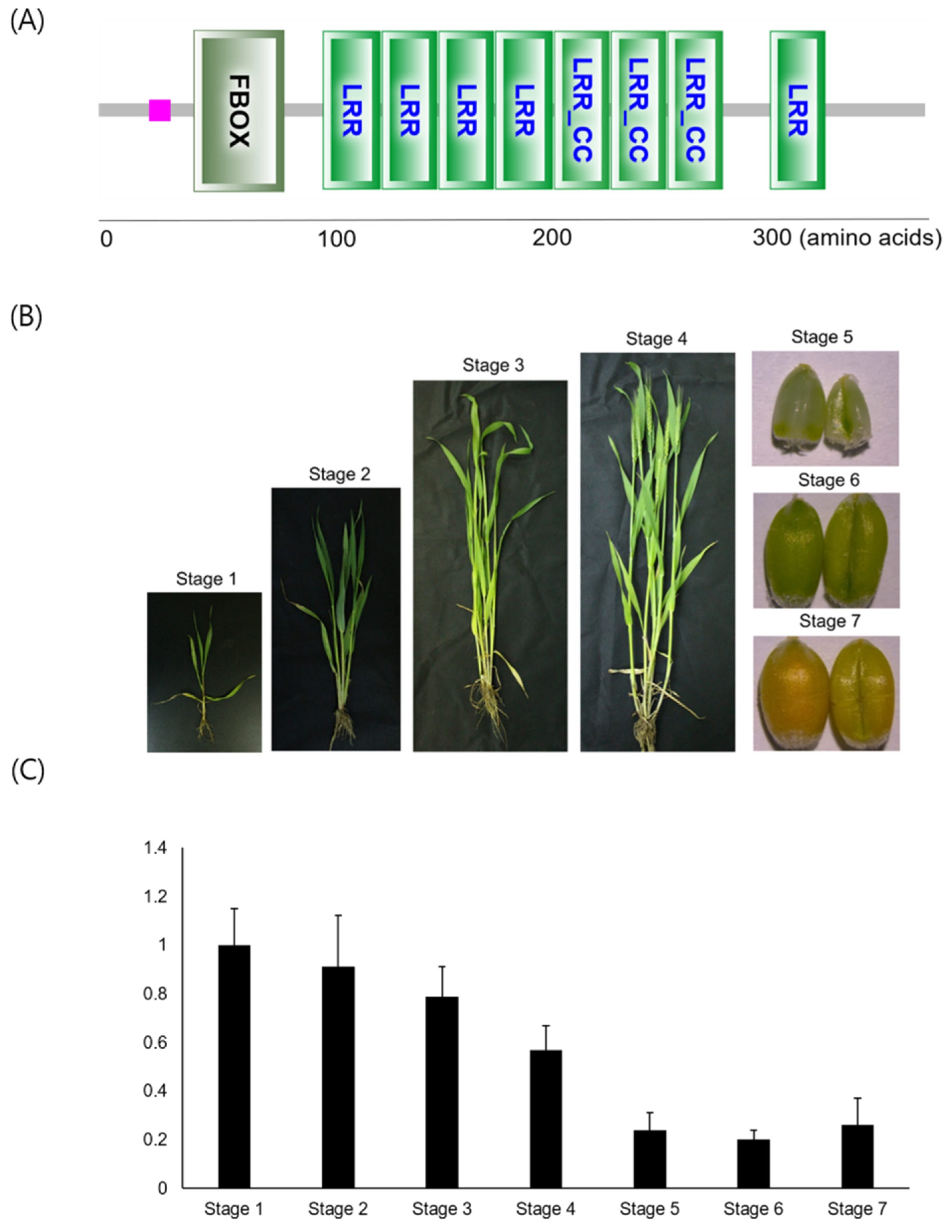
3.1. TaFBXL Isolation and RNA Accumulation Patterns
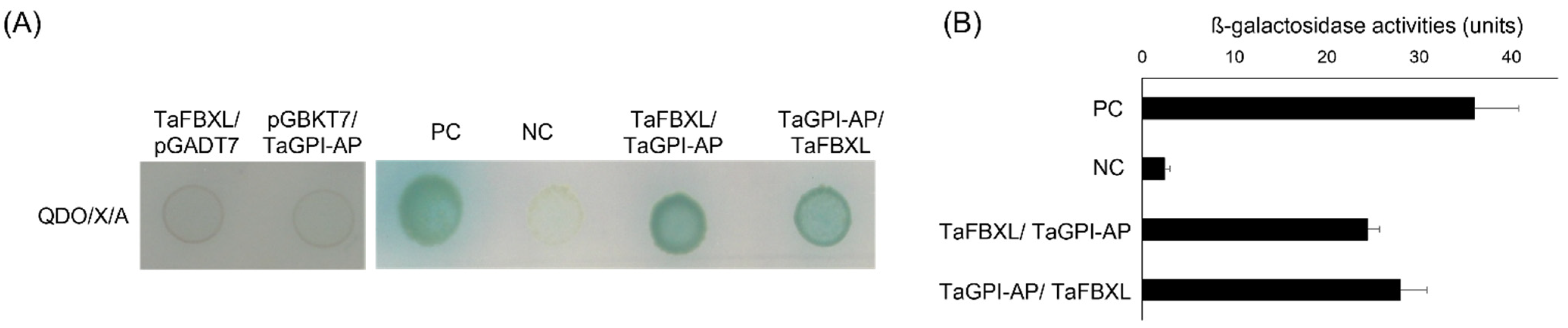
3.2. TaFBXL Is a Component of the SCF Complex
3.3. TaFBXL Interacts with TaGPI-AP
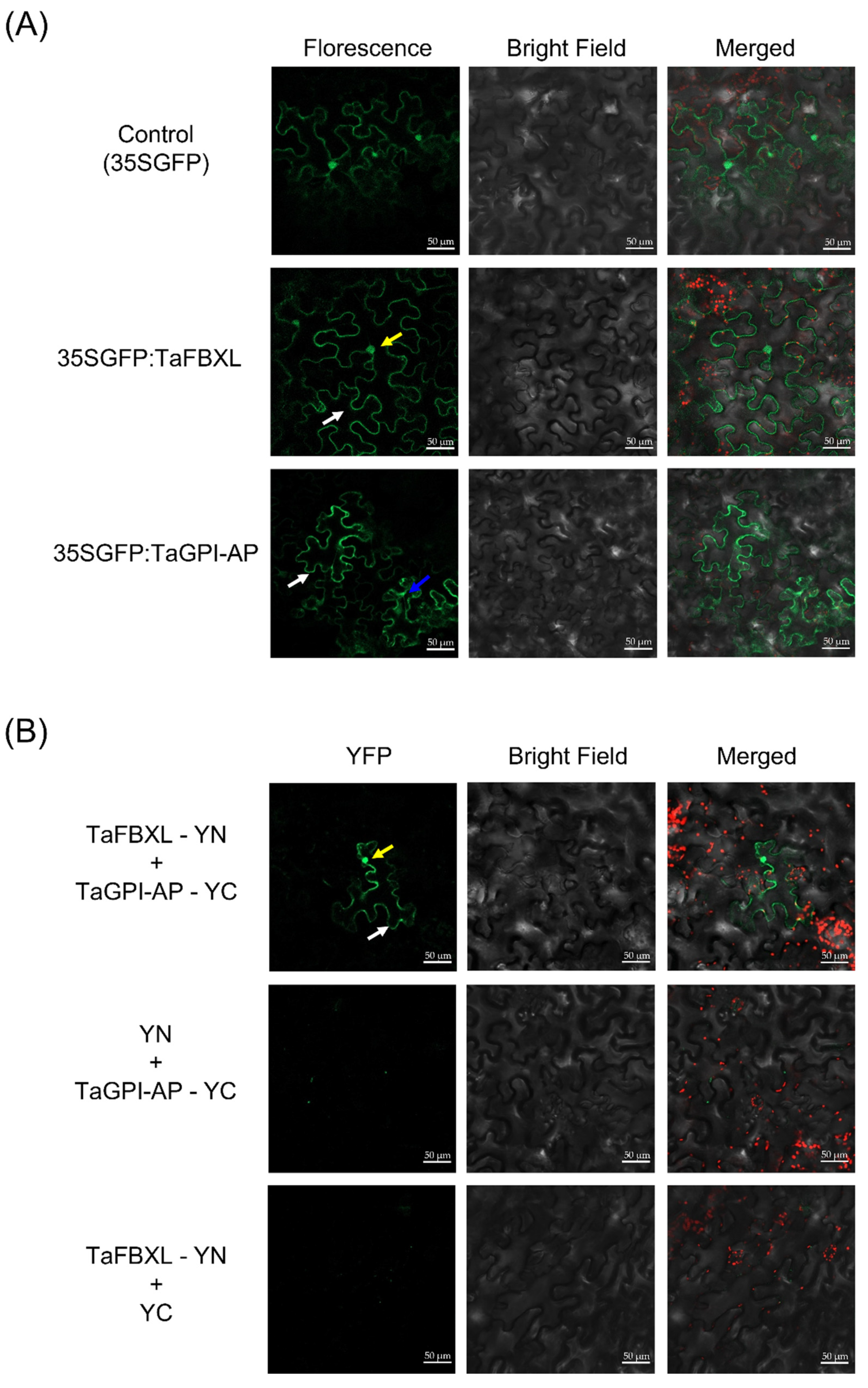
3.4. Subcellular Localization and Interaction between TaFBXL and TaGPI-AP
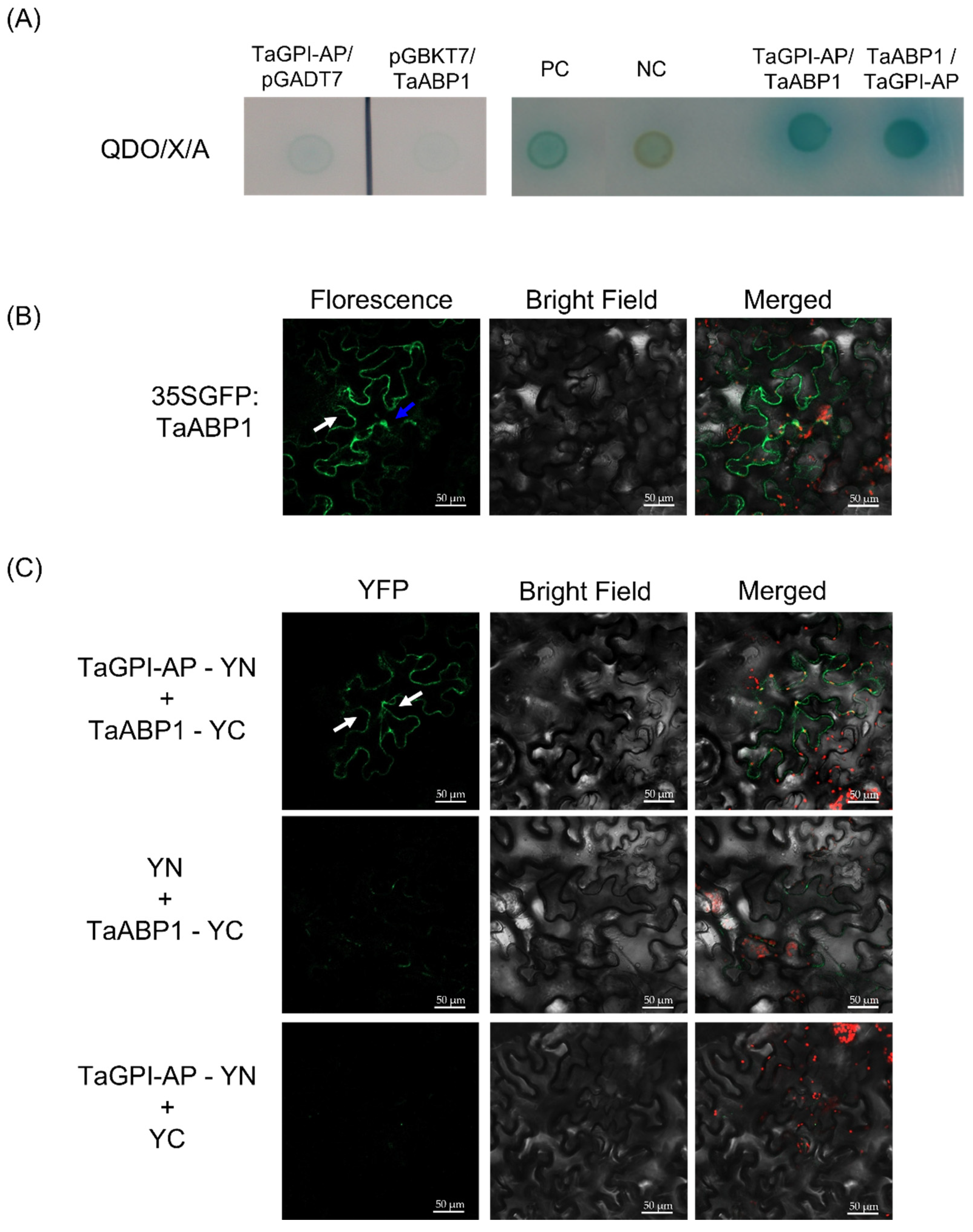
3.5. TaGPI-AP Interacts with TaABP1
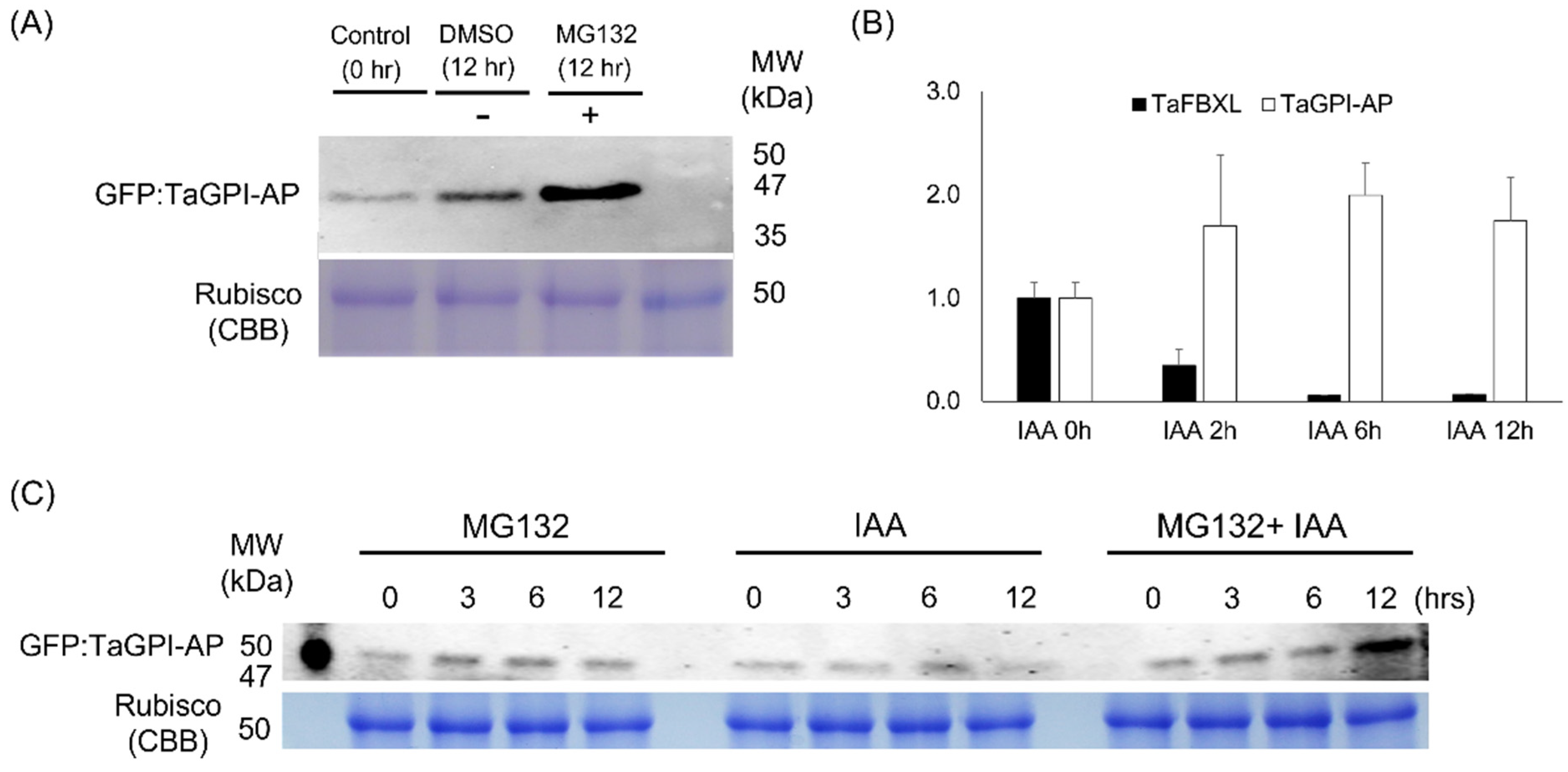
3.6. Degradation of TaGPI-AP in 26S Proteasome
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Sharma, B.; Joshi, D.; Yadav, P.K.; Gupta, A.K.; Bhatt, T.K. Role of ubiquitin-mediated degradation system in plant biology. Front. Plant Sci. 2016, 7, 806. [Google Scholar] [CrossRef]
- Dielen, A.S.; Badaoui, S.; Candresse, T.; German-Retana, S. The ubiquitin/26S proteasome system in plant–pathogen interactions: A never-ending hide-and-seek game. Mol. Plant Pathol. 2010, 11, 293–308. [Google Scholar] [CrossRef]
- Kraft, E.; Stone, S.L.; Ma, L.; Su, N.; Gao, Y.; Lau, O.S.; Deng, X.W.; Callis, J. Genome analysis and functional characterization of the E2 and RING-type E3 ligase ubiquitination enzymes of Arabidopsis. Plant Physiol. 2005, 139, 1597–1611. [Google Scholar] [CrossRef]
- Vierstra, R.D. The ubiquitin–26S proteasome system at the nexus of plant biology. Nat. Rev. Mol. Cell Biol. 2009, 10, 385. [Google Scholar] [CrossRef]
- Chen, L.; Hellmann, H. Plant E3 ligases: Flexible enzymes in a sessile world. Mol. Plant 2013, 6, 1388–1404. [Google Scholar] [CrossRef]
- Gray, W.M.; Hellmann, H.; Dharmasiri, S.; Estelle, M. Role of the Arabidopsis RING-H2 protein RBX1 in RUB modification and SCF function. Plant Cell 2002, 14, 2137–2144. [Google Scholar] [CrossRef]
- Xu, G.; Ma, H.; Nei, M.; Kong, H. Evolution of F-box genes in plants: Different modes of sequence divergence and their relationships with functional diversification. Proc. Natl. Acad. Sci. USA 2009, 106, 835–840. [Google Scholar] [CrossRef] [PubMed]
- Majee, M.; Kumar, S.; Kathare, P.K.; Wu, S.; Gingerich, D.; Nayak, N.R.; Salaita, L.; Dinkins, R.; Martin, K.; Goodin, M. KELCH F-BOX protein positively influences Arabidopsis seed germination by targeting PHYTOCHROME-INTERACTING FACTOR1. Proc. Natl. Acad. Sci. USA 2018, 115, E4120–E4129. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.M.; Yin, H.; Qiao, X.; Tan, X.; Gu, C.; Wang, B.H.; Cheng, R.; Wang, Y.Z.; Zhang, S.L. F-box genes: Genome-wide expansion, evolution and their contribution to pollen growth in pear (Pyrus bretschneideri). Plant Sci. 2016, 253, 164–175. [Google Scholar] [CrossRef] [PubMed]
- Bu, Q.; Lv, T.; Shen, H.; Luong, P.; Wang, J.; Wang, Z.; Huang, Z.; Xiao, L.; Engineer, C.; Kim, T.H. Regulation of drought tolerance by the F-box protein MAX2 in Arabidopsis. Plant Physiol. 2014, 164, 424–439. [Google Scholar] [CrossRef]
- Zhao, Z.; Zhang, G.; Zhou, S.; Ren, Y.; Wang, W. The improvement of salt tolerance in transgenic tobacco by overexpression of wheat F-box gene TaFBA1. Plant Sci. 2017, 259, 71–85. [Google Scholar] [CrossRef]
- Abel, S.; Oeller, P.W.; Theologis, A. Early auxin-induced genes encode short-lived nuclear proteins. Proc. Natl. Acad. Sci. USA 1994, 91, 326–330. [Google Scholar] [CrossRef] [PubMed]
- Song, J.B.; Wang, Y.X.; Li, H.B.; Li, B.W.; Zhou, Z.S.; Gao, S.; Yang, Z.M. The F-box family genes as key elements in response to salt, heavy mental, and drought stresses in Medicago truncatula. Funct. Integr. Genomic. 2015, 15, 495–507. [Google Scholar] [CrossRef]
- Jia, Q.; Xiao, Z.X.; Wong, F.L.; Sun, S.; Liang, K.J.; Lam, H.M. Genome-wide analyses of the soybean F-box gene family in response to salt stress. Int. J. Mol. Sci. 2017, 18, 818. [Google Scholar] [CrossRef] [PubMed]
- Dill, A.; Thomas, S.G.; Hu, J.; Steber, C.M.; Sun, T. The Arabidopsis F-box protein SLEEPY1 targets gibberellin signaling repressors for gibberellin-induced degradation. Plant Cell 2004, 16, 1392–1405. [Google Scholar] [CrossRef]
- Kepinski, S.; Leyser, O. The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 2005, 435, 446–451. [Google Scholar] [CrossRef] [PubMed]
- Binder, B.M.; Walker, J.M.; Gagne, J.M.; Emborg, T.J.; Hemmann, G.; Bleecker, A.B.; Vierstra, R.D. The Arabidopsis EIN3 binding F-Box proteins EBF1 and EBF2 have distinct but overlapping roles in ethylene signaling. Plant Cell 2007, 19, 509–523. [Google Scholar] [CrossRef]
- Villalobos, L.I.A.C.; Lee, S.; De Oliveira, C.; Ivetac, A.; Brandt, W.; Armitage, L.; Sheard, L.B.; Tan, X.; Parry, G.; Mao, H. A combinatorial TIR1/AFB–Aux/IAA co-receptor system for differential sensing of auxin. Nat. Chem. Biol. 2012, 8, 477. [Google Scholar] [CrossRef]
- Hertel, R.; Thomson, K.S.; Russo, V. In-vitro auxin binding to particulate cell fractions from corn coleoptiles. Planta 1972, 107, 325–340. [Google Scholar] [CrossRef]
- Löbler, M.; Klämbt, D. Auxin-binding protein from coleoptile membranes of corn (Zea mays L.): I. Purification by immunological methods and characterization. J. Biol. Chem. 1985, 260, 9848–9853. [Google Scholar] [CrossRef]
- Robert, S.; Kleine-Vehn, J.; Barbez, E.; Sauer, M.; Paciorek, T.; Baster, P.; Vanneste, S.; Zhang, J.; Simon, S.; Čovanová, M. ABP1 mediates auxin inhibition of clathrin-dependent endocytosis in Arabidopsis. Cell 2010, 143, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yang, Z. Novel ABP1-TMK auxin sensing system controls ROP GTPase-mediated interdigitated cell expansion in Arabidopsis. Small GTPases 2014, 30, e29711. [Google Scholar] [CrossRef]
- Shimomura, S. Identification of a glycosylphosphatidylinositol-anchored plasma membrane protein interacting with the C-terminus of auxin-binding protein 1: A photoaffinity crosslinking study. Plant Mol. Biol. 2006, 60, 663–677. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.H.; Yang, Z.B. Is ABP1 an auxin receptor yet? Mol. Plant 2011, 4, 635–640. [Google Scholar] [CrossRef]
- Hong, M.J.; Kim, J.-B.; Seo, Y.W.; Kim, D.Y. F-box genes in the wheat genome and expression profiling in wheat at different developmental stages. Genes 2020, 11, 1154. [Google Scholar] [CrossRef]
- Zadoks, J.C.; Chang, T.T.; Konzak, C.F. A decimal code for the growth stages of cereals. Weed Res. 1974, 14, 415–421. [Google Scholar] [CrossRef]
- Cai, T.; Meng, X.; Liu, X.; Liu, T.; Wang, H.; Jia, Z.; Yang, D.; Ren, X. Exogenous hormonal application regulates the occurrence of wheat tillers by changing endogenous hormones. Front. Plant Sci. 2018, 9, 1886. [Google Scholar] [CrossRef] [PubMed]
- Su, P.; Yan, J.; Li, W.; Wang, L.; Zhao, J.; Ma, X.; Wang, H.; Kong, L. A member of wheat class III peroxidase gene family, TaPRX-2A, enhanced the tolerance of salt stress. BMC Plant Biol. 2020, 20, 392. [Google Scholar] [CrossRef]
- Curtis, M.D.; Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 2003, 133, 462–469. [Google Scholar] [CrossRef]
- Hong, M.J.; Kim, D.Y.; Choi, H.; Seo, Y.W.; Kim, J.-B. Isolation and characterization of kelch repeat-containing F-box proteins from colored wheat. Mol. Biol. Rep. 2020, 47, 1129–1141. [Google Scholar] [CrossRef]
- Miller, J. Assay of B-Galactosidase: Experiments in Molecular Genetics; Cold Spring Harbor Laboratory Press: Long Island, NY, USA, 1972. [Google Scholar]
- Walter, M.; Chaban, C.; Schütze, K.; Batistic, O.; Weckermann, K.; Näke, C.; Blazevic, D.; Grefen, C.; Schumacher, K.; Oecking, C. Visualization of protein interactions in living plant cells using bimolecular fluorescence complementation. Plant J. 2004, 40, 428–438. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, D493–D496. [Google Scholar] [CrossRef] [PubMed]
- Hermand, D. F-box proteins: More than baits for the SCF? Cell Div. 2006, 1, 30. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hong, M.J.; Kim, D.Y.; Kang, S.Y.; Kim, D.S.; Kim, J.B.; Seo, Y.W. Wheat F-box protein recruits proteins and regulates their abundance during wheat spike development. Mol. Biol. Rep. 2012, 39, 9681–9696. [Google Scholar] [CrossRef] [PubMed]
- Hong, M.J.; Kim, D.Y.; Seo, Y.W. SKP1-like-related genes interact with various F-box proteins and may form SCF complexes with Cullin-F-box proteins in wheat. Mol. Biol. Rep. 2013, 40, 969–981. [Google Scholar] [CrossRef] [PubMed]
- Yeats, T.H.; Bacic, A.; Johnson, K.L. Plant glycosylphosphatidylinositol anchored proteins at the plasma membrane-cell wall nexus. J. Integr. Plant Biol. 2018, 60, 649–669. [Google Scholar] [CrossRef]
- Zhou, K. Glycosylphosphatidylinositol-anchored proteins in Arabidopsis and one of their common roles in signaling transduction. Front. Plant Sci. 2019, 10, 1022. [Google Scholar] [CrossRef]
- Jain, M.; Nijhawan, A.; Arora, R.; Agarwal, P.; Ray, S.; Sharma, P.; Kapoor, S.; Tyagi, A.K.; Khurana, J.P. F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol. 2007, 143, 1467–1483. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Gonzalez-Carranza, Z.H.; Zhang, S.; Miao, Y.; Liu, C.J.; Roberts, J.A. F-Box Proteins in Plants. Annu. Plant Rev. 2018, 2, 1–21. [Google Scholar]
- Kipreos, E.T.; Pagano, M. The F-box protein family. Genome Biol. 2000, 1, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Gagne, J.M.; Downes, B.P.; Shiu, S.H.; Durski, A.M.; Vierstra, R.D. The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc. Natl. Acad. Sci. USA 2002, 99, 11519–11524. [Google Scholar] [CrossRef]
- Hellmann, H.; Estelle, M. Plant development: Regulation by protein degradation. Science 2002, 297, 793–797. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, H.; Takahashi, N.; Shimada, H.; Seki, M.; Shinozaki, K.; Matsui, M. Classification and expression analysis of Arabidopsis F-box-containing protein genes. Plant Cell Physiol. 2002, 43, 1073–1085. [Google Scholar] [CrossRef] [PubMed]
- Kajava, A. Structural diversity of leucine-rich repeat proteins. J. Mol. Biol. 1998, 277, 519–527. [Google Scholar] [CrossRef]
- Kobe, B.; Kajava, A.V. The leucine-rich repeat as a protein recognition motif. Curr. Opin. Struct. Biol. 2001, 11, 725–732. [Google Scholar] [CrossRef]
- Forsthoefel, N.R.; Cutler, K.; Port, M.D.; Yamamoto, T.; Vernon, D.M. PIRLs: A novel class of plant intracellular leucine-rich repeat proteins. Plant Cell Physiol. 2005, 46, 913–922. [Google Scholar] [CrossRef] [PubMed]
- Teale, W.D.; Paponov, I.A.; Palme, K. Auxin in action: Signalling, transport and the control of plant growth and development. Nat. Rev. Mol. Cell Biol. 2006, 7, 847–859. [Google Scholar] [CrossRef]
- Löbler, M.; Klämbt, D. Auxin-binding protein from coleoptile membranes of corn (Zea mays L.): II. Localization of a putative auxin receptor. J. Biol. Chem. 1985, 260, 9854–9859. [Google Scholar] [CrossRef]
- Napier, R.M.; David, K.M.; Perrot-Rechenmann, C. A short history of auxin-binding proteins. Plant Mol. Biol. 2002, 49, 339–348. [Google Scholar] [CrossRef]
- Chen, X.; Naramoto, S.; Robert, S.; Tejos, R.; Löfke, C.; Lin, D.; Yang, Z.; Friml, J. ABP1 and ROP6 GTPase signaling regulate clathrin-mediated endocytosis in Arabidopsis roots. Curr. Biol. 2012, 22, 1326–1332. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Grandont, L.; Li, H.; Hauschild, R.; Paque, S.; Abuzeineh, A.; Rakusová, H.; Benkova, E.; Perrot-Rechenmann, C.; Friml, J. Inhibition of cell expansion by rapid ABP1-mediated auxin effect on microtubules. Nature 2014, 516, 90–93. [Google Scholar] [CrossRef]
- Kinoshita, T. Glycosylphosphatidylinositol (GPI) anchors: Biochemistry and cell biology: Introduction to a thematic review series. J. Lipid Res. 2016, 57, 4–5. [Google Scholar] [CrossRef]
- Stevens, V.L. Biosynthesis of glycosylphosphatidylinositol membrane anchors. Biochem. J. 1995, 310, 361. [Google Scholar] [CrossRef] [PubMed]
- Eisenhaber, B.; Bork, P.; Eisenhaber, F. Sequence properties of GPI-anchored proteins near the omega-site: Constraints for the polypeptide binding site of the putative transamidase. Protein Eng. 1998, 11, 1155–1161. [Google Scholar] [CrossRef]
- Xu, T.; Dai, N.; Chen, J.; Nagawa, S.; Cao, M.; Li, H.; Zhou, Z.; Chen, X.; De Rycke, R.; Rakusová, H. Cell surface ABP1-TMK auxin-sensing complex activates ROP GTPase signaling. Science 2014, 343, 1025–1028. [Google Scholar] [CrossRef] [PubMed]
- Cao, M.; Chen, R.; Li, P.; Yu, Y.; Zheng, R.; Ge, D.; Zheng, W.; Wang, X.; Gu, Y.; Gelová, Z. TMK1-mediated auxin signalling regulates differential growth of the apical hook. Nature 2019, 568, 240–243. [Google Scholar] [CrossRef]
- Gao, Y.; Zhang, Y.; Zhang, D.; Dai, X.; Estelle, M.; Zhao, Y. Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development. Proc. Natl. Acad. Sci. USA 2015, 112, 2275–2280. [Google Scholar] [CrossRef]
- Paponov, I.A.; Dindas, J.; Król, E.; Friz, T.; Budnyk, V.; Teale, W.; Paponov, M.; Hedrich, R.; Palme, K. Auxin-induced plasma membrane depolarization is regulated by auxin transport and not by AUXIN BINDING PROTEIN1. Front. Plant Sci. 2019, 9, 1953. [Google Scholar] [CrossRef] [PubMed]
- Zhou, K.; Fiume, E.; De-Vos, D.; Magnani, E. GPI-anchored SKU5/SKS are maternally required for integument development in Arabidopsis. Genome Biol. 2000. [Google Scholar]

| Clone No. | Putative Identification | Organism | e-Value |
|---|---|---|---|
| 25, 28 | WESR2 (wheat early salt-stress responding gene 2) | Triticum aestivum | 0.00 |
| 34 | Signal recognition particle 9-kDa protein | Aegilops tauschii | 0.00 |
| 38 | Chlorophyll a-b binding protein of LHCII type 1 | Aegilops tauschii | 0.00 |
| 42 | 40S ribosomal protein S3a | Aegilops tauschii | 0.00 |
| 45 | Glycosylphosphatidylinositol (GPI)-anchored proteins, At3g06035-like | Aegilops tauschii | 0.00 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hong, M.J.; Kim, J.-B.; Seo, Y.W.; Kim, D.Y. Regulation of Glycosylphosphatidylinositol-Anchored Protein (GPI-AP) Expression by F-Box/LRR-Repeat (FBXL) Protein in Wheat (Triticum aestivum L.). Plants 2021, 10, 1606. https://doi.org/10.3390/plants10081606
Hong MJ, Kim J-B, Seo YW, Kim DY. Regulation of Glycosylphosphatidylinositol-Anchored Protein (GPI-AP) Expression by F-Box/LRR-Repeat (FBXL) Protein in Wheat (Triticum aestivum L.). Plants. 2021; 10(8):1606. https://doi.org/10.3390/plants10081606
Chicago/Turabian StyleHong, Min Jeong, Jin-Baek Kim, Yong Weon Seo, and Dae Yeon Kim. 2021. "Regulation of Glycosylphosphatidylinositol-Anchored Protein (GPI-AP) Expression by F-Box/LRR-Repeat (FBXL) Protein in Wheat (Triticum aestivum L.)" Plants 10, no. 8: 1606. https://doi.org/10.3390/plants10081606
APA StyleHong, M. J., Kim, J.-B., Seo, Y. W., & Kim, D. Y. (2021). Regulation of Glycosylphosphatidylinositol-Anchored Protein (GPI-AP) Expression by F-Box/LRR-Repeat (FBXL) Protein in Wheat (Triticum aestivum L.). Plants, 10(8), 1606. https://doi.org/10.3390/plants10081606







