Molecular Cytogenetic Identification of the Wheat–Dasypyrum villosum T3DL·3V#3S Translocation Line with Resistance against Stripe Rust
Abstract
:1. Introduction
2. Results
2.1. Generation and Screening for Wheat–D. villosum 3V#3.3D Translocation Stocks
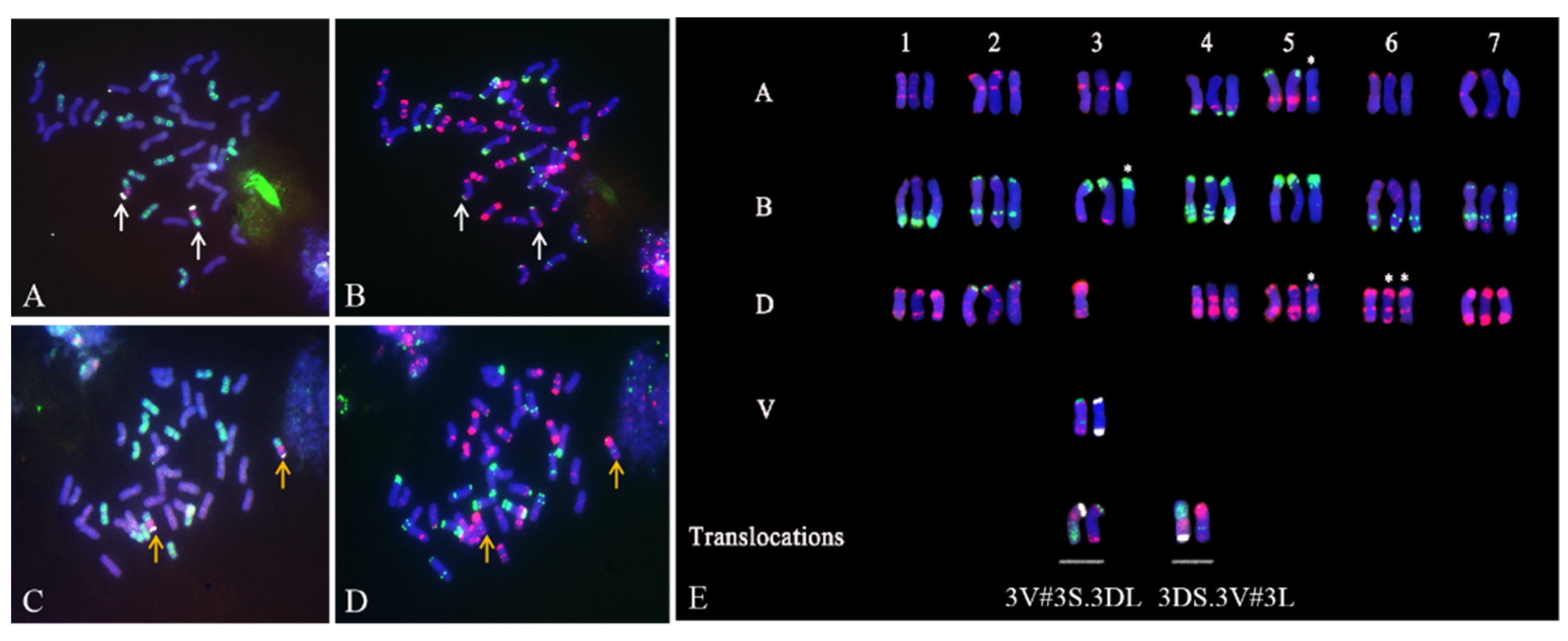
2.2. Identification of T. aestivum–D. villosum Structural Aberrations by ND-FISH
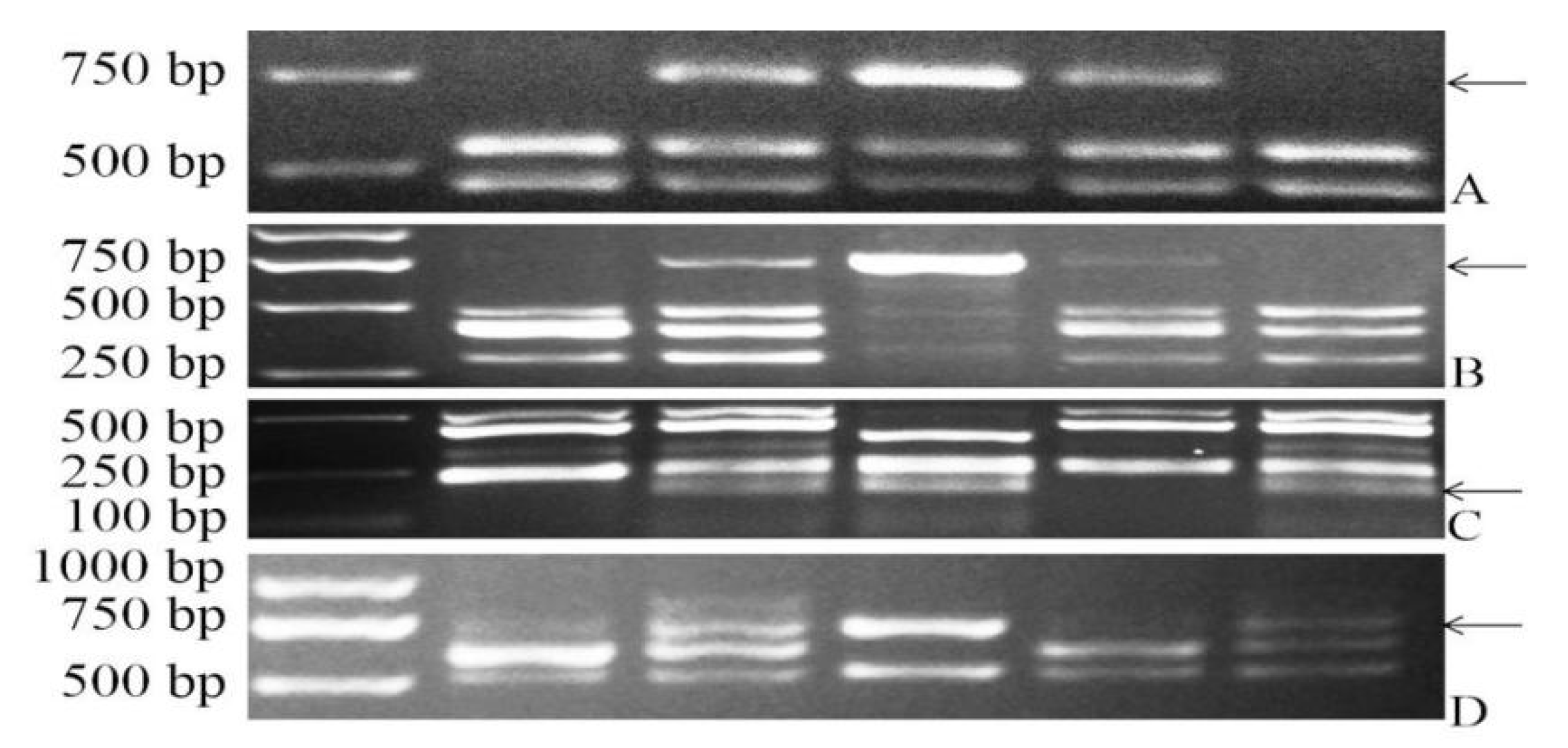
2.3. Molecular Marker Analysis
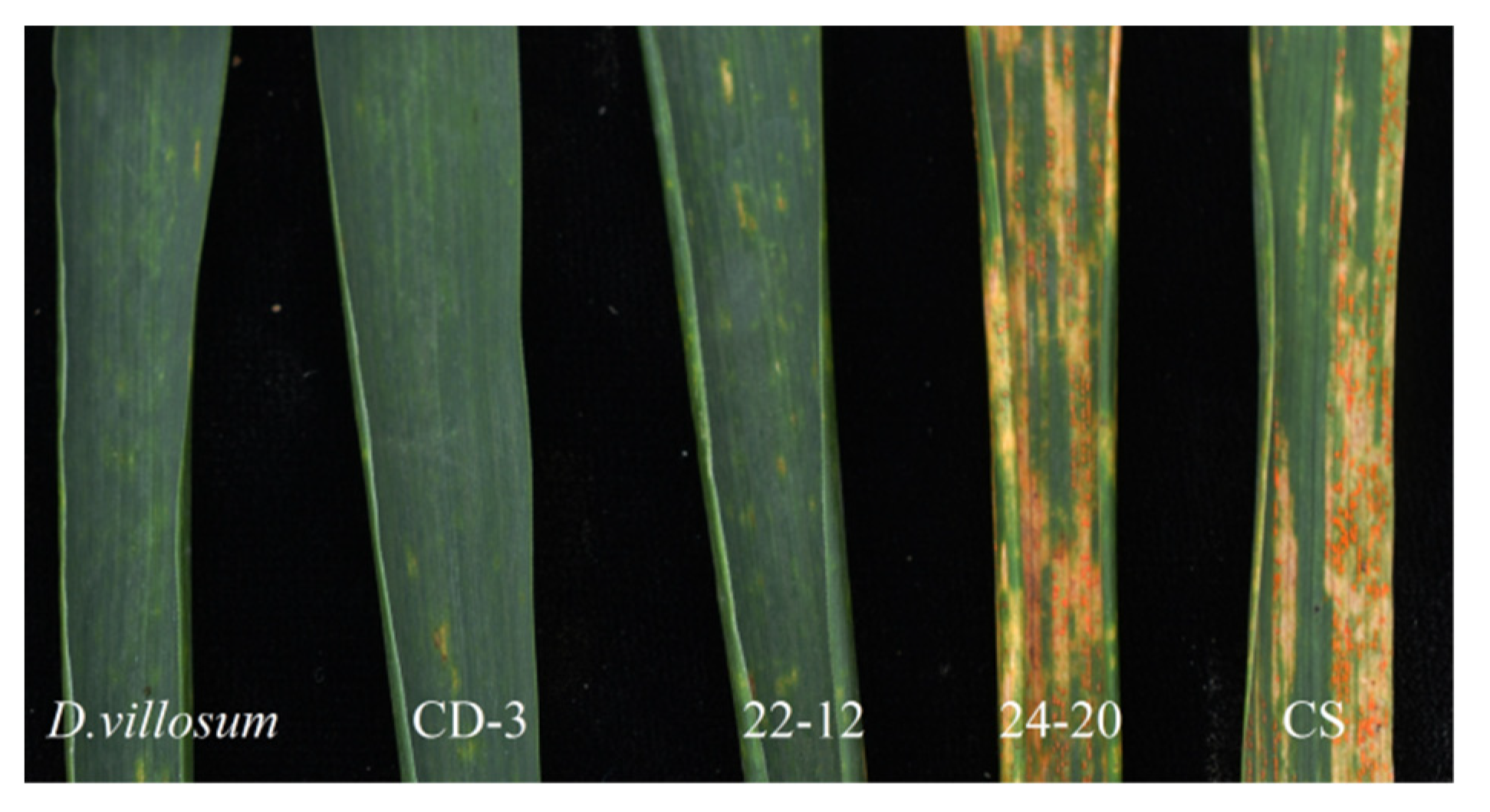
2.4. Stripe Rust Response
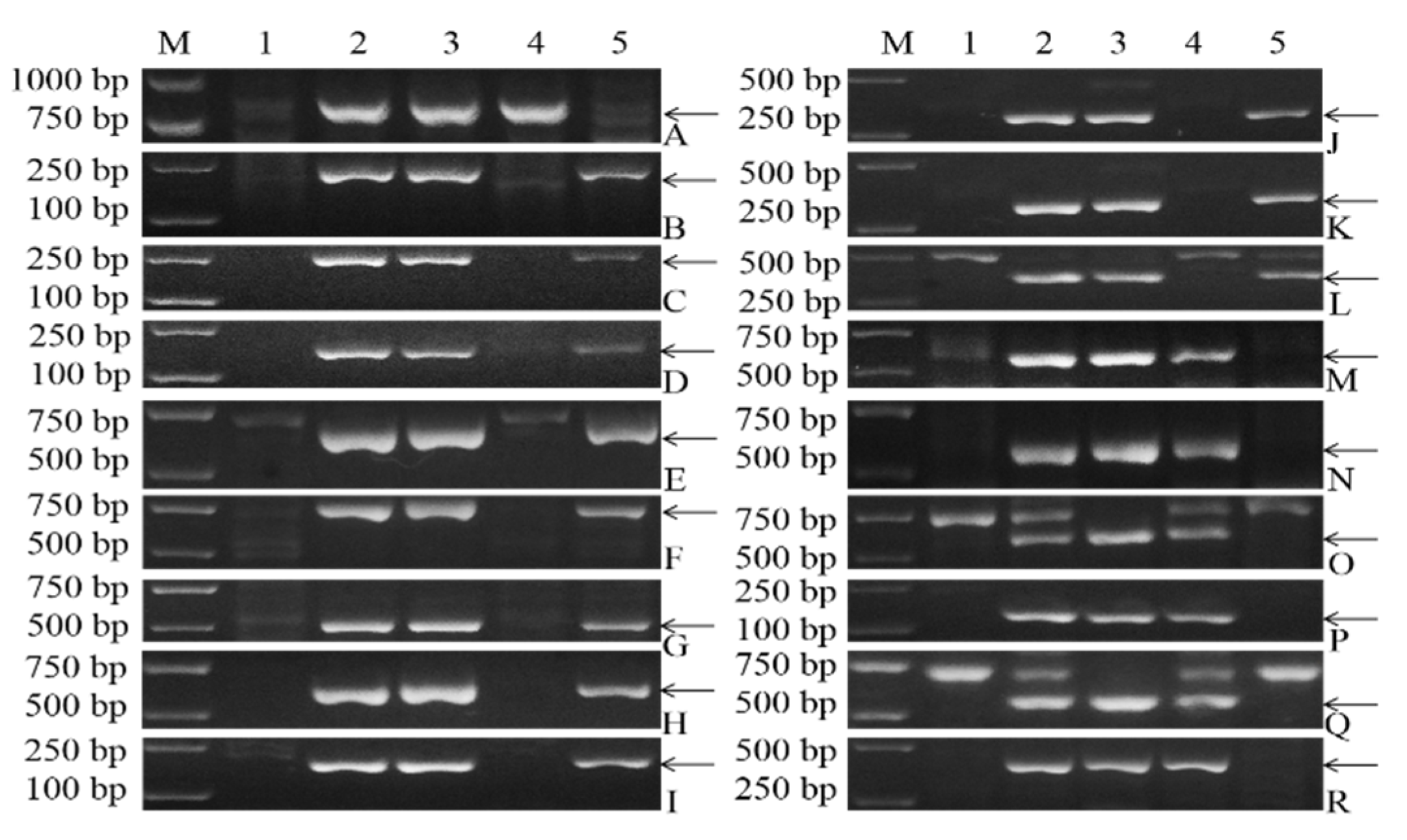
2.5. Development and Verification of 3V#3-Specific EST Markers
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. 60Co γ-Irradiation Treatment of CD-3 Seeds
4.3. Mitotic Analysis and Survival Rate Investigation of the M0 Generation
4.4. Statistics Analysis
4.5. FISH Analysis
4.6. Molecular Marker Analysis
4.7. Assessment of Adult Plant Resistance to Stripe Rust
4.8. RNA-Seq and Transcriptome Assembly
4.9. Development and Validation of 3V#3 Chromosome-Specific EST Markers
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, X. Review Article: High-Temperature Adult-Plant Resistance, Key for Sustainable Control of Stripe Rust. Am. J. Plant Sci. 2013, 4, 608–627. [Google Scholar] [CrossRef] [Green Version]
- Zadoks, T.C. Epidemiology of Wheat Rust in Europe. Glob. Chang. Bio. 1967, 13, 29–46. [Google Scholar] [CrossRef]
- Milus, E.A.; Kristensen, K.; Hovmøller, M.S. Evidence for increased aggressiveness in a recent widespread strain of Puccinia striiformis f. sp. Tritici causing stripe rust of wheat. Phytopathology 2009, 99, 89–94. [Google Scholar] [PubMed] [Green Version]
- Hovmøller, M.S.; Walter, S.; Justesen, A.F. Escalating Threat of Wheat Rusts. Science 2010, 329, 369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wellings, C.R. Global status of stripe rust: A review of historical and current threats. Euphytica 2011, 179, 129–141. [Google Scholar] [CrossRef]
- Li, J.; Dundas, I.; Dong, C.; Li, G.; Trethowan, R.; Yang, Z.; Hoxha, S.; Zhang, P. Identification and characterization of a new stripe rust resistance gene Yr83 on rye chromosome 6R in wheat. Theor. Appl. Genet. 2020, 133, 1095–1107. [Google Scholar] [CrossRef]
- McIntosh, R.A.; Mu, J.M.; Han, D.J.; Kang, Z.S. Wheat stripe rust resistance gene Yr24/Yr26: A retrospective review. Crop J. 2018, 6, 3–11. [Google Scholar] [CrossRef]
- Liu, T.G.; Peng, Y.L.; Chen, W.Q.; Zhang, Z.Y. First detection of virulence in Puccinia striiformis f. sp. Tritici in China to re-sistance genes Yr24 (=Yr26) present in wheat cultivar Chuanmai 42. Plant Dis. 2010, 94, 1163. [Google Scholar] [CrossRef]
- Zeng, Q.-D.; Han, D.-J.; Wang, Q.-L.; Yuan, F.-P.; Wu, J.; Zhang, L.; Wang, X.-J.; Huang, L.-L.; Chen, X.-M.; Kang, Z.-S. Stripe rust resistance and genes in Chinese wheat cultivars and breeding lines. Euphytica 2013, 196, 271–284. [Google Scholar] [CrossRef] [Green Version]
- Danilova, T.V.; Zhang, G.; Liu, W.; Friebe, B.; Gill, B.S. Homoeologous recombination-based transfer and molecular cytogenetic mapping of a wheat streak mosaic virus and Triticum mosaic virus resistance gene Wsm3 from Thinopyrum intermedium to wheat. Theor. Appl. Genet. 2016, 130, 549–556. [Google Scholar] [CrossRef]
- Zhang, R.; Feng, Y.; Li, H.; Yuan, H.; Dai, J.; Cao, A.; Xing, L.; Li, H. Cereal cyst nematode resistance gene CreV effective against Heterodera filipjevi transferred from chromosome 6VL of Dasypyrum villosum to bread wheat. Mol. Breed. 2016, 36, 122. [Google Scholar] [CrossRef]
- Zhang, R.Q.; Sun, B.X.; Chen, J.; Cao, A.Z.; Xing, L.P.; Feng, Y.G.; Lan, C.X.; Chen, P.D. Pm55, a developmental-stage and tissue-specifc powdery mildew resistance gene introgressed from Dasypyrum villosum into common wheat. Theor. Appl. Genet. 2016, 129, 1975–1984. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Si, B.; Zhu, G.; Xu, X.; Li, W.; Chen, S.; Zhao, J.; Li, T. Race and virulence of asexual and sexual populations of Puccinia graminis f. sp. Tritici in China from 2009 to 2015. Eur. J. Plant Pathol. 2018, 153, 545–555. [Google Scholar] [CrossRef]
- Zhao, X.L.; Zheng, T.C.; Xia, X.C.; He, Z.H.; Liu, D.Q.; Yang, W.X.; Yin, G.H.; Li, Z.F. Molecular mapping of leaf rust resistance gene LrZH84 in Chinese wheat line Zhou 8425B. Theor. Appl. Genet. 2008, 117, 1069–1075. [Google Scholar] [CrossRef] [PubMed]
- De Pace, C.; Montebove, L.; Delre, V.; Jan, C.C.; Qualset, C.O.; Scarascia Mugnozza, G.T. Biochemical versatility of amphiploids derived from crossing Dasypyrum villosum (L.) Candargy and wheat: Genetic control and phenotypical aspects. Theor. Appl. Genet. 1988, 76, 513–529. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.D.; Qi, L.L.; Zhou, B.; Zhang, S.Z.; Liu, D.J. Development and molecular cytogenetic analysis of wheat-Haynaldia villosa 6VS/6AL translocation lines specifying resistance to powdery mildew. Theor. Appl. Genet. 1995, 91, 1125–1128. [Google Scholar] [CrossRef] [PubMed]
- Yildirim, A.; Jones, S.S.; Murray, T.; Line, R.F. Evaluation of Dasypyrum villosum Populations for Resistance to Cereal Eyespot and Stripe Rust Pathogens. Plant Dis. 2000, 84, 40–44. [Google Scholar] [CrossRef] [Green Version]
- Zhang, R.; Fan, Y.; Kong, L.; Wang, Z.; Wu, J.; Xing, L.; Cao, A.; Feng, Y. Pm62, an adult-plant powdery mildew resistance gene introgressed from Dasypyrum villosum chromosome arm 2VL into wheat. Theor. Appl. Genet. 2018, 131, 2613–2620. [Google Scholar] [CrossRef]
- McFadden, E.S.; Sears, E.R. The Genome Approach in Radical Wheat Breeding. Agron. J. 1947, 39, 1011–1025. [Google Scholar] [CrossRef]
- Sears, E.R. Addition of the genome of Haynaldia villosa to Triticum aestivum. Am. J. Bot. 1953, 40, 168–174. [Google Scholar] [CrossRef]
- Liu, D.J.; Chen, P.D.; Pei, G.Z.; Wang, Y.N.; Qiu, B.X.; Wang, S.L. Transfer of Haynaldia villosa chromosomes into Triticum aestivum. In Proceedings of the 7th International Wheat Genet Symposium, Cambridge, UK, 13–19 July 1988; Miller, T.E., Koebner, R.M.D., Eds.; Institute of Plant Science Research, Cambridge Laboratory: Cambridge, UK, 1988; pp. 355–361. [Google Scholar]
- Lukaszewski, A.J.; Cowger, C. Re-Engineering of the Pm21 transfer from Haynaldia villosa to bread wheat by induced homoeologous Recombination. Crop. Sci. 2017, 57, 2590–2594. [Google Scholar] [CrossRef]
- Zhao, W.C.; Qi, L.L.; Gao, X.; Zhang, G.S.; Dong, J.; Chen, Q.J.; Friebe, B.; Gill, B.S. Development and characterization of two new Triticum aestivum-Dasypyrum villosum Robertsonian translocation lines T1DS·1V#3L and T1DL·1V#3S and their effect on grain quality. Euphytica 2010, 175, 343–350. [Google Scholar]
- Chen, P.D.; Liu, D.J. Identification of Haynaldia villosa chromosomes in wheat alien addition lines. In Proceedings of the 1st International Symposium on Chromosome Engineering in Plants, Xian, China, 20–25 October 1986; Zhensheng, L., Swaminathan, M.S., Eds.; pp. 31–32. [Google Scholar]
- Zhang, R.Q.; Zhang, M.Y.; Wang, X.E.; Chen, P.D. Introduction of chromosome segment carrying the seed storage protein genes from chromosome 1V of Dasypyrum villosum showed positive effect on bread-making quality of common wheat. Theor. Appl. Genet. 2014, 127, 523–533. [Google Scholar]
- Qi, L.L.; Wang, S.L.; Chen, P.D.; Liu, D.J.; Gill, B.S. Identification and physical mapping of three Haynaldia villosa chromo-some-6V deletion lines. Theor. Appl. Genet. 1998, 97, 1042–1046. [Google Scholar] [CrossRef]
- He, H.G.; Zhu, S.Y.; Zhao, R.H.; Jiang, Z.N.; Ji, Y.Y.; Ji, J.; Liu, D.; Li, H.J.; Bie, T.D. Pm21, encoding a typical CC-NBS-LRR protein, confers broad-spectrum resistance to wheat powdery mildew disease. Mol. Plant. 2018, 11, 879–882. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xing, L.; Hu, P.; Liu, J.; Witek, K.; Zhou, S.; Xu, J.; Zhou, W.; Gao, L.; Huang, Z.; Zhang, R.; et al. Pm21 from Haynaldia villosa Encodes a CC-NBS-LRR Protein Conferring Powdery Mildew Resistance in Wheat. Mol. Plant 2018, 11, 874–878. [Google Scholar] [CrossRef] [Green Version]
- Li, G.R.; Zhao, J.M.; Li, D.H.; Yang, E.N.; Huang, Y.F.; Liu, C.; Yang, Z.J. A novel Wheat-Dasypyrum breviaristatum substitution line with stripe rust resistance. Cytogenet. Genome Res. 2014, 143, 280–287. [Google Scholar] [CrossRef]
- Zhang, J.; Jiang, Y.; Wang, Y.; Guo, Y.; Long, H.; Deng, G.; Chen, Q.; Xuan, P. Molecular markers and cytogenetics to characterize a wheat–Dasypyrum villosum 3V (3D) substitution line conferring resistance to stripe rust. PLoS ONE 2018, 13, e0202033. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Long, H.; Pan, Z.; Liang, J.; Yu, S.; Deng, G.; Yu, M. Characterization of a genome-specific Gypsy-like retrotransposon sequence and development of a molecular marker specific for Dasypyrum villosum (L.). J. Genet. 2013, 92, 103–108. [Google Scholar] [CrossRef]
- Wang, H.; Dai, K.; Jin, X.; Yuan, C.; Zhao, R.; Doležel, J.; Wu, Y.F.; Cao, A.Z.; Chen, P.D.; Zhang, S.Z.; et al. Development of intron targeting (IT) markers specific for chromosome arm 4VS of Haynaldia villosa by chromosome sorting and next-generation sequencing. BMC Genom. 2017, 18, 167. [Google Scholar] [CrossRef] [Green Version]
- Cao, A.Z.; Wang, X.E.; Chen, Y.P.; Zou, X.W.; Chen, P.D. A sequence-specific PCR marker linked with Pm21 distinguishes chromosomes 6AS, 6BS, 6DS of Triticum aestivum and 6VS of Haynaldia villosa. Plant Breed. 2006, 125, 201–205. [Google Scholar] [CrossRef]
- Song, W.; Xie, C.; Du, J.; Xie, H.; Liu, Q.; Ni, Z.; Yang, T.; Sun, Q.; Liu, Z. A “one-marker-for-two-genes” approach for efficient molecular discrimination of Pm12 and Pm21 conferring resistance to powdery mildew in wheat. Mol. Breed. 2008, 23, 357–363. [Google Scholar] [CrossRef]
- Zhang, J.; Liang, S.; Duan, J.; Wang, J.; Chen, S.; Cheng, Z.; Zhang, Q.; Liang, X.; Li, Y. De novo assembly and Characterisation of the Transcriptome during seed development, and generation of genic-SSR markers in Peanut (Arachis hypogaea L.). BMC Genom. 2012, 13, 90. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Druka, A.; Muehlbauer, G.; Druka, I.; Caldo, R.; Baumann, U.; Rostoks, N.; Schreiber, A.; Wise, R.; Close, T.; Kleinhofs, A.; et al. An atlas of gene expression from seed to seed through barley development. Funct. Integr. Genom. 2006, 6, 202–211. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Wang, J.; Wang, K.; Chen, J.; Wang, K.; Du, L.; Ni, Z.; Lin, Z.; Ye, X. Development of PCR markers specific to Dasypyrum villosum genome based on transcriptome data and their application in breeding Triticum aestivum-D. villosum#4 alien chromosome lines. BMC Genom. 2019, 20, 289. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Jiang, Y.; Guo, Y.; Li, G.; Yang, Z.; Xu, D.; Xuan, P. Identification of Novel Chromosomal Aberrations Induced by 60Co-γ-Irradiation in Wheat-Dasypyrum villosum Lines. Int. J. Mol. Sci. 2015, 16, 29787–29796. [Google Scholar] [CrossRef] [Green Version]
- Friebe, B.; Hatchett, J.H.; Mukai, Y.; Gill, B.S.; Sebesta, E.E. Transfer of Hessian fly resistance from rye to wheat via radiation-induced terminal and intercalary chromosomal translocations. Theor. Appl. Genet. 1991, 83, 33–40. [Google Scholar] [CrossRef]
- Tomlekova, N.B. Induced mutagenesis for crop improvement in bulgaria. Plant Mutat. Rep. 2010, 2, 4–27. [Google Scholar]
- Chen, P.; You, C.; Hu, Y.; Chen, S.; Zhou, B.; Cao, A.; Wang, X. Radiation-induced translocations with reduced Haynaldia villosa chromatin at the Pm21 locus for powdery mildew resistance in wheat. Mol. Breed. 2012, 31, 477–484. [Google Scholar] [CrossRef]
- Tang, Z.X.; Yang, Z.J.; Fu, S.L. Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J. Appl. Genet. 2014, 55, 313–318. [Google Scholar] [CrossRef]
- Ishikawa, G.; Nakamura, T.; Ashida, T.; Saito, M.; Nasuda, S.; Endo, T.R.; Wu, J.; Matsumoto, T. Localization of anchor loci representing five hundred annotated rice genes to wheat chromosomes using PLUG markers. Theor. Appl. Genet. 2008, 118, 499–514. [Google Scholar] [CrossRef] [PubMed]
- De Pace, C.; Vaccino, P.; Cionini, G.; Pasquini, M.; Bizzarri, M.; Qualset, C.O. Dasypyrum. In Wild Crop Relatives: Genomic and Breeding Resources, Cereals; Kole, C., Heidelberg, F.L., Eds.; Springer Press: Berlin/Heidelberg, Germany, 2011; pp. 185–292. [Google Scholar]
- Zhang, Q.P.; Li, Q.; Wang, X.E.; Wang, H.Y.; Lang, S.P.; Wang, Y.L.; Wang, S.L.; Chen, P.D.; Liu, D.J. Development and characterization of a Triticum aestivum-Haynaldia villosa translocation line T4VS.4DL conferring resistance to wheat spindle streak mosaic virus. Euphytica 2005, 145, 317–320. [Google Scholar] [CrossRef]
- Ma, J.A.; Zhou, R.H.; Jia, J.Z. Identification of wheat Haynaldia villosa substitution lines conferring resistance to powdery mildew using genomic in situ hybridization (GISH) and RFLP markers. Acta Genet. Sin. 1997, 24, 449–451. [Google Scholar]
- Qi, L.L.; Chen, P.D.; Liu, D.J.; Zhou, B.; Zhang, S.Z. Development of translocation lines of Triticum aestivum with powdery mildew resistance introduced from Haynaldia villosa. In Proceedings of the 8th International Wheat Genetics Symposium, Beijing, China, 20–25 July 1993; Li, Z.S., Xin, Z.Y., Eds.; China Agriculture Scientech Press: Beijing, China, 1995; Volume 1, pp. 333–337. [Google Scholar]
- Chen, Q.-Z.; Qi, Z.-J.; Feng, Y.-G.; Wang, S.-L.; Chen, P.-D. Structural changes of 4V chromosome of Haynaldia villosa induced by gametocidal chromosome 3C of Aegilops triuncialis. Acta Genet. Sin. 2002, 29, 355–358. [Google Scholar]
- Chen, S.; Chen, P.; Wang, X. Inducement of chromosome translocation with small alien segments by irradiating mature female gametes of the whole arm translocation line. Sci. China Ser. C Life Sci. 2008, 51, 346–352. [Google Scholar] [CrossRef]
- Zhu, S.; Ji, Y.; Ji, J.; Bie, T.; Gao, A.; He, H. Fine Physical Bin Mapping of the Powdery Mildew Resistance Gene Pm21 Based on Chromosomal Structural Variations in Wheat. Int. J. Mol. Sci. 2018, 19, 643. [Google Scholar] [CrossRef] [Green Version]
- Qi, L.L.; Pumphrey, M.Q.; Friebe, B.; Zhang, P.; Qian, C.; Bowden, R.L.; Rouse, M.N.; Jin, Y.; Gill, B.S. A novel Robertsonian translocation event leads to transfer of a stem rust resistance gene (Sr52) effective against race Ug99 from Dasypyrum villosum into bread wheat. Theor. Appl. Genet. 2011, 123, 159–167. [Google Scholar] [CrossRef]
- Zhao, W.; Gao, X.; Dong, J.; Zhao, Z.; Chen, Q.; Chen, L.; Shi, Y.; Li, X. Stripe rust resistance and dough quality of new wheat–Dasypyrum villosum translocation lines. Genet. Mol. Res. 2015, 14, 8077–8083. [Google Scholar] [CrossRef]
- Wang, H.J.; Yu, Z.H.; Li, G.R.; Yang, Z.J. Diversified chromosome rearrangements detected in a wheat–Dasypyrum brevia-ristatum substitution line induced by gamma-ray irradiation. Plants 2019, 8, 175. [Google Scholar] [CrossRef] [Green Version]
- Marcu, D.; Damian, G.; Cosma, C.; Cristea, V. Gamma radiation effects on seed germination, growth and pigment content, and ESR study of induced free radicals in maize (Zea mays). J. Biol. Phys. 2013, 39, 625–634. [Google Scholar] [CrossRef] [Green Version]
- Fu, S.; Chen, L.; Wang, Y.; Li, M.; Yang, Z.; Qiu, L.; Yan, B.; Ren, Z.; Tang, Z. Oligonucleotide Probes for ND-FISH Analysis to Identify Rye and Wheat Chromosomes. Sci. Rep. 2015, 5, 10552. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, S.; Tang, Z.; Qiu, L.; Yang, Z.; Li, G.; Lang, T.; Zhu, W.; Zhang, J.; Fu, S. Developing New Oligo Probes to Distinguish Specific Chromosomal Segments and the A, B, D Genomes of Wheat (Triticum aestivum L.) Using ND-FISH. Front. Plant Sci. 2018, 9, 1104. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Z.; Tang, S.; Qiu, L.; Tang, Z.; Fu, S. Oligonucleotides and ND-FISH Displaying Different Arrangements of Tandem Repeats and Identification of Dasypyrum villosum Chromosomes in Wheat Backgrounds. Molecules 2017, 22, 973. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xi, W.; Tang, Z.; Tang, S.; Yang, Z.; Luo, J.; Fu, S. New ND-FISH-Positive Oligo Probes for Identifying Thinopyrum Chromosomes in Wheat Backgrounds. Int. J. Mol. Sci. 2019, 20, 2031. [Google Scholar] [CrossRef] [Green Version]
- Cui, Y.; Bao, Y.G.; Wang, H.G.; Li, X.F. Development of specific molecular markers for Thinopyrum intermedium using RNA-seq data. J. Triticeae Crops 2016, 36, 699–707. [Google Scholar]
- Dai, C.; Zhang, J.P.; Wu, X.Y.; Yang, X.M.; Li, L.H. Development of EST markers specifc to Agropyron cristatum chromosome 6P in common wheat background. Acta Agron. Sin. 2013, 38, 1791–1801. [Google Scholar] [CrossRef]
- Wang, K.; Lin, Z.; Wang, L.; Wang, K.; Shi, Q.; Du, L.; Ye, X. Development of a set of PCR markers specific to Aegilops longissima chromosome arms and application in breeding a translocation line. Theor. Appl. Genet. 2017, 131, 13–25. [Google Scholar] [CrossRef]
- Wu, N.; Li, M.; Sun, H.X.; Cao, Z.L.; Liu, P.; Ding, T.C.; Xu, H.B.; Chu, C.G.; Zhuang, L.F.; Qi, Z.J. RNA-seq facilitates development of chromosome-specific markers and transfer of rye chromatin to wheat. Mol. Breed. 2018, 38, 6. [Google Scholar] [CrossRef]
- Li, S.J.; Lin, Z.S.; Liu, C.; Wang, K.; Du, L.P.; Ye, X.G. Development and comparative genomic mapping of Dasypyrum villosum 6V#4S-specifc PCR markers using transcriptome data. Theor. Appl. Genet. 2017, 130, 2057–2068. [Google Scholar]
- Blanco, A.; Simeone, R.; Resta, P. The addition of Dasypyrum villosum (L.) Candargy chromosomes to durum wheat (Triticum durum Desf.). Theor. Appl. Genet. 1987, 74, 328–333. [Google Scholar] [CrossRef]
- Kato, A.; Lamb, J.C.; Birchler, J.A. Chromosome painting using repetitive DNA sequences as probes for somatic chromosome identification in maize. Proc. Natl. Acad. Sci. USA 2004, 101, 13554–13559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, G.; Gao, D.; Zhang, H.; Li, J.; Wang, H.; La, S.; Ma, J.; Yang, Z. Molecular cytogenetic characterization of Dasypyrum breviaristatum chromosomes in wheat background revealing the genomic divergence between Dasypyrum species. Mol. Cytogenet. 2016, 9, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4326. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, F.; Liu, Q.; Wang, Q.; Yang, N.; Li, J.; Wan, H.; Liu, Z.; Yang, S.; Wang, Y.; Zhang, J.; et al. Characterization of the Durum Wheat-Aegilops tauschii 4D(4B) Disomic Substitution Line YL-443 with Superior Characteristics of High Yielding and Stripe Rust Resistance. Front. Plant Sci. 2021, 12, 745290. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.D.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [Green Version]
- Pertea, G.; Huang, X.; Liang, F.; Antonescu, V.; Sultana, R.; Karamycheva, S.; Lee, Y.; White, J.; Cheung, F.; Parvizi, B.; et al. TIGR Gene Indices clustering tools (TGICL): A software system for fast clustering of large EST datasets. Bioinformatics 2003, 19, 651–652. [Google Scholar] [CrossRef] [Green Version]


| Marker Name | Primer Sequence (5′–3′) | Wheat Bin Map Location | Wheat Chromosomal Location | Restriction Enzymes | Length of 3V Bands, bp |
|---|---|---|---|---|---|
| TNAC1248 | F: ATGATGCAGCAGCAAATTACA | 3AS4-0.45-1.00 | 3AS-211.50 | TaqI | 1000 |
| R: CTGAGGAGCCTCTCCAACTCT | C-3BS1-0.33 | 3BS-251.65 | |||
| 3DS3-0.24-0.31 | 3DS-172.90 | ||||
| TNAC1326 | F:ACAGATCGAGATGTTTATTGAAA | 3AS4-0.45-1.00 | 3AS-137.91 | TaqI | 750 |
| R: GATCAAAGAGATGCGCTGAAG | 3BS1-0.33-0.57 | 3BS-181.49 | |||
| 3DS10-0.31-0.44 | 3DS-127.34 | ||||
| TNAC1267 | F: GAGAGGCAGCTTCACTAGCAG | 3AL3-0.42-0.61 | 3AL-522.17 | TaqI | 200 |
| R: CGTCAGGATCAGCTCTCATGT | 3BL2-0.22-0.41 | 3BL-527.29 | |||
| 3DL1-0.23-0.81 | 3DL-401.87 | ||||
| TNAC1359 | F: GTAAATAGCGCCATCTGCGTA | 3AL3-0.42-0.61 | 3AL-531.08 | TaqI | 750 |
| R: CTCTGGATGCAGTTGGAATGT | 3BL3-0.41-0.50 | 3BL-547.12 | |||
| 3DL1-0.23-0.81 | 3DL-419.90 |
| Primers No. | Primer Sequences(5’–3’) | Localization in D. villosum#3 | Localization in Wheat | Product Size in D. villosum#3 (bp) | Product Size in D. villosum#4 (bp) | |
|---|---|---|---|---|---|---|
| Forward | Reverse | |||||
| 3V#3-1 | GCCTCATGCGGCTGTTGG | GAGGTCATGGTGAGCACGAGA | 3V#3S | 3BS | 148 | - |
| 3V#3-2 | CGGCAAGAGGTCGATGGT | TGACCCACGCACGCACTA | 3V#3L | 1DS | 210 | - |
| 3V#3-3 | GCGTCTTGGATGTCCTG | CGATTTGCTGCCCTACAT | 3V#3L | 3AL | 177 | - |
| 3V#3-4 | GCATGATAGAGAGGTTAGCCAT | CACTGGTGATGTTGTTCAGTACT | 3V#3S | 1AL | 90 | - |
| 3V#3-5 | TCGATACAATTGTTCTTGAGATATG | ACTGGTGCCCTCTTGACG | 3V#3L | 3AL | 560 | - |
| 3V#3-6 | AAACAATCTAGCACTACCCAGAGG | AAGAGGAAGAGAAATAAGCGAGG | 3V#3L | 3DL | 166 | - |
| 3V#3-7 | AGGTCCTTGTCCGAGGTGAT | ATGTTACCGATACTGATGCCACT | 3V#3S | 3AS | 982 | - |
| 3V#3-8 | ATGCTGAACGCAAGGTCAAATA | TGCTGAAGCCCATCACGAAG | 3V#3L | 3DL | 1500 | - |
| 3V#3-9 | CGACTGGTCCACCGTTTC | CGCTGCCTAGTTACCTCTGTT | 3V#3L | 3DL | 1700 | - |
| 3V#3-10 | CTTTCAAGGTAATCCCAGAACT | TGGAAAGCAAACAGGATACG | 3V#3S | 3AS | 126 | - |
| 3V#3-11 | TACGAATAACAACTGCAAGCAGAAT | TGCTAATGGCATCAGCGTCA | 3V#3L | 3DL | 1302 | - |
| 3V#3-12 | CGACTGGTCCACCGTTTC | CGCTGCCTAGTTACCTCTGTT | 3V#3L | 3AL | 1700 | - |
| 3V#4-1 * | TCCATCATAGCACCTTCAGACTCAAG | GACAACTCGGCAATCACCAAGGA | 3V#3S | 5BL | 850 | 377 |
| 3V#4-2 * | GGCAACTCAAATTATAGGATCACGAC | GCAAGGCGGAGTAGCTCACA | 3V#3L | 5DL | 243 | 243 |
| 3V#4-3 * | TTCGTCATCTTTGTTGACATGGCAA | GCAAGGCGGAGTAGCTCACA | 3V#3L | 5DL | 240 | 263 |
| 3V#4-5 * | TTAGAGCGACGACAACTATGC | CAGTATAAGATAGGAGAAGCGACAG | 3V#3L | 3AL | 175 | 175 |
| 3V#4-7 * | GTTATATCGGTTGAGGCGTCTATAC | AACAGTGAGTTCTTCAGGACAGA | 3V#3L | 3DL | 700 | 341 |
| 3V#4-11 * | CTCGTCGGTCTCAGAAGTCAA | TCCACAGAATCATCGGCTCTC | 3V#3L | 3DL | 760 | 760 |
| 3V#4-12 * | CCTCCTCTTCCTCCTCTTCC | TCGCACCATCACCGTACTT | 3V#3L | 3DL | 499 | 499 |
| 3V#4-14 * | GGACGGATGTAGTCTTGTTCAA | CTCGTATCGTACTGCTACTCA | 3V#3L | 3DL | 593 | 593 |
| 3V#4-16 * | GTCCACCAAATCACATCAAACA | GCTCTCACAAGTCACAACAATT | 3V#3L | 3DL | 180 | 180 |
| 3V#4-17 * | ACCATATACTTCGGTGGAACATAC | GCATAGTTACTCTATCACAGACTCA | 3V#3L | 3DL | 304 | 304 |
| 3V#4-18 * | TACCATATACTTCGGTGGAACATAC | GCATAGTTACTCTATCACAGACTCA | 3V#3L | 3DL | 305 | 305 |
| 3V#4-19 * | TGCTCTTCACAGTTCATCTCCT | AGACAAGTTCAGTTCCACACTC | 3V#3L | 3DL | 364 | 364 |
| 3V#4-20 * | TGGTTGCTTCTCAGTTGTGTTG | TACTCGGATAGTGCCTTGTTGA | 3V#3S | 3AS | 600 | 237 |
| 3V#4-21 * | GTTGCTTCTCAGTTGTGTTGGA | TACTCGGATAGTGCCTTGTTGA | 3V#3S | 3AS | 600 | 235 |
| 3V#4-22 * | TGGTTGCTTCTCAGTTGTGTTG | CGGATAGTGCCTTGTTGATGAC | 3V#3S | 3AS | 600 | 233 |
| 3V#4-24 * | GAGAACTGCTCAACATGACAATAAG | CAACAGTATCATCAATGGAGGTCTT | 3V#3S | 3DS | 144 | 144 |
| 3V#4-26 * | TCGCCAGCACCAACCAAT | CAGCACAGCACACCAATGAA | 3V#3S | 3DS | 686 | 686 |
| 3V#4-27 * | GTGACACCAATAGAAGGCAGAA | GGAGGAGCATACCGTGGAA | 3V#3S | 3AS | 403 | 403 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Tang, S.; Lang, T.; Wang, Y.; Long, H.; Deng, G.; Chen, Q.; Guo, Y.; Xuan, P.; Xiao, J.; et al. Molecular Cytogenetic Identification of the Wheat–Dasypyrum villosum T3DL·3V#3S Translocation Line with Resistance against Stripe Rust. Plants 2022, 11, 1329. https://doi.org/10.3390/plants11101329
Zhang J, Tang S, Lang T, Wang Y, Long H, Deng G, Chen Q, Guo Y, Xuan P, Xiao J, et al. Molecular Cytogenetic Identification of the Wheat–Dasypyrum villosum T3DL·3V#3S Translocation Line with Resistance against Stripe Rust. Plants. 2022; 11(10):1329. https://doi.org/10.3390/plants11101329
Chicago/Turabian StyleZhang, Jie, Shuyao Tang, Tao Lang, Ying Wang, Hai Long, Guangbing Deng, Qian Chen, Yuanlin Guo, Pu Xuan, Jun Xiao, and et al. 2022. "Molecular Cytogenetic Identification of the Wheat–Dasypyrum villosum T3DL·3V#3S Translocation Line with Resistance against Stripe Rust" Plants 11, no. 10: 1329. https://doi.org/10.3390/plants11101329





