Genome-Wide Characterization, Identification and Expression Profile of MYB Transcription Factor Gene Family during Abiotic and Biotic Stresses in Mango (Mangifera indica)
Abstract
:1. Introduction
2. Results
2.1. Identification and Bioinformatics Analysis of MYB Transcription Factor Genes in Mango
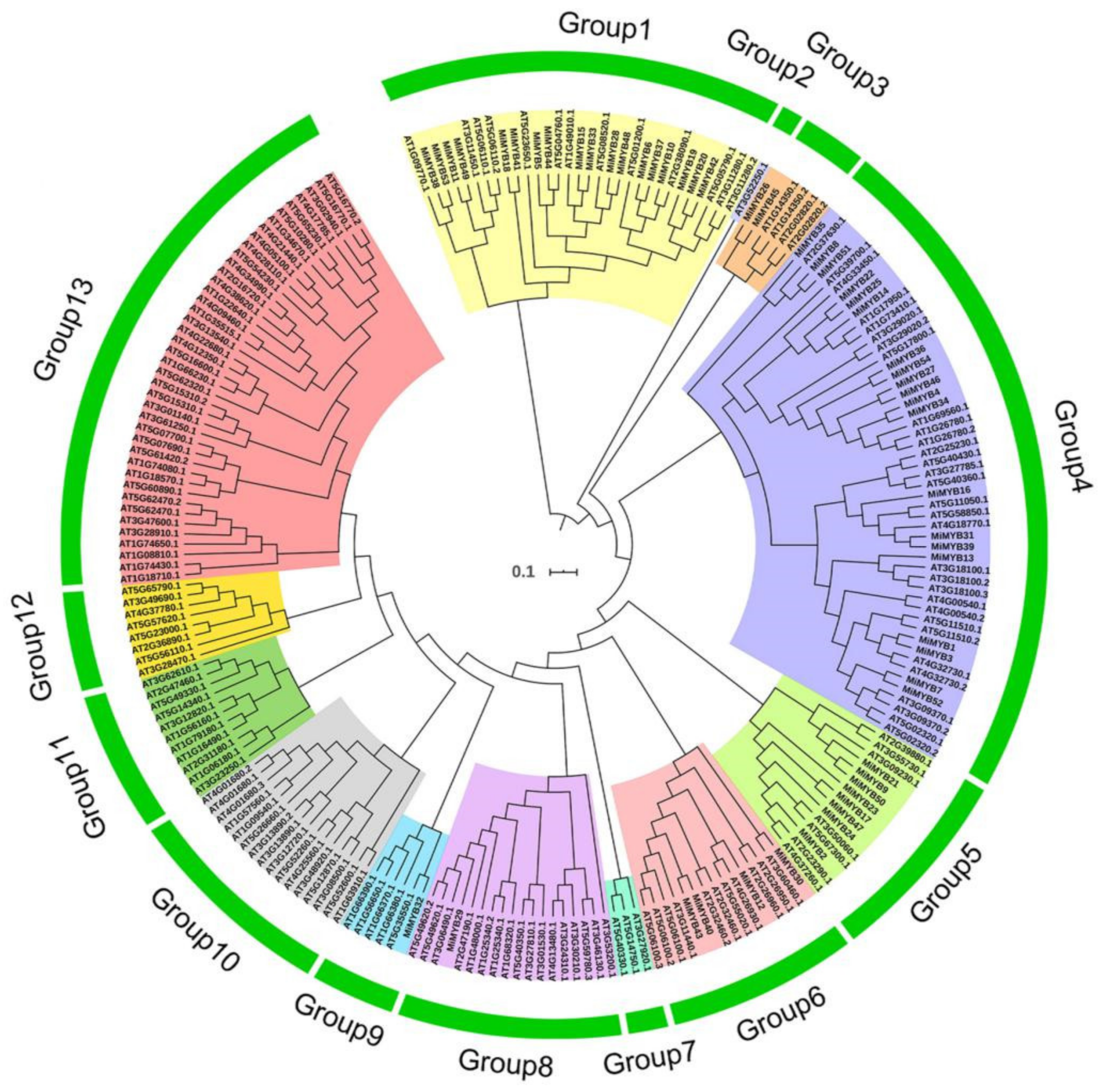
2.2. Phylogenetic, Ka/Ks Analysis, and Synteny Analysis of MYB Transcription Factor Genes in Mango
2.3. The Conserved Domains and Motifs Analysis of MYB Transcription Factors in Mango
2.4. Differential Expression of MiMYBs in Response to SA, MeJA, and H2O2 Treatments
2.5. Differential Expression of MiMYBs Profiles in Response to Pathogen Infection
3. Discussion
4. Materials and Methods
4.1. Identification of MYB Transcription Factor Genes in Mango
4.2. Multiple Sequence Alignments, Phylogenetic Analysis, Ka/Ks Calculation, and Synteny Analysis
4.3. Plant Materials, Growth Conditions, and Treatments
4.4. RNA Extraction and qRT-PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Warschefsky, E.; von Wettberg, E. Population genomic analysis of mango (Mangifera indica) suggests a complex history of domestication. New Phytol. 2019, 222, 2023–2037. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Luo, Y.F.; Huang, J.F.; Gao, S.H.; Zhu, G.P.; Dang, Z.G.; Gai, J.T.; Yang, M.; Zhu, M.; Zhang, H.K.; et al. The genome evolution and domestication of tropical fruit mango. Genome Biol. 2020, 21, 60. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.; Jiang, C.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.J.; Samaha, R.R.; et al. Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef] [PubMed]
- Feller, A.; Machemer, K.; Braun, E.L.; Grotewold, E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J. 2011, 66, 94–116. [Google Scholar] [CrossRef] [PubMed]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010, 15, 573–581. [Google Scholar] [CrossRef]
- Paz-Ares, J.; Ghosal, D.; Wienand, U.; Peterson, P.A.; Saedler, H. The regulatory c1 locus of Zea mays encodes a protein with homology to myb proto-oncogene products and with structural similarities to transcriptional activators. EMBO J. 1987, 6, 3553–3558. [Google Scholar] [CrossRef] [PubMed]
- Campos, J.F.; Cara, B.; Pérez-Martín, F.; Pineda, B.; Egea, I.; Flores, F.B.; Fernandez-Garcia, N.; Capel, J.; Moreno, V.; Angosto, T.; et al. The tomato mutant ars1 (altered response to salt stress 1) identifies an R1-type MYB transcription factor involved in stomatal closure under salt acclimation. Plant Biotechnol. J. 2016, 14, 1345–1356. [Google Scholar] [CrossRef] [Green Version]
- Chezem, W.R.; Memon, A.; Li, F.S.; Weng, J.K.; Clay, N.K. SG2-Type R2R3-MYB Transcription factor MYB15 controls defense-induced lignification and basal immunity in Arabidopsis. Plant Cell. 2017, 29, 1907–1926. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, X.Y.; Yin, Q.G.; Liu, J.Y.; Jiang, W.B.; Di, S.K.; Pang, Y.Z. GmMYB58 and GmMYB205 are seed-specific activators for isoflavonoid biosynthesis in Glycine max. Plant Cell Rep. 2017, 36, 1889–1902. [Google Scholar] [CrossRef] [PubMed]
- Salih, H.; Gong, W.F.; He, S.P.; Sun, G.F.; Sun, J.L.; Du, M.M. Genome-wide characterization and expression analysis of MYB transcription factors in Gossypium hirsutum. BMC Genet. 2016, 17, 129. [Google Scholar] [CrossRef]
- Wang, L.N.; Gao, W.; Wu, X.L.; Zhao, M.R.; Qu, J.B.; Huang, C.Y.; Zhang, J.X. Genome-wide characterization and expression analyses of Pleurotus ostreatus MYB transcription factors during developmental stages and under heat stress based on de novo sequenced genome. Int. J. Mol. Sci. 2018, 19, 2052. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katiyar, A.; Smita, S.; Lenka, S.K.; Rajwanshi, R.; Chinnusamy, V.; Bansal, K.C. Genome-wide classification and expression analysis of MYB transcription factor families in rice and Arabidopsis. BMC Genom. 2012, 13, 544. [Google Scholar] [CrossRef] [Green Version]
- Bian, S.H.; Jin, D.H.; Sun, G.Q.; Shan, B.H.; Zhou, H.N.; Wang, J.Y.; Zhai, L.L.; Li, X.Y. Characterization of the soybean R2R3-MYB transcription factor GmMYB81 and its functional roles under abiotic stresses. Gene 2020, 753, 144803. [Google Scholar] [CrossRef]
- An, J.P.; Wang, X.F.; Zhang, X.W.; Xu, H.F.; Bi, S.Q.; You, C.X.; Hao, Y.J. An apple MYB transcription factor regulates cold tolerance and anthocyanin accumulation and undergoes MIEL1-mediated degradation. Plant Biotechnol. J. 2020, 18, 337–353. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ruan, M.B.; Guo, X.; Wang, B.; Yang, Y.L.; Li, W.Q.; Yu, X.L.; Zhang, P.; Peng, M. Genome-wide characterization and expression analysis enables identification of abiotic stress-responsive MYB transcription factors in cassava (Manihot esculenta). J. Exp. Bot. 2017, 68, 3657–3672. [Google Scholar] [CrossRef] [Green Version]
- Wei, Q.H.; Chen, R.; Wei, X.; Liu, Y.H.; Zhao, S.J.; Yin, X.P.; Xie, T. Genome-wide identification of R2R3-MYB family in wheat and functional characteristics of the abiotic stress responsive gene TaMYB344. BMC Genom. 2020, 21, 792. [Google Scholar] [CrossRef] [PubMed]
- Wan, S.Z.; Li, C.F.; Ma, X.D.; Luo, K.M. PtrMYB57 contributes to the negative regulation of anthocyanin and proanthocyanidin biosynthesis in poplar. Plant Cell Rep. 2017, 36, 1263–1276. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.P.; Luo, Y.; Wang, W.F.; Fan, N.B.; Wang, D.B.; Yang, C.; Jia, H.F. NtMYB12 positively regulates flavonol biosynthesis and enhances tolerance to low pi stress in Nicotiana tabacum. Front. Plant Sci. 2020, 10, 1683. [Google Scholar] [CrossRef]
- Li, W.; Zhu, X.G.; Zhang, Q.J.; Li, K.; Zhang, D.; Shi, C.; Gao, L.Z. SMRT sequencing generates the chromosome-scale reference enome of tropical fruit mango, Mangifera indica. bioRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Tian, F.; Yang, D.C.; Meng, Y.Q.; Jin, J.; Gao, G. PlantRegMap: Charting functional regulatory maps in plants. Nucleic Acids Res. 2020, 48, D1104–D1113. [Google Scholar] [CrossRef]
- Kim, S.H.; Kim, H.S.; Bahk, S.; An, J.; Yoo, Y.; Kim, J.Y.; Chung, W.S. Phosphorylation of the transcriptional repressor MYB15 by mitogen-activated protein kinase 6 is required for freezing tolerance in Arabidopsis. Nucleic Acids Res. 2017, 45, 6613–6627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, B.; Wu, H.; Wang, S.; Ma, X.; Luo, W.; Yao, Q. Identification and analysis of MYB transcription factor family based on transcriptome data in mango (Mangifera indica L.). Chin. J. Trop. Crops 2017, 38, 1285–1294. [Google Scholar] [CrossRef]
- Tafolla-Arellano, J.C.; Zheng, Y.; Sun, H.G.; Jiao, C.; Ruiz-May, E.; Hernández-Oñate, M.A.; González-León, A.; Báez-Sañudo, R.; Fei, Z.J.; Domozych, D.; et al. Transcriptome analysis of mango (Mangifera indica L.) fruit epidermal peel to identify putative cuticle-associated genes. Sci. Rep. 2017, 7, 46163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Xu, Y.; Huang, D.; Xing, W.; Wu, B.; Wei, Q.; Xu, Y.; Zhan, R.; Ma, F.; Song, S. Research progress on the MYB transcription factors in tropical fruit. Trop. Plants 2022, 1, 5. [Google Scholar] [CrossRef]
- Kanzaki, S.; Ichihi, A.; Tanaka, Y.; Fujishige, S.; Koeda, S.; Shimizu, K. The R2R3-MYB transcription factor MiMYB1 regulates light dependent red coloration of ‘Irwin’ mango fruit skin. Sci. Hortic. 2020, 272, 109567. [Google Scholar] [CrossRef]
- Sivankalyani, V.; Sela, N.; Feygenberg, O.; Zemach, H.; Maurer, D.; Alkan, N. Transcriptome dynamics in mango fruit peel reveals mechanisms of chilling stress. Front. Plant Sci. 2016, 7, 1579. [Google Scholar] [CrossRef] [Green Version]
- Dautt-Castro, M.; López-Virgen, A.G.; Ochoa-Leyva, A.; Contreras-Vergara, C.A.; Sortillón-Sortillón, A.P.; Martínez-Téllez, M.A.; González-Aguilar, G.A.; Casas-Flores, J.S.; Sañudo-Barajas, A.; Kuhn, D.N.; et al. Genome-wide identification of mango (Mangifera indica L.) polygalacturonases: Expression analysis of family members and total enzyme activity during fruit ripening. Front. Plant Sci. 2019, 10, 969. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.H.; Yang, X.Y.; He, K.; Liu, M.H.; Li, J.G.; Gao, Z.F.; Lin, Z.Q.; Zhang, Y.F.; Wang, X.X.; Qiu, X.M.; et al. The MYB transcription factor superfamily of Arabidopsis: Expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol. Biol. 2006, 60, 107–124. [Google Scholar] [CrossRef]
- Du, H.; Yang, S.S.; Liang, Z.; Feng, B.R.; Liu, L.; Huang, Y.B.; Tang, Y.X. Genome-wide analysis of the MYB transcription factor superfamily in soybean. BMC Plant Biol. 2012, 12, 106. [Google Scholar] [CrossRef] [Green Version]
- Sun, W.; Ma, Z.; Chen, H.; Liu, M. MYB gene family in Potato (Solanum tuberosum L.): Genome-wide identification of hormone-responsive reveals their potential functions in growth and development. Int. J. Mol. Sci. 2019, 20, 4847. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Li, X. Genome-wide characterization and expression analysis of MYB transcription factors in Lotus japonicas and Medicago truncatula. Genes Genom. 2017, 39, 831–842. [Google Scholar] [CrossRef]
- Li, H.Y.; Yue, Y.Z.; Ding, W.J.; Chen, G.W.; Li, L.; Li, Y.L.; Shi, T.T.; Yang, X.L.; Wang, L.G. Genome-wide identification, classification, and expression profiling reveals R2R3-MYB transcription factors related to monoterpenoid biosynthesis in Osmanthus fragrans. Genes 2020, 11, 353. [Google Scholar] [CrossRef] [Green Version]
- Wu, Z.; Shen, S.; Wang, Y.; Tao, W.; Zhao, Z.; Hu, X.; Yu, P. Genome-wide identification and expression analysis of the Zinc Finger Protein gene subfamilies under drought stress in Triticum aestivum. Plants 2022, 11, 2511. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Crabbe, M.J.C.; Ren, Z. Detoxification gene families at the genome-wide level of rhus gall aphid Schlechtendalia chinensis. Genes 2022, 13, 1627. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Niu, Y.; Zheng, Y. Multiple functions of MYB transcription factors in abiotic stress responses. Int. J. Mol. Sci. 2021, 22, 6125. [Google Scholar] [CrossRef] [PubMed]
- Gong, X.X.; Yan, B.Y.; Hu, J.; Yang, C.P.; Li, Y.J.; Liu, J.P.; Liao, W.B. Transcriptome profiling of rubber tree (Hevea brasiliensis) discovers candidate regulators of the cold stress response. Genes Genom. 2018, 40, 1181–1197. [Google Scholar] [CrossRef] [PubMed]
- Roy, S. Function of MYB domain transcription factors in abiotic stress and epigenetic control of stress response in plant genome. Plant Signal. Behav. 2016, 11, e1117723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Connell, R.J.; Thon, M.R.; Hacquard, S.; Amyotte, S.G.; Kleemann, J.; Torres, M.F.; Damm, U.; Buiate, E.A.; Epstein, L.; Alkan, N.; et al. Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses. Nat. Genet. 2012, 44, 1060–1065. [Google Scholar] [CrossRef] [PubMed]
- Alkan, N.; Friedlander, G.; Ment, D.; Prusky, D.; Fluhr, R. Simultaneous transcriptome analysis of Colletotrichum gloeosporioides and tomato fruit pathosystem reveals novel fungal pathogenicity and fruit defense strategies. New Phytol. 2015, 205, 801–815. [Google Scholar] [CrossRef]
- Lei, Y.; Yuan, X.; Chen, T.; Yuan, Y.; Liu, X.; Tang, X.; Chen, Q. Transcriptome analysis of berries of spine grape (Vitis davidii Föex) infected by Colletotrichum viniferum during symptom development. Horticulturae 2022, 8, 843. [Google Scholar] [CrossRef]
- Zhu, X.; Li, X.; He, Q.; Guo, D.; Liu, C.; Cao, J.; Wu, Z.; Kang, Z.; Wang, X. TaMYB29: A novel R2R3-MYB transcription factor involved inwheat defense against stripe rust. Front. Plant Sci. 2021, 12, 783388. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Deng, R.; Bai, Y.; Wu, H.; Li, C.; Wu, Q.; Zhao, H. Tartary buckwheat R2R3-MYB gene FtMYB3 negatively regulates anthocyanin and proanthocyanin biosynthesis. Int. J. Mol. Sci. 2022, 23, 2775. [Google Scholar] [CrossRef] [PubMed]
- Yi, G.E.; Robin, A.H.K.; Yang, K.; Park, J.I.; Hwang, B.H.; Nou, I. Exogenous methyljasmonate and salicylic acid induce subspeciess-specific patterns of glucosinolate accumulation and gene expression in Brassica oleracea L. Molecules 2016, 21, 1417. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rubel, M.H.; Abuyusuf, M.; Nath, U.K.; Robin, A.H.K.; Jung, H.J.; Kim, H.T.; Park, J.I.; Nou, I. Glucosinolate profile and glucosinolate biosynthesis and breakdown gene expression manifested by black rot disease infection in cabbage. Plants 2020, 9, 1121. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Kesawat, M.S.; Kherawat, B.S.; Singh, A.; Dey, P.; Routray, S.; Mohapatra, C.; Saha, D.; Ram, C.; Siddique, K.H.M.; Kumar, A.; et al. Genome-wide analysis and characterization of the proline-rich extensin-like receptor kinases (PERKs) gene family reveals their role in different developmental stages and stress conditions in wheat (Triticum aestivum L.). Plants 2022, 11, 496. [Google Scholar] [CrossRef]
- Winter, D.; Vinegar, B.; Nahal, H.; Ammar, R.; Wilson, G.V.; Provart, N.J. An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2007, 2, e718. [Google Scholar] [CrossRef]
- Jiang, Y.; Duan, Y.; Yin, J.; Ye, S.; Zhu, J.; Zhang, F.; Lu, W.; Fan, D.; Luo, K. Genome-wide identification and characterization of the Populus WRKY transcription factor family and analysis of their expression in response to biotic and abiotic stresses. J. Exp. Bot. 2014, 65, 6629–6644. [Google Scholar] [CrossRef]
- Zhang, H.; Kang, H.; Su, C.; Qi, Y.; Liu, X.; Pu, J. Genome-wide identification and expression profile analysis of the NAC transcription factor family during abiotic and biotic stress in woodland strawberry. PLoS ONE 2018, 13, e0197892. [Google Scholar] [CrossRef]
- Untergasser, A.; Nijveen, H.; Rao, X.; Bisseling, T.; Geurts, R.; Leunissen, J.A.M. Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res. 2007, 35, W71–W74. [Google Scholar] [CrossRef]
- Luo, C.; He, X.H.; Chen, H.; Hu, Y.; Ou, S.J. Molecular cloning and expression analysis of four actin genes (MiACT) from mango. Biol. Plantarum 2013, 57, 238–244. [Google Scholar] [CrossRef]








Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Liu, Z.; Luo, R.; Sun, Y.; Yang, C.; Li, X.; Gao, A.; Pu, J. Genome-Wide Characterization, Identification and Expression Profile of MYB Transcription Factor Gene Family during Abiotic and Biotic Stresses in Mango (Mangifera indica). Plants 2022, 11, 3141. https://doi.org/10.3390/plants11223141
Zhang H, Liu Z, Luo R, Sun Y, Yang C, Li X, Gao A, Pu J. Genome-Wide Characterization, Identification and Expression Profile of MYB Transcription Factor Gene Family during Abiotic and Biotic Stresses in Mango (Mangifera indica). Plants. 2022; 11(22):3141. https://doi.org/10.3390/plants11223141
Chicago/Turabian StyleZhang, He, Zhixin Liu, Ruixiong Luo, Yu Sun, Cuifeng Yang, Xi Li, Aiping Gao, and Jinji Pu. 2022. "Genome-Wide Characterization, Identification and Expression Profile of MYB Transcription Factor Gene Family during Abiotic and Biotic Stresses in Mango (Mangifera indica)" Plants 11, no. 22: 3141. https://doi.org/10.3390/plants11223141




