Fine Mapping of Stripe-Rust-Resistance Gene YrJ22 in Common Wheat by BSR-Seq and MutMap-Based Sequencing
Abstract
1. Introduction
2. Results
2.1. Phenotypic Characterization of Stripe-Rust Resistance
2.2. BSR-Seq and MutMap Analysis
2.3. Collinearity Analysis of 2A, 2B, and 2D
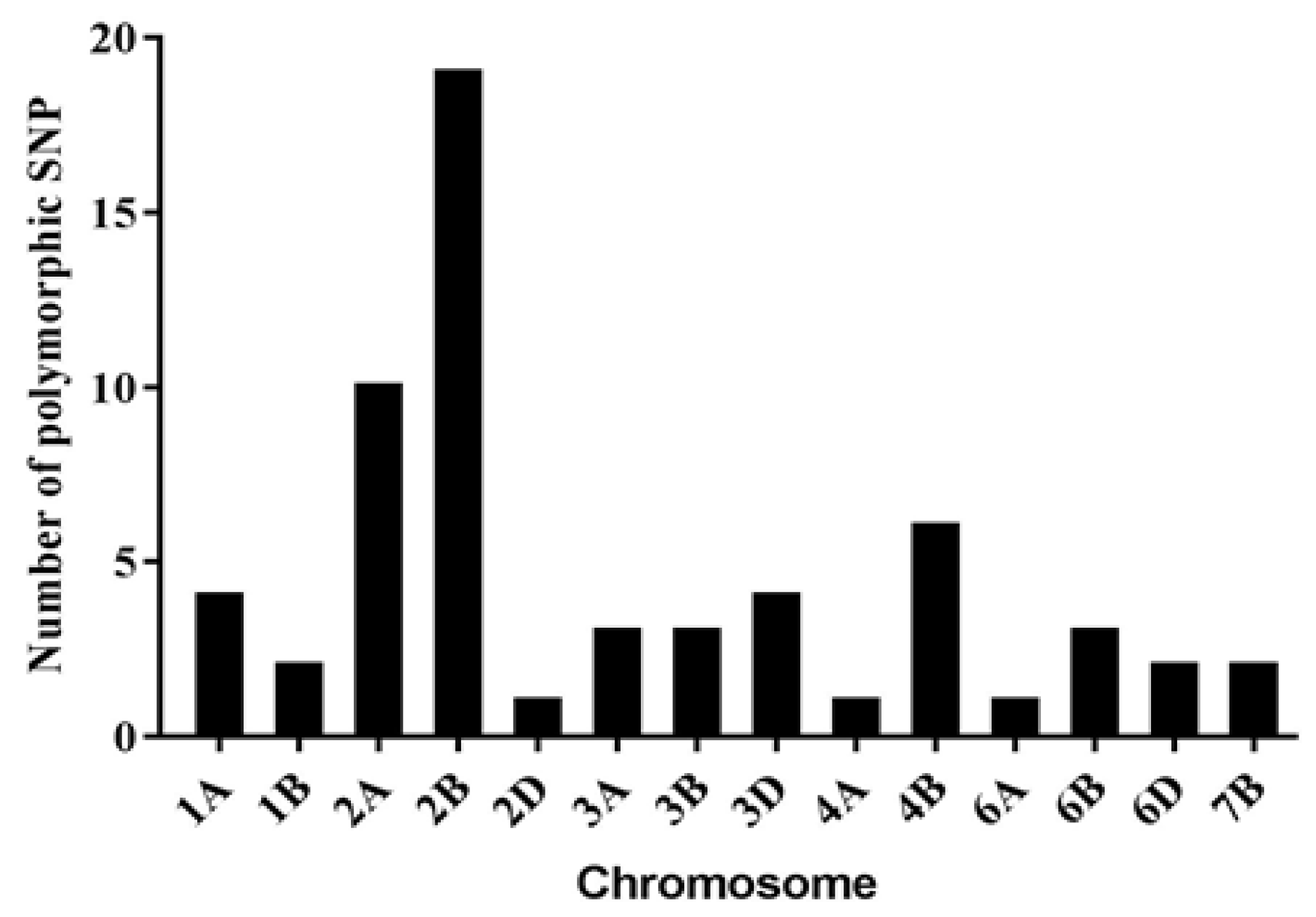
2.4. Development of Polymorphic Markers
2.5. Linkage-Map Construction and Candidate-Genes Analysis
3. Discussion
3.1. Combining Mutmap and BSR-Seq Is an Effective Approach to Fine-Mapping
3.2. Genetic and Physical Mapping of YrJ22
3.3. Analysis of the Candidate Region of YrJ22
4. Materials and Methods
4.1. Plant Materials
4.2. Evaluation of Stripe-Rust Reaction
4.3. BSR-Seq Analysis
4.4. MutMap Analysis
4.5. Genomic DNA Isolation
4.6. Development of Polymorphic CAPS and STS Markers
4.7. Quantitative Real-Time PCR Analysis
4.8. Statical Analysis and Genetic Map Construction
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- FAO. Despite a Cut in World Cereal Production, this Year’s Forecast Output Remains an All-Time High. 2020. Available online: https://uga.ua/en/news/despite-a-cut-in-world-cereal-production-this-year-s-forecast-output-remains-an-all-time-high/ (accessed on 20 October 2021).
- Wellings, C.R. Global status of stripe rust: A review of historical and current threats. Euphytica 2011, 179, 129–141. [Google Scholar] [CrossRef]
- Chen, X.M.; Can, J. Epidemiology and control of stripe rust (Puccinia striiformis f. sp. Tritici) on wheat. Plant Pathol. 2005, 27, 314–337. [Google Scholar]
- Wu, J.; Yu, R.; Wang, H.; Zhou, C.; Huang, S.; Jiao, H.; Yu, S.; Nie, X.; Wang, Q.; Liu, S.; et al. A large-scale genomic association analysis identifies the candidate causal genes conferring stripe rust resistance under multiple field environments. Plant Biotechnol. J. 2020, 19, 177–191. [Google Scholar] [CrossRef] [PubMed]
- The National Agro-Tech Extension and Service Center (NATESC). Forecast of occurrence trend of major crop diseases and insect pests in China in 2020. China Plant Prot. 2020, 2, 37–39. [Google Scholar]
- Zeng, Q.; Zhao, J.; Wu, J.; Zhan, G.; Han, D.; Kang, Z. Wheat Stripe Rust and Integration of Sustainable Control Strategies in China. Front. Agric. Sci. Eng. 2022, 9, 37–51. [Google Scholar] [CrossRef]
- McIntosh, R.A.; Dubcovsky, J.; Rogers, W.J.; Xia, X.C.; Raupp, W.J. Catalogue of gene symbols for wheat. Annu. Wheat Newsl. 2021, 67, 104–113. [Google Scholar]
- McIntosh, R.A.; Dubcovsky, J.; Rogers, W.J.; Morris, C.; Appels, R.; Xia, X.C. Catalogue of Gene Symbols for Wheat: 2015–2016 Supplement. 2017. Available online: https://wheat.pw.usda.gov/GG3/wgc (accessed on 20 October 2021).
- Dong, Z.; Hegarty, J.M.; Zhang, J.; Zhang, W.; Chao, S.; Chen, X.; Zhou, Y.; Dubcovsky, J. Validation and characterization of a QTL for adult plant resistance to stripe rust on wheat chromosome arm 6BS (Yr78). Theor. Appl. Genet. 2017, 130, 2127–2137. [Google Scholar] [CrossRef]
- Feng, J.Y.; Wang, M.N.; See, D.R.; Chao, S.; Zheng, Y.; Chen, X.M. Characterization of novel gene Yr79 and four additional quantitative trait loci for all-stage and high-temperature adult-plant resistance to strip-e rust in spring wheat PI182103. Phytopathology 2018, 108, 737–747. [Google Scholar] [CrossRef]
- Gessese, M.; Bariana, H.; Wong, D.; Hayden, M.; Bansal, U. Molecular Mapping of Stripe Rust Resistance Gene Yr81 in a Common Wheat Landrace Aus27430. Plant Dis. 2019, 103, 1166–1171. [Google Scholar] [CrossRef]
- Pakeerathan, K.; Bariana, H.; Qureshi, N.; Wong, D.; Hayden, M.; Bansal, U. Identification of a new source of stripe rust resistance Yr82 in wheat. Theor. Appl. Genet. 2019, 132, 3169–3176. [Google Scholar] [CrossRef]
- Li, J.; Dundas, I.; Dong, C.; Li, G.; Trethowan, R.; Yang, Z.; Hoxha, S.; Zhang, P. Identification and characterization of a new stripe rust resistance gene Yr83 on rye chromosome 6R in wheat. Theor. Appl. Genet. 2020, 133, 1095–1107. [Google Scholar] [CrossRef]
- Wang, B.; Hu, X.; Li, Q.; Hao, B.; Zhang, B.; Li, G.; Kang, Z. Development of Race-Specific SCAR Markers for Detection of Chinese Races CYR32 and CYR33 of Puccinia striiformis f. sp. tritici. Plant Dis. 2010, 94, 221–228. [Google Scholar] [CrossRef]
- Wang, L.; Tang, X.; Wu, J.; Shen, C.; Dai, M.; Wang, Q.; Zeng, Q.; Kang, Z.; Wu, Y.; Han, D. Stripe rust resistance to a burgeoning Puccinia striiformis f. sp. tritici race CYR34 in current Chinese wheat cultivars for breeding and research. Euphytica 2019, 215, 68. [Google Scholar] [CrossRef]
- Cheng, P.; Xu, L.S.; Wang, M.N.; See, D.R.; Chen, X.M. Molecular mapping of genes Yr64 and Yr65 for stripe rust resistance in hexaploid derivatives of durum wheat accessions PI 331260 and PI 480016. Theor. Appl. Genet. 2014, 127, 2267–2277. [Google Scholar] [CrossRef]
- Zegeye, H.; Rasheed, A.; Makdis, F.; Badebo, A.; Ogbonnaya, F.C. Genome-Wide Association Mapping for Seedling and Adult Plant Resistance to Stripe Rust in Synthetic Hexaploid Wheat. PLoS ONE 2014, 9, e105593. [Google Scholar] [CrossRef]
- The International Wheat Genome Sequencing Consortium (IWGSC). A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 2014, 345, 1251788. [Google Scholar]
- Zheng, S.; Wu, Y.; Zhou, M.; Zeng, L.; Liu, R.; Li, Y.; Liu, Z.; Zhang, C.; Lu, L.; Zhang, L. Characterization and diagnostic marker development for Yr28-rga1 conferring stripe rust resistance in wheat. Eur. J. Plant Pathol. 2019, 156, 623–634. [Google Scholar] [CrossRef]
- Zhang, C.; Huang, L.; Zhang, H.; Hao, Q.; Lyu, B.; Wang, M.; Epstein, L.; Liu, M.; Kou, C.; Qi, J.; et al. An ancestral NB-LRR with duplicated 3′UTRs confers stripe rust resistance in wheat and barley. Nat. Commun. 2019, 10, 4023. [Google Scholar] [CrossRef]
- Liu, W.; Frick, M.; Huel, R.; Nykiforuk, C.L.; Wang, X.; Gaudet, D.A.; Eudes, F.; Conner, R.L.; Kuzyk, A.; Chen, Q.; et al. The Stripe Rust Resistance Gene Yr10 Encodes an Evolutionary-Conserved and Unique CC–NBS–LRR Sequence in Wheat. Mol. Plant 2014, 7, 1740–1755. [Google Scholar] [CrossRef]
- Krattinger, S.G.; Lagudah, E.S.; Spielmeyer, W.; Singh, R.P.; Huerta-Espino, J.; McFadden, H.; Bossolini, E.; Selter, L.L.; Keller, B. A Putative ABC Transporter Confers Durable Resistance to Multiple Fungal Pathogens in Wheat. Science 2009, 323, 1360–1363. [Google Scholar] [CrossRef]
- Klymiuk, V.; Yaniv, E.; Huang, L.; Raats, D.; Fatiukha, A.; Chen, S.; Feng, L.; Frenkel, Z.; Krugman, T.; Lidzbarsky, G.; et al. Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase-pseudo kinase family. Nat. Commun. 2018, 9, 3735. [Google Scholar] [CrossRef]
- Fu, D.; Uauy, C.; Distelfeld, A.; Blechl, A.; Epstein, L.; Chen, X.; Sela, H.; Fahima, T.; Dubcovsky, J. A Kinase-START Gene Confers Temperature-Dependent Resistance to Wheat Stripe Rust. Science 2009, 323, 1357–1360. [Google Scholar] [CrossRef] [PubMed]
- Moore, J.W.; Herrera-Foessel, S.; Lan, C.; Schnippenkoetter, W.; Ayliffe, M.; Huerta-Espino, J.; Lillemo, M.; Viccars, L.; Milne, R.; Periyannan, S.; et al. A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat. Genet. 2015, 47, 1494–1498. [Google Scholar] [CrossRef] [PubMed]
- Marchal, C.; Zhang, J.; Zhang, P.; Fenwick, P.; Steuernagel, B.; Adamski, N.M.; Boyd, L.; McIntosh, R.; Wulff, B.B.H.; Berry, S.; et al. BED-domain-containing immune receptors confer diverse resistance spectra to yellow rust. Nat. Plants 2018, 4, 662–668. [Google Scholar] [CrossRef] [PubMed]
- Zegeye, W.A.; Zhang, Y.; Cao, L.; Cheng, S. Whole Genome Resequencing from Bulked Populations as a Rapid QTL and Gene Identification Method in Rice. Int. J. Mol. Sci. 2018, 19, 4000. [Google Scholar] [CrossRef] [PubMed]
- Francis, Y.L.C.; Henry, C.M.L.; Yiu, S.M. Sequence assembly using next generation sequencing data—Challenges and solutions. Sci. China (Life Sci.) 2014, 57, 1140–1148. [Google Scholar]
- Zou, C.; Wang, P.; Xu, Y. Bulked sample analysis in genetics, genomics and crop improvement. Plant Biotechnol. J. 2016, 14, 1941–1955. [Google Scholar] [CrossRef]
- Wang, Y.; Xie, J.; Zhang, H.; Guo, B.; Ning, S.; Chen, Y.; Lu, P.; Wu, Q.; Li, M.; Zhang, D.; et al. Mapping stripe rust resistance gene YrZH22 in Chinese wheat cultivar Zhoumai 22 by bulked segregant RNA-Seq (BSR-Seq) and comparative genomics analyses. Theor. Appl. Genet. 2017, 130, 2191–2201. [Google Scholar] [CrossRef]
- Li, Y.; Shi, X.; Hu, J.; Wu, P.; Qiu, D.; Qu, Y.; Xie, J.; Wu, Q.; Zhang, H.; Yang, L.; et al. Identification of a Recessive Gene PmQ Conferring Resistance to Powdery Mildew in Wheat Landrace Qingxinmai Using BSR-Seq Analysis. Plant Dis. 2020, 104, 743–751. [Google Scholar] [CrossRef]
- Xie, J.Z. Establishment and application of BSR-Seq for gene mapping in wheat and sequence analysis of Aegilops tauschii chromosome arm 3DS. Ph.D. Thesis, China University of Agricultural, Beijing, China, 2016. [Google Scholar]
- Liang, Y.; Zhang, D.-Y.; Ouyang, S.; Xie, J.; Wu, Q.; Wang, Z.; Cui, Y.; Lu, P.; Zhang, D.; Liu, Z.-J.; et al. Dynamic evolution of resistance gene analogs in the orthologous genomic regions of powdery mildew resistance gene MlIW170 in Triticum dicoccoides and Aegilops tauschii. Theor. Appl. Genet. 2015, 128, 1617–1629. [Google Scholar] [CrossRef]
- Nakata, M.; Miyashita, T.; Kimura, R.; Nakata, Y.; Yamakawa, H. MutMapPlus identified novel mutant alleles of a rice starch branching enzyme IIb gene for fine-tuning of cooked rice texture. Plant Biotechnol. J. 2017, 16, 1–13. [Google Scholar]
- Mo, Y.; Howell, T.; Vasquez-Gross, H.; de Haro, L.A.; Dubcovsky, J.; Pearce, S. Mapping causal mutations by exome sequencing in a wheat TILLING population: A tall mutant case study. Mol. Genet. Genom. 2017, 293, 463–477. [Google Scholar] [CrossRef]
- Li, M.; Dong, L.; Li, B.; Wang, Z.; Xie, J.; Qiu, D.; Li, Y.; Shi, W.; Yang, L.; Wu, Q.; et al. A CNL protein in wild emmer wheat confers powdery mildew resistance. New Phytol. 2020, 228, 1027–1037. [Google Scholar] [CrossRef]
- Abe, A.; Kosugi, S.; Yoshida, K.; Natsume, S.; Takagi, H.; Kanzaki, H.; Matsumura, H.; Yoshida, K.; Mitsuoka, C.; Tamiru, M.; et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 2012, 30, 174–178. [Google Scholar] [CrossRef]
- Nordström, K.J.V.; Albani, M.C.; James, G.V.; Gutjahr, C.; Hartwig, B.; Turck, F.; Paszkowski, U.; Coupland, G.; Schneeberger, K. Mutation identification by direct comparison of whole-genome sequencing data from mutant and wild-type individuals using k-mers. Nat. Biotechnol. 2013, 31, 325–330. [Google Scholar] [CrossRef]
- Fekih, R.; Takagi, H.; Tamiru, M.; Abe, A.; Natsume, S.; Yaegashi, H.; Sharma, S.; Sharma, S.; Kanzaki, H.; Matsumura, H.; et al. MutMap+: Genetic mapping and muta-nt identification without crossing in rice. PLoS ONE 2013, 8, e68529. [Google Scholar] [CrossRef]
- Schneeberger, K.; Ossowski, S.; Lanz, C.; Juul, T.; Petersen, A.H.; Nielsen, K.L.; Jorgensen, J.E.; Weigel, D.; Andersen, S.U. SHORE map: Simultaneous mapping and mutation identification by deep sequencing. Nat. Methods 2009, 6, 550–551. [Google Scholar] [CrossRef]
- Wang, Z.; Li, J.; Chen, S.; Heng, Y.; Chen, Z.; Yang, J.; Zhou, K.; Pei, J.; He, H.; Deng, X.W.; et al. Poaceae-specific MS1 encodes a phospholipid-binding protein for male fertility in bread wheat. Proc. Natl. Acad. Sci. USA 2017, 114, 12614–12619. [Google Scholar] [CrossRef]
- Singh, V.K.; Khan, A.W.; Saxena, R.K.; Kumar, V.; Kale, S.M.; Sinha, P.; Chitikineni, A.; Pazhamala, L.T.; Garg, V.; Sharma, M.; et al. Next-generation sequencing for identification of candidate genes for Fusarium wilt and sterility mosaic disease in pigeonpea (Cajanus cajan). Plant Biotechnol. J. 2016, 14, 1183–1194. [Google Scholar] [CrossRef]
- Appels, R. Wheat research and breeding in the new era of a high-quality reference genome. Front. Agric. Sci. Eng. 2019, 6, 225. [Google Scholar] [CrossRef]
- Yang, Z.Z.; Wang, Z.H.; Hu, Z.R.; Xin, M.M.; Guo, W.L. Comparative analysis of the genomic sequences between commercial wheat varieties jimai 22 and liangxing 99. Acta Agron. Sinica. 2020, 46, 1870–1883. [Google Scholar]
- Chen, C.; He, Z.; Lu, J.; Li, J.; Ren, Y.; Ma, C.; Xia, X. Molecular mapping of stripe rust resistance gene YrJ22 in Chinese wheat cultivar Jimai 22. Mol. Breed. 2016, 36, 118. [Google Scholar] [CrossRef]
- Ma, S.; Wang, M.; Wu, J.; Guo, W.; Chen, Y.; Li, G.; Wang, Y.; Shi, W.; Xia, G.; Fu, D.; et al. WheatOmics: A platform combining multiple omics data to accelerate functional genomics studies in wheat. Mol. Plant 2021, 14, 1965–1968. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.-H.; Meng, J.-L. MapDraw: A microsoft excel macro for drawing genetic linkage maps based on given genetic linkage data. Hereditas 2003, 25, 317–321. [Google Scholar] [PubMed]
- Luo, M.; Xie, L.; Chakraborty, S.; Wang, A.; Matny, O.; Jugovich, M.; Kolmer, J.A.; Richardson, T.; Bhatt, D.; Hoque, M.; et al. A five-transgene cassette confers broad-spectrum resistance to a fungal rust pathogen in wheat. Nat. Biotechnol. 2021, 39, 561–566. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xia, C.; Wang, M.; Yin, C.; Chen, X. Whole-genome sequencing of Puccinia striiformis f. sp. tritici mutant isolates identifies avirulence gene candidates. BMC Genom. 2020, 21, 247. [Google Scholar] [CrossRef]
- Wu, J.; Zeng, Q.; Wang, Q.; Liu, S.; Yu, S.; Mu, J.; Huang, S.; Sela, H.; Distelfeld, A.; Huang, L.; et al. SNP-based pool genotyping and haplotype analysis accelerate fine-mapping of the wheat genomic region containing stripe rust resistance gene Yr26. Theor. Appl. Genet. 2018, 131, 1481–1496. [Google Scholar] [CrossRef]
- Wu, Q.; Chen, Y.; Xie, J.; Dong, L.; Wang, Z.; Lu, P.; Wang, R.; Yuan, C.; Zhang, Y.; Liu, Z. A 36 Mb terminal deletion of chromosome 2BL is responsible for a wheat semi-dwarf mutation. Crop J. 2020, 9, 873–881. [Google Scholar] [CrossRef]
- Wu, J.; Xu, D.; Fu, L.; Wu, L.; Hao, W.; Li, J.; Dong, Y.; Wang, F.; Wu, Y.; He, Z.; et al. Fine mapping of a stripe rust resistance gene YrZM175 in bread wheat. Theor. Appl. Genet. 2022, 135, 3485–3496. [Google Scholar] [CrossRef]
- Yang, M.Y.; Li, G.G.; Wan, H.S.; Li, L.P.; Li, J.; Yang, W.Y.; Pu, Z.J.; Yang, Z.J.; Yang, E.N. Identification of QTLs for stripe rust r-esistance in a recombinant inbred line population. Inter. J. Mol. Sci. 2019, 20, 3410. [Google Scholar]
- Wang, Z.; Ren, J.; Du, Z.; Che, M.; Zhang, Y.; Quan, W.; Jiang, X.; Ma, Y.; Zhao, Y.; Zhang, Z. Identification of a major QTL on chromosome arm 2AL for reducing yellow rust severity from a Chinese wheat landrace with evidence for durable resistance. Theor. Appl. Genet. 2018, 132, 457–471. [Google Scholar] [CrossRef]
- Liu, J.; He, Z.; Wu, L.; Bai, B.; Wen, W.; Xie, C.; Xia, X. Genome-Wide Linkage Mapping of QTL for Adult-Plant Resistance to Stripe Rust in a Chinese Wheat Population Linmai 2 × Zhong 892. PLoS ONE 2015, 10, e0145462. [Google Scholar] [CrossRef]
- Zeng, Q.; Wu, J.; Liu, S.; Chen, X.; Yuan, F.; Su, P.; Wang, Q.; Huang, S.; Mu, J.; Han, D.; et al. Genome-wide Mapping for Stripe Rust Resistance Loci in Common Wheat Cultivar Qinnong 142. Plant Dis. 2019, 103, 439–447. [Google Scholar] [CrossRef]
- Naruoka, Y.; Garland-Campbell, K.A.; Carter, A.H. Genome-wide association mapping for stripe rust (Puccinia striiformis F. sp. tritici) in US Pacific Northwest winter wheat (Triticum aestivum L.). Theor. Appl. Genet. 2015, 128, 1083–1101. [Google Scholar] [CrossRef]
- Beat, K.; Thomas, W.; Krattinger, S.G. Advances in wheat and pathogen genomics: Implications for disease control. Annu. Rev. Phytopathol. 2018, 56, 67–87. [Google Scholar]
- Walkowiak, S.; Gao, L.; Monat, C.; Haberer, G.; Kassa, M.T.; Brinton, J.; Ramirez-Gonzalez, R.H.; Kolodziej, M.C.; Delorean, E.; Thambugala, D.; et al. Multiple wheat genomes reveal global variation in modern breeding. Nature 2020, 588, 277–283. [Google Scholar] [CrossRef]
- Andersen, E.J.; Nepal, M.P.; Purintun, J.M.; Nelson, D.; Mermigka, G.; Sarris, P.F. Wheat Disease Resistance Genes and Their Diversification Through Integrated Domain Fusions. Front. Genet. 2020, 11, 898. [Google Scholar] [CrossRef]
- Wang, Y.; Huang, L.; Luo, W.; Jin, Y.; Gong, F.; He, J.; Liu, D.; Zheng, Y.; Wu, B. Transcriptome analysis provides insights into the mechanisms underlying wheat cultivar Shumai126 responding to stripe rust. Gene 2020, 768, 145290. [Google Scholar] [CrossRef]
- Li, J.; Yang, J.; Li, Y.; Ma, L. Current strategies and advances in wheat biology. Crop J. 2020, 8, 879–891. [Google Scholar] [CrossRef]
- Huang, L.; Sela, H.; Feng, L.; Chen, Q.; Krugman, T.; Yan, J.; Dubcovsky, J.; Fahima, T. Distribution and haplotype diversity of WKS resistance genes in wild emmer wheat natural populations. Theor. Appl. Genet. 2016, 129, 921–934. [Google Scholar] [CrossRef]
- Catherine, F.; Silvia, T.; Nils, S.; Laurence, A.; Aurélie, N.; Beat, K. Map-based isolation of the leaf rust disease resistance gene Lr10 from the hexaploid wheat (Triticum aestivum L.) genome. Proc. Natl. Acad. Sci. USA 2003, 25, 15253–15258. [Google Scholar]
- Bariana, H.S.; McIntosh, R.A. Cytogenetic studies in wheat. XV. Location of rust resistance genes in VPM1 and their genetic linkage with other disease resistance genes in chromosome 2A. Genome 1993, 36, 476–482. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.F.; Zheng, T.C.; He, Z.H.; Li, G.Q.; Xu, S.C.; Li, X.P.; Yang, G.Y.; Singh, R.P.; Xia, X.C. Molecular tagging of stripe rust resistance gene YrZH84 in Chinese wheat line Zhou 8425B. Theor. Appl. Genet. 2006, 112, 1098–1103. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Walker, M.A.; Pedamallu, C.S.; I Ojesina, A.; Bullman, S.; Sharpe, T.; Whelan, C.W.; Meyerson, M. GATK PathSeq: A customizable computational tool for the discovery and identification of microbial sequences in libraries from eukaryotic hosts. Bioinformatics 2018, 34, 4287–4289. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Tian, X.; Wen, W.; Xie, L.; Fu, L.; Xu, D.; Fu, C.; Wang, D.; Chen, X.; Xia, X.; Chen, Q.; et al. Molecular Mapping of Reduced Plant Height Gene Rht24 in Bread Wheat. Front. Plant Sci. 2017, 8, 1379. [Google Scholar] [CrossRef]
- Ling, H.-Q.; Zhao, S.; Liu, D.; Wang, J.; Sun, H.; Zhang, C.; Fan, H.; Li, D.; Dong, L.; Tao, Y.; et al. Draft genome of the wheat A-genome progenitor Triticum urartu. Nature 2013, 496, 87–90. [Google Scholar] [CrossRef]
- Singh, V.K.; Khan, A.W.; Jaganathan, D.; Thudi, M.; Roorkiwal, M.; Takagi, H.; Garg, V.; Kumar, V.; Chitikineni, A.; Gaur, P.M.; et al. QTL-seq for rapid identification of candidate genes for 100-seed weight and root/total plant dry weight ratio under rainfed conditions in chickpea. Plant Biotechnol. J. 2016, 14, 2110–2119. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]





| Plant Material | Total | Infection Type | Expected Ratio | χ2 | p | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 1 | 2 | 3 | 4 | |||||
| Jimai 22 | 10 | 10 | ||||||||
| Avocet S | 10 | 3 | 7 | |||||||
| F1 | 15 | 15 | ||||||||
| F2 | 2273 | 3 | 1482 | 169 | 66 | 128 | 425 | 3:1 | 0.46 | 0.55 |
| Marker | Forward Primer Sequence (5′–3′) | Reverse Primer Sequence (5′–3′) | Physical Pos.* (Mb) | Restriction Enzyme | Product Size (bp) |
|---|---|---|---|---|---|
| H796 | AGTGGGGCTATTTTGTCGG | CCTTGTTCATGGCTGGTTG | 773.2 | BSSHII | 739 |
| H896 | TCTAGTTGCTGCCGTGGTTA | AACCAAACGGACACACATGG | 764.1 | HpyCH4V | 718 |
| H804 | GTGCCGTGAGCACCCTGCTG | TGGACGTCTTTTGCCCTCTTGAA | 772.6 | FoKI | 869 |
| H690 | GGATTCTCACGGTCACTCAA | CTCATTCCCGCAACAGG | 773.0 | HaeIII | 546 |
| H805 | GAGCAGCCGGAGGAGTTG | CAGGAGGAGATGGAGAGCAT | 772.7 | FoKI | 1013 |
| H034 | AGTAGGGTAAATGGCGAGCAG | GTCCCAAGGAACAAACACG | 750.4 | Hpy188III | 720 |
| H727 | GGGTGGTCACATCCAGGTCC | GAACATGCCTCAGAACAATGGA | 768.7 | TspRI | 443 |
| H400 | CGATTGCTGCTTTCCTTCAT | ATCCCATCGGTCCCGTGTT | 768.7 | BbvI | 1677 |
| H732 | GCATCGCAGCAACACTCG | GGAGACAATGGGCGGTTT | 768.9 | BstUI | 1128 |
| H736 | GTGCTCCTTACAGGGAACAAC | ATCCACAGCCGAACCAAA | 769.0 | HpyCH4III | 1573 |
| H801 | GGACAAATACAAGGGTTCG | TGTCGTCGGGATTCAAGG | 772.4 | - | 279 |
| Name | Length (bp) | Position on Chr2A * (bp) | Functional Annotation * |
|---|---|---|---|
| TraesCS2A01G572700 | 1455 | 768,663,628–768,665,082 | Peroxidase |
| TraesCS2A01G572800 | 1625 | 768,676,637–768,678,261 | Peroxidase |
| TraesCS2A01G740000LC | 1200 | 768,691,235–768,692,434 | F-box domain containing protein |
| TraesCS2A01G572900(R) | 5771 | 768,729,058–768,734,828 | Disease resistance protein |
| TraesCS2A01G740100LC | 483 | 768,882,506–768,882,988 | Endonuclease/exonuclease/ phosphatase family protein |
| TraesCS2A01G573000 | 1497 | 768,923,600–768,925,096 | Retrovirus-related Pol polyprotein from transposon TNT 1-94 |
| TraesCS2A01G573100(NLR) | 5239 | 768,926,410–768,931,648 | Disease resistance protein (TIR-NBS-LRR class) |
| TraesCS2A01G573200 | 1655 | 768,981,429–768,983,083 | Succinate dehydrogenase subunit |
| TraesCS2A01G740200LC | 1251 | 768,992,115–768,993,365 | F-box domain containing protein |
| TraesCS2A01G573300 | 1463 | 768,997,070–768,998,532 | Peroxidase |
| TraesCS2A01G573400 | 1706 | 769,003,029–769,004,734 | Peroxidase |
| TraesCS2A01G573500 | 1136 | 769,020,358–769,021,493 | Peroxidase |
| TraesCS2A01G573600 | 1129 | 769,032,902–769,034,030 | Peroxidase |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, C.; Hao, W.; Wu, J.; Si, H.; Xia, X.; Ma, C. Fine Mapping of Stripe-Rust-Resistance Gene YrJ22 in Common Wheat by BSR-Seq and MutMap-Based Sequencing. Plants 2022, 11, 3244. https://doi.org/10.3390/plants11233244
Chen C, Hao W, Wu J, Si H, Xia X, Ma C. Fine Mapping of Stripe-Rust-Resistance Gene YrJ22 in Common Wheat by BSR-Seq and MutMap-Based Sequencing. Plants. 2022; 11(23):3244. https://doi.org/10.3390/plants11233244
Chicago/Turabian StyleChen, Can, Weihao Hao, Jingchun Wu, Hongqi Si, Xianchun Xia, and Chuanxi Ma. 2022. "Fine Mapping of Stripe-Rust-Resistance Gene YrJ22 in Common Wheat by BSR-Seq and MutMap-Based Sequencing" Plants 11, no. 23: 3244. https://doi.org/10.3390/plants11233244
APA StyleChen, C., Hao, W., Wu, J., Si, H., Xia, X., & Ma, C. (2022). Fine Mapping of Stripe-Rust-Resistance Gene YrJ22 in Common Wheat by BSR-Seq and MutMap-Based Sequencing. Plants, 11(23), 3244. https://doi.org/10.3390/plants11233244





