MIX-NET: Deep Learning-Based Point Cloud Processing Method for Segmentation and Occlusion Leaf Restoration of Seedlings
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Materials
2.2. Mix-Net Based Seedling Point Cloud Processing Method
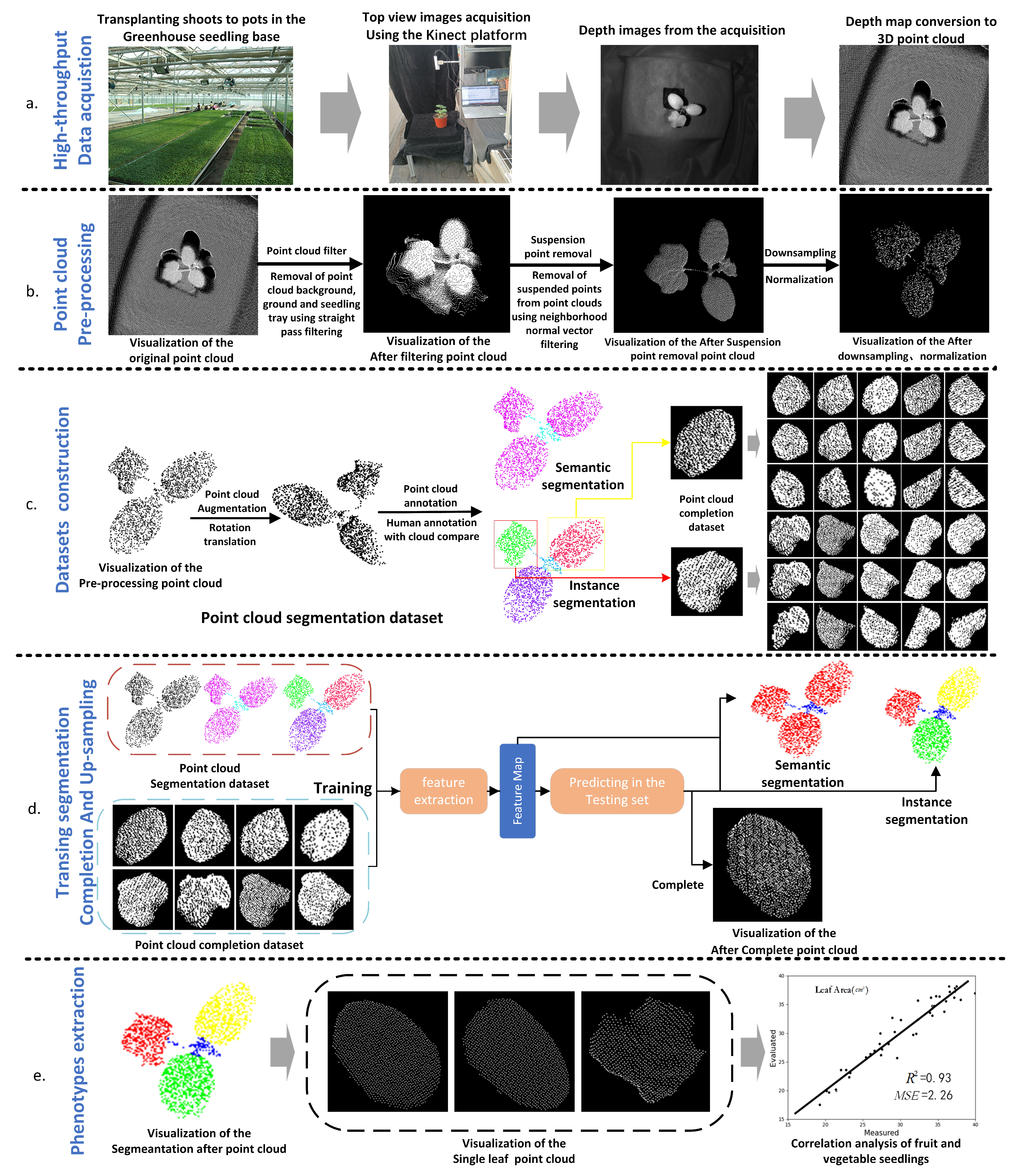
2.2.1. Overview
2.2.2. Data Acquisition
2.2.3. Point Cloud Preprocessing
- The original point cloud is filtered by (1) to obtain the point cloud containing only the plant area.
- Set a threshold N, find N neighborhoods around each centroid using KNN, and find the average value D of the Euclidean distance between the centroid and the neighborhoods.
- The angle W between the normal vector and z-axis is solved by fitting the plane with least squares to predict the normal vector of each centroid through the set neighborhood threshold N.
- Repeat the above operations First and second, if D ≥ d or W ≥ c, it is judged to be a hover point, and the point is deleted. Iterate through the whole point cloud to eliminate all the hover points.
2.2.4. Datasets Construction
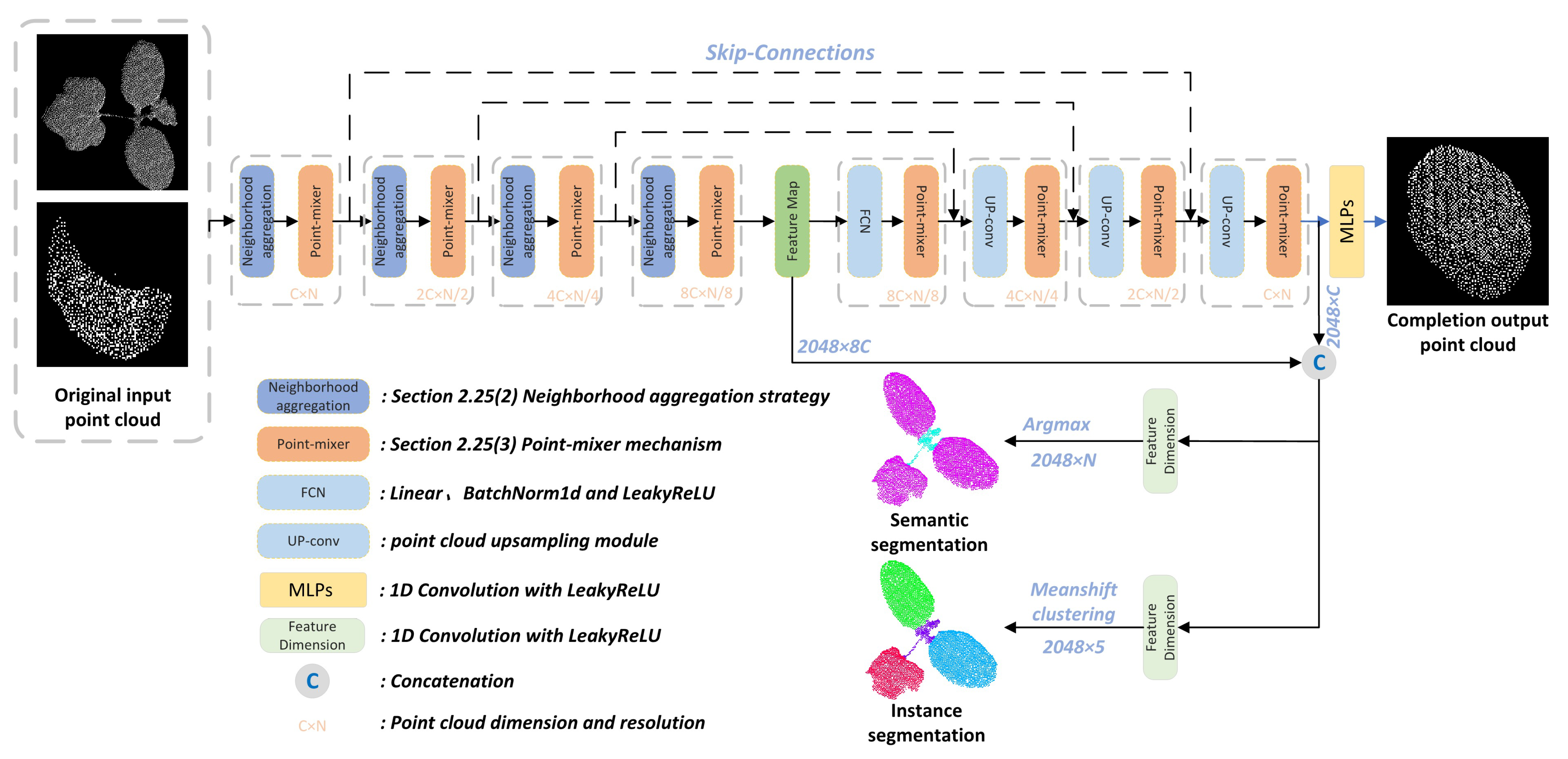
2.2.5. MIX-NET Network for Segmenting and Completing Point Clouds
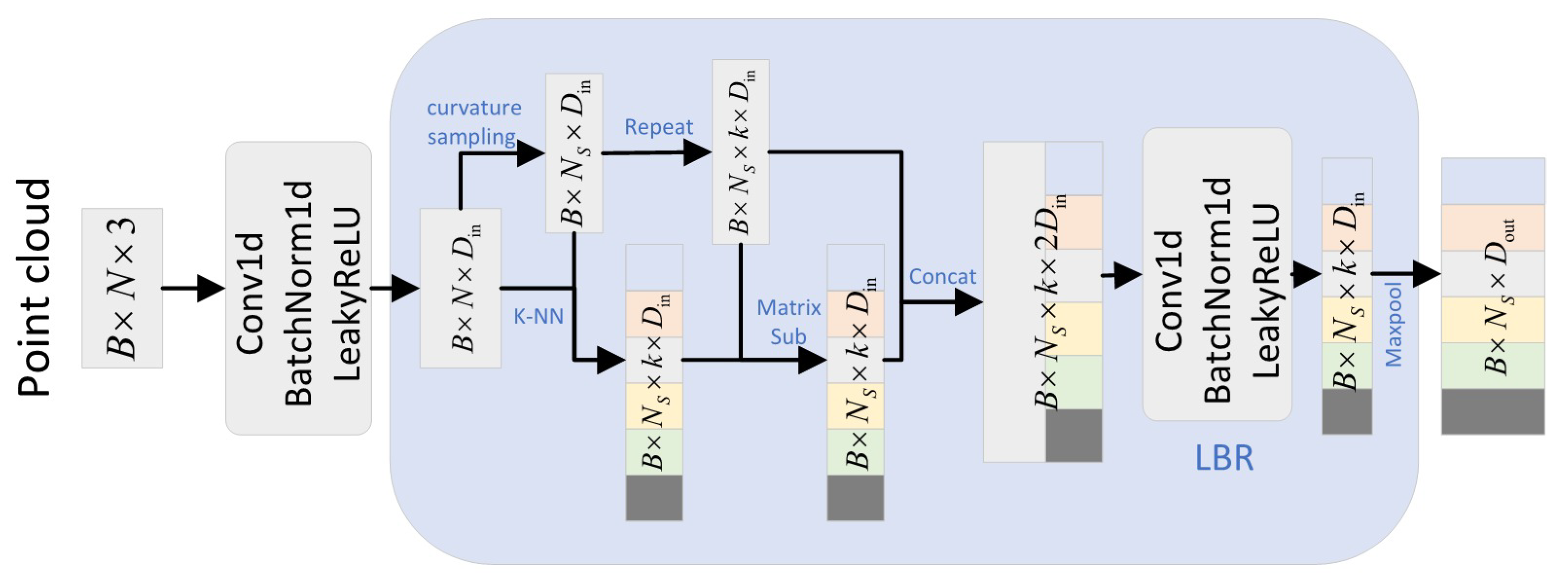
2.2.6. Neighborhood Aggregation Strategy
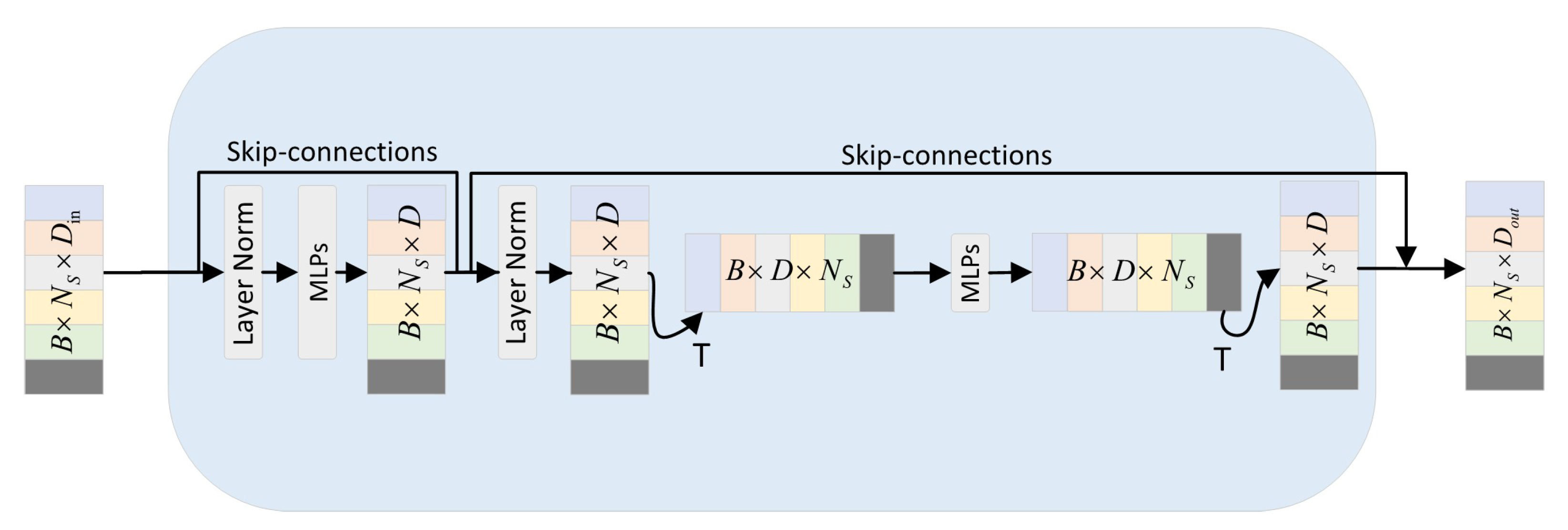
2.2.7. Point-Mixer Mechanism
2.2.8. Loss Function
3. Results
3.1. Point-Cloud Noise Removing
3.2. Evaluation Metrics
3.3. Effectiveness of MIX-Net Network on Seedling Datasets
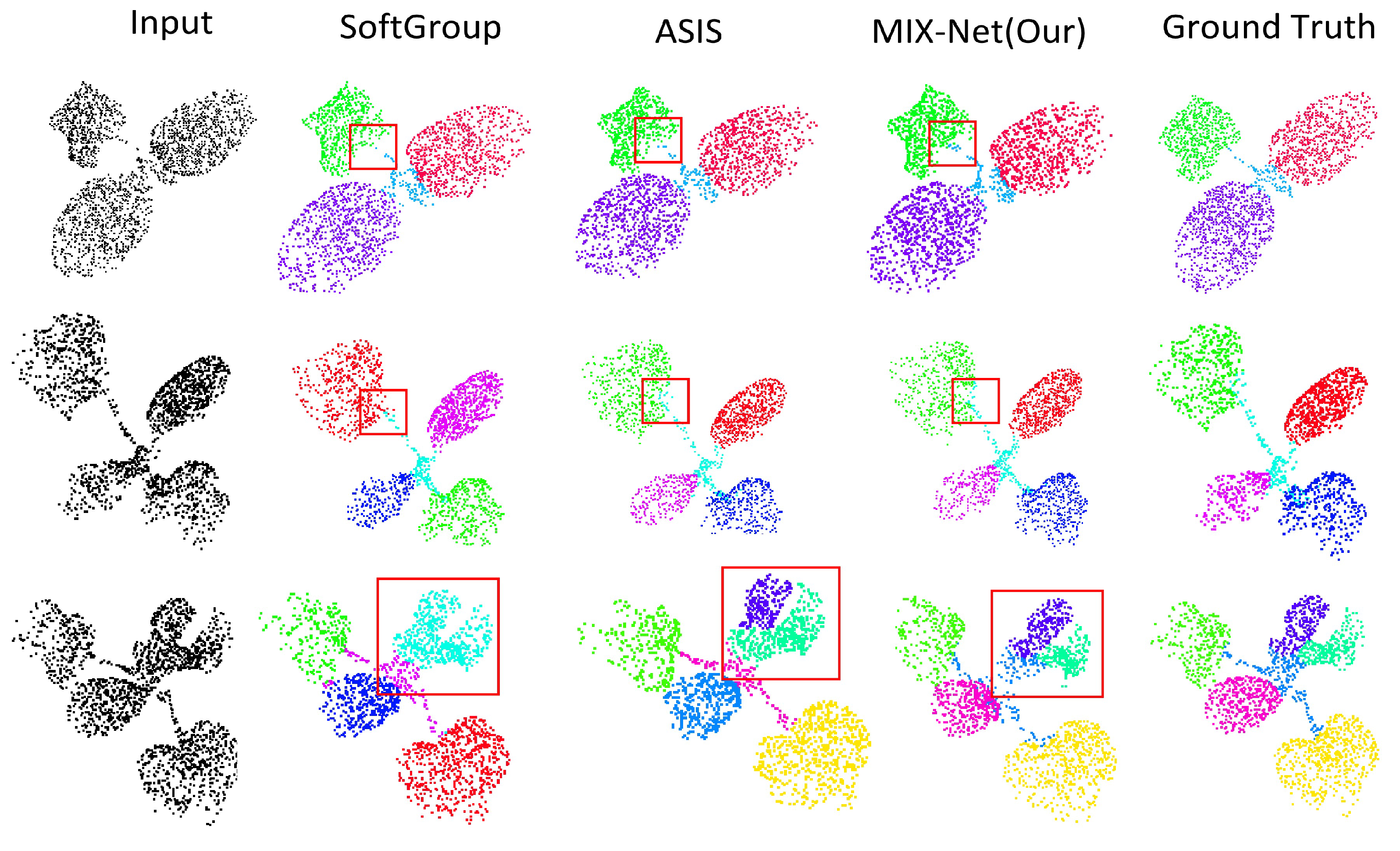
Results of Seedling Leaf Segmentation
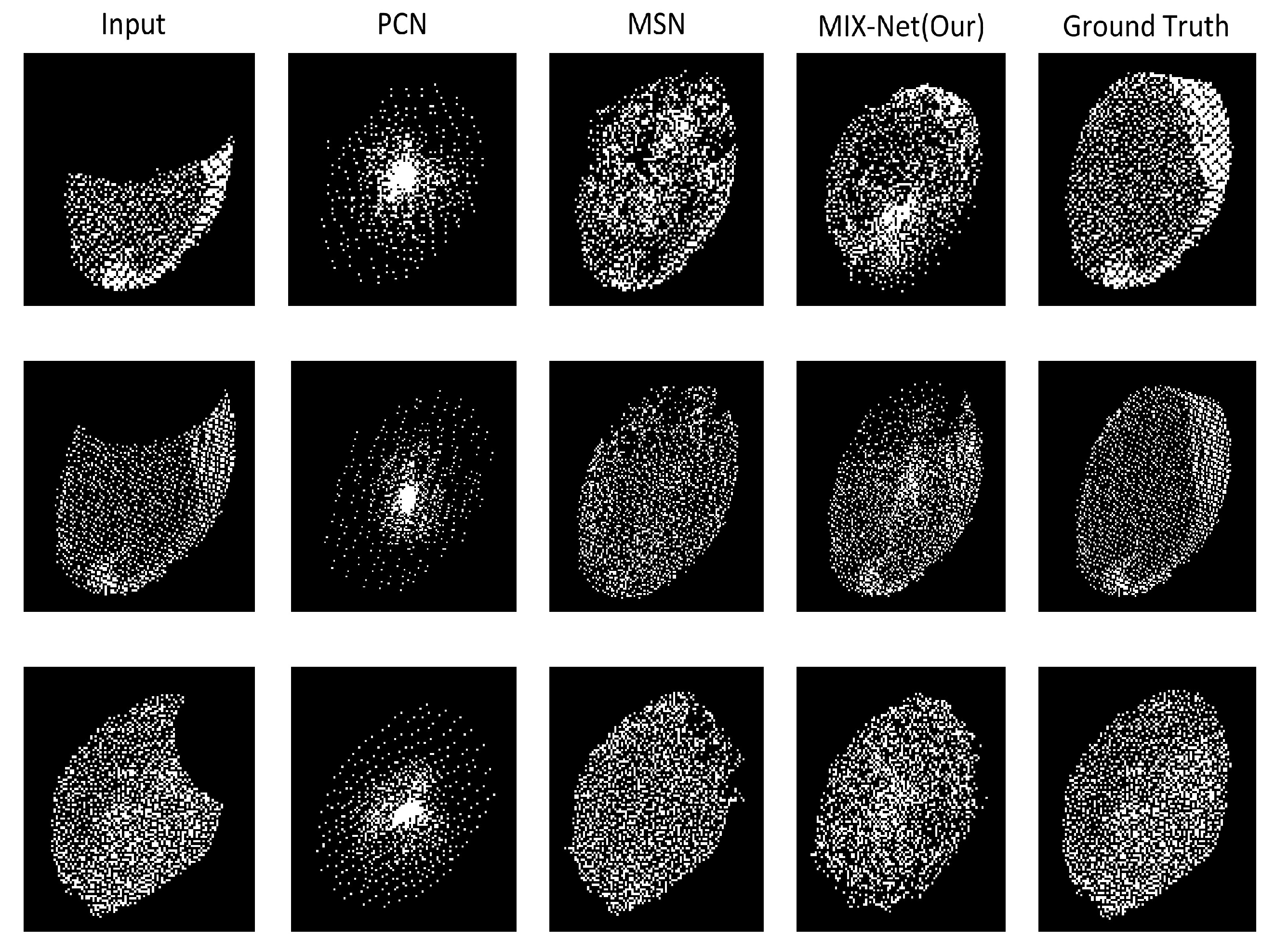
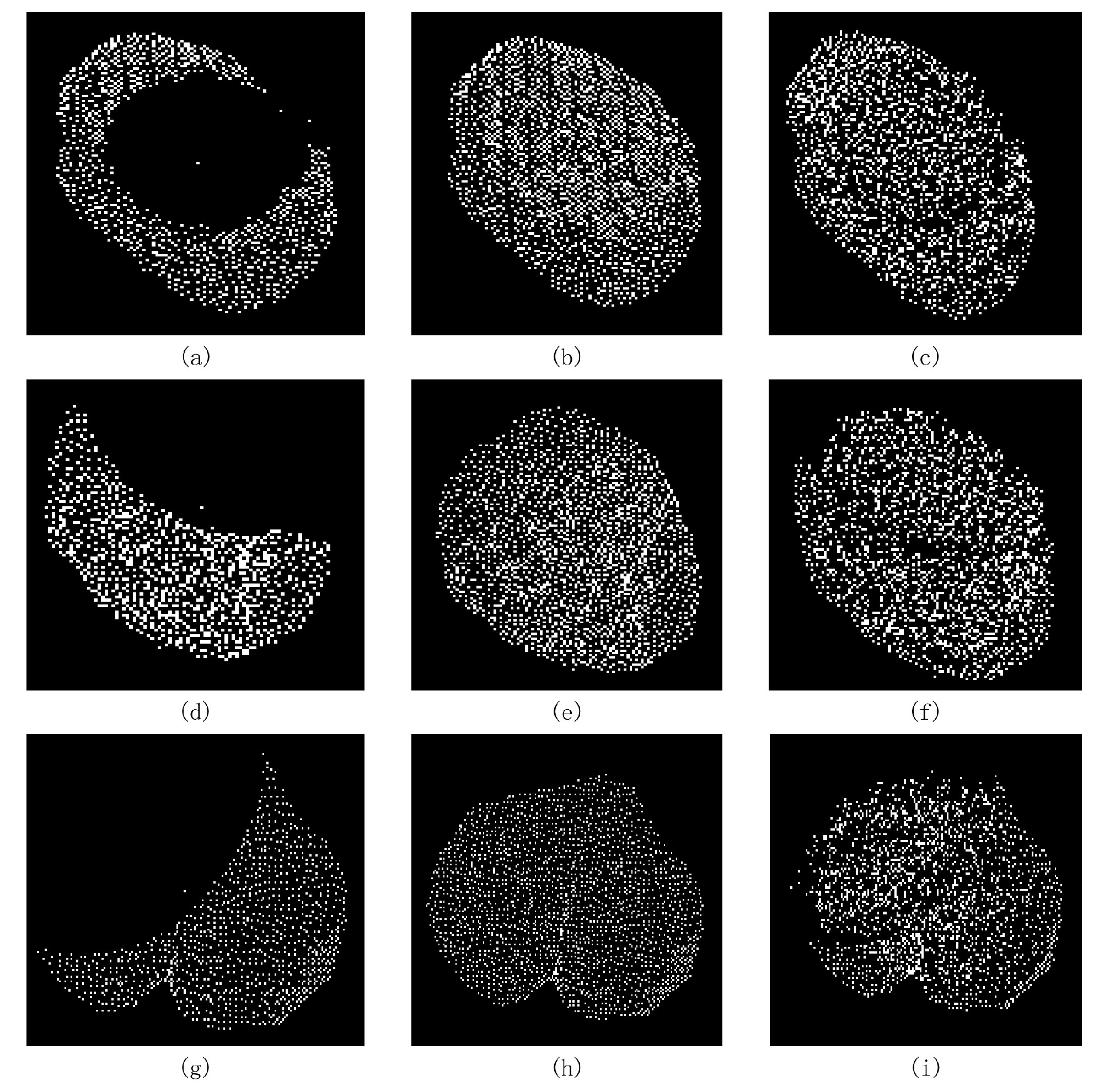
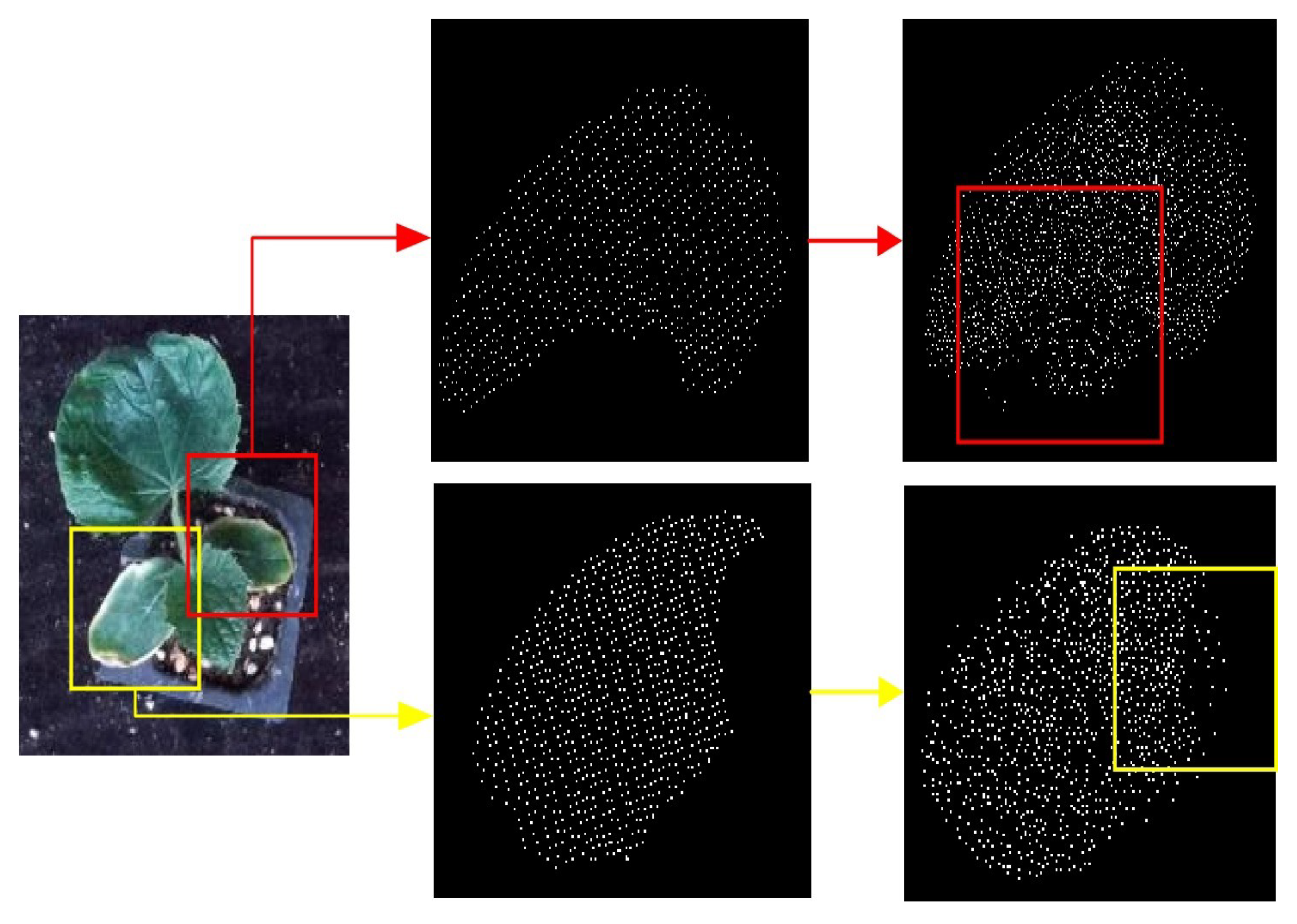
3.4. Results of MIX-Net Applied to Leaf Completion under Self-Supervised Learning
3.5. Results of MIX-Net Applied to Leaf Completion under Supervised Learning
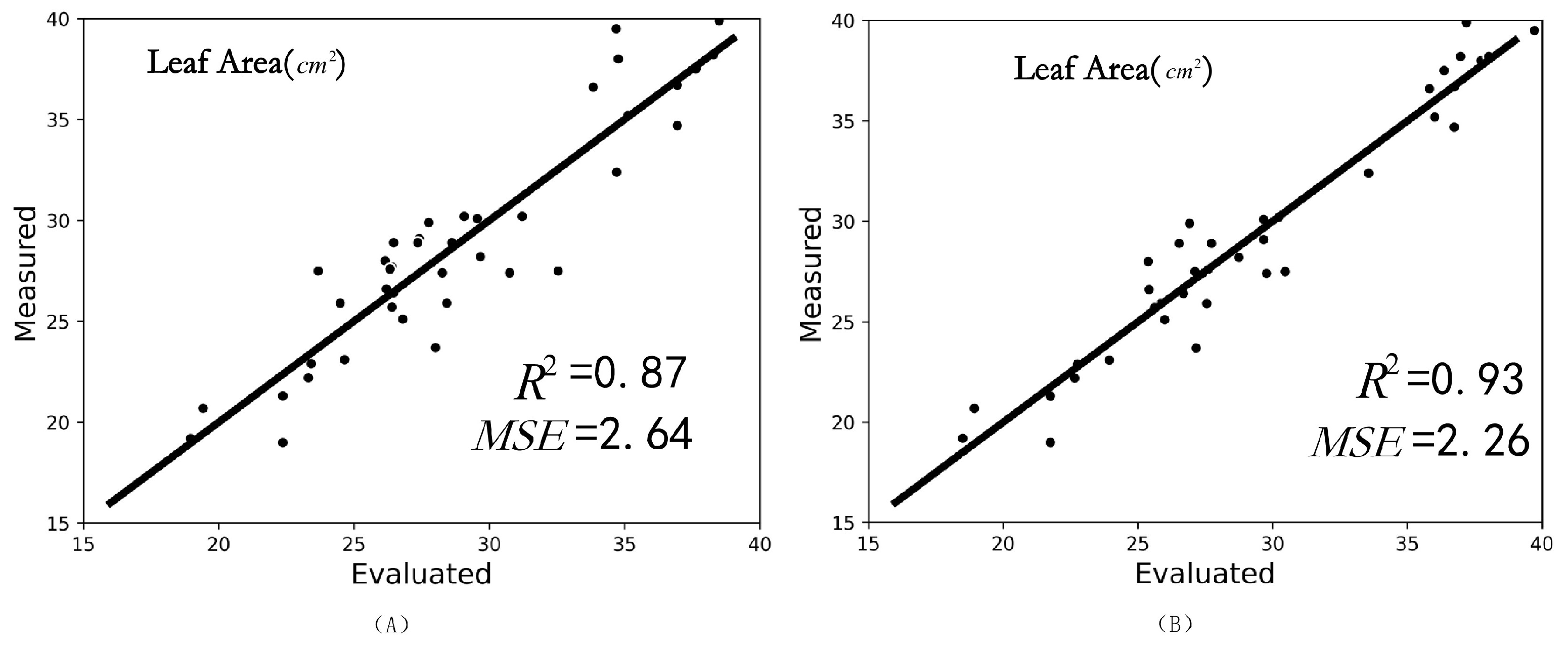
3.6. Nondestructive Leaf Area Measurement Results Using MIX-Net
4. Discussion
4.1. Point Cloud Classification Results on the Modelnet40 Datasets
4.2. Point Cloud Segmentation Results on the ShapeNet-Part Dataset
4.3. Point Cloud Completion Validated on a ShapeNet-Part Dataset
4.4. Ablation Experiments
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Feng, L.; Raza, M.A.; Li, Z.; Chen, Y.; Khalid, M.H.B.; Du, J.; Liu, W.; Wu, X.; Song, C.; Yu, L.; et al. The influence of light intensity and leaf movement on photosynthesis characteristics and carbon balance of soybean. Front. Plant Sci. 2019, 9, 1952. [Google Scholar] [CrossRef]
- Ninomiya, S.; Baret, F.; Cheng, Z.M.M. Plant phenomics: Emerging transdisciplinary science. Plant Phenomics 2019, 2019, 2765120. [Google Scholar] [CrossRef]
- Liu, H.J.; Yan, J. Crop genome-wide association study: A harvest of biological relevance. Plant J. 2019, 97, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Gara, T.W.; Skidmore, A.K.; Darvishzadeh, R.; Wang, T. Leaf to canopy upscaling approach affects the estimation of canopy traits. GIScience Remote Sens. 2019, 56, 554–575. [Google Scholar] [CrossRef]
- Fu, L.; Tola, E.; Al-Mallahi, A.; Li, R.; Cui, Y. A novel image processing algorithm to separate linearly clustered kiwifruits. Biosyst. Eng. 2019, 183, 184–195. [Google Scholar] [CrossRef]
- Sapoukhina, N.; Samiei, S.; Rasti, P.; Rousseau, D. Data augmentation from RGB to chlorophyll fluorescence imaging application to leaf segmentation of Arabidopsis thaliana from top view images. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, Long Beach, CA, USA, 16–17 June 2019. [Google Scholar]
- Panjvani, K.; Dinh, A.V.; Wahid, K.A. LiDARPheno—A low-cost lidar-based 3D scanning system for leaf morphological trait extraction. Front. Plant Sci. 2019, 10, 147. [Google Scholar] [CrossRef]
- Hu, C.; Li, P.; Pan, Z. Phenotyping of poplar seedling leaves based on a 3D visualization method. Int. J. Agric. Biol. Eng. 2018, 11, 145–151. [Google Scholar] [CrossRef]
- Wu, S.; Wen, W.; Wang, Y.; Fan, J.; Wang, C.; Gou, W.; Guo, X. MVS-Pheno: A portable and low-cost phenotyping platform for maize shoots using multiview stereo 3D reconstruction. Plant Phenomics 2020, 2020, 1848437. [Google Scholar] [CrossRef]
- Wang, Y.; Wen, W.; Wu, S.; Wang, C.; Yu, Z.; Guo, X.; Zhao, C. Maize plant phenotyping: Comparing 3D laser scanning, multi-view stereo reconstruction, and 3D digitizing estimates. Remote Sens. 2018, 11, 63. [Google Scholar] [CrossRef]
- Xu, H.; Hou, J.; Yu, L.; Fei, S. 3D Reconstruction system for collaborative scanning based on multiple RGB-D cameras. Pattern Recognit. Lett. 2019, 128, 505–512. [Google Scholar] [CrossRef]
- Teng, X.; Zhou, G.; Wu, Y.; Huang, C.; Dong, W.; Xu, S. Three-dimensional reconstruction method of rapeseed plants in the whole growth period using RGB-D camera. Sensors 2021, 21, 4628. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.E.; Park, R.H. Segmentation with saliency map using colour and depth images. IET Image Process. 2015, 9, 62–70. [Google Scholar] [CrossRef]
- Hu, Y.; Wu, Q.; Wang, L.; Jiang, H. Multiview point clouds denoising based on interference elimination. J. Electron. Imaging 2018, 27, 023009. [Google Scholar] [CrossRef]
- Ma, Z.; Sun, D.; Xu, H.; Zhu, Y.; He, Y.; Cen, H. Optimization of 3D Point Clouds of Oilseed Rape Plants Based on Time-of-Flight Cameras. Sensors 2021, 21, 664. [Google Scholar] [CrossRef] [PubMed]
- Hazirbas, C.; Ma, L.; Domokos, C.; Cremers, D. Fusenet: Incorporating depth into semantic segmentation via fusion-based cnn architecture. In Proceedings of the Asian Conference on Computer Vision, Taipei, Taiwan, 20–24 November 2016; pp. 213–228. [Google Scholar]
- van Dijk, A.D.J.; Kootstra, G.; Kruijer, W.; de Ridder, D. Machine learning in plant science and plant breeding. Iscience 2021, 24, 101890. [Google Scholar] [CrossRef] [PubMed]
- Hesami, M.; Jones, A.M.P. Application of artificial intelligence models and optimization algorithms in plant cell and tissue culture. Appl. Microbiol. Biotechnol. 2020, 104, 9449–9485. [Google Scholar] [CrossRef]
- Singh, A.; Ganapathysubramanian, B.; Singh, A.K.; Sarkar, S. Machine learning for high-throughput stress phenotyping in plants. Trends Plant Sci. 2016, 21, 110–124. [Google Scholar] [CrossRef]
- Grinblat, G.L.; Uzal, L.C.; Larese, M.G.; Granitto, P.M. Deep learning for plant identification using vein morphological patterns. Comput. Electron. Agric. 2016, 127, 418–424. [Google Scholar] [CrossRef]
- Duan, T.; Chapman, S.; Holland, E.; Rebetzke, G.; Guo, Y.; Zheng, B. Dynamic quantification of canopy structure to characterize early plant vigour in wheat genotypes. J. Exp. Bot. 2016, 67, 4523–4534. [Google Scholar] [CrossRef]
- Itakura, K.; Hosoi, F. Automatic leaf segmentation for estimating leaf area and leaf inclination angle in 3D plant images. Sensors 2018, 18, 3576. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Li, C.; Takeda, F.; Kramer, E.A.; Ashrafi, H.; Hunter, J. 3D point cloud data to quantitatively characterize size and shape of shrub crops. Hortic. Res. 2019, 6, 43. [Google Scholar] [CrossRef] [PubMed]
- Qi, C.R.; Yi, L.; Su, H.; Guibas, L.J. Pointnet++: Deep hierarchical feature learning on point sets in a metric space. In Proceedings of the 1st Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, CA, USA, 4–9 December 2017; Volume 30. [Google Scholar]
- Masuda, T. Leaf area estimation by semantic segmentation of point cloud of tomato plants. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, BC, Canada, 11–17 October 2021; pp. 1381–1389. [Google Scholar]
- Li, D.; Li, J.; Xiang, S.; Pan, A. PSegNet: Simultaneous Semantic and Instance Segmentation for Point Clouds of Plants. Plant Phenomics 2022, 2022, 9787643. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Sun, Y.; Liu, Z.; Sarma, S.E.; Bronstein, M.M.; Solomon, J.M. Dynamic graph cnn for learning on point clouds. Acm Trans. Graph. (tog) 2019, 38, 1–12. [Google Scholar] [CrossRef]
- Tolstikhin, I.O.; Houlsby, N.; Kolesnikov, A.; Beyer, L.; Zhai, X.; Unterthiner, T.; Yung, J.; Steiner, A.; Keysers, D.; Uszkoreit, J.; et al. Mlp-mixer: An all-mlp architecture for vision. Adv. Neural Inf. Process. Syst. 2021, 34, 24261–24272. [Google Scholar]
- Pan, L.; Chew, C.M.; Lee, G.H. PointAtrousGraph: Deep hierarchical encoder-decoder with point atrous convolution for unorganized 3D points. In Proceedings of the 2020 IEEE International Conference on Robotics and Automation (ICRA), Paris, France, 31 May–31 August 2020; pp. 1113–1120. [Google Scholar]
- Kazhdan, M.; Hoppe, H. Screened poisson surface reconstruction. ACM Trans. Graph. (ToG) 2013, 32, 1–13. [Google Scholar] [CrossRef]
- Mitra, N.J.; Pauly, M.; Wand, M.; Ceylan, D. Symmetry in 3d geometry: Extraction and applications. Comput. Graphics Forum 2013, 32, 1–23. [Google Scholar] [CrossRef]
- Yang, B.; Wen, H.; Wang, S.; Clark, R.; Markham, A.; Trigoni, N. 3d object reconstruction from a single depth view with adversarial learning. In Proceedings of the IEEE international Conference on Computer Vision Workshops, Venice, Italy, 22–29 October 2017; pp. 679–688. [Google Scholar]
- Yuan, W.; Khot, T.; Held, D.; Mertz, C.; Hebert, M. Pcn: Point completion network. In Proceedings of the IEEE 2018 International Conference on 3D Vision (3DV), Verona, Italy, 5–8 September 2018; pp. 728–737. [Google Scholar]
- Pan, L.; Chen, X.; Cai, Z.; Zhang, J.; Zhao, H.; Yi, S.; Liu, Z. Variational relational point completion network. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Nashville, TN, USA, 20–25 June 2021; pp. 8524–8533. [Google Scholar]
- Li, Y.; Bu, R.; Sun, M.; Wu, W.; Di, X.; Chen, B. Pointcnn: Convolution on x-transformed points. In Proceedings of the 32nd Conference on Neural Information Processing Systems (NeurIPS 2018), Montreal, QC, Canada, 3–8 December 2018. [Google Scholar]
- Lin, T.Y.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature pyramid networks for object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2117–2125. [Google Scholar]
- Pagani, L.; Scott, P.J. Curvature based sampling of curves and surfaces. Comput. Aided Geom. Des. 2018, 59, 32–48. [Google Scholar] [CrossRef]
- Fan, H.; Su, H.; Guibas, L.J. A point set generation network for 3d object reconstruction from a single image. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 605–613. [Google Scholar]
- Vu, T.; Kim, K.; Luu, T.M.; Nguyen, T.; Yoo, C.D. SoftGroup for 3D Instance Segmentation on Point Clouds. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, New Orleans, LA, USA, 18–24 June 2022; pp. 2708–2717. [Google Scholar]
- Wang, X.; Liu, S.; Shen, X.; Shen, C.; Jia, J. Associatively segmenting instances and semantics in point clouds. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 15–20 June 2019; pp. 4096–4105. [Google Scholar]
- Liu, M.; Sheng, L.; Yang, S.; Shao, J.; Hu, S.M. Morphing and sampling network for dense point cloud completion. In Proceedings of the AAAI Conference on Artificial Intelligence, New York, NY, USA, 7–12 February 2020; Volume 34, pp. 11596–11603. [Google Scholar]
- Huang, Z.; Yu, Y.; Xu, J.; Ni, F.; Le, X. Pf-net: Point fractal network for 3d point cloud completion. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, 13–19 June 2020; pp. 7662–7670. [Google Scholar]
- Wu, Z.; Song, S.; Khosla, A.; Yu, F.; Zhang, L.; Tang, X.; Xiao, J. 3d shapenets: A deep representation for volumetric shapes. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1912–1920. [Google Scholar]
- Li, R.; Li, X.; Heng, P.A.; Fu, C.W. Point cloud upsampling via disentangled refinement. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Nashville, TN, USA, 20–25 June 2021; pp. 344–353. [Google Scholar]
- Guo, M.H.; Cai, J.X.; Liu, Z.N.; Mu, T.J.; Martin, R.R.; Hu, S.M. Pct: Point cloud transformer. Comput. Vis. Media 2021, 7, 187–199. [Google Scholar] [CrossRef]
- Yi, L.; Kim, V.G.; Ceylan, D.; Shen, I.C.; Yan, M.; Su, H.; Lu, C.; Huang, Q.; Sheffer, A.; Guibas, L. A scalable active framework for region annotation in 3d shape collections. ACM Trans. Graph. (ToG) 2016, 35, 1–12. [Google Scholar] [CrossRef]
- Tchapmi, L.P.; Kosaraju, V.; Rezatofighi, H.; Reid, I.; Savarese, S. Topnet: Structural point cloud decoder. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 15–20 June 2019; pp. 383–392. [Google Scholar]



| Numder of Training Point Clouds | Number of Testing Point Clouds | Points | Number of Training Point Clouds after Augmention | Number of Testing Point Clouds after Augmention | |
|---|---|---|---|---|---|
| Number of seedlings point cloud | 130 | 20 | 2048 | 520 | 80 |
| Number of leaf point clouds | 500 | 100 | 2048 | 1800 | 300 |
| Methods | Input | Points | mIoU (%) |
|---|---|---|---|
| PointNet++ [24] | P | 2048 | 91.5 |
| DGCNN [21] | P | 2048 | 92.9 |
| MIX-Net (Our) | P | 2048 | 94.6 |
| Methods | Input | Points | mPrec (%) | mRec (%) |
|---|---|---|---|---|
| Soft-Group [39] | P | 2048 | 74.26 | 68.04 |
| ASIS [40] | P | 2048 | 79.13 | 75.64 |
| MIX-Net (Our) | P | 2048 | 82.31 | 77.46 |
| Methods | Input | Points | m-Value | CD × | EMD |
|---|---|---|---|---|---|
| PCN [33] | P | 2048 | 50%, 25%, 15% | 1.947 | 0.106 |
| MSN [41] | P | 2048 | 50%, 25%, 15% | 0.870 | 0.072 |
| PF-Net [42] | P | 2048 | 50%, 25%, 15% | 1.947 | – |
| Vrc-Net [34] | P | 2048 | 50%, 25%, 15% | 1.783 | 0.107 |
| MIX-Net (Our) | P | 2048 | 50%, 25%, 15% | 1.679 | 0.071 |
| Methods | Input | Points | m-Value | CD × | EMD |
|---|---|---|---|---|---|
| PCN [33] | P | 2048 | 50%, 25%, 15% | 1.773 | 0.113 |
| MSN [41] | P | 2048 | 50%, 25%, 15% | 1.914 | 0.065 |
| MIX-Net (Our) | P | 2048 | 50%, 25%, 15% | 1.276 | 0.063 |
| Methods | Input | Points | Accuracy (%) |
|---|---|---|---|
| PointNet++ [24] | P | 1024 | 90.7 |
| PointNet++ [24] | P, N | 1024 | 91.9 |
| PointCNN [35] | P | 1024 | 92.5 |
| DGCNN [27] | P | 1024 | 92.9 |
| PCT [45] | P | 1024 | 93.2 |
| MIX-Net (Our) | P | 1024 | 93.4 |
| Methods | Input | Points | mIoU (%) |
|---|---|---|---|
| PointNet++ [24] | P | 2048 | 85.1 |
| DGCNN [27] | P | 2048 | 85.2 |
| MIX-Net (Our) | P | 2048 | 85.7 |
| Methods | Input | Points | m-Value | CD × | F-Score@1% |
|---|---|---|---|---|---|
| PCN [33] | P | 2048 | 50% | 2.929 | 0.29 |
| TopNet [47] | P | 2048 | 50% | 3.805 | 0.38 |
| MSN [41] | P | 2048 | 50% | 2.376 | 0.41 |
| PF-Net [42] | P | 2048 | 50% | 3.037 | – |
| Vrc-Net [34] | P | 2048 | 50% | 2.881 | 0.42 |
| MIX-Net (Our) | P | 2048 | 50% | 2.111 | 0.45 |
| classification (modelnet40 dataset) | PointNet++ (encoder) [24] | Nas | Point-mixer | Accuracy (%) | ||
| ✓ | 90.7 | |||||
| ✓ | 89.4 | |||||
| ✓ | ✓ | 92.7 | ||||
| ✓ | ✓ | 93.4 | ||||
| semantic segmentation (seedling semantic segmentation dataset) | PointNet++ (encoder) [24] | PointNet++ (decoder) [24] | Nas | Point-mixer | mIoU (%) | |
| ✓ | ✓ | 91.5 | ||||
| ✓ | ✓ | 92.4 | ||||
| ✓ | ✓ | ✓ | 93.7 | |||
| ✓ | ✓ | 91.8 | ||||
| ✓ | ✓ | 94.6 | ||||
| instance segmentation (seedling instance segmentation dataset) | ASIS (encoder) [40] | ASIS (decoder) [40] | Nas | Point-mixer | mPrec (%) | mRec (%) |
| ✓ | ✓ | 79.13 | 75.64 | |||
| ✓ | ✓ | 77.41 | 72.36 | |||
| ✓ | ✓ | ✓ | 81.32 | 79.56 | ||
| ✓ | ✓ | 78.44 | 76.54 | |||
| ✓ | ✓ | 82.31 | 77.46 | |||
| Leaf completion (seedling leaf completion dataset) | PCN (encoder) [33] | PCN (decoder) [33] | Nas | Point-mixer | CD × | EMD |
| ✓ | ✓ | 1.773 | 0.113 | |||
| ✓ | ✓ | 1.345 | 0.094 | |||
| ✓ | ✓ | ✓ | 1.254 | 0.061 | ||
| ✓ | ✓ | 1.493 | 0.108 | |||
| ✓ | ✓ | 1.276 | 0.059 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Han, B.; Li, Y.; Bie, Z.; Peng, C.; Huang, Y.; Xu, S. MIX-NET: Deep Learning-Based Point Cloud Processing Method for Segmentation and Occlusion Leaf Restoration of Seedlings. Plants 2022, 11, 3342. https://doi.org/10.3390/plants11233342
Han B, Li Y, Bie Z, Peng C, Huang Y, Xu S. MIX-NET: Deep Learning-Based Point Cloud Processing Method for Segmentation and Occlusion Leaf Restoration of Seedlings. Plants. 2022; 11(23):3342. https://doi.org/10.3390/plants11233342
Chicago/Turabian StyleHan, Binbin, Yaqin Li, Zhilong Bie, Chengli Peng, Yuan Huang, and Shengyong Xu. 2022. "MIX-NET: Deep Learning-Based Point Cloud Processing Method for Segmentation and Occlusion Leaf Restoration of Seedlings" Plants 11, no. 23: 3342. https://doi.org/10.3390/plants11233342
APA StyleHan, B., Li, Y., Bie, Z., Peng, C., Huang, Y., & Xu, S. (2022). MIX-NET: Deep Learning-Based Point Cloud Processing Method for Segmentation and Occlusion Leaf Restoration of Seedlings. Plants, 11(23), 3342. https://doi.org/10.3390/plants11233342







