Silencing of MsD14 Resulted in Enhanced Forage Biomass through Increasing Shoot Branching in Alfalfa (Medicago sativa L.)
Abstract
:1. Introduction
2. Results
2.1. Identification of the Strigolactone Receptor Gene (D14) in Alfalfa
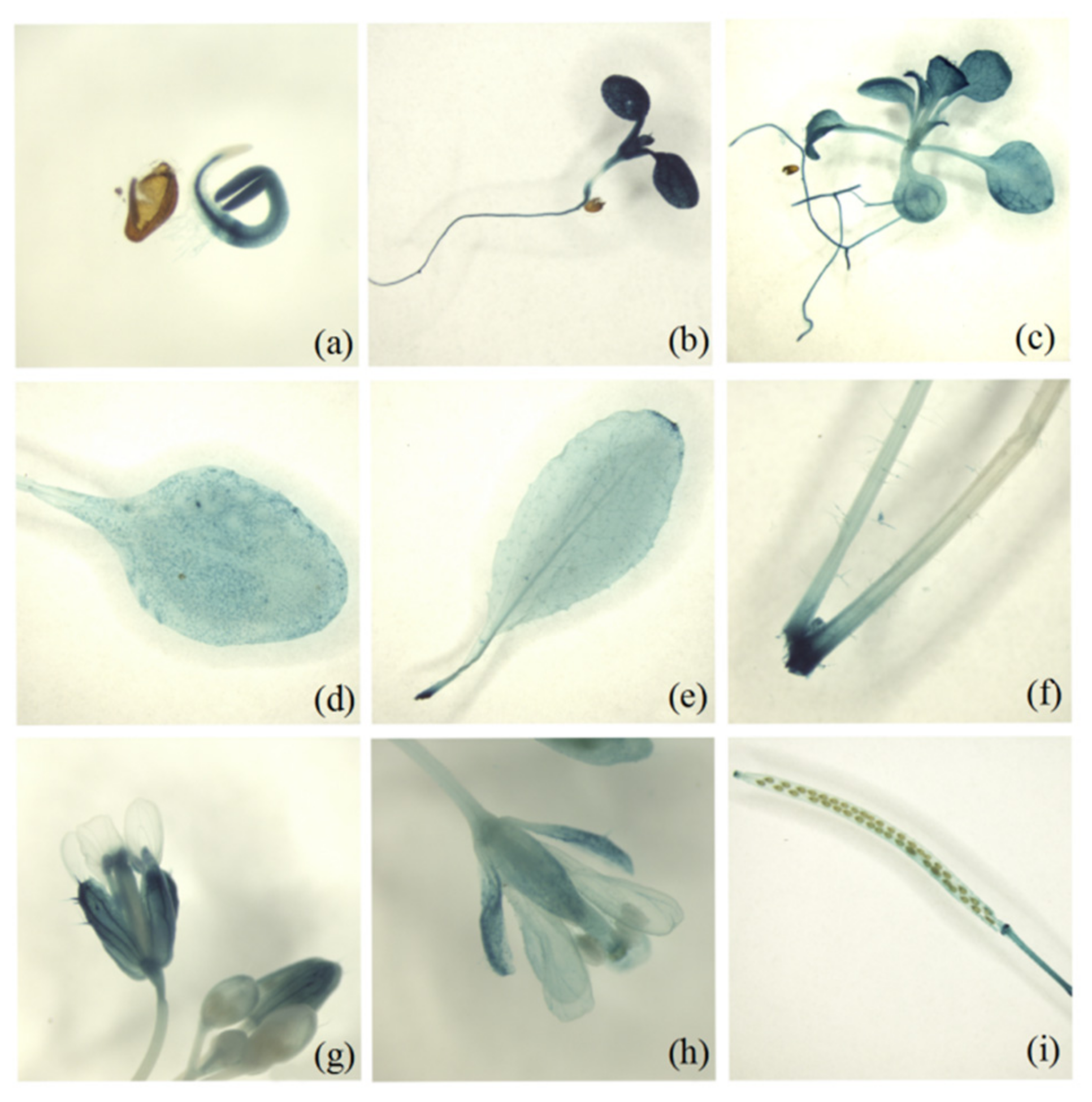
2.2. Temporal and Spatial Expression Patterns of MsD14
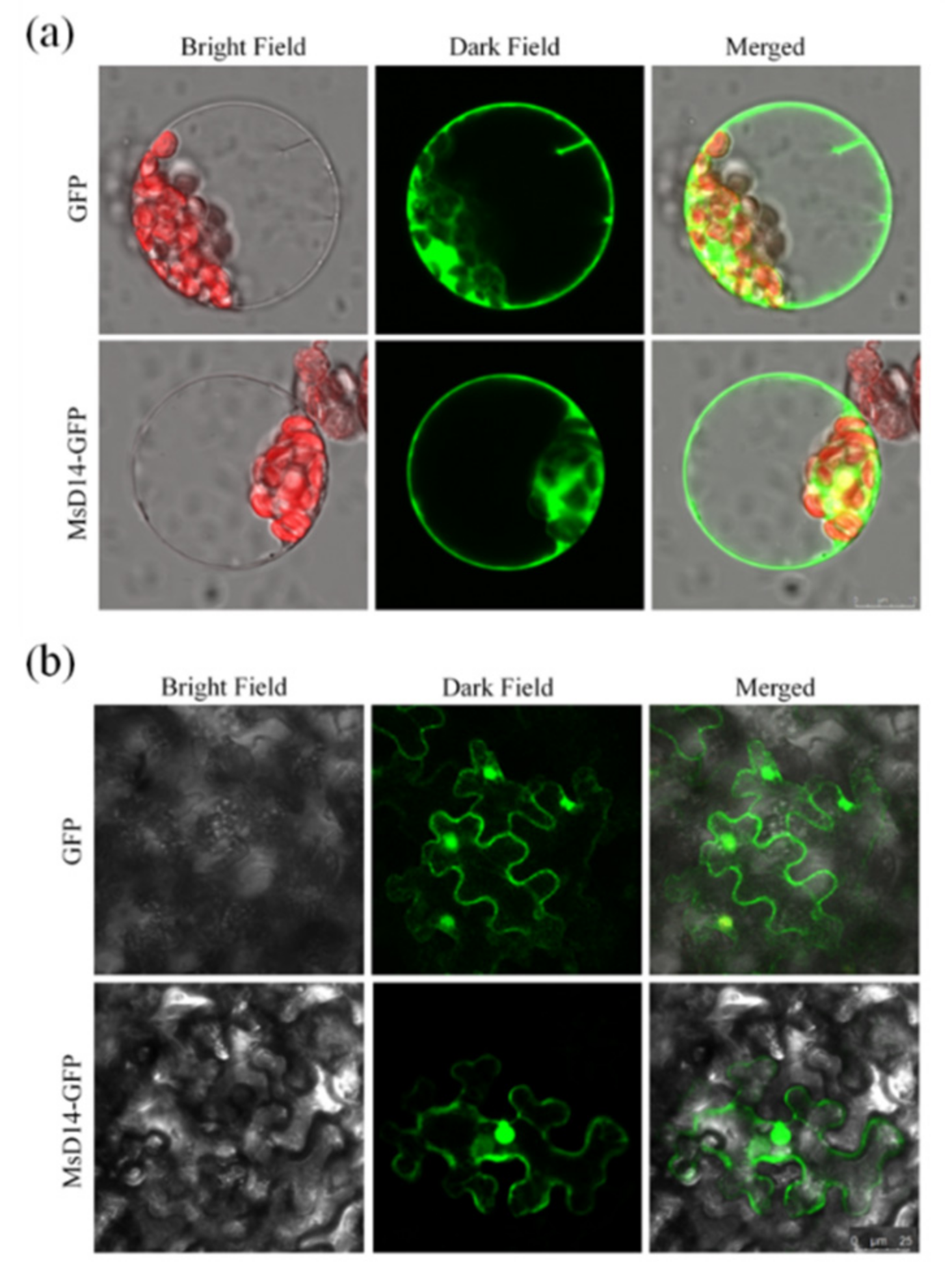
2.3. Subcellular Localization of MsD14
2.4. Silencing of MsD14 Increased the Branching Number in Alfalfa
2.5. Interaction of MsD14 with MAX2
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Nucleic Acid Extraction and PCR
4.3. MsD14 Cloning and Informatics Analysis
4.4. GUS Histochemical Assay
4.5. Subcellular Localization Analysis
4.6. Alfalfa Expression Vector Construction and Genetic Transformation
4.7. Yeast Two-Hybrid Assays
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lorenzo, C.D.; Garcia-Gagliardi, P.; Antonietti, M.S.; Sanchez-Lamas, M.; Mancini, E.; Dezar, C.A.; Vazquez, M.; Watson, G.; Yanovsky, M.J.; Cerdan, P.D. Improvement of alfalfa forage quality and management through the down-regulation of MsFTa1. Plant Biotechnol. J. 2019, 18, 944–954. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adhikari, L.; Makaju, S.O.; Missaoui, A.M. QTL mapping of flowering time and biomass yield in tetraploid alfalfa (Medicago sativa L.). BMC Plant Biol. 2019, 19, 359. [Google Scholar] [CrossRef] [PubMed]
- Gou, J.Q.; Debnath, S.; Sun, L.; Flanagan, A.; Tang, Y.H.; Jiang, Q.Z.; Wen, J.Q.; Wang, Z.Y. From model to crop: Functional characterization of SPL8 in M.truncatula led to genetic improvement of biomass yield and abiotic stress tolerance in alfalfa. Plant Biotechnol. J. 2018, 16, 951–962. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ward, S.P.; Leyser, O. Shoot branching. Curr. Opin. Plant Biol. 2004, 7, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Kotov, A.A.; Kotova, L.M.; Romanov, G.A. Signaling network regulating plant branching: Recent advances and new challenges. Plant Sci. 2021, 307, 110880. [Google Scholar] [CrossRef] [PubMed]
- Kebrom, T.H.; Spielmeyer, W.; Finnegan, E.J. Grasses provide new insights into regulation of shoot branching. Trends Plant Sci. 2013, 18, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.H.; Li, J.Y. Branching in rice. Curr. Opin. Plant Biol. 2011, 14, 94–99. [Google Scholar] [CrossRef]
- Harrison, C.J. Auxin transport in the evolution of branching forms. New Phytol. 2017, 215, 545–551. [Google Scholar] [CrossRef] [Green Version]
- Waldie, T.; Leyser, O. Cytokinin targets auxin transport to promote shoot branching. Plant Physiol. 2018, 177, 803–818. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Sun, S.Y.; Wang, X.L. Regulation of shoot branching by strigolactones and brassinosteroids: Conserved and specific functions of Arabidopsis BES1 and rice BZR1. Mol. Plant 2020, 13, 808–810. [Google Scholar] [CrossRef]
- Booker, J.; Chatfield, S.; Leyser, O. Auxin acts in xylem-associated or medullary cells to mediate apical dominance. Plant Cell 2003, 15, 495–507. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vernoux, T.; Besnard, F.; Traas, J. Auxin at the shoot apical meristem. C.S.H. Perspect. Biol. 2010, 2, a1001487. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nordstrom, A.; Tarkowski, P.; Tarkowska, D.; Norbaek, R.; Astot, C.; Dolezal, K.; Sandberg, G. Auxin regulation of cytokinin biosynthesis in Arabidopsis thaliana: A factor of potential importance for auxin-cytokinin-regulated development. Proc. Natl. Acad. Sci. USA 2004, 101, 8039–8044. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dun, E.A.; de Saint Germain, A.; Rameau, C.; Beveridge, C.A. Antagonistic action of strigolactone and cytokinin in bud outgrowth control. Plant Physiol. 2012, 158, 487–498. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, X.J.; Dong, H.; Yin, Y.L.; Song, X.W.; Gu, X.H.; Sang, K.Q.; Zhou, J.; Shi, K.; Zhou, Y.H.; Foyer, C.H.; et al. Brassinosteroid signaling integrates multiple pathways to release apical dominance in tomato. Proc. Natl. Acad. Sci. USA 2021, 118, e2004384118. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Roldan, V.; Fermas, S.; Brewer, P.B.; Puech-Pages, V.; Dun, E.A.; Pillot, J.P.; Letisse, F.; Matusova, R.; Danoun, S.; Portais, J.C.; et al. Strigolactone inhibition of shoot branching. Nature 2008, 455, 189–194. [Google Scholar] [CrossRef]
- Yao, R.F.; Li, J.Y.; Xie, D.X. Recent advances in molecular basis for strigolactone action. Sci. China Life Sci. 2018, 61, 277–284. [Google Scholar] [CrossRef]
- Qu, B.Y.; Qin, Y.; Bai, Y. From signaling to function: How strigolactones regulate plant development. Sci. China Life Sci. 2020, 63, 1768–1770. [Google Scholar] [CrossRef]
- Jiang, L.; Liu, X.; Xiong, G.S.; Liu, H.H.; Chen, F.L.; Wang, L.; Meng, X.B.; Liu, G.F.; Yu, H.; Yuan, Y.D.; et al. DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature 2013, 504, 401–405. [Google Scholar] [CrossRef] [Green Version]
- Zhou, F.; Lin, Q.B.; Zhu, L.H.; Ren, Y.L.; Zhou, K.N.; Shabek, N.; Wu, F.Q.; Mao, H.B.; Dong, W.; Gan, L.; et al. D14-SCFD3-dependent degradation of D53 regulates strigolactone signalling. Nature 2013, 504, 406–410. [Google Scholar] [CrossRef] [Green Version]
- Yao, R.F.; Ming, Z.H.; Yan, L.M.; Li, S.H.; Wang, F.; Ma, S.; Yu, C.T.; Yang, M.; Chen, L.; Chen, L.H.; et al. DWARF14 is a non-canonical hormone receptor for strigolactone. Nature 2016, 536, 469–473. [Google Scholar] [CrossRef] [PubMed]
- Soundappan, I.; Bennett, T.; Morffy, N.; Liang, Y.Y.; Stang, J.P.; Abbas, A.; Leyser, O.; Nelson, D.C. SMAX1-LIKE/D53 family members enable distinct MAX2-dependent responses to strigolactones and karrikins in Arabidopsis. Plant Cell 2015, 27, 3143–3159. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, B.; Jiang, L.; Liu, X.; Li, X.L.; Lu, Z.F.; Meng, X.B.; Wang, Y.H.; Smith, S.M.; Li, J.Y. Strigolactone signaling in Arabidopsis regulates shoot development by targeting D53-Like SMXL repressor proteins for ubiquitination and degradation. Plant Cell 2015, 27, 3128–3142. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takeda, T.; Suwa, Y.; Suzuki, M.; Kitano, H.; Ueguchi-Tanaka, M.; Ashikari, M.; Matsuoka, M.; Ueguchi, C. The OsTB1 gene negatively regulates lateral branching in rice. Plant J. 2003, 33, 513–520. [Google Scholar] [CrossRef]
- Aguilar-Martinez, J.A.; Poza-Carrion, C.; Cubas, P. Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell 2007, 19, 458–472. [Google Scholar] [CrossRef]
- Wang, L.; Wang, B.; Yu, H.; Guo, H.Y.; Lin, T.; Kou, L.Q.; Wang, A.Q.; Shao, N.; Ma, H.Y.; Xiong, G.S.; et al. Transcriptional regulation of strigolactone signalling in Arabidopsis. Nature 2020, 583, 277–282. [Google Scholar] [CrossRef]
- Liu, R.F.; Hou, J.; Li, H.F.; Xu, P.; Zhang, Z.B.; Zhang, X.Y. Association of TaD14-4D, a gene involved in strigolactone signaling, with yield contributing traits in wheat. Int. J. Mol. Sci. 2021, 22, 3748. [Google Scholar] [CrossRef]
- Yang, Y.Y.; Ren, Y.R.; Zheng, P.F.; Zhao, L.L.; You, C.X.; Wang, X.F.; Hao, Y.J. Cloning and functional identification of a strigolactone receptor gene MdD14 in apple. Plant Cell Tiss. Org. 2020, 140, 197–208. [Google Scholar] [CrossRef]
- Marzec, M.; Gruszka, D.; Tylec, P.; Szarejko, I. Identification and functional analysis of the HvD14 gene involved in strigolactone signaling in Hordeum vulgare. Physiol. Plant. 2016, 158, 341–355. [Google Scholar] [CrossRef]
- Zhang, J.; Mazur, E.; Balla, J.; Gallei, M.; Kalousek, P.; Medvedova, Z.; Li, Y.; Wang, Y.P.; Prat, T.; Vasileva, M.; et al. Strigolactones inhibit auxin feedback on PIN-dependent auxin transport canalization. Nat. Commun. 2020, 11, 3508. [Google Scholar] [CrossRef]
- Ahmad, M.Z.; Rehman, N.U.; Yu, S.W.; Zhou, Y.Z.; ul Haq, B.; Wang, J.J.; Li, P.H.; Zeng, Z.X.; Zhao, J. GmMAX2-D14 and -KAI interaction-mediated SL and KAR signaling play essential roles in soybean root nodulation. Plant J. 2020, 101, 334–351. [Google Scholar] [CrossRef] [PubMed]
- Aung, B.; Gruber, M.Y.; Amyot, L.; Omari, K.; Bertrand, A.; Hannoufa, A. MicroRNA156 as a promising tool for alfalfa improvement. Plant Biotechnol. J. 2015, 13, 779–790. [Google Scholar] [CrossRef] [PubMed]
- Arshad, M.; Feyissa, B.A.; Amyot, L.; Aung, B.; Hannoufa, A. MicroRNA156 improves drought stress tolerance in alfalfa (Medicago sativa) by silencing SPL13. Plant Sci. 2017, 258, 122–136. [Google Scholar] [CrossRef] [PubMed]
- Arite, T.; Umehara, M.; Ishikawa, S.; Hanada, A.; Maekawa, M.; Yamaguchi, S.; Kyozuka, J. d14, a strigolactone-insensitive mutant of rice, shows an accelerated outgrowth of tillers. Plant Cell Physiol. 2009, 50, 1416–1424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chevalier, F.; Nieminen, K.; Sanchez-Ferrero, J.C.; Rodriguez, M.L.; Chagoyen, M.; Hardtke, C.S.; Cubas, P. Strigolactone promotes degradation of DWARF14, an alpha/beta hydrolase essential for strigolactone signaling in Arabidopsis. Plant Cell 2014, 26, 1134–1150. [Google Scholar] [CrossRef] [Green Version]
- Dun, E.A.; Brewer, P.B.; Beveridge, C.A. Strigolactones: Discovery of the elusive shoot branching hormone. Trends Plant Sci. 2009, 14, 364–372. [Google Scholar] [CrossRef]
- Yoneyama, K.; Brewer, P. Strigolactones, how are they synthesized to regulate plant growth and development? Curr. Opin. Plant Biol. 2021, 63, 102072. [Google Scholar] [CrossRef]
- Hamiaux, C.; Drummond, R.S.M.; Janssen, B.J.; Ledger, S.E.; Cooney, J.M.; Newcomb, R.D.; Snowden, K.C. DAD2 is an alpha/beta hydrolase likely to be involved in the perception of the plant branching hormone, strigolactone. Curr. Biol. 2012, 22, 2032–2036. [Google Scholar] [CrossRef] [Green Version]
- Zhang, F.; Kang, J.M.; Long, R.C.; Yu, L.X.; Wang, Z.; Zhao, Z.X.; Zhang, T.J.; Yang, Q.C. High-density linkage map construction and mapping QTL for yield and yield components in autotetraploid alfalfa using RAD-seq. BMC Plant Biol. 2019, 19, 165. [Google Scholar] [CrossRef] [Green Version]
- Ray, I.M.; Monteros, M.J.; Julier, B.; Sledge, M.K.; Brummer, E.C. Identification of consensus regions associated with shoot biomass production in the Medicago genome. Crop Sci. 2018, 58, 1037–1060. [Google Scholar] [CrossRef]
- Wolabu, T.W.; Cong, L.L.; Park, J.J.; Bao, Q.Y.; Chen, M.; Sun, J.; Xu, B.; Ge, Y.X.; Chai, M.F.; Liu, Z.P.; et al. Development of a highly efficient multiplex genome editing system in outcrossing tetraploid alfalfa (Medicago sativa). Front. Plant Sci. 2020, 11, 1063. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.L.; Ishak, A.; Yu, J.; Zhao, R.Y.; Zhao, L.J. Identification and functional analysis of three MAX2 orthologs in Chrysanthemum. J. Integr. Plant Biol. 2013, 55, 434–442. [Google Scholar] [CrossRef] [PubMed]
- An, J.P.; Li, R.; Qu, F.J.; You, C.X.; Wang, X.F.; Hao, Y.J. Apple F-Box protein MdMAX2 regulates plant photomorphogenesis and stress response. Front. Plant Sci. 2016, 7, 1685. [Google Scholar] [CrossRef] [Green Version]
- Cheng, T.T.; Wang, D.H.; Wang, Y.F.; Zhang, S.M.; Zhang, C.; Liu, S.D.; Xi, Y.J.; Sun, F.L. Identification and functional characterization of a MAX2 ortholog from switchgrass (Panicum virgatum L.). Plant Physiol. Bioch. 2018, 128, 106–114. [Google Scholar] [CrossRef]
- Cardon, G.; Hohmann, S.; Klein, J.; Nettesheim, K.; Saedler, H.; Huijser, P. Molecular characterisation of the Arabidopsis SBP-box genes. Gene 1999, 237, 91–104. [Google Scholar] [CrossRef]
- Si, L.Z.; Chen, J.Y.; Huang, X.H.; Gong, H.; Luo, J.H.; Hou, Q.Q.; Zhou, T.Y.; Lu, T.T.; Zhu, J.J.; Shangguan, Y.Y.; et al. OsSPL13 controls grain size in cultivated rice. Nat. Genet. 2016, 48, 447–457. [Google Scholar] [CrossRef] [PubMed]
- Brun, G.; Braem, L.; Thoiron, S.; Gevaert, K.; Goormachtig, S.; Delavault, P. Seed germination in parasitic plants: What insights can we expect from strigolactone research? J. Exp. Bot. 2018, 69, 2265–2280. [Google Scholar] [CrossRef]
- Yamada, Y.; Otake, M.; Furukawa, T.; Shindo, M.; Shimomura, K.; Yamaguchi, S.; Umehara, M. Effects of strigolactones on grain yield and seed development in rice. J. Plant Growth Regul. 2019, 38, 753–764. [Google Scholar] [CrossRef]
- de Saint Germain, A.; Ligerot, Y.; Dun, E.A.; Pillot, J.P.; Ross, J.J.; Beveridge, C.A.; Rameau, C. Strigolactones stimulate internode elongation independently of gibberellins. Plant Physiol. 2013, 163, 1012–1025. [Google Scholar] [CrossRef] [Green Version]
- Hu, Q.N.; Zhang, S.X.; Huang, B.R. Strigolactones promote leaf elongation in tall fescue through upregulation of cell cycle genes and downregulation of auxin transport genes in tall fescue under different temperature regimes. Int. J. Mol. Sci. 2019, 20, 1836. [Google Scholar] [CrossRef] [Green Version]
- Yamada, Y.; Umehara, M. Possible roles of strigolactones during leaf senescence. Plants 2015, 4, 664–677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ha, C.V.; Leyva-Gonzalez, M.A.; Osakabe, Y.; Tran, U.T.; Nishiyama, R.; Watanabe, Y.; Tanaka, M.; Seki, M.; Yamaguchi, S.; Dong, N.V.; et al. Positive regulatory role of strigolactone in plant responses to drought and salt stress. Proc. Natl. Acad. Sci. USA 2014, 111, 851–856. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Soto, M.J.; Fernandez-Aparicio, M.; Castellanos-Morales, V.; Garcia-Garrido, J.M.; Ocampo, J.A.; Delgado, M.J.; Vierheilig, H. First indications for the involvement of strigolactones on nodule formation in alfalfa (Medicago sativa). Soil Biol. Biochem. 2010, 42, 383–385. [Google Scholar] [CrossRef]
- Ma, L.; Hao, C.Y.; Liu, H.X.; Hou, J.; Li, T.; Zhang, X.Y. Diversity and sub-functionalization of TaGW8 homoeologs hold potential for genetic yield improvement in wheat. Crop J. 2019, 7, 830–844. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [Green Version]
- Li, T.; Wu, X.Y.; Li, H.; Song, J.H.; Liu, J.Y. A dual-function transcription factor, AtYY1, is a novel negative regulator of the Arabidopsis ABA response network. Mol. Plant 2016, 9, 650–661. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.M.; Li, J.; Ban, L.P.; Wu, Y.D.; Wu, X.M.; Wang, Y.Q.; Wen, H.Y.; Chapurin, V.; Dzyubenko, N.; Li, Z.Y.; et al. Functional characterization of a gibberellin receptor and its application in alfalfa biomass improvement. Sci. Rep. 2017, 7, 41296. [Google Scholar] [CrossRef] [Green Version]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, L.; Zhang, Y.; Wen, H.; Liu, W.; Zhou, Y.; Wang, X. Silencing of MsD14 Resulted in Enhanced Forage Biomass through Increasing Shoot Branching in Alfalfa (Medicago sativa L.). Plants 2022, 11, 939. https://doi.org/10.3390/plants11070939
Ma L, Zhang Y, Wen H, Liu W, Zhou Y, Wang X. Silencing of MsD14 Resulted in Enhanced Forage Biomass through Increasing Shoot Branching in Alfalfa (Medicago sativa L.). Plants. 2022; 11(7):939. https://doi.org/10.3390/plants11070939
Chicago/Turabian StyleMa, Lin, Yongchao Zhang, Hongyu Wen, Wenhui Liu, Yu Zhou, and Xuemin Wang. 2022. "Silencing of MsD14 Resulted in Enhanced Forage Biomass through Increasing Shoot Branching in Alfalfa (Medicago sativa L.)" Plants 11, no. 7: 939. https://doi.org/10.3390/plants11070939





