Comparative Microscopic, Transcriptome and IAA Content Analyses Reveal the Stem Growth Variations in Two Cultivars Ilex verticillata
Abstract
1. Introduction
2. Results
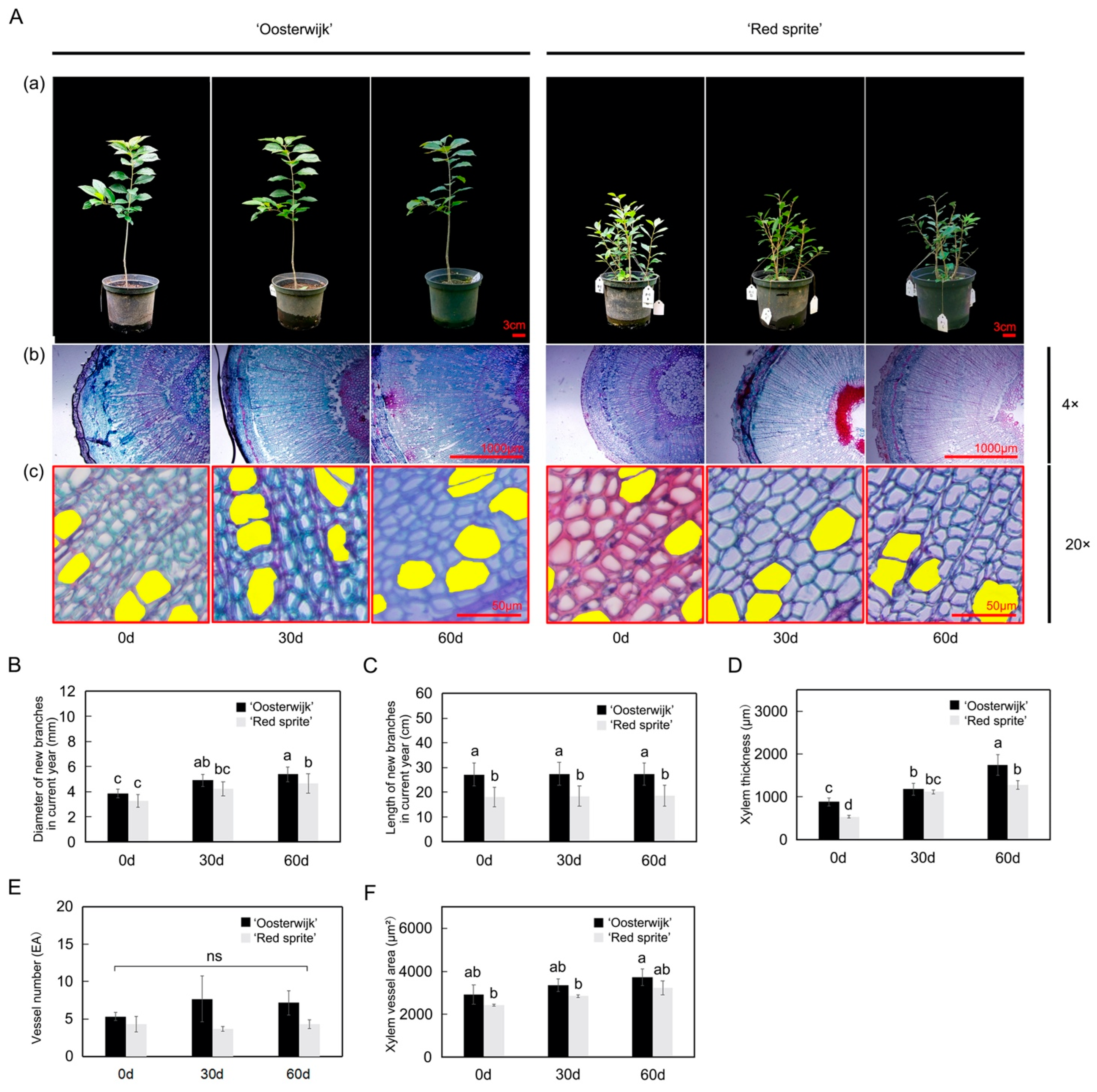
2.1. Morphological and Histological Analysis of the Stem in ‘Oosterwijk’ and ‘Red Sprite’
2.2. Analysis of Differentially Expressed Genes
2.3. GO Classification and KEGG Analysis of Differentially Expressed Genes
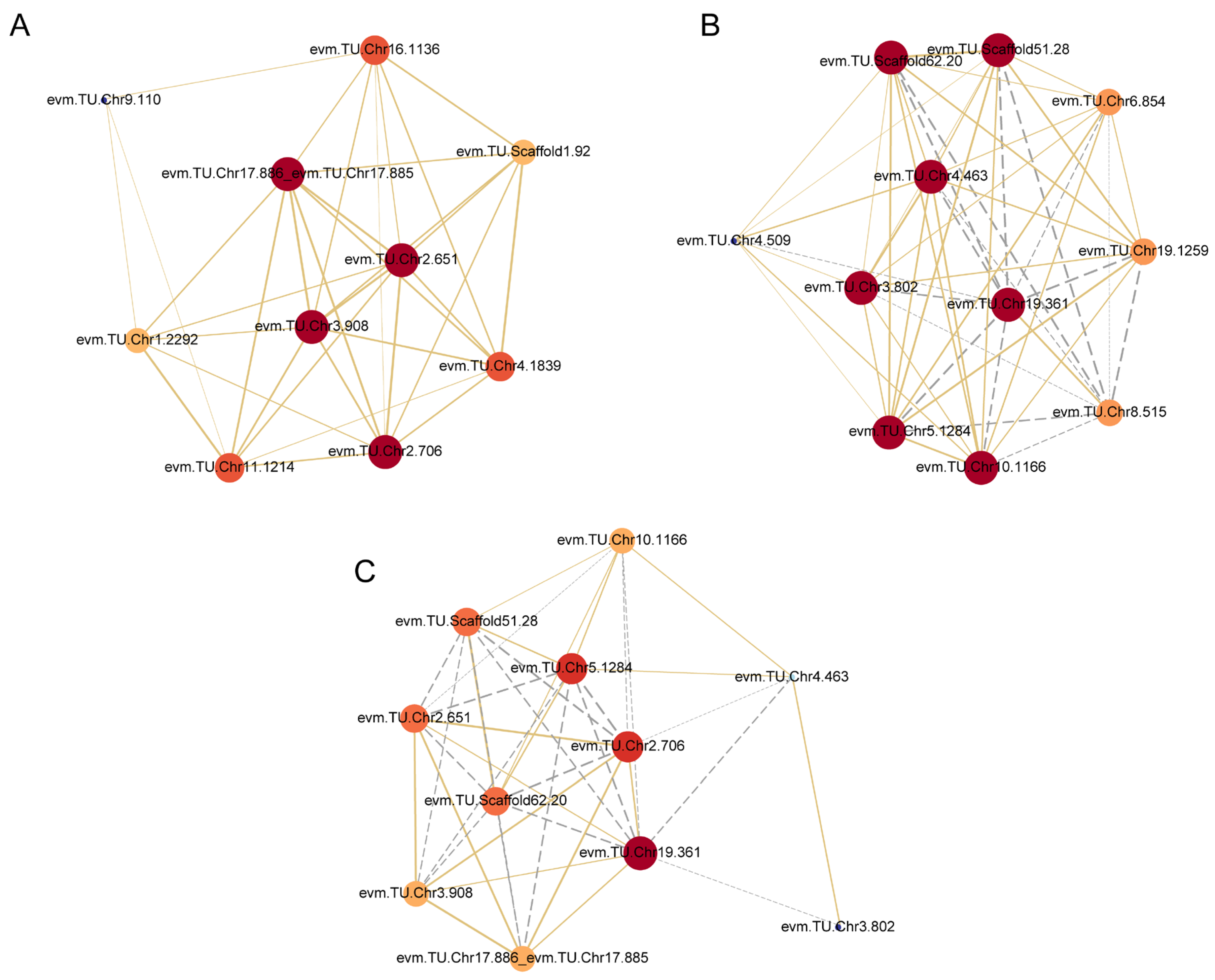
2.4. Co-Expression Network Analysis of Genes in the Module
2.5. Construction of the Correlation Network of DEGs in the Module
2.6. Analysis of DEGs Related to the Auxin Signaling Pathway and Endogenous Auxin Content Determination
2.7. Analysis of Auxin Content Determination
3. Discussion
3.1. Water Transport Capacity of the ‘Oosterwijk’ Stem May Be Stronger Than ‘Red Sprite’
3.2. AST, HCT and bHLH94 May Be the Key Genes to the Regulation of Stem Growth
3.3. Auxin Affected the Growth and Development of the Stem
4. Materials and Methods
4.1. Plant Growth Conditions
4.2. Histological Analysis
4.3. RNA Isolation and Illumina Sequencing
4.4. De Novo Assembly and Functional
4.5. Identification of Differentially Expressed Genes
4.6. Weighted Gene Co-Expression Network Analysis
4.7. IAA Content Determination
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Islam, N.; Evans, E.J. Influence of lodging and nitrogen rate on the yield and yield attributes of oilseed rape (Brassica-napus L.). Theor. Appl. Genet. 1994, 88, 530–534. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Smith, S.M.; Li, J. Genetic regulation of shoot architecture. Annu. Rev. Plant Biol. 2018, 69, 437–468. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, J. Molecular basis of plant architecture. Annu. Rev. Plant Biol. 2008, 59, 253–279. [Google Scholar] [CrossRef] [PubMed]
- Agusti, J.; Blazquez, M.A. Plant vascular development: Mechanisms and environmental regulation. Cell. Mol. Life Sci. 2020, 77, 3711–3728. [Google Scholar] [CrossRef]
- Shi, Q.; Tian, D.; Wang, J.; Chen, A.; Miao, Y.; Chen, Y.; Li, J.; Wu, X.; Zheng, B.; Guo, W.; et al. Overexpression of miR390b promotes stem elongation and height growth in Populus. Hortic. Res. 2023, 10, uhac258. [Google Scholar] [CrossRef]
- Leguay, J.J.; Guern, J. Quantitative effects of 2,4-dichlo-rophenoxyacetic acid on growth of suspension-cultured acer pseudoplatanus cells. Plant Physiol. 1975, 56, 356–359. [Google Scholar] [CrossRef]
- Campanoni, P.; Nick, P. Auxin-dependent cell division and cell elongation. 1-naphthaleneacetic acid and 2,4-dichlorophenoxyacetic acid activate different pathways. Plant Physiol. 2005, 137, 939–948. [Google Scholar] [CrossRef]
- Perrot, R.C. Cellular responses to auxin: Division versus expansion. Cold Spring Harb. Perspect. Biol. 2010, 2, a001446. [Google Scholar] [CrossRef]
- Liscum, E.; Reed, J.W. Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol. Biol. 2002, 49, 387–400. [Google Scholar] [CrossRef]
- Mockaitis, K.; Estelle, M. Auxin receptors and plant development: A new signaling paradigm. Annu. Rev. Cell Dev. Biol. 2008, 24, 55–80. [Google Scholar] [CrossRef]
- Heyn, A.N. Molecular basis of auxin-regulated extension growth and role of dextranase. Proc. Natl. Acad. Sci. USA 1981, 78, 6608–6612. [Google Scholar] [CrossRef]
- Wei, T.; Zhang, L.; Zhu, R.; Jiang, X.; Yue, C.; Su, Y.; Ren, H.; Wang, M. A gain-of-function mutant of IAA7 inhibits stem elongation by transcriptional repression of EXPA5 genes in Brassica napus. Int. J. Mol. Sci. 2021, 22, 9018. [Google Scholar] [CrossRef]
- Palm, D.; Streit, D.; Shanmugam, T.; Weis, B.L.; Ruprecht, M.; Simm, S.; Schleiff, E. Plant-specific ribosome biogenesis factors in Arabidopsis thaliana with essential function in rRNA processing. Nucleic Acids Res. 2019, 47, 1880–1895. [Google Scholar] [CrossRef]
- Edelmann, H.; Schopfer, P. Role of protein and RNA synthesis in the initiation of auxin-mediated growth in coleoptiles of Zea mays L. Planta 1989, 179, 475–485. [Google Scholar] [CrossRef] [PubMed]
- Shu, W.; Liu, Y.; Guo, Y.; Zhou, H.; Zhang, J.; Zhao, S.; Lu, M. A populus TIR1 gene family survey reveals differential expression patterns and responses to 1-naphthaleneacetic acid and stress treatments. Front. Plant Sci. 2015, 6, 719. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Guan, R.; Li, G.; Su, K.; Duan, L.; Sun, W.; Meng, X.; Wan, H.; Wang, S.; Chen, S.; et al. Genomic identification of ARF transcription factors and expression analysis in Cannabis sativa L. Ind. Crops Prod. 2022, 186, 115118. [Google Scholar] [CrossRef]
- Nagpal, P.; Ellis, C.M.; Weber, H.; Ploense, S.E.; Barkawi, L.S.; Guilfoyle, T.J.; Hagen, G.; Alonso, J.M.; Cohen, J.D.; Farmer, E.E.; et al. Auxin response factors ARF6 and ARF8 promote jasmonic acid production and flower maturation. Development 2005, 132, 4107–4118. [Google Scholar] [CrossRef]
- Wu, M.; Tian, Q.; Reed, J.W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef]
- Yu, H.; Soler, M.; Mila, I.; Clemente, H.S.; Savelli, B.; Dunand, C.; Paiva, J.A.P.; Myburg, A.A.; Bouzayen, M.; Pettenati, J.G.; et al. Genome-wide characterization and expression profiling of the auxin response factor (ARF) gene family in Eucalyptus grandis. PLoS ONE 2014, 9, e108906. [Google Scholar] [CrossRef]
- Xu, Z.; Ji, A.; Song, J.; Chen, S. Genome-wide analysis of auxin response factor gene family members in medicinal model plant Salvia miltiorrhiza. Biol. Open 2016, 5, 848–857. [Google Scholar] [CrossRef]
- Liu, N.; Dong, L.; Deng, X.; Liu, D.; Liu, Y.; Li, M.; Hu, Y.; Yan, Y. Genome-wide identification, molecular evolution, and expression analysis of auxin response factor (ARF) gene family in Brachypodium distachyon L. BMC Plant Biol. 2018, 18, 336. [Google Scholar] [CrossRef] [PubMed]
- An, W.; Gong, W.; He, S.; Pan, Z.; Sun, J.; Du, X. MicroRNA and mRNA expression profiling analysis revealed the regulation of plant height in Gossypium hirsutum. BMC Genom. 2015, 16, 886. [Google Scholar] [CrossRef] [PubMed]
- Ross, J.J.; O’Neill, D.P.; Rathbone, D.A. Auxin-gibberellin interactions in pea: Integrating the old with the new. Plant Growth Regul. 2003, 22, 99–108. [Google Scholar] [CrossRef]
- Nemhauser, J.L.; Mockler, T.C.; Chory, J. Interdependency of brassinosteroid and auxin signaling in Arabidopsis. PLoS Biol. 2004, 2, 1460–1471. [Google Scholar] [CrossRef]
- Luo, L.; Zeng, J.; Wu, H.; Tian, Z.; Zhao, Z. A molecular framework for auxin-controlled homeostasis of shoot stem cells in Arabidopsis. Mol. Plant 2018, 11, 899–913. [Google Scholar] [CrossRef]
- Thelander, M.; Landberg, K.; Sundberg, E. Minimal auxin sensing levels in vegetative moss stem cells revealed by a ratiometric reporter. New Phytol. 2019, 224, 775–788. [Google Scholar] [CrossRef] [PubMed]
- Roodbarkelari, F.; Du, F.; Truernit, E.; Laux, T. ZLL/AGO10 maintains shoot meristem stem cells during Arabidopsis embryogenesis by down-regulating ARF2-mediated auxin response. BMC Biol. 2015, 13, 74. [Google Scholar] [CrossRef]
- Wu, J.; Wang, X.; Zhang, R.; Fu, Q.; Tang, F.; Shi, F.; Temuer, B.; Zhang, Z. Comparative transcriptome and anatomic characteristics of stems in two alfalfa genotypes. Plants 2022, 11, 2601. [Google Scholar] [CrossRef]
- Zhang, B.; Xiao, X.; Zong, J.; Chen, J.; Li, J.; Guo, H.; Liu, J. Comparative transcriptome analysis provides new insights into erect and prostrate growth in bermudagrass (Cynodon dactylon L.). Plant Physiol. Biochem. 2017, 121, 31–37. [Google Scholar] [CrossRef]
- Zhao, D.; Luan, Y.; Xia, X.; Shi, W.; Tang, Y.; Tao, J. Lignin provides mechanical support to herbaceous peony (Paeonia lactiflora Pall.) stems. Hortic. Res. 2021, 7, 213. [Google Scholar] [CrossRef]
- Pan, R.; Melvin, T.T. How does water flow from vessel to vessel? Further investigation of the tracheid bridge concept. Tree Physiol. 2019, 39, 1019–1031. [Google Scholar] [CrossRef] [PubMed]
- Roig, P.F.; Berli, F.; Roig, F.A.; Tomazello, M.; Mastrantonio, L.; Piccoli, P. Wood hydrosystem of three cultivars of Vitis vinifera L. is modified in response to contrasting soils. Plant Soil 2021, 463, 573–588. [Google Scholar] [CrossRef]
- Jiang, X. Xylem Hydraulic Structure and Function in Mangroves. Ph.D. Thesis, Guangxi University, Nanning, China, 2021. [Google Scholar]
- Pan, S. The Xylem Structure and Relationship between Water Relations and Photosynthesis in Three Phyllostachys Bamboo. Master’s Thesis, Zhejiang Normal University, Jinhua, China, 2016. [Google Scholar]
- Jotan, P.; Maycock, C.R.; Burslem, D.F.R.P.; Berhaman, A.; Both, S. Comparative vessel traits of Macaranga gigantea and Vatica dulitensis from malaysian borneo. J. Trop. For. Sci. 2020, 32, 25–34. [Google Scholar] [CrossRef]
- Kaur, G.; Vikal, Y.; Kaur, L.; Kalia, A.; Mittal, A.; Kaur, D.; Yadav, I. Elucidating the morpho-physiological adaptations and molecular responses under long-term waterlogging stress in maize through gene expression analysis. Plant Sci. 2021, 304, 110823. [Google Scholar] [CrossRef] [PubMed]
- Cui, K.; Wang, H.; Liao, S.; Tang, Q.; Li, L.; Cui, Y.; He, Y. Transcriptome sequencing and analysis for culm elongation of the world’s largest bamboo (Dendrocalamus sinicus). PLoS ONE 2016, 11, e0157362. [Google Scholar] [CrossRef]
- Noh, S.A.; Choi, Y.I.; Cho, J.S.; Lee, H. The poplar basic helix-loop-helix transcription factor BEE3-like gene affects biomass production by enhancing proliferation of xylem cells in poplar. Biochem. Biophys. Res. Commun. 2015, 462, 64–70. [Google Scholar] [CrossRef]
- Zhu, Z.; Chen, G.; Guo, X.; Yin, W.; Yu, X.; Hu, J.; Hu, Z. Overexpression of SlPRE2, an atypical bHLH transcription factor, affects plant morphology and fruit pigment accumulation in tomato. Sci. Rep. 2017, 7, 5786. [Google Scholar] [CrossRef]
- Jiang, Z.; Liu, D.; Wang, T.; Liang, X.; Cui, Y.; Liu, Z.; Li, W. Concentration difference of auxin involved in stem development in soybean. J. Integr. Agric. 2020, 19, 953–964. [Google Scholar] [CrossRef]
- Xu, X.; Hu, Q.; Wang, J.; Wang, X.; Lou, L.; Xu, J.; Yang, X.; Chen, X. A 2-bp deletion in the protein kinase domain region of the ERECTA-like receptor kinase gene in cucumber results in short internode phenotype. Plant Sci. 2023, 327, 111536. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Y.; Hassan, M.; Li, Z.; Peng, Y. Indole-3-acetic acid improves drought tolerance of white clover via activating auxin, abscisic acid and jasmonic acid related genes and inhibiting senescence genes. BMC Plant Biol. 2020, 20, 150. [Google Scholar] [CrossRef]
- Da, C.C.T.; Gaeta, M.L.; Mariath, J.E.D.; Offringa, R.; Fett, N.A.G. Comparative adventitious root development in pre-etiolated and flooded Arabidopsis hypocotyls exposed to different auxins. Plant Physiol. Biochem. 2018, 127, 161–168. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Y.; Yang, R.; Wang, F.; Fu, J.; Yang, W.; Bai, T.; Wang, S.; Yin, H. Effects of gibberellin priming on seedling emergence and transcripts involved in mesocotyl elongation in rice under deep direct-seeding conditions. J. Zhejiang Univ.-Sci. B 2022, 22, 1002–1021. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Li, H.; Zhai, L.; Xie, X.; Li, X.; Bian, S. Identification and functional characterization of the Aux/IAA gene VcIAA27 in Blueberry. Plant Signal. Behav. 2019, 15, 1700327. [Google Scholar] [CrossRef]
- Shahzad, Z.; Eaglesfield, R.; Carr, C.; Amtmann, A. Cryptic variation in RNA-directed DNA methylation controls lateral root development when auxin signaling is perturbed. Nat. Commun. 2020, 11, 218. [Google Scholar] [CrossRef]
- Stamm, P.; Kumar, P.P. Auxin and gibberellin responsive Arabidopsis SMALL AUXIN UP RNA36 regulates hypocotyl elongation in the light. Plant Cell Rep. 2013, 32, 759–769. [Google Scholar] [CrossRef]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-level expression analysis of RNA-seq experiments with hisat, stringtie and ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef] [PubMed]
- Kovaka, S.; Zimin, A.V.; Pertea, G.M.; Razaghi, R.; Salzberg, S.L.; Pertea, M. Transcriptome assembly from long-read RNA-seq alignments with stringtie2. Genome Biol. 2019, 20, 278. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. Stringtie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Xin, P.; Guo, Q.; Li, B.; Cheng, S.; Yan, J.; Chu, J. A tailored high-efficiency sample pretreatment method for simultaneous quantification of 10 classes of known endogenous phytohormones. Plant Commun. 2020, 1, 100047. [Google Scholar] [CrossRef]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qin, S.; Fu, S.; Yang, Y.; Sun, Q.; Wang, J.; Dong, Y.; Gu, X.; Wang, T.; Xie, X.; Mo, X.; et al. Comparative Microscopic, Transcriptome and IAA Content Analyses Reveal the Stem Growth Variations in Two Cultivars Ilex verticillata. Plants 2023, 12, 1941. https://doi.org/10.3390/plants12101941
Qin S, Fu S, Yang Y, Sun Q, Wang J, Dong Y, Gu X, Wang T, Xie X, Mo X, et al. Comparative Microscopic, Transcriptome and IAA Content Analyses Reveal the Stem Growth Variations in Two Cultivars Ilex verticillata. Plants. 2023; 12(10):1941. https://doi.org/10.3390/plants12101941
Chicago/Turabian StyleQin, Sini, Siyi Fu, Ying Yang, Qiumin Sun, Jingqi Wang, Yanling Dong, Xinyi Gu, Tao Wang, Xiaoting Xie, Xiaorong Mo, and et al. 2023. "Comparative Microscopic, Transcriptome and IAA Content Analyses Reveal the Stem Growth Variations in Two Cultivars Ilex verticillata" Plants 12, no. 10: 1941. https://doi.org/10.3390/plants12101941
APA StyleQin, S., Fu, S., Yang, Y., Sun, Q., Wang, J., Dong, Y., Gu, X., Wang, T., Xie, X., Mo, X., Jiang, H., Yu, Y., Yan, J., Chu, J., Zheng, B., & He, Y. (2023). Comparative Microscopic, Transcriptome and IAA Content Analyses Reveal the Stem Growth Variations in Two Cultivars Ilex verticillata. Plants, 12(10), 1941. https://doi.org/10.3390/plants12101941






