Chromosome Rearrangement in Elymus dahuricus Revealed by ND-FISH and Oligo-FISH Painting
Abstract
:1. Introduction
2. Results
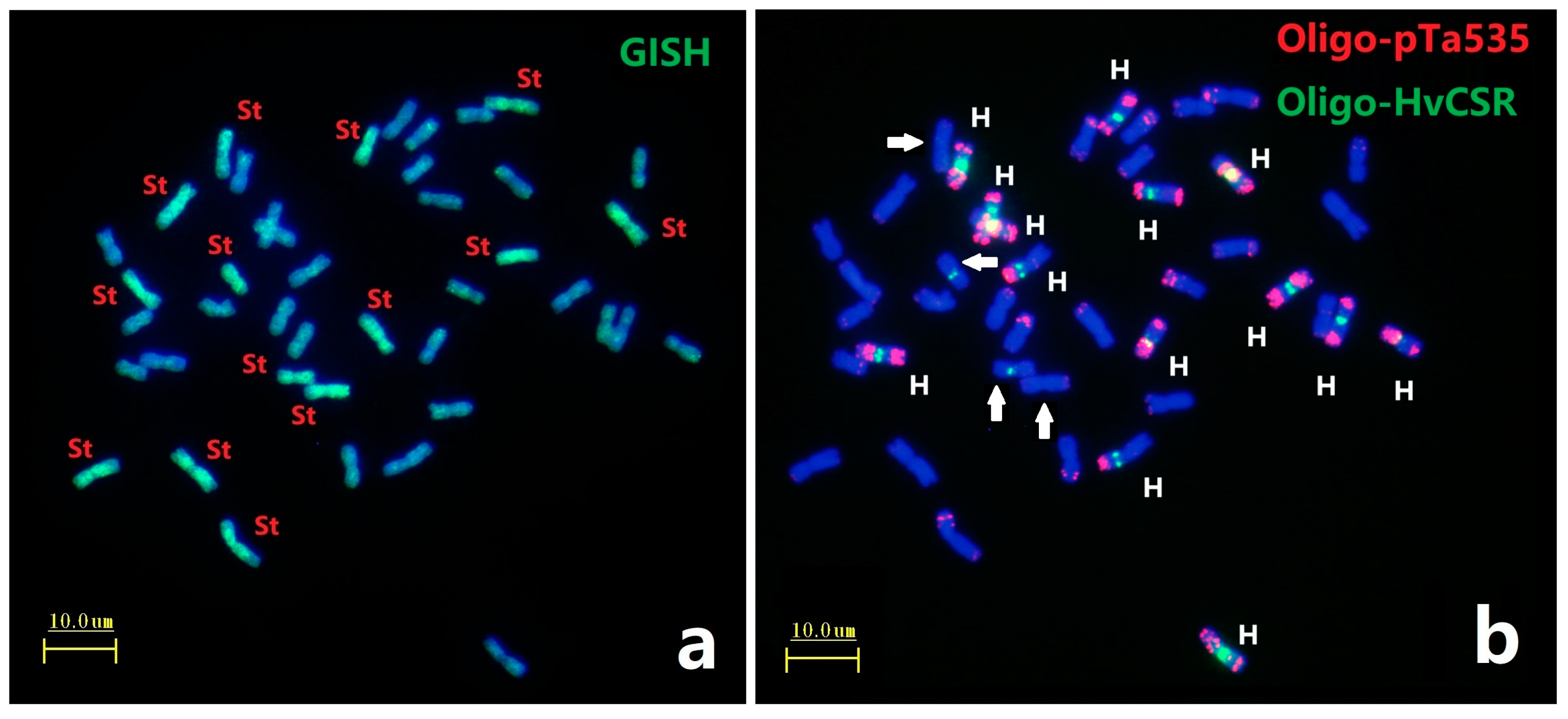
2.1. Genome Identification Using Combined FISH and GISH Analyses
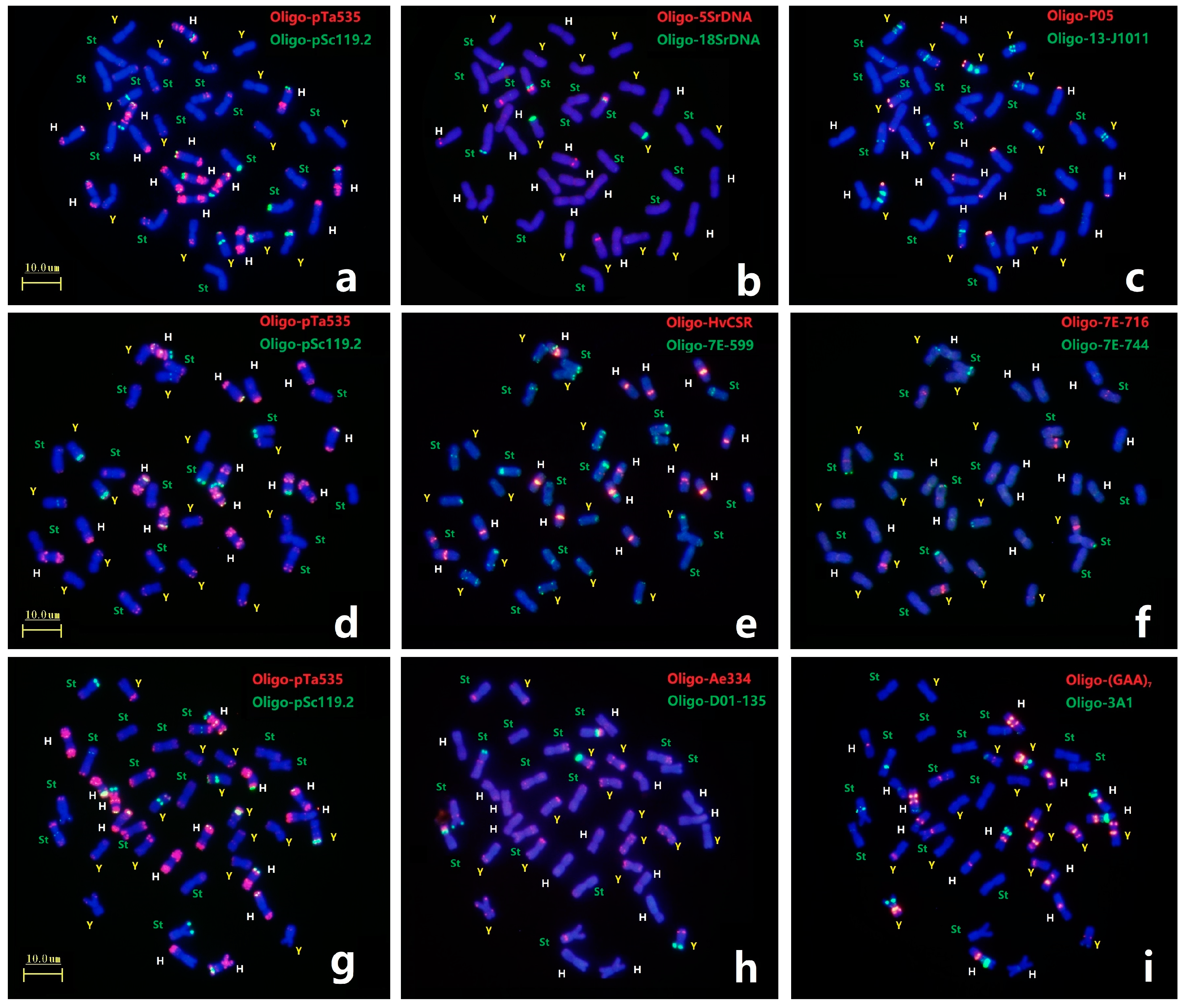
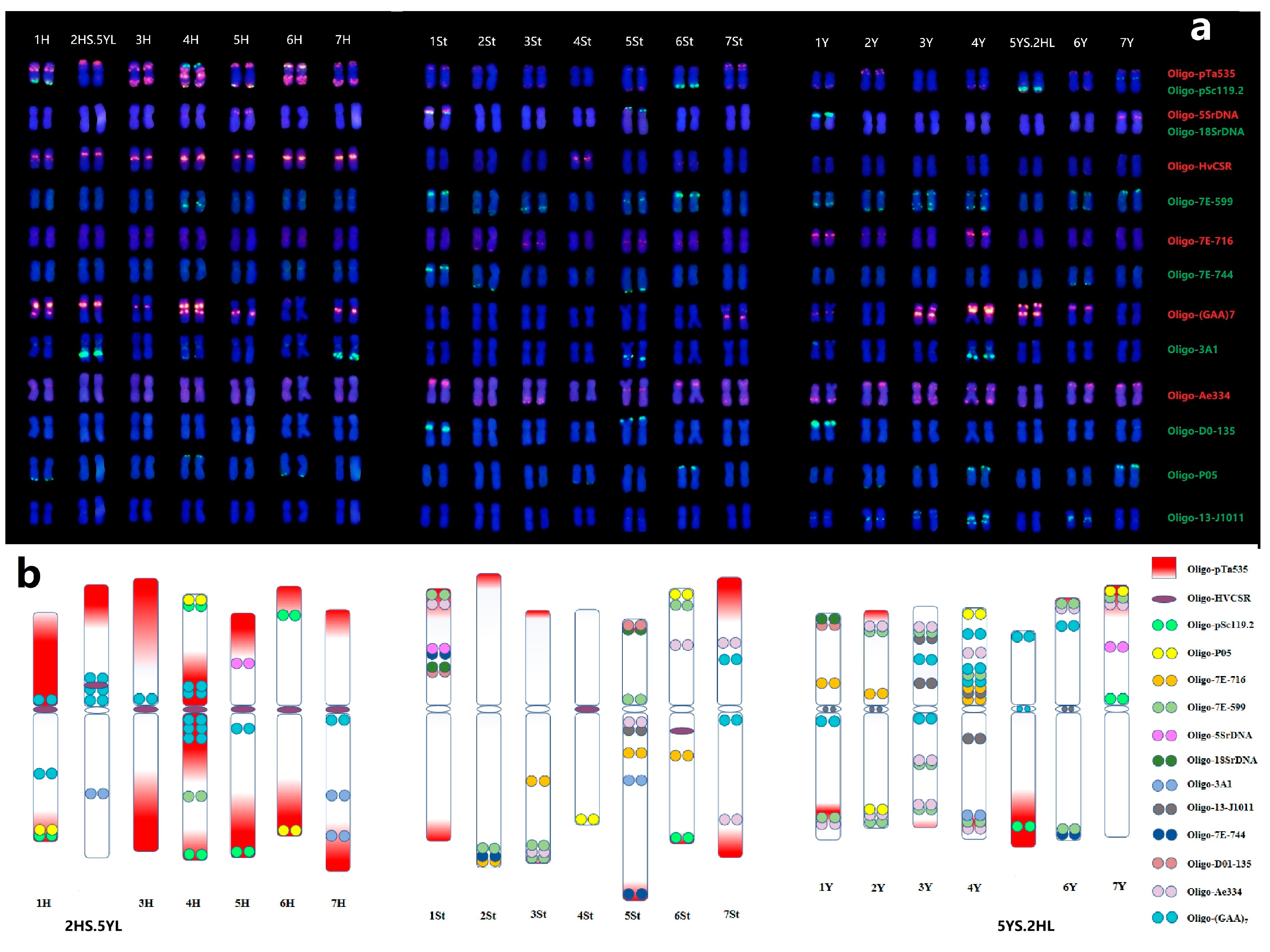
2.2. Precise Chromosome Identification by ND-FISH
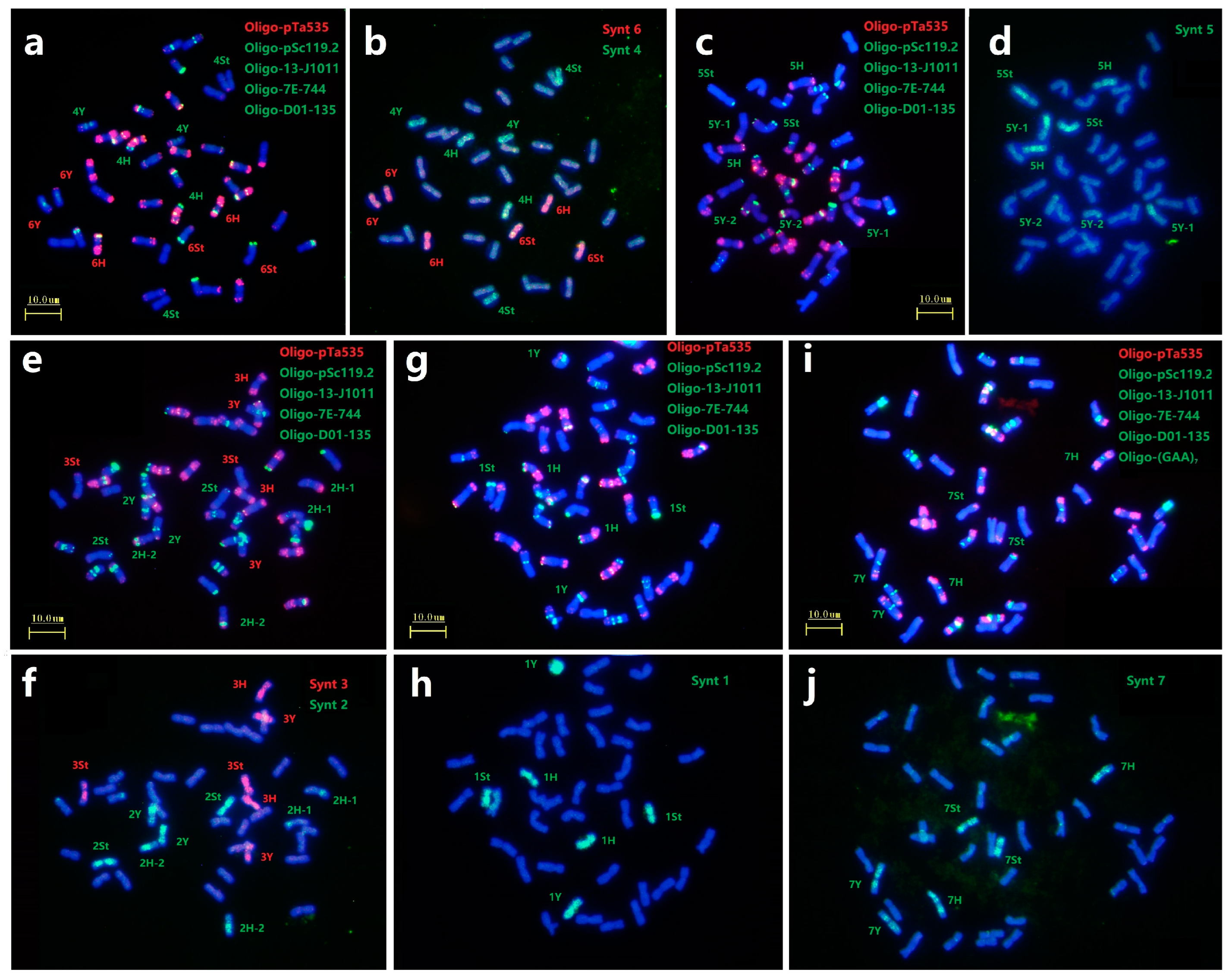
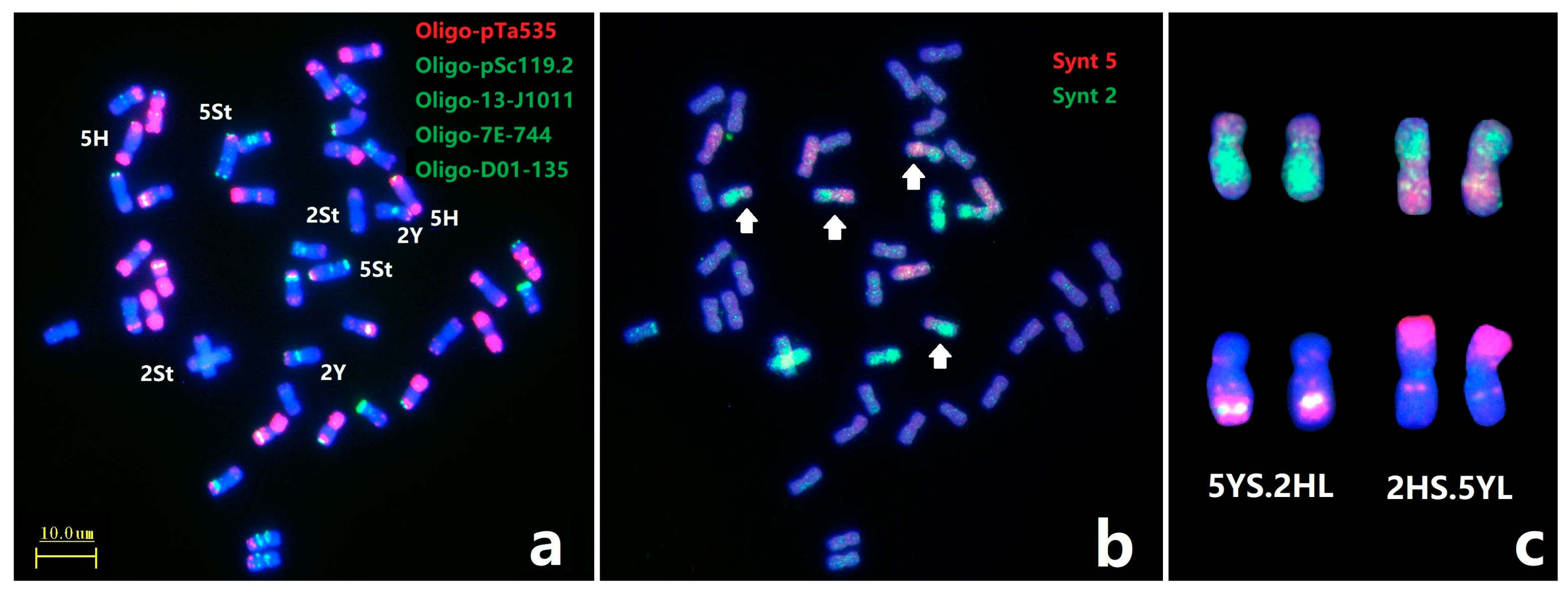
2.3. Identification of Chromosome Linkage Groups
2.4. Establishment of Universal Karyotyping Nomenclature System
2.5. Comparison Analysis of Centromere Specific Repeat Sequences and CENH3
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Chromosome Preparation and ND-FISH Analysis
4.3. GISH and Oligo-FISH Painting
4.4. Chromosomal Immunolocalization
4.5. Drawing Karyotype Ideogram
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Agafonov, A.V.; Baum, B.R.; Bailey, L.G.; Agafonova, O.V. Differentiation in the Elymus dahuricus complex (Poaceae): Evidence from grain proteins, DNA, and crossability. Hereditas 2001, 135, 277–289. [Google Scholar] [CrossRef] [PubMed]
- Cui, G.W.; Ji, G.X.; Liu, S.Y.; Zhang, P. Physiological adaptations of Elymus dahuricus to high altitude on the Qinghai-Tibetan Plateau. Acta Physiol. Plant 2019, 41, 115. [Google Scholar] [CrossRef]
- Barkworth, M.E.; Dewey, D.R. The genomic system of classification as a guide to intergeneric hybridization with the perennial Triticeae. Taxon 1986, 35, 202. [Google Scholar] [CrossRef]
- Löve, Á. Conspectus of the Triticeae. Feddes Repert. 1984, 95, 425–521. [Google Scholar] [CrossRef]
- Baum, B.R.; Yang, J.L.; Agafonov, A.V. A taxonomic synopsis of the genus Campeiostachys Drobov. J. Syst. Evol. 2011, 49, 146–159. [Google Scholar] [CrossRef]
- Dou, Q.W.; Zhang, T.L.; Tsujimoto, H. Physical mapping of repetitive sequences and genome analysis in six Elymus species by in situ hybridization. J. Syst. Evol. 2011, 49, 347–352. [Google Scholar] [CrossRef]
- Wang, R.R.C.; Liu, B. Biosystematic and evolutionary relationships of perennial Triticeae species revealed by genomic analyses. J. Syst. Evol. 2014, 52, 697–705. [Google Scholar] [CrossRef]
- Matic, I. Chromosomal rearrangements and speciation. Trends Microbiol. 2000, 8, 353. [Google Scholar] [CrossRef]
- Faria, R.; Navarro, A. Chromosomal speciation revisited: Rearranging theory with pieces of evidence. Trends Ecol. Evol. 2010, 25, 660–669. [Google Scholar] [CrossRef]
- Li, W.L.; Challa, G.S.; Zhu, H.L.; Wei, W.J. Recurrence of chromosome rearrangements and reuse of DNA breakpoints in the evolution of the Triticeae genomes. G3 Genes Genomes Genet. 2016, 6, 3837–3847. [Google Scholar] [CrossRef]
- Tang, Z.X.; Li, M.; Chen, L.; Fu, S.L. New types of wheat chromosomal structural variations in derivatives of wheat-rye hybrids. PLoS ONE 2014, 9, e110282. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.P.; Bie, T.D.; Wang, X.E.; Chen, P.D. Induction and transmission of wheat-Haynaldia villosa chromosomal translocations. J. Genet. Genom. 2009, 36, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.X.; Chen, P.D.; Liu, D.J. Development of Triticum aestivum-Leymus racemosus translocation lines by irradiating adult plants at meiosis. Acta Bot. Sin. 1999, 41, 463–467. [Google Scholar]
- Yang, C.R.; Zhang, H.Q.; Chen, W.H.; Zhou, Y.H. Genomic constitution and intergenomic translocations in Elymus dahuricus complex revealed by multicolor GISH. Genome 2017, 60, 510–517. [Google Scholar] [CrossRef]
- Jiang, J.M. Fluorescence in situ hybridization in plants: Recent developments and future applications. Chromosome Res. 2019, 27, 153–165. [Google Scholar] [CrossRef]
- Han, F.P.; Liu, B.; Liu, Z.H. Chromosomal variation, constitution of five partial amphiploids of wheat-Thinopyrum intermedium detected by GISH, seed storage protein marker and multicolor GISH. Theor. Appl. Genet. 2004, 109, 1070–1076. [Google Scholar] [CrossRef]
- Cuadrado, A.; Golczyk, H.; Jouve, N. A novel, simple and rapid nondenaturing FISH (ND-FISH) technique for the detection of plant telomeres. Potential used and possible target structures detected. Chromosome Res. 2009, 17, 755–762. [Google Scholar] [CrossRef]
- Jiang, M.; Xiao, Z.Q.; Fu, S.L.; Tang, Z.X. FISH karyotype of 85 common wheat cultivars/lines displayed by ND-FISH using oligonucleotide probes. Cereal Res. Commun. 2017, 45, 549–563. [Google Scholar] [CrossRef]
- Cheng, M.H.; Li, X.Y.; Cui, H.M. FISH-based “pan” and “core” karyotypes reveal genetic diversification of Roegneria ciliaris. J. Genet. Genom. 2023, 50, 135–138. [Google Scholar] [CrossRef]
- Li, G.R.; Zhang, T.; Yu, Z.H.; Yang, Z.J. An efficient Oligo-FISH painting system for revealing chromosome rearrangements and polyploidization in Triticeae. Plant J. 2020, 105, 978–993. [Google Scholar] [CrossRef]
- Sakamoto, S. Cytogenetical studies on artificial hybrids among Elymus sibiricus, E. dahuricus and Agropyron tsukushiense in the Tribe Triticeae, Gramineae. J. Plant Res. 1982, 95, 375–383. [Google Scholar] [CrossRef]
- Tsujimoto, H.; Mukai, Y.; Akagawa, K.; Sakakuma, T. Identification of individual barley chromosomes based on repetitive sequences: Conservative distribution of Afa-family repetitive sequences on the chromosomes of barley and wheat. Genes Genet. Syst. 1997, 72, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Lang, T.; Li, G.R.; Wang, H.J.; Yang, Z.J. Physical location of tandem repeats in the wheat genome and application for chromosome identification. Planta 2019, 249, 663–675. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.X.; Yang, Z.J.; Fu, S.L. Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J. Appl. Genet. 2014, 55, 313–318. [Google Scholar] [CrossRef]
- Yu, Z.H.; Wang, H.J.; Jiang, W.X.; Yang, Z.J. Karyotyping Dasypyrum breviaristatum chromosomes with multiple oligonucleotide probes reveals the genomic divergence in Dasypyrum. Genome 2021, 64, 789–800. [Google Scholar] [CrossRef]
- Lang, T.; La, S.X.; Li, B.; Yang, Z.J. Precise identification of wheat-Thinopyrum intermedium translocation chromosomes carrying resistance to wheat stripe rust in line Z4 and its derived progenies. Genome 2018, 61, 177–185. [Google Scholar] [CrossRef]
- Cuadrado, A.; Jouve, N. Chromosomal detection of simple sequence repeats (SSRs) using nondenaturing FISH (ND-FISH). Chromosoma 2010, 119, 495–503. [Google Scholar] [CrossRef]
- Gupta, P.; Kulwal, P.; Rustgi, S. Wheat cytogenetics in the genomics era and its relevance to breeding. Cytogenet. Genome Res. 2005, 109, 315–327. [Google Scholar] [CrossRef]
- Liu, B.; Chen, J.; Yang, Y.; Dou, Q.W. Single-gene FISH maps and major chromosomal rearrangements in Elymus sibiricus and E. nutans. BMC Plant Biol. 2023, 23, 98. [Google Scholar] [CrossRef]
- Huang, X.Y.; Zhu, M.Q.; Zhuang, L.F.; Chen, P.D. Structural chromosome rearrangements and polymorphisms identified in Chinese wheat cultivars by high-resolution multiplex oligonucleotide FISH. Theor. Appl. Genet. 2018, 131, 1967–1986. [Google Scholar] [CrossRef]
- Shi, P.Y.; Sun, H.J.; Liu, G.Q.; Wang, X.E. Chromosome painting reveals inter-chromosomal rearrangements and evolution of the subgenome D of wheat. Plant J. 2022, 112, 55–68. [Google Scholar] [CrossRef] [PubMed]
- Lu, B.R.; Bothmer, R.V. Genomic constitution of Elymus parviglumis and E. pseudonutans: Triticeae (Poaceae). Hereditas 2008, 113, 109–119. [Google Scholar] [CrossRef]
- Jiang, J.C.; Gill, B. Different species-specific chromosome translocations in Triticum timopheevii and T. turgidum support the diphyletic origin of polyploid wheats. Chromosome Res. 1994, 2, 59–64. [Google Scholar] [CrossRef]
- Jiang, J.C.; Gill, B. New 18S-26S ribosomal RNA gene loci: Chromosomal landmarks for the evolution of polyploid wheats. Chromosoma 1994, 103, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Murata, M.; Heslop-Harrison, J.S.; Motoyoshi, F. Physical mapping of the 5S ribosomal RNA genes in Arabidopsis thaliana by multi-color fluorescence in situ hybridization with cosmid clones. Plant J. 1997, 12, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Taketa, S.; Harrison, G.E. Comparative physical mapping of the 5S and 18S-25S rDNA in nine wild Hordeum species and cytotypes. Theor. Appl. Genet. 1999, 98, 1–9. [Google Scholar] [CrossRef]
- Lermontova, I.; Koroleva, O.; Rutten, T.; Schubert, I. Knockdown of CENH3 in Arabidopsis reduces mitotic divisions and causes sterility by disturbed meiotic chromosome segregation. Plant J. 2011, 68, 40–50. [Google Scholar] [CrossRef]
- Hirsch, C.D.; Wu, Y.F.; Yan, H.H.; Jiang, J.M. Lineage-specific adaptive evolution of the centromeric protein CENH3 in diploid and allotetraploid Oryza species. Mol. Biol. Evol. 2009, 26, 2877–2885. [Google Scholar] [CrossRef]
- Zhao, J.; Hao, W.W.; Tang, C.G.; Zhang, X.Y. Plasticity in Triticeae centromere DNA sequences: A wheat × tall wheatgrass (decaploid) model. Plant J. 2019, 100, 314–327. [Google Scholar] [CrossRef]
- Li, G.R.; Chen, Q.H.; Jiang, W.X.; Yang, Z.J. Molecular and cytogenetic identification of wheat-Thinopyrum intermedium double substitution line-derived progenies for stripe rust resistance. Plants 2022, 12, 28. [Google Scholar] [CrossRef]
- Martinez-Zapater, J.M.; Estelle, M.A.; Somerville, C.R. A highly repeated DNA sequence in Arabidopsis thaliana. MGG Mol. Gen. Genet. 1986, 204, 417–423. [Google Scholar] [CrossRef]
- Cheng, Z.K.; Dong, F.G.; Langdon, T.; Jiang, J.M. Functional rice centromeres are marked by a satellite repeat and a centromere-specific retrotransposon. Plant Cell 2022, 14, 1691–1704. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Su, H.D.; Shi, Q.H.; Han, F.P. De Novo centromere formation and centromeric sequence expansion in wheat and its wide hybrids. PLoS Genet. 2016, 12, e1005997. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Xie, Y.L.; Kong, C.Z.; Zhang, X.Y. Centromere repositioning and shifts in wheat evolution. Plant Commun. 2023, 4, 100556. [Google Scholar] [CrossRef]
- Michael, T.P.; Vanburen, R. Building near-complete plant genomes. Curr. Opin. Plant Biol. 2020, 54, 26–33. [Google Scholar] [CrossRef]
- Chen, J.; Wang, Z.J.; Tan, K.W.; Lai, J.S. A complete telomere-to-telomere assembly of the maize genome. Nat. Genet. 2023, 55, 1221–1231. [Google Scholar] [CrossRef]
- Su, H.D.; Liu, Y.L.; Liu, C. Centromere satellite repeats have undergone rapid changes in polyploid wheat subgenomes. Plant Cell 2019, 31, 2035–2051. [Google Scholar] [CrossRef]
- Han, F.P.; Lamb, J.C.; Birchler, J.A. High frequency of centromere inactivation resulting in stable dicentric chromosomes of maize. Proc. Natl. Acad. Sci. USA 2006, 103, 3238–3243. [Google Scholar] [CrossRef]
- Fu, S.L.; Chen, L.; Wang, Y.Y.; Tang, Z.X. Oligonucleotide probes for ND-FISH analysis to identify rye and wheat chromosomes. Sci. Rep. 2015, 5, 10552. [Google Scholar] [CrossRef]
- Nasuda, S.; Hudakova, S.; Schubert, I.; Endo, T.R. Stable barley chromosomes without centromeric repeats. Proc. Natl. Acad. Sci. USA 2005, 102, 9842–9847. [Google Scholar] [CrossRef]
- Li, G.R.; Yang, Z.J. Oligo-FISH paints in Triticeae. Curr. Protoc. 2022, 2, e364. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Guo, X.; Hu, J.; Han, F.P. Characterization of two CENH3 genes and their roles in wheat evolution. New Phytol. 2014, 206, 839–851. [Google Scholar] [CrossRef] [PubMed]
- Han, F.P.; Gao, Z.; Birchler, J.A. Reactivation of an inactive centromere reveals epigenetic and structural components for centromere specification in maize. Plant Cell 2009, 21, 1929–1939. [Google Scholar] [CrossRef] [PubMed]
- Mahmoudi, S.; Mirzaghaderi, G. Tools for drawing informative ideograms. In Plant Cytogenetics and Cytogenomics: Methods and Protocols, Methods in Molecular Biology; Heitkam, T., Garcia, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2023; Volume 2672, pp. 515–527. [Google Scholar] [CrossRef]

| Oligo Probes | Sequences | Reference |
|---|---|---|
| Oligo-pTa535 | AAAAACTTGACGCACGTCACGTACAAATTGGACAAACTCTTTCGGAGTATCAGGGTTTC | [24] |
| Oligo-pSc119.2 | CCGTTTTGTGGACTATTACTCACCGCTTTGGGGTCCCATAGCTAT | [24] |
| Oligo-5SrDNA | TCAGAACTCCGAAGTTAAGCGTGCTTGGGCGAGAGTAGTAC | [23] |
| Oligo-18SrDNA | GGGCAAGTCTGGTGCCAGCAGCCGCGGT | [23] |
| Oligo-7E-744 | GCCACCGTGCAGTAGACTTTTTTTGTACCCAAACCATCAGTAACAAAGTTCGTTCAC | [25] |
| Oligo-3A1 | AATAATTTTACACTAGAGTTGAACTAGCTCTATAAGCTAGTTCA | [26] |
| Oligo-(GAA)7 | GAAGAAGAAGAAGAAGAAGAA | [27] |
| Oligo-HvCSR | ACAACGACAACAACGACAATGACGAGA | This study |
| Oligo-7E-716 | GTACAGGACTGCAGCTAAGCCCCCGAGTGAGAGGGTTGCTCATCACTCGGTAGGATT | This study |
| Oligo-7E-599 | CATCGGTCAAACCTCGTCCGGCGAAAGTCAAAGGCGTAGACCGCCCGGTCAACGGTGCC | This study |
| Oligo-P05 | AATACGCTCTTGTTCTTGGCTGTCACGCACATACTTTATGGGATGTCATAGG | This study |
| Oligo-13-J1011 | CATCATGCTTGTTGTGAGAAGCTCTGGTTTGTGAGAAGCATATACCCAAACC | This study |
| Oligo-D01-135 | ACGCGCGCCATGGAAAACAGGGCAAAACCACCGACTCGTCCACGACTCGTAC | This study |
| Oligo-Ae334 | CTCCAAAGTGTTCCTATGGGCTGACCTAACACAACCGGGTGG | This study |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, C.; Liu, X.; Yang, Z.; Li, G. Chromosome Rearrangement in Elymus dahuricus Revealed by ND-FISH and Oligo-FISH Painting. Plants 2023, 12, 3268. https://doi.org/10.3390/plants12183268
Jiang C, Liu X, Yang Z, Li G. Chromosome Rearrangement in Elymus dahuricus Revealed by ND-FISH and Oligo-FISH Painting. Plants. 2023; 12(18):3268. https://doi.org/10.3390/plants12183268
Chicago/Turabian StyleJiang, Chengzhi, Xiaodan Liu, Zujun Yang, and Guangrong Li. 2023. "Chromosome Rearrangement in Elymus dahuricus Revealed by ND-FISH and Oligo-FISH Painting" Plants 12, no. 18: 3268. https://doi.org/10.3390/plants12183268





