Comparative Analysis of Circadian Transcriptomes Reveals Circadian Characteristics between Arabidopsis and Soybean
Abstract
:1. Introduction
2. Results
2.1. Comparison of Circadian Rhythmic Genes Unveils Distinctive Circadian Parameter Characteristics between Arabidopsis and Soybean
2.2. Comparative Analysis of Homologous Circadian Clock Genes Implies Intricate Circadian Regulatory Mechanisms Distinguishing Soybean from Arabidopsis
2.3. Circadian Parameter Alterations of Genes Provide Insights into the Circadian Control of Flowering and Maturity in Soybean
2.4. Circadian Control of Physiological Activities in Arabidopsis and Soybean
2.5. Differential Circadian Regulation of Physiological Activities in Arabidopsis and Soybean
3. Discussion
4. Materials and Methods
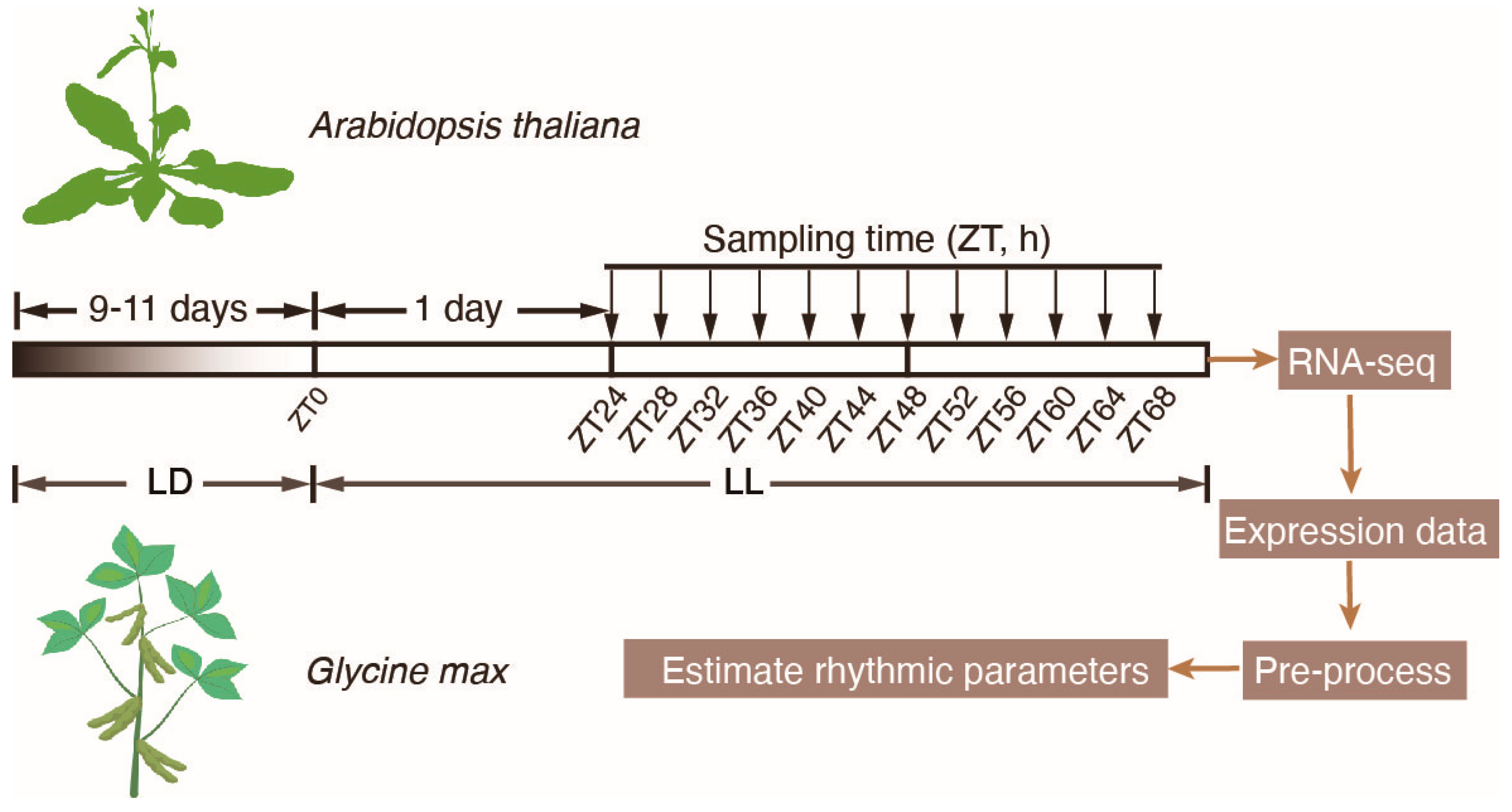
4.1. Plant Materials and Growth Conditions
4.2. Sequencing Reads Acquisition, Read Alignment, and mRNA Quantification
4.3. Pre-Processing of Expression Data
4.4. Weighted Estimation of Rhythmic Parameters of Genes
- (1)
- Remove the slope trends by fitting a linear regression model to the gene expression data and keep residuals.
- (2)
- Perform Fast Fourier Transformation (FFT) of the time series data and keep main signals.
- (3)
- Estimate the initial period using FFT transformed data.
- (4)
- Perform non-linear regression fitting. The fitting was applied with weights obtained in the pre-processing. Amplitude was constrained to be non-negative. Period was constrained to be greater than 12 h but less than 36 h. Phase24 and Constant have no constraints. Phase was normalized as phase24, which is more than 0 but less than 24. Constant indicates the average expression level of the gene.
- (5)
- Predict the best-fit data and calculate circadian oscillation correlation between observed data and predicted data.
- (6)
- Genes without convergent fits were considered arrhythmic. Genes with the resulting best-fit Amplitude, Period, Phase24, Constant and their standard error and degree of freedom were used for downstream statistical analysis.
4.5. Analysis of Homologous Genes
4.6. Phase24 Plots
4.7. Gene Set Enrichment Analysis
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yakir, E.; Hilman, D.; Harir, Y.; Green, R.M. Regulation of output from the plant circadian clock. FEBS J. 2007, 274, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-Y.; Oh, E.; Wang, T.; Wang, Z.-Y. TOC1–PIF4 interaction mediates the circadian gating of thermoresponsive growth in Arabidopsis. Nat. Commun. 2016, 7, 13692. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Gao, M.; Seitz, N.C.; Angel, W.; Hallworth, A.; Wiratan, L.; Darwish, O.; Alkharouf, N.; Dawit, T.; Lin, D.; et al. LUX ARRHYTHMO mediates crosstalk between the circadian clock and defense in Arabidopsis. Nat. Commun. 2019, 10, 2543. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; McClung, C.R.; Zhang, C. Tick Tock: Circadian Regulation of Plant Innate Immunity. Annu. Rev. Phytopathol. 2017, 55, 287–311. [Google Scholar] [CrossRef] [PubMed]
- Greenham, K.; McClung, C.R. Integrating circadian dynamics with physiological processes in plants. Nat. Rev. Genet. 2015, 16, 598–610. [Google Scholar] [CrossRef]
- Xu, H.; Wang, X.; Wei, J.; Zuo, Y.; Wang, L. The Regulatory Networks of the Circadian Clock Involved in Plant Adaptation and Crop Yield. Plants 2023, 12, 1897. [Google Scholar] [CrossRef]
- Hsu, P.Y.; Harmer, S.L. Wheels within wheels: The plant circadian system. Trends Plant Sci. 2014, 19, 240–249. [Google Scholar] [CrossRef]
- Inoue, K.; Araki, T.; Endo, M. Integration of Input Signals into the Gene Network in the Plant Circadian Clock. Plant Cell Physiol. 2017, 58, 977–982. [Google Scholar] [CrossRef]
- Millar, A.J. Input signals to the plant circadian clock. J. Exp. Bot. 2004, 55, 277–283. [Google Scholar] [CrossRef]
- Pokhilko, A.; Fernández, A.P.; Edwards, K.D.; Southern, M.M.; Halliday, K.J.; Millar, A.J. The clock gene circuit in Arabidopsis includes a repressilator with additional feedback loops. Mol. Syst. Biol. 2012, 8, 574. [Google Scholar] [CrossRef]
- Schaffer, R.; Ramsay, N.; Samach, A.; Corden, S.; Putterill, J.; Carré, I.A.; Coupland, G. The late elongated hypocotyl Mutation of Arabidopsis Disrupts Circadian Rhythms and the Photoperiodic Control of Flowering. Cell 1998, 93, 1219–1229. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.-Y.; Tobin, E.M. Constitutive Expression of the Circadian Clock Associated 1 (CCA1) Gene Disrupts Circadian Rhythms and Suppresses Its Own Expression. Cell 1998, 93, 1207–1217. [Google Scholar] [CrossRef] [PubMed]
- Somers, D.E.; Webb, A.A.; Pearson, M.; Kay, S.A. The short-period mutant, toc1-1, alters circadian clock regulation of multiple outputs throughout development in Arabidopsis thaliana. Development 1998, 125, 485–494. [Google Scholar] [CrossRef] [PubMed]
- Mizoguchi, T.; Wheatley, K.; Hanzawa, Y.; Wright, L.; Mizoguchi, M.; Song, H.-R.; Carré, I.A.; Coupland, G. LHY and CCA1 Are Partially Redundant Genes Required to Maintain Circadian Rhythms in Arabidopsis. Dev. Cell 2002, 2, 629–641. [Google Scholar] [CrossRef]
- Strayer, C.; Oyama, T.; Schultz, T.F.; Raman, R.; Somers, D.E.; Más, P.; Panda, S.; Kreps, J.A.; Kay, S.A. Cloning of the Arabidopsis Clock Gene TOC1, an Autoregulatory Response Regulator Homolog. Science 2000, 289, 768–771. [Google Scholar] [CrossRef]
- Adams, S.; Manfield, I.; Stockley, P.; Carré, I.A. Revised Morning Loops of the Arabidopsis Circadian Clock Based on Analyses of Direct Regulatory Interactions. PLoS ONE 2015, 10, e0143943. [Google Scholar] [CrossRef]
- Alabadí, D.; Oyama, T.; Yanovsky, M.J.; Harmon, F.G.; Más, P.; Kay, S.A. Reciprocal Regulation Between TOC1 and LHY/CCA1 Within the Arabidopsis Circadian Clock. Science 2001, 293, 880–883. [Google Scholar] [CrossRef]
- Nakamichi, N.; Kiba, T.; Kamioka, M.; Suzuki, T.; Yamashino, T.; Higashiyama, T.; Sakakibara, H.; Mizuno, T. Transcriptional repressor PRR5 directly regulates clock-output pathways. Proc. Natl. Acad. Sci. USA 2012, 109, 17123–17128. [Google Scholar] [CrossRef]
- Rawat, R.; Schwartz, J.; Jones, M.A.; Sairanen, I.; Cheng, Y.; Andersson, C.R.; Zhao, Y.; Ljung, K.; Harmer, S.L. REVEILLE1, a Myb-like transcription factor, integrates the circadian clock and auxin pathways. Proc. Natl. Acad. Sci. USA 2009, 106, 16883–16888. [Google Scholar] [CrossRef]
- Hicks, K.A.; Millar, A.J.; Carré, I.A.; Somers, D.E.; Straume, M.; Meeks-Wagner, D.R.; Kay, S.A. Conditional Circadian Dysfunction of the Arabidopsis early-flowering 3 Mutant. Science 1996, 274, 790–792. [Google Scholar] [CrossRef]
- Bendix, C.; Marshall, C.M.; Harmon, F.G. Circadian Clock Genes Universally Control Key Agricultural Traits. Mol. Plant 2015, 8, 1135–1152. [Google Scholar] [CrossRef]
- Li, C.; Li, Y.-H.; Li, Y.; Lu, H.; Hong, H.; Tian, Y.; Li, H.; Zhao, T.; Zhou, X.; Liu, J.; et al. A Domestication-Associated Gene GmPRR3b Regulates the Circadian Clock and Flowering Time in Soybean. Mol. Plant 2020, 13, 745–759. [Google Scholar] [CrossRef]
- Wei, H.; Xu, H.; Su, C.; Wang, X.; Wang, L. Rice Circadian Clock Associated 1 transcriptionally regulates ABA signaling to confer multiple abiotic stress tolerance. Plant Physiol. 2022, 190, 1057–1073. [Google Scholar] [CrossRef]
- Wittern, L.; Steed, G.; Taylor, L.J.; Ramirez, D.C.; Pingarron-Cardenas, G.; Gardner, K.; Greenland, A.; Hannah, M.A.; Webb, A.A.R. Wheat EARLY FLOWERING 3 affects heading date without disrupting circadian oscillations. Plant Physiol. 2023, 191, 1383–1403. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Zhao, B.; Xie, Y.; Jia, H.; Li, Y.; Xu, M.; Wu, G.; Ma, X.; Li, Q.; Hou, M.; et al. The evening complex promotes maize flowering and adaptation to temperate regions. Plant Cell 2023, 35, 369–389. [Google Scholar] [CrossRef]
- Li, M.; Cao, L.; Mwimba, M.; Zhou, Y.; Li, L.; Zhou, M.; Schnable, P.S.; O’rourke, J.A.; Dong, X.; Wang, W. Comprehensive mapping of abiotic stress inputs into the soybean circadian clock. Proc. Natl. Acad. Sci. USA 2019, 116, 23840–23849. [Google Scholar] [CrossRef] [PubMed]
- James, A.B.; Syed, N.H.; Bordage, S.; Marshall, J.; Nimmo, G.A.; Jenkins, G.I.; Herzyk, P.; Brown, J.W.; Nimmo, H.G. Alternative Splicing Mediates Responses of the Arabidopsis Circadian Clock to Temperature Changes. Plant Cell 2012, 24, 961–981. [Google Scholar] [CrossRef] [PubMed]
- Braun, R.; Kath, W.L.; Iwanaszko, M.; Kula-Eversole, E.; Abbott, S.M.; Reid, K.J.; Zee, P.C.; Allada, R. Universal method for robust detection of circadian state from gene expression. Proc. Natl. Acad. Sci. USA 2018, 115, E9247–E9256. [Google Scholar] [CrossRef] [PubMed]
- Marcolino-Gomes, J.; Rodrigues, F.A.; Fuganti-Pagliarini, R.; Bendix, C.; Nakayama, T.J.; Celaya, B.; Molinari, H.B.C.; de Oliveira, M.C.N.; Harmon, F.G.; Nepomuceno, A. Diurnal Oscillations of Soybean Circadian Clock and Drought Responsive Genes. PLoS ONE 2014, 9, e86402. [Google Scholar] [CrossRef]
- Romanowski, A.; Schlaen, R.G.; Perez-Santangelo, S.; Mancini, E.; Yanovsky, M.J. Global transcriptome analysis reveals circadian control of splicing events in Arabidopsis thaliana. Plant J. 2020, 103, 889–902. [Google Scholar] [CrossRef]
- Rees, H.; Rusholme-Pilcher, R.; Bailey, P.; Colmer, J.; White, B.; Reynolds, C.; Ward, S.J.; Coombes, B.; Graham, C.A.; Dantas, L.L.d.B.; et al. Circadian regulation of the transcriptome in a complex polyploid crop. PLoS Biol. 2022, 20, e3001802. [Google Scholar] [CrossRef] [PubMed]
- Schlueter, J.A.; Lin, J.Y.; Schlueter, S.D.; Vasylenko-Sanders, I.F.; Deshpande, S.; Yi, J.; O’Bleness, M.; Roe, B.A.; Nelson, R.T.; Scheffler, B.E.; et al. Gene duplication and paleopolyploidy in soybean and the implications for whole genome sequencing. BMC Genom. 2007, 8, 330. [Google Scholar] [CrossRef]
- Ovacik, M.A.; Sukumaran, S.; Almon, R.R.; DuBois, D.C.; Jusko, W.J.; Androulakis, I.P. Circadian signatures in rat liver: From gene expression to pathways. BMC Bioinform. 2010, 11, 540. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Wai, C.M.; Ming, R.; Yu, Q. Diurnal Cycling Transcription Factors of Pineapple Revealed by Genome-Wide Annotation and Global Transcriptomic Analysis. Genome Biol. Evol. 2017, 9, 2170–2190. [Google Scholar] [CrossRef] [PubMed]
- Green, R.M.; Tobin, E.M. The Role of CCA1 and LHY in the Plant Circadian Clock. Dev. Cell 2002, 2, 516–518. [Google Scholar] [CrossRef] [PubMed]
- Tsubokura, Y.; Watanabe, S.; Xia, Z.; Kanamori, H.; Yamagata, H.; Kaga, A.; Katayose, Y.; Abe, J.; Ishimoto, M.; Harada, K. Natural variation in the genes responsible for maturity loci E1, E2, E3 and E4 in soybean. Ann. Bot. 2014, 113, 429–441. [Google Scholar] [CrossRef]
- Watanabe, S.; Hideshima, R.; Xia, Z.; Tsubokura, Y.; Sato, S.; Nakamoto, Y.; Yamanaka, N.; Takahashi, R.; Ishimoto, M.; Anai, T.; et al. Map-Based Cloning of the Gene Associated With the Soybean Maturity Locus E3. Genetics 2009, 182, 1251–1262. [Google Scholar] [CrossRef]
- Watanabe, S.; Xia, Z.; Hideshima, R.; Tsubokura, Y.; Sato, S.; Yamanaka, N.; Takahashi, R.; Anai, T.; Tabata, S.; Kitamura, K.; et al. A Map-Based Cloning Strategy Employing a Residual Heterozygous Line Reveals that the GIGANTEA Gene Is Involved in Soybean Maturity and Flowering. Genetics 2011, 188, 395–407. [Google Scholar] [CrossRef]
- Liu, B.; Kanazawa, A.; Matsumura, H.; Takahashi, R.; Harada, K.; Abe, J. Genetic Redundancy in Soybean Photoresponses Associated With Duplication of the Phytochrome A Gene. Genetics 2008, 180, 995–1007. [Google Scholar] [CrossRef]
- Lu, S.; Zhao, X.; Hu, Y.; Liu, S.; Nan, H.; Li, X.; Fang, C.; Cao, D.; Shi, X.; Kong, L.; et al. Natural variation at the soybean J locus improves adaptation to the tropics and enhances yield. Nat. Genet. 2017, 49, 773–779. [Google Scholar] [CrossRef]
- Xia, Z.; Watanabe, S.; Yamada, T.; Tsubokura, Y.; Nakashima, H.; Zhai, H.; Anai, T.; Sato, S.; Yamazaki, T.; Lü, S.; et al. Positional cloning and characterization reveal the molecular basis for soybean maturity locus E1 that regulates photoperiodic flowering. Proc. Natl. Acad. Sci. USA 2012, 109, E2155–E2164. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Song, W.; Wang, L.; Sun, X.; Qi, Y.; Wu, T.; Sun, S.; Jiang, B.; Wu, C.; Hou, W.; et al. Allele combinations of maturity genes E1-E4 affect adaptation of soybean to diverse geographic regions and farming systems in China. PLoS ONE 2020, 15, e0235397. [Google Scholar] [CrossRef] [PubMed]
- Alexa, A.; Rahnenführer, J. Topgo: Enrichment Analysis for Gene Ontology. R Package Version 2023, 2. Available online: https://bioconductor.org/packages/release/bioc/html/topGO.html (accessed on 15 August 2023).
- Ferrari, C.; Proost, S.; Janowski, M.; Becker, J.; Nikoloski, Z.; Bhattacharya, D.; Price, D.; Tohge, T.; Bar-Even, A.; Fernie, A.; et al. Kingdom-wide comparison reveals the evolution of diurnal gene expression in Archaeplastida. Nat. Commun. 2019, 10, 737. [Google Scholar] [CrossRef] [PubMed]
- Mure, L.S.; Le, H.D.; Benegiamo, G.; Chang, M.W.; Rios, L.; Jillani, N.; Ngotho, M.; Kariuki, T.; Dkhissi-Benyahya, O.; Cooper, H.M.; et al. Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 2018, 359, eaao0318. [Google Scholar] [CrossRef]
- Zhang, R.; Lahens, N.F.; Ballance, H.I.; Hughes, M.E.; Hogenesch, J.B. A circadian gene expression atlas in mammals: Implications for biology and medicine. Proc. Natl. Acad. Sci. USA 2014, 111, 16219–16224. [Google Scholar] [CrossRef]
- Muller, N.A.; Wijnen, C.L.; Srinivasan, A.; Ryngajllo, M.; Ofner, I.; Lin, T.; Ranjan, A.; West, D.; Maloof, J.N.; Sinha, N.R.; et al. Domestication selected for deceleration of the circadian clock in cultivated tomato. Nat. Genet. 2016, 48, 89–93. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.C.; Wang, L.-G.; Han, Y.Y.; He, Q.-Y. clusterProfiler: An R Package for Comparing Biological Themes Among Gene Clusters. OMICS J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Yu, G. Enrichplot: Visualization of Functional Enrichment Result. R Package Version 2021. Available online: https://rdrr.io/bioc/enrichplot/ (accessed on 15 August 2023).








| Arabidopsis thaliana | Glycine max | |||||||
|---|---|---|---|---|---|---|---|---|
| Type | GO ID | Description | NES 1 | p Value | Rank 2 | NES | p Value | Rank |
| BP | GO:0042274 | ribosomal small subunit biogenesis | 2.7329 | 0.0001 | 5671 | −1.5852 | 0.0405 | 20,587 |
| CC | GO:0005852 | eukaryotic translation initiation factor 3 complex | 2.3767 | 0.0001 | 5596 | −1.6797 | 0.0131 | 15,219 |
| BP | GO:0055075 | potassium ion homeostasis | 1.7333 | 0.0126 | 9684 | −1.6109 | 0.0496 | 8780 |
| CC | GO:0000502 | proteasome complex | 1.5905 | 0.0146 | 10,956 | −2.2152 | 0.0120 | 16,030 |
| CC | GO:0005839 | proteasome core complex | 1.6584 | 0.0166 | 11,279 | −2.2368 | 0.0013 | 16,579 |
| CC | GO:1905369 | endopeptidase complex | 1.5202 | 0.0229 | 10,956 | −2.4388 | 0.0169 | 16,030 |
| BP | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process | 1.5683 | 0.0364 | 11,279 | −2.0977 | 0.0033 | 16,579 |
| CC | GO:0070993 | translation preinitiation complex | 1.4947 | 0.0488 | 5486 | −1.7524 | 0.0045 | 15,219 |
| Arabidopsis thaliana | Glycine max | |||||||
|---|---|---|---|---|---|---|---|---|
| Term | NES 1 | p Value | FDR 2 | Rank 3 | NES | p Value | FDR | Rank |
| ribonucleoprotein complex subunit organization | 1.965 | 0.000 | 0.003 | 5744 | −1.933 | 0.333 | 0.674 | 20,531 |
| ribonucleoprotein complex assembly | 1.995 | 0.000 | 0.003 | 5744 | −1.943 | 0.333 | 0.674 | 20,531 |
| ribosomal small subunit biogenesis | 2.745 | 0.000 | 0.003 | 5671 | −1.633 | 0.037 | 0.199 | 20,587 |
| cytoplasmic translation | 2.741 | 0.000 | 0.003 | 5696 | −1.311 | 0.185 | 0.494 | 18,680 |
| maturation of SSU-rRNA from tricistronic rRNA transcript | 2.491 | 0.000 | 0.003 | 7533 | −1.331 | 0.083 | 0.323 | 20,587 |
| cytidine to uridine editing | −0.743 | 0.842 | 0.944 | 17,268 | 2.032 | 0.001 | 0.022 | 5953 |
| phosphogluconate dehydrogenase (decarboxylating) activity | −0.882 | 0.598 | 0.819 | 11,493 | 1.780 | 0.008 | 0.075 | 5914 |
| amyloplast | −0.939 | 0.520 | 0.769 | 10,958 | 1.752 | 0.013 | 0.104 | 11,820 |
| carbon-oxygen lyase activity, acting on phosphates | −0.732 | 0.847 | 0.948 | 15,240 | 1.721 | 0.015 | 0.112 | 8267 |
| protein kinase CK2 complex | −1.005 | 0.422 | 0.703 | 13,202 | 1.684 | 0.019 | 0.131 | 4358 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, X.; Hu, Y.; Wang, W. Comparative Analysis of Circadian Transcriptomes Reveals Circadian Characteristics between Arabidopsis and Soybean. Plants 2023, 12, 3344. https://doi.org/10.3390/plants12193344
Wang X, Hu Y, Wang W. Comparative Analysis of Circadian Transcriptomes Reveals Circadian Characteristics between Arabidopsis and Soybean. Plants. 2023; 12(19):3344. https://doi.org/10.3390/plants12193344
Chicago/Turabian StyleWang, Xingwei, Yanfei Hu, and Wei Wang. 2023. "Comparative Analysis of Circadian Transcriptomes Reveals Circadian Characteristics between Arabidopsis and Soybean" Plants 12, no. 19: 3344. https://doi.org/10.3390/plants12193344





