Comparative Transcriptome and Metabolome Profiling Reveal Mechanisms of Red Leaf Color Fading in Populus × euramericana cv. ‘Zhonghuahongye’
Abstract
:1. Introduction
2. Results
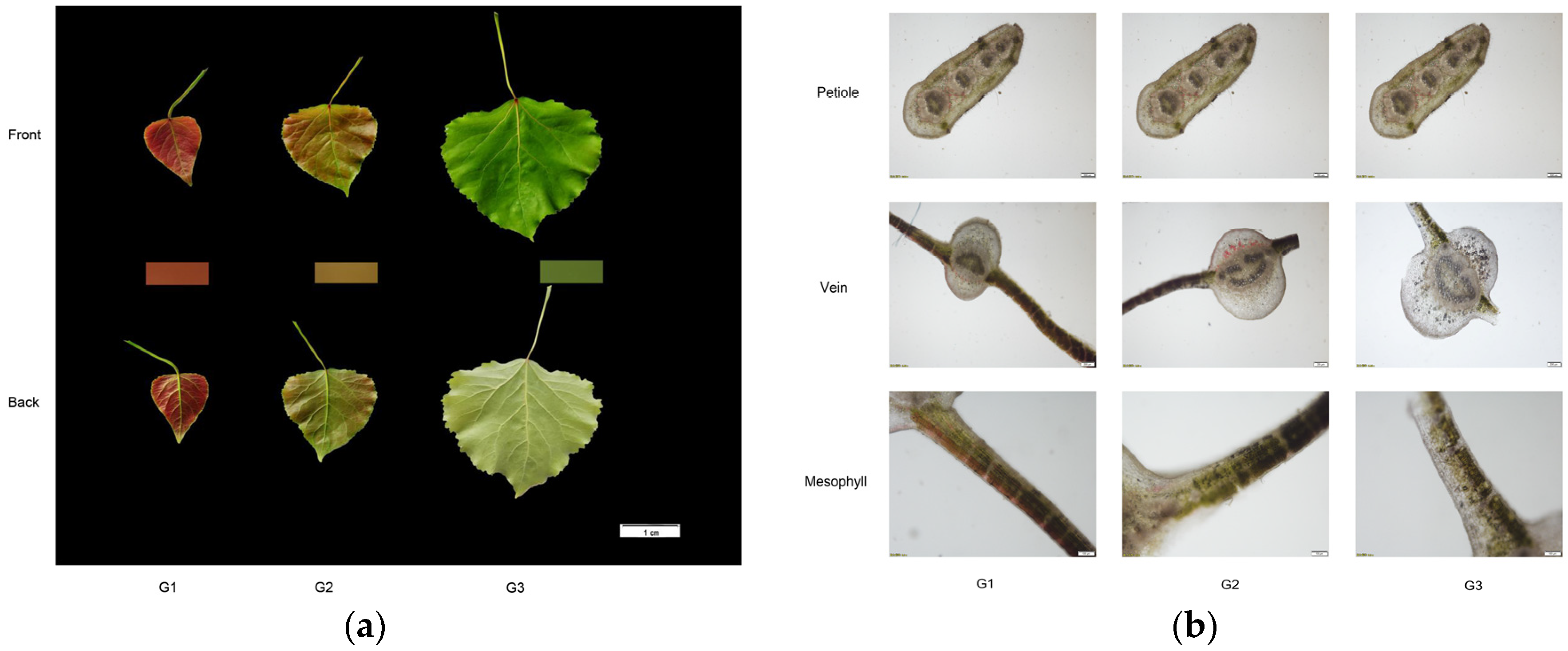
2.1. Comparison of Color and Phenotype and Changes in Leaf Pigment Content
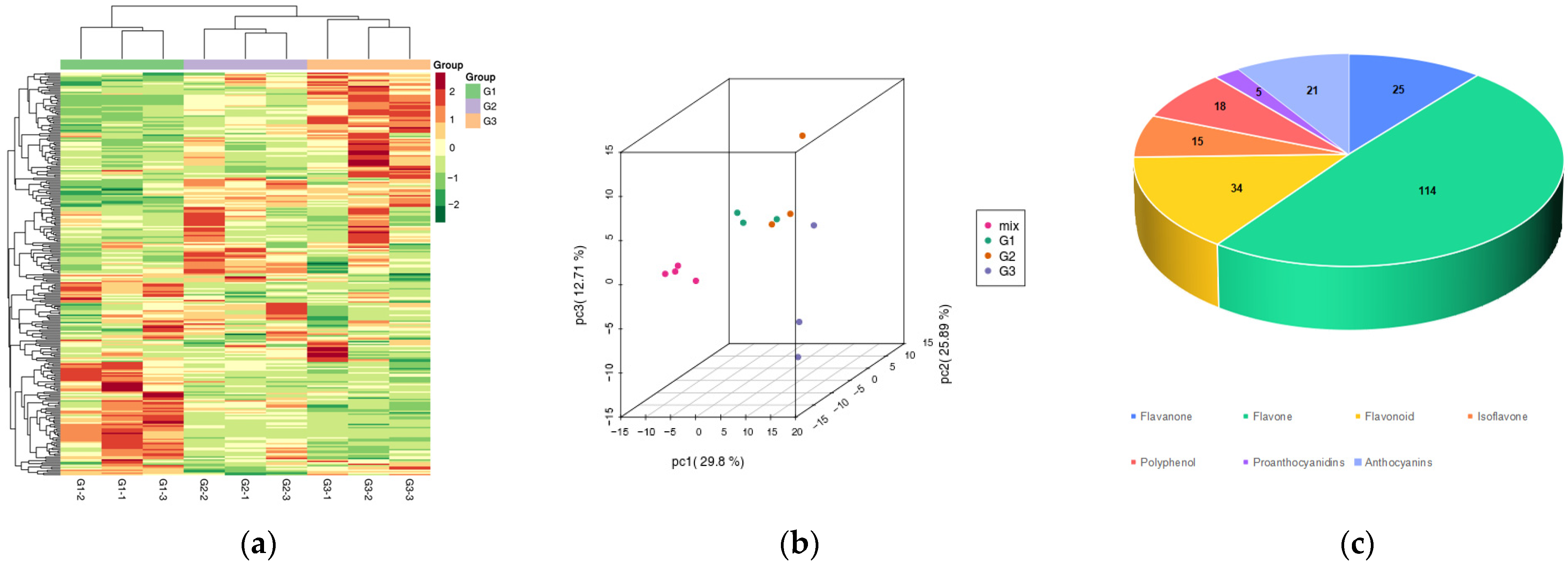
2.2. Qualitative and Quantitative Analysis of Leaf Metabolites in Different Periods
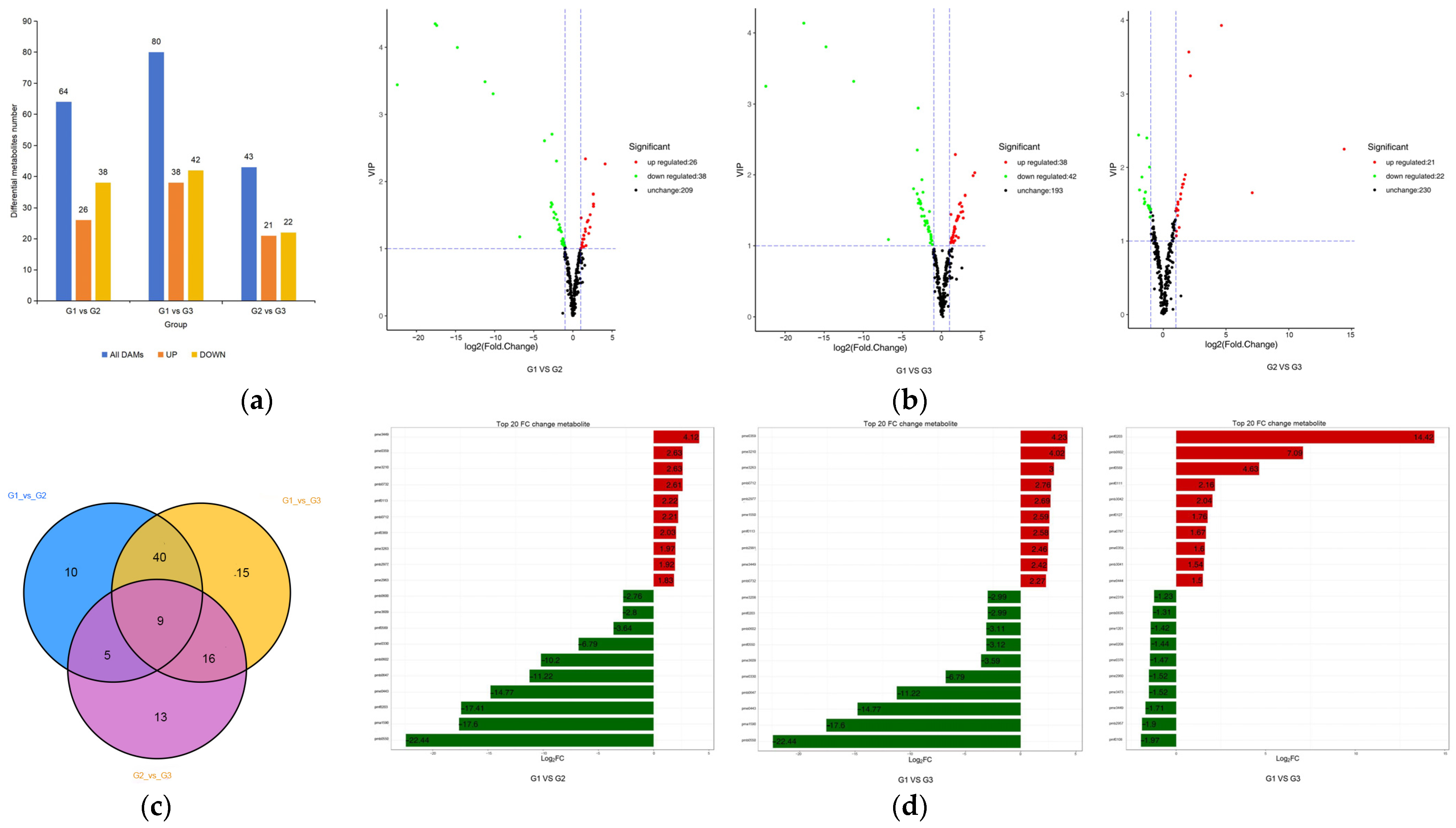
2.3. Analysis of Leaf Metabolites Difference in Different Periods
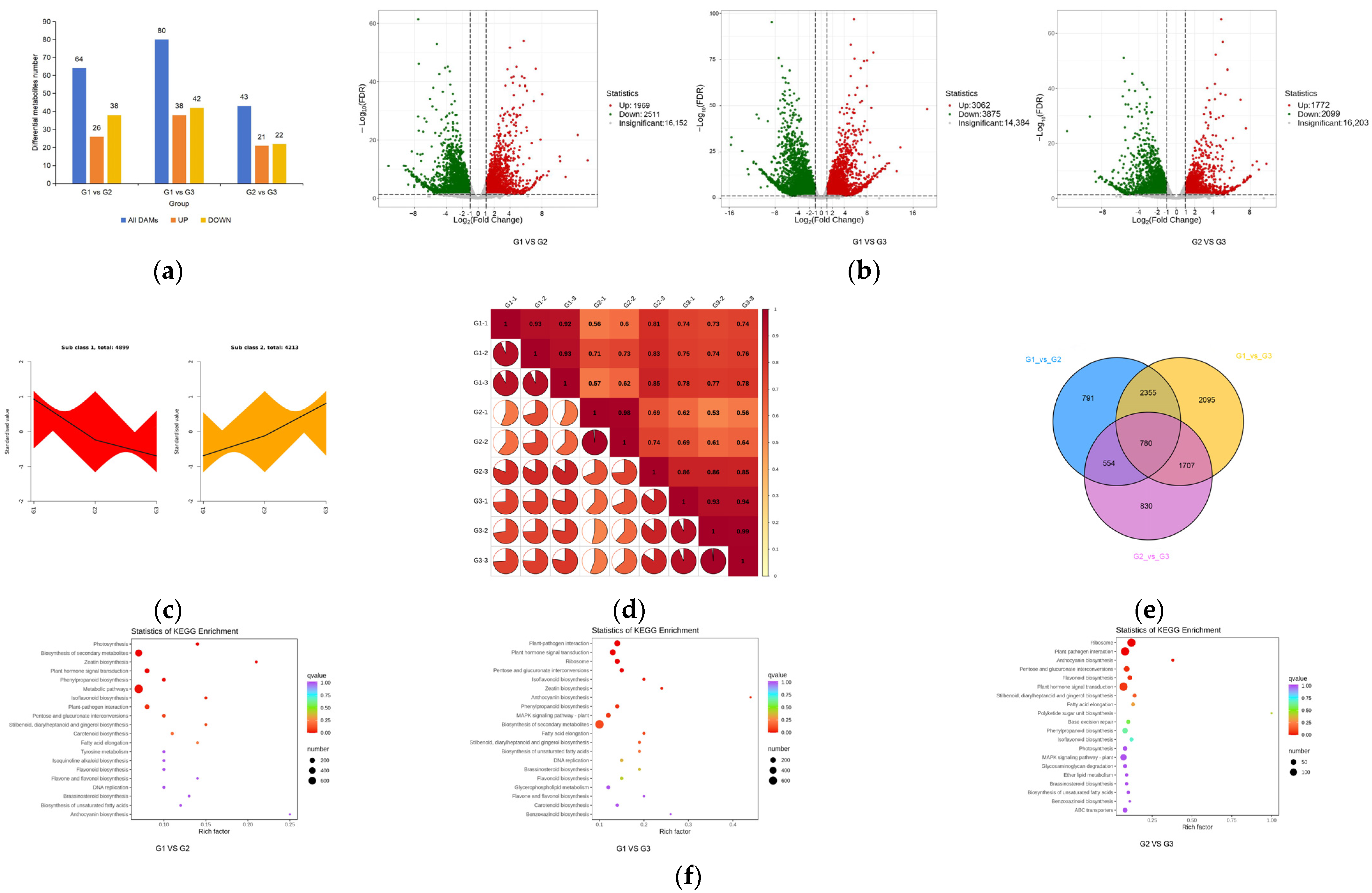
2.4. Functional Annotation and Enrichment Analysis of Differential Metabolite KEGG
2.5. Metabolome and Transcriptome Association Analysis
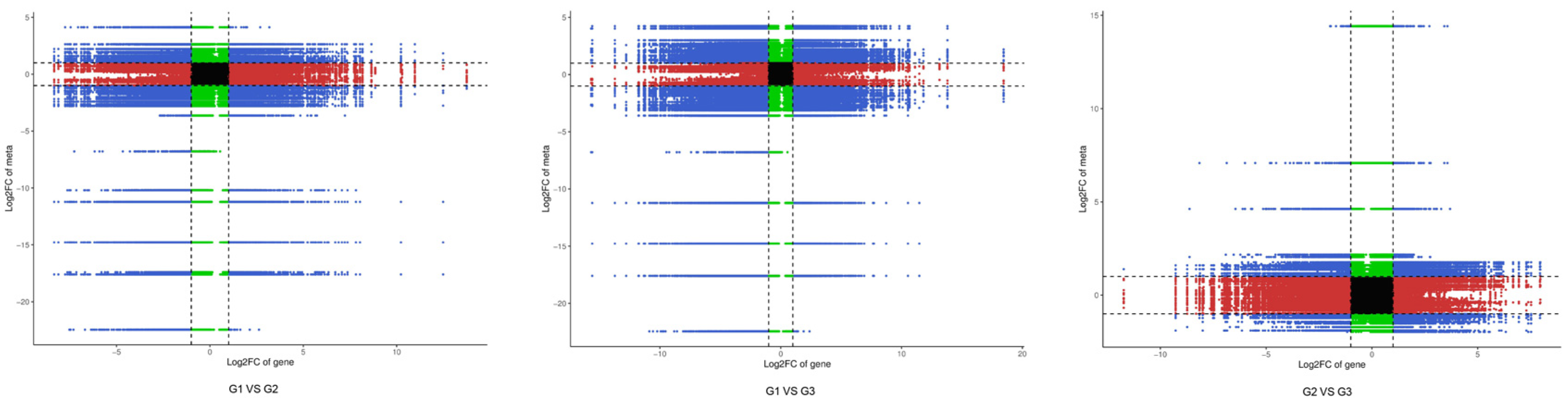
2.5.1. Model-Based Association Analysis
2.5.2. Association Analysis Based on KEGG
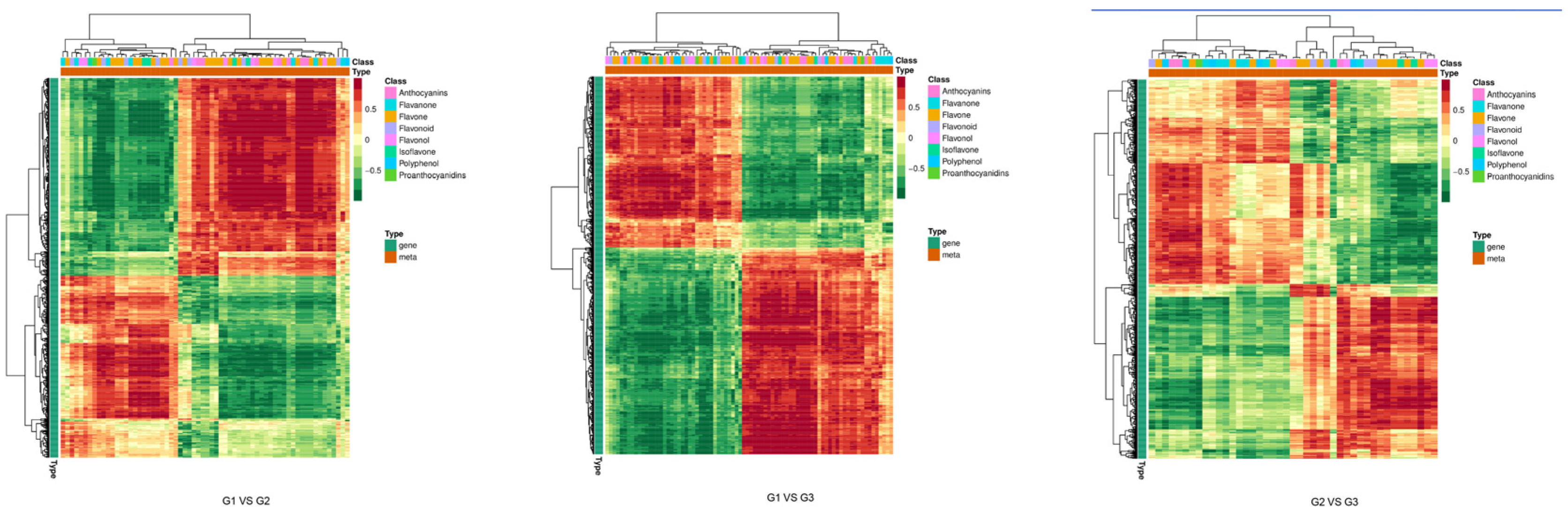
2.5.3. Correlation Analysis Based on Correlation
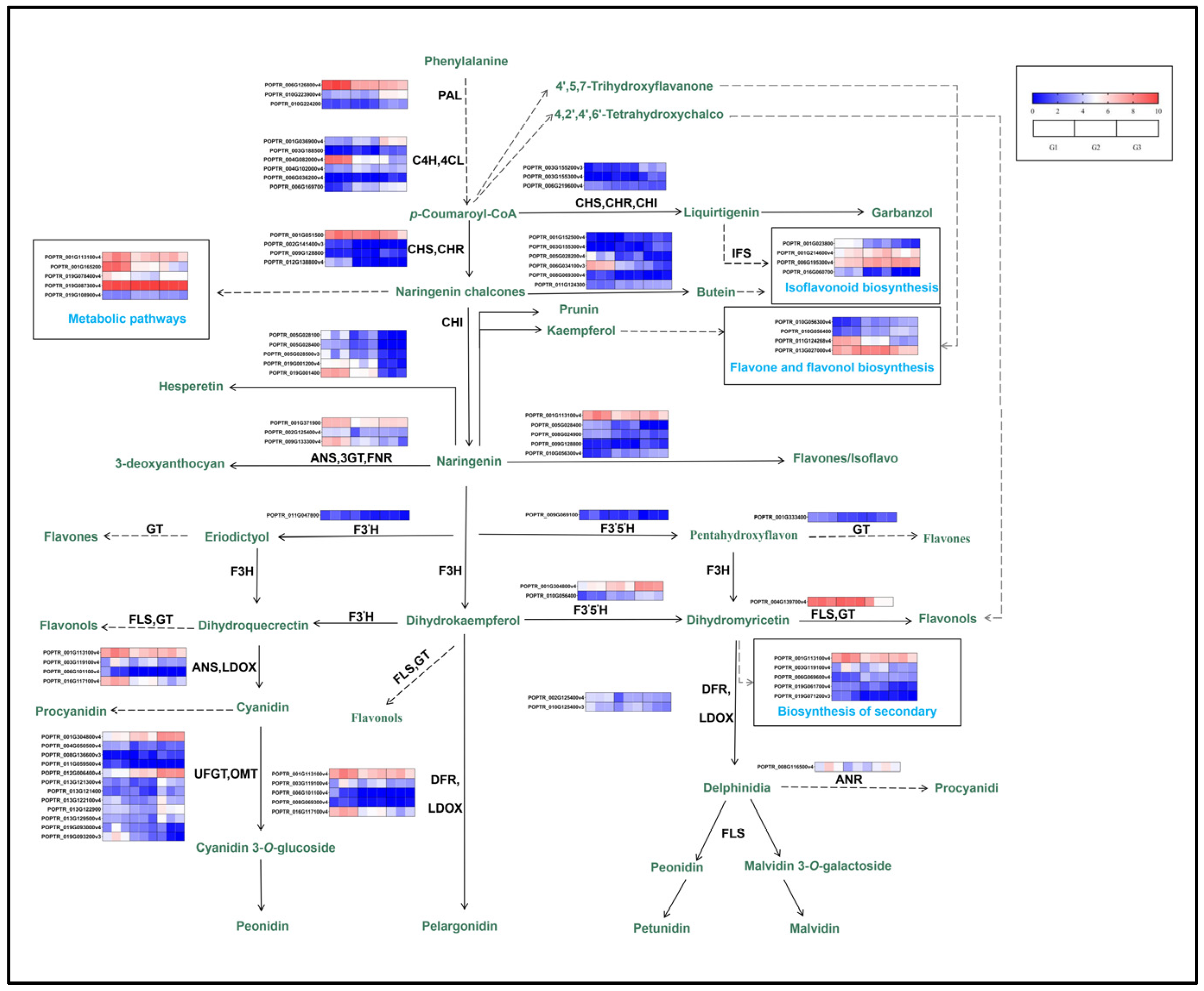
2.5.4. Influence on Differential Metabolites and Differential Genes Regulating Anthocyanin Synthesis of ‘Zhonghuahongye’
2.5.5. Screening of Related Transcription Factors and Related Metabolites Affecting Anthocyanin Synthesis in Leaves of ‘Zhonghuahongye’
2.5.6. Key Genes Were Verified Using qRT-PCR
2.5.7. Study on the Synthesis and Regulation of Anthocyanins in Leaves of ‘Zhonghuahongye’
3. Discussion
3.1. Primary Study on the Main Substances of Leaf Color and the Cause of Color Fading of Red Leaves of ‘Zhonghuahongye’
3.2. Changes in Metabolome and Transcriptome in Leaves of ‘Zhonghuahongye’ in Different Periods
3.3. Atavism of Zhonghuahongye’s Leaves
4. Materials and Methods
4.1. Plant Material Treatments
4.2. Determination of Color Phenotype
4.3. Determination of Pigment Content
4.4. Metabolite Analysis
- (1)
- If the metabolite difference is more than 2 times or less than 0.5 times, the difference is considered to be significant.
- (2)
- On the basis of the above, select metabolites with VIP ≥ 1. This indicates that the difference between groups of metabolites has a strong influence on the classification and discrimination of each group of samples, and that the difference is considered significant.
4.5. Transcriptomic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, Q. Study on the Resistance to Competition and Leaf Xerophytic Structure of 2 New Poplar Clones. Master’s Thesis, Northwest A&F University, Xianyang, China, 2012. [Google Scholar]
- Michalak, M.; Plitta-Michalak, B.P.; Naskręt-Barciszewska, M.Z. DNA Methylation as an early indicator of aging in stored seeds of “exceptional” species Populus nigra L. Cells 2022, 11, 2080. [Google Scholar] [CrossRef]
- Cheng, X. A Study on the Phenomenon of Red Leaf Poplar Returning to the Ancestor. Master’s Thesis, Henan Agricultural University, Zhengzhou, China, 2012. [Google Scholar]
- Wu, D. Metabolomic Analysis and Molecular Mechanism Exploration of the Fading Phenomenon of Young Red in Sweet Potato Leaves. Master’s Thesis, Hainan University, Haikou, China, 2020. [Google Scholar]
- Hall, B.K. Developmental mechanisms underlying the formation of atavisms. Biol. Rev. 1984, 59, 89–122. [Google Scholar] [CrossRef]
- Chen, X.; Wang, N. Research on atavism of plants. Acta Agric. Shanghai 2012, 28, 102–105. [Google Scholar]
- Song, B.; Stöcklin, J.; Armbruster, W.S. Reversible colour change in leaves enhances pollinator attraction and reproductive success in Saururus chinensis (Saururaceae). Ann. Bot. 2018, 14, 641–650. [Google Scholar] [CrossRef]
- Li, X. The Effect of different temperature differences between day and night on the leaf color of ‘Zhonghuahongye’. J. Henan Agric. Sci. 2011, 40, 119–122. [Google Scholar]
- Zhang, X.; Cheng, Y. Plants are also “atavism”—Exploring the Atavism of plants with the example of Calamagrostis fragrans. Fujian J. Agric. Sci. 2014, 2, 19–21. [Google Scholar]
- Hou, M. Preliminary Study on Physiology, Biochemistry and Structure Related to Leaf Color Change of ‘Zhonghuahongye’. Master’s Thesis, Huazhong Agricultural University, Wuhan, China, 2008. [Google Scholar]
- Chen, M.J.; Chang, C.F. Metabolome analysis reveals flavonoid changes during the leaf color transition in Populus×euramerican ‘Zhonghuahongye’. Front. Plant Sci. 2023, 14, 1162893. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhao, X. Research progress on the regulation of biosynthesis of plant flavonoids. Sci. Technol. Food Ind. 2021, 42, 454–463. [Google Scholar]
- Wu, X.X.; Zhang, A. Study on bioanabolic pathway and regulatory factors of plant anthocyanins. Acta. Agric. Shanghai 2018, 34, 127–132. [Google Scholar]
- Baudry, A.; Heim, M. TT2, TT8, and TTG1 synergistically specify the expression of BANYULS and proanthocyanidin biosynthesis in Arabidopsis thaliana. Plant J. 2004, 39, 366–380. [Google Scholar] [CrossRef]
- Kundan, M.; Gani, U. Two R2R3-MYB transcription factors, CsMYB33 and CsMYB78 are involved in the regulation of anthocyanin biosynthesis in Cannabis sativa L. Ind. Crops Prod. 2022, 188, 115546. [Google Scholar] [CrossRef]
- Boldt, J.; Erwin, J. Foliar anthocyanins: A horticultural review. Hortic. Rev. (Am. Soc. Hortic. Sci.) 2014, 42, 209–251. [Google Scholar]
- Wang, J. Study on the Production of Flavonoids by Suspension Cell Culture of Hypericum perforatum. Master’s Thesis, East China University of Science and Technology, Shanghai, China, 2015. [Google Scholar]
- Jin, J.J.; Zhang, H.; Zhang, J.F. Integrated transcriptomics and metabolomics analysis to characterize cold stress responses in Nicotiana tabacum. BMC Genom. 2017, 18, 496. [Google Scholar] [CrossRef]
- Liu, J.Q.; Zhou, H.B.; Song, L. Anthocyanins: Promising natural products with diverse pharmacological activities. Molecules 2021, 26, 3807–3832. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Li, Y. Molecular and Metabolic Insights into Anthocyanin Biosynthesis for Leaf Color Change in Chokecherry (Padus virginiana). Int. J. Mol. Sci. 2021, 22, 10697. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Yu, Y. Research progress on pigment composition, structure and photosynthetic characteristics of colorful leaf plants. Plant Physiol. 2016, 52, 1–7. [Google Scholar]
- Hu, C.; Shang, H. A brief description of the coloration mechanism of color-leaved plants. J. Huazhong Agric. Univ. 2014, 8, 162–163. [Google Scholar]
- Yao, X.H.; An, L.K.; Li, X. Accumulation and regulation of anthocyanins in white and purple Tibetan Hulless Barley (Hordeum vulgare L. var. nudum Hook. f.) revealed by combined de novo transcriptomics and metabolomics. BMC Plant Biol. 2022, 22, 391. [Google Scholar] [CrossRef]
- Li, X. Physiological and Biochemical Studies on Leaf Color Changes of Populus deltoids. Master’s Thesis, Henan Agricultural University, Zhengzhou, China, 2008. [Google Scholar]
- Hong, Y. Study on Anthocyanin Changes and Differential Expression of Related Genes during Strawberry Petal Development. Master’s Thesis, Fujian Agriculture and Forestry University, Fuzhou, China, 2020. [Google Scholar]
- Zhang, Q.; Wang, L.L.; Liu, Z.G. Transcriptome and metabolome profiling unveil the mechanisms of Ziziphus jujuba Mill. peel coloration. Food Chem. 2020, 15, 125903. [Google Scholar] [CrossRef]
- Cheng, S.; Wang, Y. Study on the relationship between pigment content and flavonoid content in Ginkgo biloba leaves. For. Sci. 2001, 5, 31–34. [Google Scholar]
- Ku, Y.S.; Ng, M.S.; Cheng, S.S. Understanding the Composition, Biosynthesis, Accumulation and transport of flavonoids in crops for the promotion of crops as healthy sources of flavonoids for human consumption. Nutrients 2020, 12, 1717. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y. Cloning and Expression Differences of Anthocyanin Synthesis Genes in Tulips and Their Relationship with Flower Color Changes. Master’s Thesis, Shanghai Jiao Tong University, Shanghai, China, 2015. [Google Scholar]
- Fogelman, E.; Tanami, S. Anthocyanin synthesis in native and wound peridermsof potato. Physiol. Plant. 2015, 153, 616–626. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.L.; Pei, X.N.; Zhang, H. MYB-Mediated Regulation of Anthocyanin Biosynthesis. Int. J. Mol. Sci. 2021, 22, 3103. [Google Scholar] [CrossRef]
- Wang, L.; Lu, W. R2R3-MYB transcription factor MYB6 promotes anthocyanin and proanthocyanidin biosynthesis but inhibits secondary cell wall formation in Populus tomentosa. Plant J. 2019, 99, 733–751. [Google Scholar] [CrossRef] [PubMed]
- Fan, D.; Wang, X. Histone H3K9 demethylase JMJ25 epigenetically modulates anthocyanin biosynthesis in poplar. Plant J. 2018, 96, 1121–1136. [Google Scholar] [CrossRef]
- Zhang, Y.; Butelli, E.; Martin, C. Engineering anthocyanin biosynthesis in plants. Curr. Opin. Plant Biol. 2014, 19, 81–90. [Google Scholar] [CrossRef]
- Tan, J.; Liu, M. Preliminary study on the types and characteristics of mulberry leaf color mutants. Sci. Sericul. 2003, 3, 286–290. [Google Scholar]
- Tian, Y.; Li, Q. Metabolic profiling and gene expression analysis provides insights into flavonoid and anthocyanin metabolism in poplar. Tree Physiol. 2020, 41, 1046–1064. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B. HISAT:A fast spliced aligner with low memory requirements. Nat. Nanotechnol. 2015, 12, 357. [Google Scholar]
- Chetri, H.; Furches, A. Genome-Wide Association Study of Wood Anatomical and Morphological Traits in Populus trichocarpa. Front. Plant Sci. 2020, 9, 545748. [Google Scholar] [CrossRef]
- Love, M.; Huber, W. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar]
- Varet, H.; Brillet-Guéguen, L. SARTools: A DESeq2- and edgeR-based R pipeline for comprehensive differential analysis of RNA-Seq data. PLoS ONE 2016, 11, e157022. [Google Scholar]
- Kanehisa, M.; Sato, Y.; Kawashima, M. KEGG mapping tools for uncovering hidden features in biological data. Protein Sci. 2022, 31, 47–53. [Google Scholar]
- Crécy-Lagard, V.; Amorin, H.R.; Arighi, C.; Babor, J. A roadmap for the functional annotation of protein families: A community perspective. Database 2022, 12, baac062. [Google Scholar]
- Tatusov, R.; Galperin, M. The COG database: A tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000, 28, 33–36. [Google Scholar] [CrossRef]
- Gong, W.F.; Song, Q.L.; Ji, K. Fulllength transcriptome from Camellia oleifera seed provides insight into the transcript variants involved in oil biosynthesis. J. Agric. Food Chem. 2020, 68, 14670–14683. [Google Scholar]
- Schmittgen, T.; Livak, K. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [PubMed]


| Pigment | G1 | G2 | G3 |
|---|---|---|---|
| Anthocyanin | 0.24 ± 0.00 a | 0.10 ± 0.01 b | 0.09 ± 0.00 c |
| Chlorophyll a | 0.23 ± 0.00 b | 0.71 ± 0.03 a | 0.70 ± 0.03 a |
| Chlorophyll b | 0.13 ± 0.01 b | 0.22 ± 0.01 a | 0.22 ± 0.01 a |
| Total chlorophyll | 0.36 ± 0.01 b | 0.93 ± 0.04 a | 0.92 ± 0.04 a |
| Carotenoid | 0.15 ± 0.02 b | 0.25 ± 0.02 a | 0.24 ± 0.02 a |
| KEGG Pathway | G1 vs. G2 | G1 vs. G3 | G2 vs. G3 |
|---|---|---|---|
| ko00944 | 2 | 1 | 1 |
| ko00943 | 7 | 7 | 6 |
| ko00942 | 2 | 3 | 2 |
| ko00941 | 9 | 11 | 10 |
| KEGG_Map | Description | p-Value_Meta | Count_Meta | p-Value_Gene | Count_Gene |
|---|---|---|---|---|---|
| ko01110 | Biosynthesis of secondary metabolites | 0.14436567 | 11 | 4.200984 × 10−6 | 470 |
| ko01100 | Metabolic pathways | 0.76832403 | 3 | 5.180148 × 10−5 | 793 |
| ko00943 | Isoflavonoid biosynthesis | 0.04405258 | 7 | 2.393100 × 10−4 | 19 |
| ko00942 | Anthocyanin biosynthesis | 0.45535157 | 2 | 1.330260 × 10−2 | 4 |
| ko00941 | Flavonoid biosynthesis | 0.25504294 | 9 | 1.581373 × 10−2 | 24 |
| ko00944 | Flavone and flavonol biosynthesis | 0.93960917 | 2 | 2.652742 × 10−2 | 7 |
| ID | Family | Category | log2 FC | Type |
|---|---|---|---|---|
| POPTR_017G082500v4 | MYB | TF | 2.952 | up |
| POPTR_017G125600 | MYB | TF | −3.548 | down |
| POPTR_015G082700v4 | MYB | TF | 6.074 | up |
| POPTR_014G081200v4 | MYB | TF | 3.203 | up |
| POPTR_013G149200v4 | MYB | TF | −3.054 | down |
| POPTR_012G080400v4 | MYB | TF | −3.344 | down |
| POPTR_010G141000v4 | MYB | TF | −5.133 | down |
| POPTR_009G053900v4 | MYB | TF | 4.733 | up |
| POPTR_003G114100 | MYB | TF | 3.972 | up |
| POPTR_001G118800v4 | MYB | TF | 3.289 | up |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, S.; Yu, X.; Chen, M.; Chang, C.; Zhu, J.; Zhao, H. Comparative Transcriptome and Metabolome Profiling Reveal Mechanisms of Red Leaf Color Fading in Populus × euramericana cv. ‘Zhonghuahongye’. Plants 2023, 12, 3511. https://doi.org/10.3390/plants12193511
Zhang S, Yu X, Chen M, Chang C, Zhu J, Zhao H. Comparative Transcriptome and Metabolome Profiling Reveal Mechanisms of Red Leaf Color Fading in Populus × euramericana cv. ‘Zhonghuahongye’. Plants. 2023; 12(19):3511. https://doi.org/10.3390/plants12193511
Chicago/Turabian StyleZhang, Shaowei, Xinran Yu, Mengjiao Chen, Cuifang Chang, Jingle Zhu, and Han Zhao. 2023. "Comparative Transcriptome and Metabolome Profiling Reveal Mechanisms of Red Leaf Color Fading in Populus × euramericana cv. ‘Zhonghuahongye’" Plants 12, no. 19: 3511. https://doi.org/10.3390/plants12193511







