Genome-Wide Identification and Expression Analysis of Auxin Response Factor (ARF) Gene Family in Panax ginseng Indicates Its Possible Roles in Root Development
Abstract
:1. Introduction
2. Results
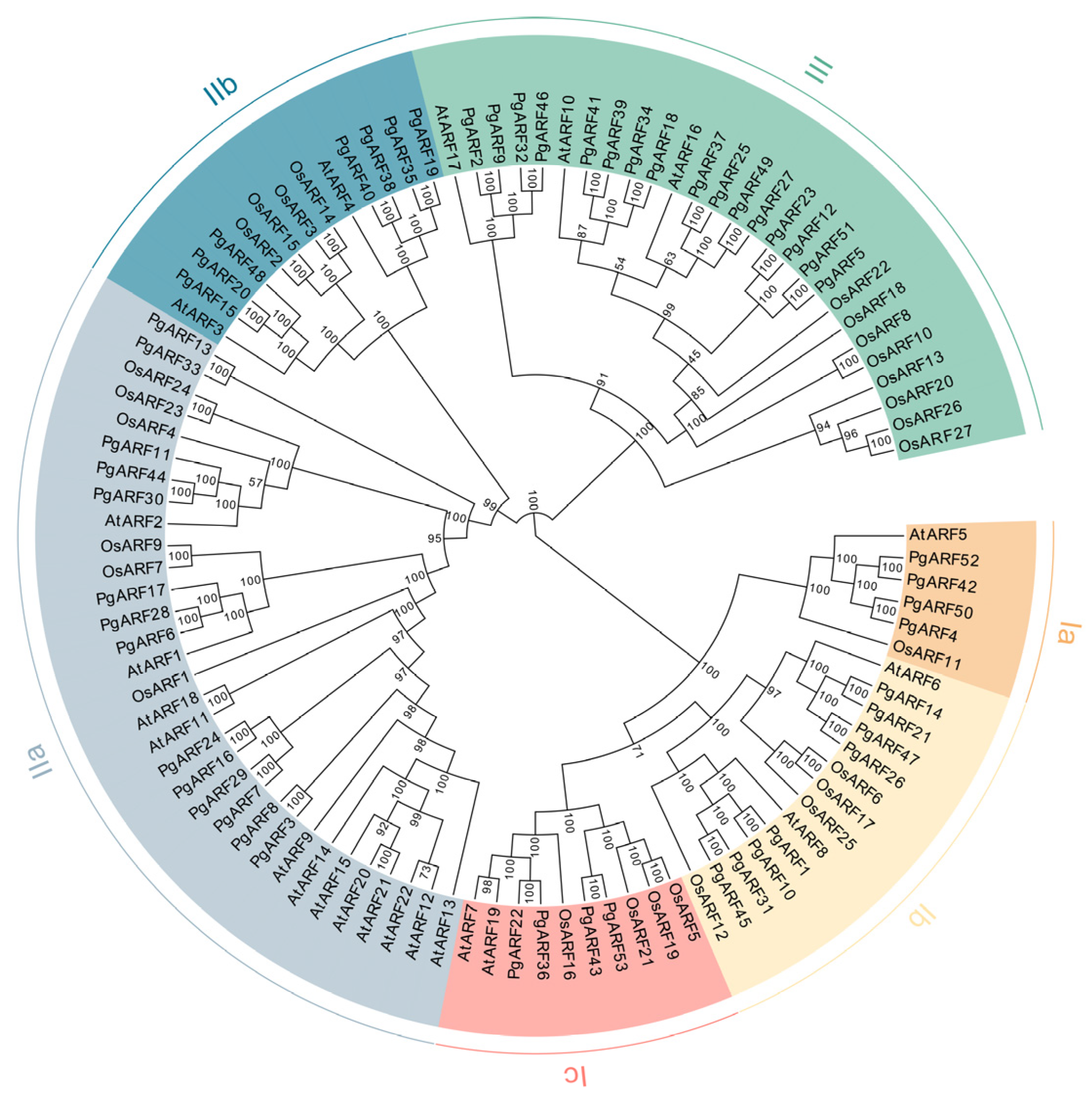
2.1. Identification, Classification, and Phylogenetic Analysis of PgARF
2.2. Protein Conservation Motif Analysis of PgARF
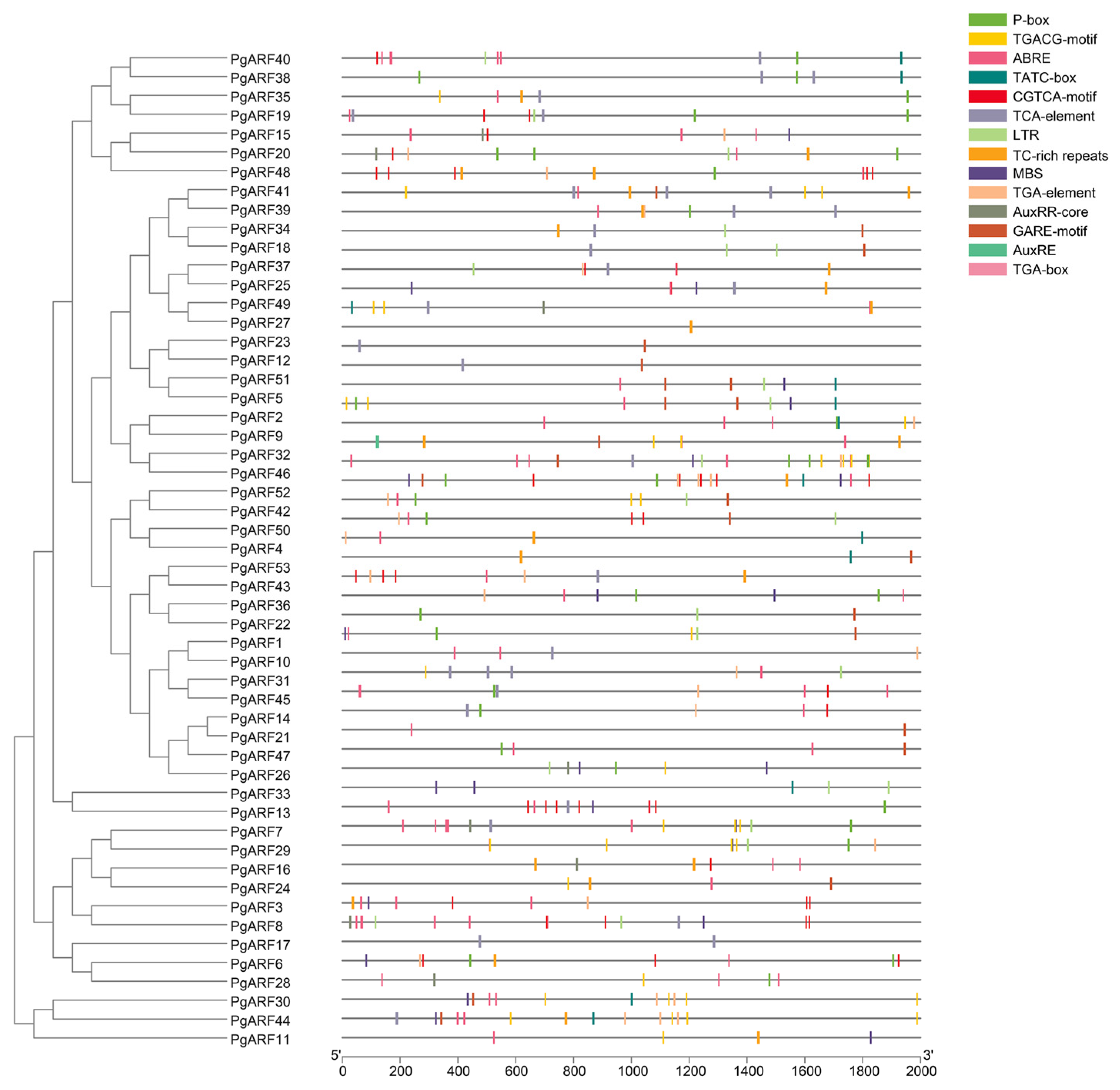
2.3. Analysis of Cis-Elements of PgARF Family Promoters
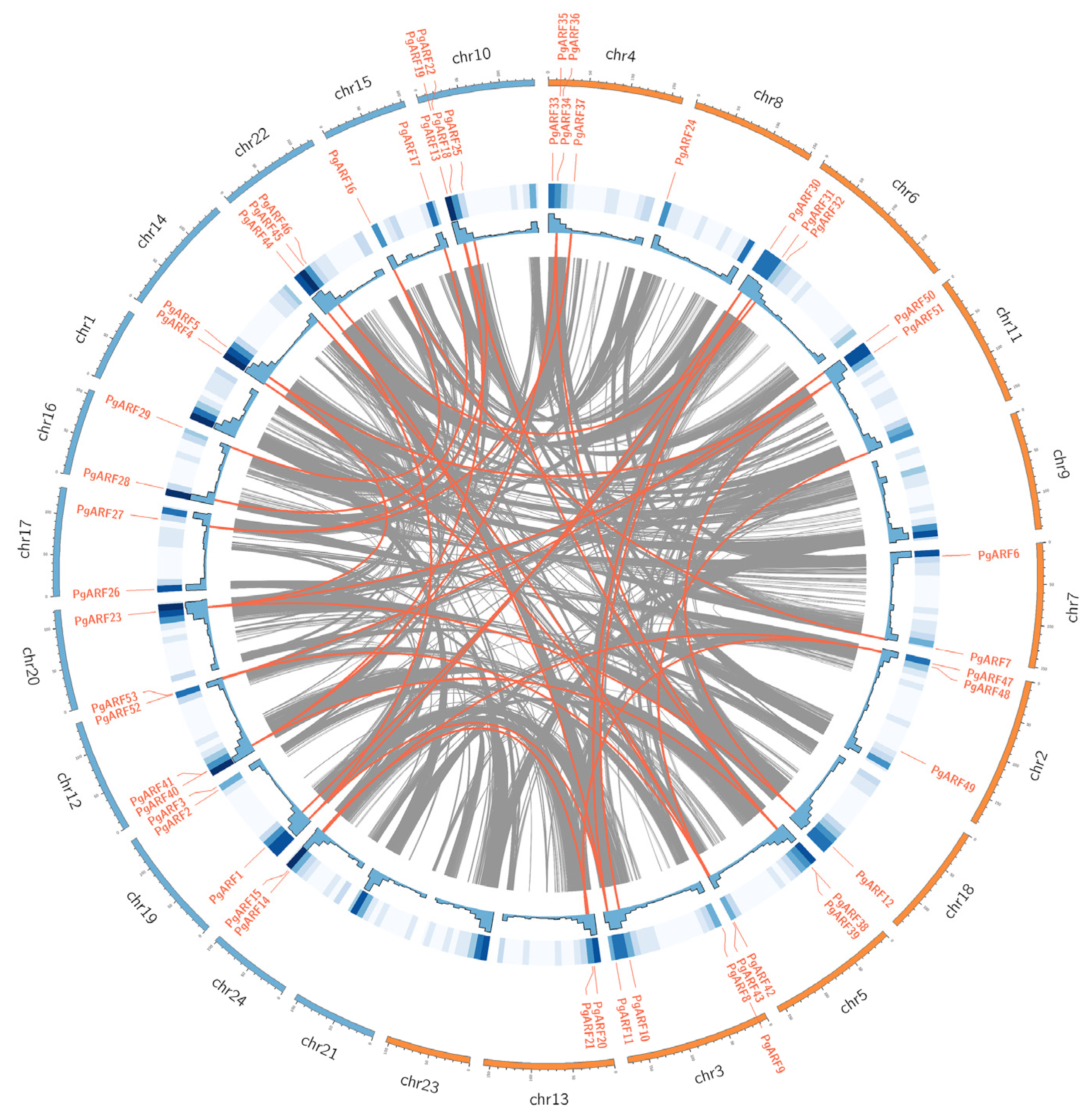
2.4. Chromosomal Distribution and Collinearity Analysis of PgARF
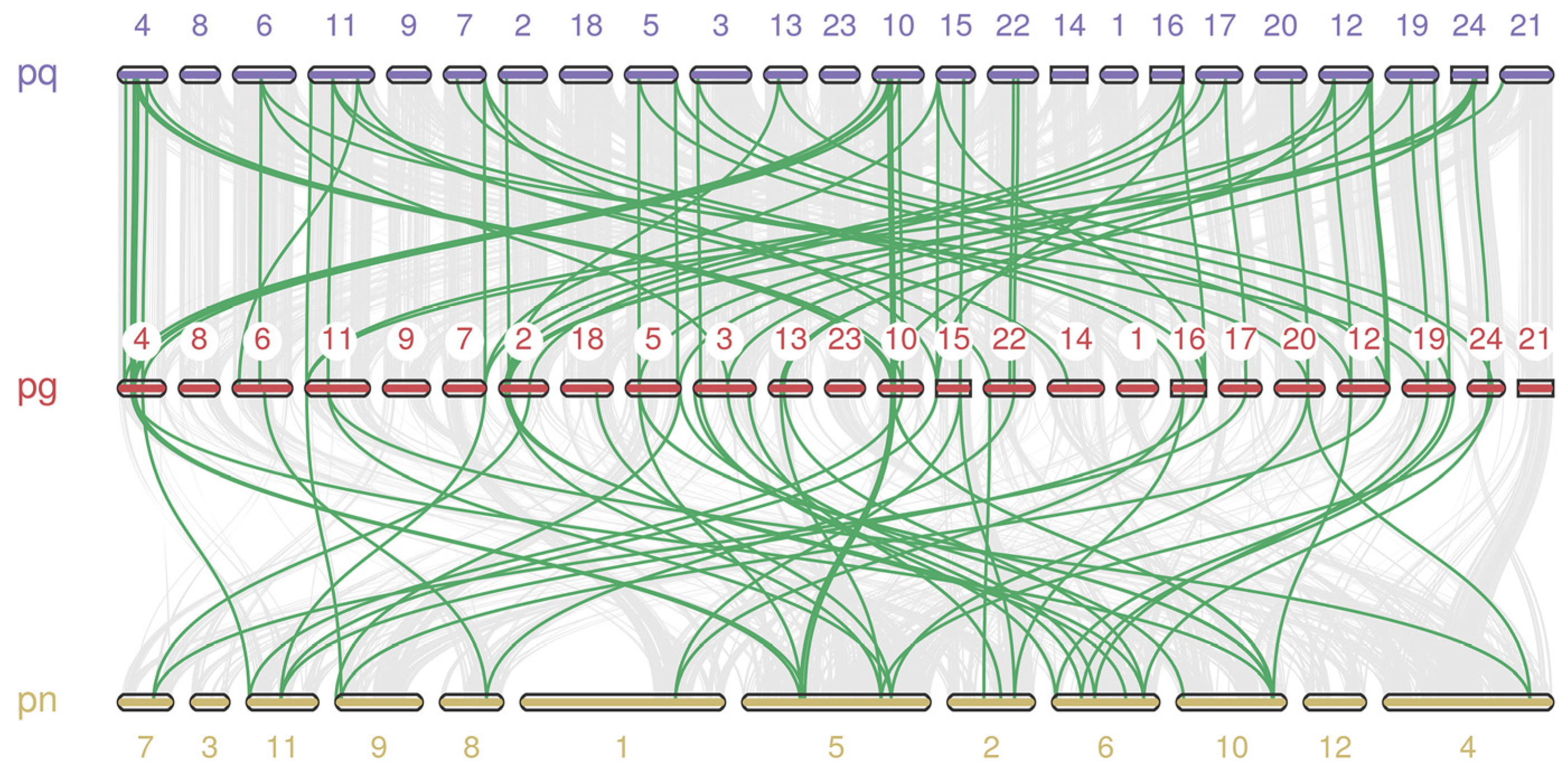
2.5. Synteny and Evolutionary Analyses of PgARF and Other Plants’ ARF
2.6. Expression Profiles of PgARF in Different Tissues and Growth Years
2.7. Expression Profiles of PgARF under Different Environmental Stresses and Hormone Treatments
2.8. Subcellular Localization of PgARF14, PgARF42, and PgARF53
2.9. PgARF14 and PgARF53 Increase the Root Length in Arabidopsis
3. Discussion
4. Materials and Methods
4.1. Identification and Annotation of ARF Genes in Panax ginseng
4.2. Phylogenetic analysis of PgARF with AtARFs and OsARFs
4.3. Exon–Intron Structure, Conserved Motif, and Cis-Acting Element Analysis
4.4. Chromosomal Location, Duplication, Synteny, and Evolution Analyses PgARF
4.5. PgARF Gene Expression Analysis
4.6. Subcellular Localization
4.7. Plant Materials
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Lai, X.; Chahtane, H.; Martin-Arevalillo, R.; Zubieta, C.; Parcy, F. Contrasted evolutionary trajectories of plant transcription factors. Curr. Opin. Plant Biol. 2015, 37, 33. [Google Scholar] [CrossRef] [PubMed]
- Benfey, P.N. Toward a systems analysis of the root. Cold Spring Harb. Symp. Quant. Biol. 2012, 77, 91–96. [Google Scholar] [CrossRef]
- Ayaz, A.; Saqib, S.; Huang, H.; Zaman, W.; Lü, S.; Zhao, H. Genome-wide comparative analysis of long-chain acyl-CoA synthetases (LACSs) gene family: A focus on identification, evolution and expression profiling related to lipid synthesis. Plant Physiol. Bioch. 2021, 161, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Soltis, D.E.; Soltis, P.S.; Albert, V.A.; Oppenheimer, D.G.; DePamphilis, C.W.; Ma, H.; Frohlich, M.W.; Theißen, G. Missing links: The genetic architecture of flower and floral diversification. Trends Plant Sci. 2002, 1, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Xiong, H.; Lu, D.; Li, Z.; Wu, J.; Ning, X.; Lin, W.; Bai, Z.; Zheng, C.; Sun, Y.; Chi, W.; et al. The DELLA-ABI4-HY5 module integrates light and gibberellin signals to regulate hypocotyl elongation. Plant Commun. 2023, 4, 100597. [Google Scholar] [CrossRef]
- Lu, L.; Zhai, X.; Li, X.; Wang, S.; Zhang, L.; Wang, L.; Jin, X.; Liang, L.; Deng, Z.; Li, Z.; et al. Met1-specific motifs conserved in OTUB subfamily of green plants enable rice OTUB1 to hydrolyse Met1 ubiquitin chains. Nat. Commun. 2022, 13, 4672. [Google Scholar] [CrossRef]
- Boer, D.R.; Freire-Rios, A.; van den Berg, W.A.M.; Saaki, T.; Manfield, I.W.; Kepinski, S.; López-Vidrieo, I.; FrancoZorrilla, J.M.; de Vries, S.C.; Solano, R.; et al. Structural Basis for DNA Binding Specificity by the Auxin-Dependent ARF Transcription Factors. Cell 2014, 156, 577–589. [Google Scholar] [CrossRef]
- Strader, L.; Weijers, D.; Wagner, D. Plant transcription factors-being in the right place with the right company. Curr. Opin. Plant Biol. 2022, 65, 102136. [Google Scholar] [CrossRef]
- Ulmasov, T.; Hagen, G.; Guilfoyle, T.J. Dimerization and DNA binding of auxin response factors. Plant J. Cell Mol. Biol. 1999, 19, 309–319. [Google Scholar] [CrossRef]
- Ulmasov, T.; Hagen, G.; Guilfoyle, T.J. Activation and Repression of Transcription by Auxin-Response Factors. Proc. Natl. Acad. Sci. USA 1999, 96, 5844–5849. [Google Scholar] [CrossRef]
- Nanao, M.H.; Vinos-Poyo, T.; Brunoud, G.; Thevenon, E.; Mazzoleni, M.; Mast, D.; Laine, S.; Wang, S.; Hagen, G.; Li, H.; et al. Structural basis for oligomerization of auxin transcriptional regulators. Nat. Commun. 2014, 5, 3617. [Google Scholar] [CrossRef]
- Li, W.; Chen, F.; Wang, Y.; Zheng, H.; Yi, Q.; Ren, Y.; Gao, J. Genome-wide identification and functional analysis of ARF transcription factors in Brassica juncea var. tumida. PLoS ONE 2020, 15, e232039. [Google Scholar] [CrossRef] [PubMed]
- Hagen, T.J.G.A. Auxin response factors. Curr. Opin. Plant Biol. 2007, 10, 453–460. [Google Scholar]
- Szemenyei, H.; Hannon, M.; Long, J.A. TOPLESS Mediates Auxin-Dependent Transcriptional Repression During Arabidosis Embryogenesis. Science 2008, 319, 1384–1386. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.; Wang, S.; Bai, Y.; Wu, Y.; Zhang, S.; Chen, M.; Guilfoyle, T.J.; Wu, P.; Qi, Y. Functional analysis of the structural domain of ARF proteins in rice (Oryza sativa L.). J. Exp. Bot. 2010, 61, 3971–3981. [Google Scholar] [CrossRef] [PubMed]
- Okushima, Y.; Overvoorde, P.J.; Arima, K.; Alonso, J.M.; Chan, A.; Chang, C.; Ecker, J.R.; Hughes, B.; Lui, A.; Nguyen, D.; et al. Functional Genomic Analysis of the AUXIN RESPONSE FACTOR Gene Family Members in Arabidopsis thaliana: Unique and Overlapping Functions ofARF7 andARF19. Plant Cell 2005, 17, 444–463. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Harter, K.; Theologis, A. Protein–protein interactions among the Aux/IAA proteins. Proc. Natl. Acad. Sci. USA 1997, 94, 11786–11791. [Google Scholar] [CrossRef] [PubMed]
- Cancé, C.; Martin Arevalillo, R.; Boubekeur, K.; Dumas, R. Auxin response factors are keys to the many auxin doors. New Phytol. 2022, 235, 402–419. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.S.; Seo, M.; Cho, H.T. Two TPL-Binding Motifs of ARF2 Are Involved in Repression of Auxin Responses. Front. Plant Sci. 2018, 9, 372. [Google Scholar] [CrossRef]
- Chandler, J.W. Auxin response factors. Plant Cell Environ. 2016, 39, 1014–1028. [Google Scholar] [CrossRef]
- Wang, D.; Pei, K.; Fu, Y.; Sun, Z.; Li, S.; Liu, H.; Tang, K.; Han, B.; Tao, Y. Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 2007, 394, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Ha, C.V.; Le, D.T.; Nishiyama, R.; Watanabe, Y.; Sulieman, S.; Tran, U.T.; Mochida, K.; Dong, N.V.; Yamagu-chi-Shinozaki, K.; Shinozaki, K.; et al. The auxin response factor transcription factor family in soybean: Genome-wide identification and expression analyses during development and water stress. DNA Res. 2013, 20, 511–524. [Google Scholar] [PubMed]
- Diao, D.; Hu, X.; Guan, D.; Wang, W.; Yang, H.; Liu, Y. Genome-wide identification of the ARF (auxin response factor) gene family in peach and their expression analysis. Mol. Biol. Rep. 2020, 47, 4331–4344. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Guo, Z.; Lu, S. Genome-Wide Identification and Expression Analysis of the Aux/IAA and Auxin Response Factor Gene Family in Medicago truncatula. Int. J. Mol. Sci. 2021, 22, 10494. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Wan, Q.; Yu, Z.; Zhang, K.; Zhang, X.; Zhu, S.; Wan, Y.; Ding, Z.; Liu, F. Genome-Wide Identification of Auxin Response Factors in Peanut (Arachis hypogaea L.) and Functional Analysis in Root Morphology. Int. J. Mol. Sci. 2022, 23, 5309. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Wang, S.; Wu, F.; Wei, M.; Li, J.; Yang, F. Genome-Wide Identification and Functional Characterization of Auxin Response Factor (ARF) Genes in Eggplant. Int. J. Mol. Sci. 2022, 23, 6219. [Google Scholar] [CrossRef] [PubMed]
- Tim Ulmasov, G.H.T.J. ARF1, a Transcription Factor That Binds to Auxin Response Elements. Science 1997, 276, 1864–1868. [Google Scholar] [CrossRef]
- Christine, M.; Ellis, P.N.J.C. AUXIN RESPONSE FACTOR1and AUXIN RESPONSE FACTOR2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 2005, 132, 4563–4574. [Google Scholar]
- Guilfoyle, T.J.; Ulmasov, T.; Hagen, G. The ARF family of transcription factors and their role in plant hormone-responsive transcription. Cell. Mol. Life Sci. 1998, 54, 619–627. [Google Scholar] [CrossRef]
- Pekker, I.; Alvarez, J.P.; Eshed, Y. Auxin Response Factors Mediate Arabidopsis Organ Asymmetry via Modulation of KANADI Activity. Plant Cell 2005, 17, 2899–2910. [Google Scholar] [CrossRef]
- Hardtke, C.S.; Berleth, T. The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J. 1998, 17, 1405–1411. [Google Scholar] [CrossRef] [PubMed]
- Hardtke, C.S.; Ckurshumova, W.; Vidaurre, D.P.; Singh, S.A.; Stamatiou, G.; Tiwari, S.B.; Hagen, G.; Guilfoyle, T.J.; Berleth, T. Overlapping and non-redundant functions of the Arabidopsis auxin response factors MONOPTEROS and NONPHOTOTROPIC HYPOCOTYL 4. Development 2004, 131, 1089–1100. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.F.; Tian, Q.; Reed, J.W. Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 2006, 133, 4211–4218. [Google Scholar] [CrossRef] [PubMed]
- Wilmoth, J.C.; Wang, S.; Tiwari, S.B.; Joshi, A.D.; Hagen, G.; Guilfoyle, T.J.; Alonso, J.M.; Ecker, J.R.; Reed, J.W. NPH4/ARF7 and ARF19 promote leaf expansion and auxin-induced lateral root formation. Plant J. 2005, 43, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Xing, H.; Pudake, R.N.; Guo, G.; Xing, G.; Hu, Z.; Zhang, Y.; Sun, Q.; Ni, Z. Genome-wide identification and expression profiling of auxin response factor (ARF) gene family in maize. BMC Genom. 2011, 12, 178. [Google Scholar] [CrossRef]
- Ren, Z.; Liu, R.; Gu, W.; Dong, X. The Solanum lycopersicum auxin response factor SlARF2 participates in regulating lateral root formation and flower organ senescence. Plant Sci. 2017, 256, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Bao, X.; Liu, K.; Wang, J.; Zhang, J.; Feng, Y.; Wang, Y.; Lin, L.; Feng, J.; Li, C. Genome-wide identification and expression profiling of the auxin response factor (ARF) gene family in physic nut. PLoS ONE 2018, 13, e201024. [Google Scholar] [CrossRef]
- Liu, B.; Zhu, J.; Lin, L.; Yang, Q.; Hu, B.; Wang, Q.; Zou, X.X.; Zou, S.Q. Genome-Wide Identification and Co-Expression Analysis of ARF and IAA Family Genes in Euscaphis konishii: Potential Regulators of Triterpenoids and Anthocyanin Biosynthesis. Front. Genet. 2021, 12, 737293. [Google Scholar] [CrossRef]
- Wang, Z.H.; Wang, X.F.; Lu, T.; Li, M.R.; Jiang, P.; Zhao, J.; Liu, S.T.; Fu, X.Q.; Wendel, J.F.; Van de Peer, Y.; et al. Reshuffling of the ancestral core-eudicot genome shaped chromatin topology and epigenetic modification in Panax. Nat. Commun. 2022, 13, 1902. [Google Scholar] [CrossRef]
- Wang, K.; Jiang, S.; Sun, C.; Lin, Y.; Yin, R.; Wang, Y.; Zhang, M. The Spatial and Temporal Transcriptomic Land-scapes of Ginseng, Panax ginseng C. A. Meyer. Sci. Rep.-UK 2015, 5, 18283. [Google Scholar] [CrossRef]
- Kim, N.H.; Jayakodi, M.; Lee, S.C.; Choi, B.S.; Jang, W.; Lee, J.; Kim, H.H.; Waminal, N.E.; Lakshmanan, M.; van Nguyen, B.; et al. Genome and evolution of the shade-requiring medicinal herb Panax ginseng. Plant Biotechnol. J. 2018, 16, 1904–1917. [Google Scholar] [CrossRef]
- Wu, P.; Zhou, C.; Cheng, S.; Wu, Z.; Lu, W.; Han, J.; Chen, Y.; Chen, Y.; Ni, P.; Wang, Y.; et al. Integrated genome sequence and linkage map of physic nut (Jatropha curcas L.), a biodiesel plant. Plant J. 2015, 81, 810–821. [Google Scholar] [CrossRef]
- Ayaz, A.; Huang, H.; Zheng, M.; Zaman, W.; Li, D.; Saqib, S.; Zhao, H.; Lü, S. Molecular Cloning and Functional Analysis of GmLACS2-3 Reveals Its Involvement in Cutin and Suberin Biosynthesis along with Abiotic Stress Tolerance. Int. J. Mol. Sci. 2021, 22, 9175. [Google Scholar] [CrossRef] [PubMed]
- Marin, E.; Jouannet, V.; Herz, A.; Lokerse, A.S.; Weijers, D.; Vaucheret, H.; Nussaume, L.; Crespi, M.D.; Maizel, A. miR390, Arabidopsis TAS3 tasiRNAs, and Their AUXIN RESPONSE FACTOR Targets Define an Autoregulatory Network Quantitatively Regulating Lateral Root Growth. Plant Cell 2010, 22, 1104–1117. [Google Scholar] [CrossRef] [PubMed]
- Kazan, K. Auxin and the integration of environmental signals into plant root development. Ann. Bot.-Lond. 2013, 112, 1655–1665. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Zuo, J.; Hou, X.; Yan, Y.; Wei, Y.; Liu, J.; Li, M.; Xu, B.; Jin, Z. The auxin response factor gene family in banana: Genome-wide identification and expression analyses during development, ripening, and abiotic stress. Front. Plant Sci. 2015, 6, 742. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Khurana, J.P. Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J. 2009, 276, 3148–3162. [Google Scholar] [CrossRef] [PubMed]
- Xi, D.; Chen, X.; Wang, Y.; Zhong, R.; He, J.; Shen, J.; Ming, F. Arabidopsis ANAC092 regulates auxin-mediated root development by binding to the ARF8 and PIN4 promoters. J. Integr. Plant Biol. 2019, 61, 1015–1031. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Yamamuro, C. Interplays between auxin and GA signaling coordinate early fruit development. Hortic. Res.-Engl. 2022, 9, uhab78. [Google Scholar] [CrossRef]
- Hu, J.; Su, H.; Cao, H.; Wei, H.; Fu, X.; Jiang, X.; Song, Q.; He, X.; Xu, C.; Luo, K. AUXIN RESPONSE FACTOR7 integrates gibberellin and auxin signaling via interactions between DELLA and AUX/IAA proteins to regulate cambial activity in poplar. Plant Cell 2022, 34, 2688–2707. [Google Scholar] [CrossRef]
- Dastidar, M.G.; Scarpa, A.; Mägele, I.; Ruiz Duarte, P.; Born, P.; Bald, L.; Jouannet, V.; Maizel, A. ARF5/MONOPTEROS directly regulates miR390 expression in the Arabidopsis thaliana primary root meristem. Plant Direct 2019, 3, e116. [Google Scholar] [CrossRef] [PubMed]
- He, F.; Xu, C.; Fu, X.; Shen, Y.; Guo, L.; Leng, M.; Luo, K. The MicroRNA390/TRANS-ACTING SHORT INTERFER-ING RNA3 Module Mediates Lateral Root Growth under Salt Stress via the Auxin Pathway1. Plant Physiol. (Bethesda) 2018, 177, 775–791. [Google Scholar] [CrossRef] [PubMed]
- Okushima, Y.; Fukaki, H.; Onoda, M.; Theologis, A.; Tasaka, M. ARF7 and ARF19 Regulate Lateral Root Formation via Direct Activation of LBD/ASL Genes in Arabidopsis. Plant Cell 2007, 19, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Stothard, P. The Sequence Manipulation Suite: JavaScript Programs for Analyzing and Formatting Protein and DNA Sequences. Biotechniques 2000, 28, 1102–1104. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for es-timating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, B.; Gao, S.; Lercher, M.J.; Hu, S.; Chen, W.H. Evolview v3: A webserver for visualization, annotation, and management of phylogenetic trees. Nucleic Acids Res. 2019, 47, W270–W275. [Google Scholar] [CrossRef] [PubMed]
- Timothy, L.; Bailey, J.J.C.E. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Rombauts, S.; Dehais, P.; Van Montagu, M.; Rouze, P. PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res. 1999, 27, 295–296. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An in-formation aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A Toolkit Incorporating Gamma-Series Methods and Sliding Window Strategies. Genom. Proteom. Bioinform. 2010, 1, 77–80. [Google Scholar] [CrossRef]
- Fang, X.; Wang, M.; Zhou, X.; Wang, H.; Wang, H.; Xiao, H. Effects of growth years on ginsenoside biosynthesis of wild ginseng and cultivated ginseng. BMC Genom. 2022, 23, 325. [Google Scholar] [CrossRef]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yan, M.; Yan, Y.; Wang, P.; Wang, Y.; Piao, X.; Di, P.; Yang, D.-C. Genome-Wide Identification and Expression Analysis of Auxin Response Factor (ARF) Gene Family in Panax ginseng Indicates Its Possible Roles in Root Development. Plants 2023, 12, 3943. https://doi.org/10.3390/plants12233943
Yan M, Yan Y, Wang P, Wang Y, Piao X, Di P, Yang D-C. Genome-Wide Identification and Expression Analysis of Auxin Response Factor (ARF) Gene Family in Panax ginseng Indicates Its Possible Roles in Root Development. Plants. 2023; 12(23):3943. https://doi.org/10.3390/plants12233943
Chicago/Turabian StyleYan, Min, Yan Yan, Ping Wang, Yingping Wang, Xiangmin Piao, Peng Di, and Deok-Chun Yang. 2023. "Genome-Wide Identification and Expression Analysis of Auxin Response Factor (ARF) Gene Family in Panax ginseng Indicates Its Possible Roles in Root Development" Plants 12, no. 23: 3943. https://doi.org/10.3390/plants12233943






