A 14-3-3 Protein-Encoding Gene, BdGF14g, Confers Better Drought Tolerance by Regulating ABA Biosynthesis and Signaling
Abstract
1. Introduction
2. Results
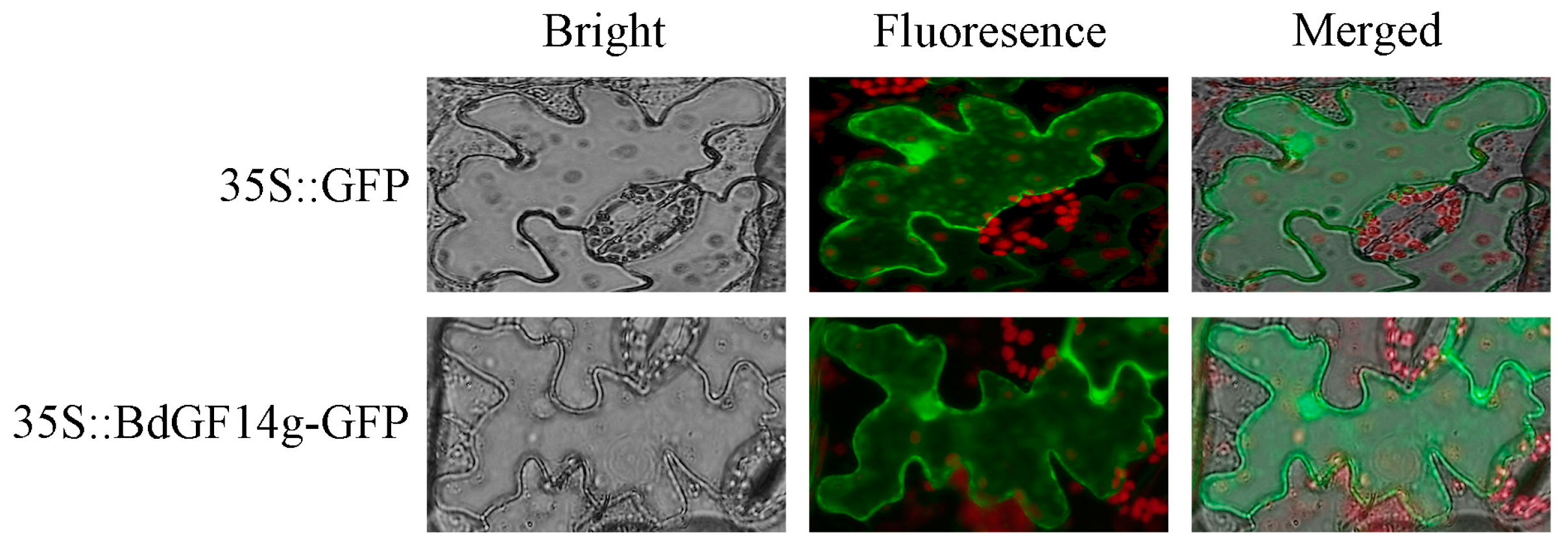
2.1. BdGF14g Is Located in the Cytoplasm and Nucleus
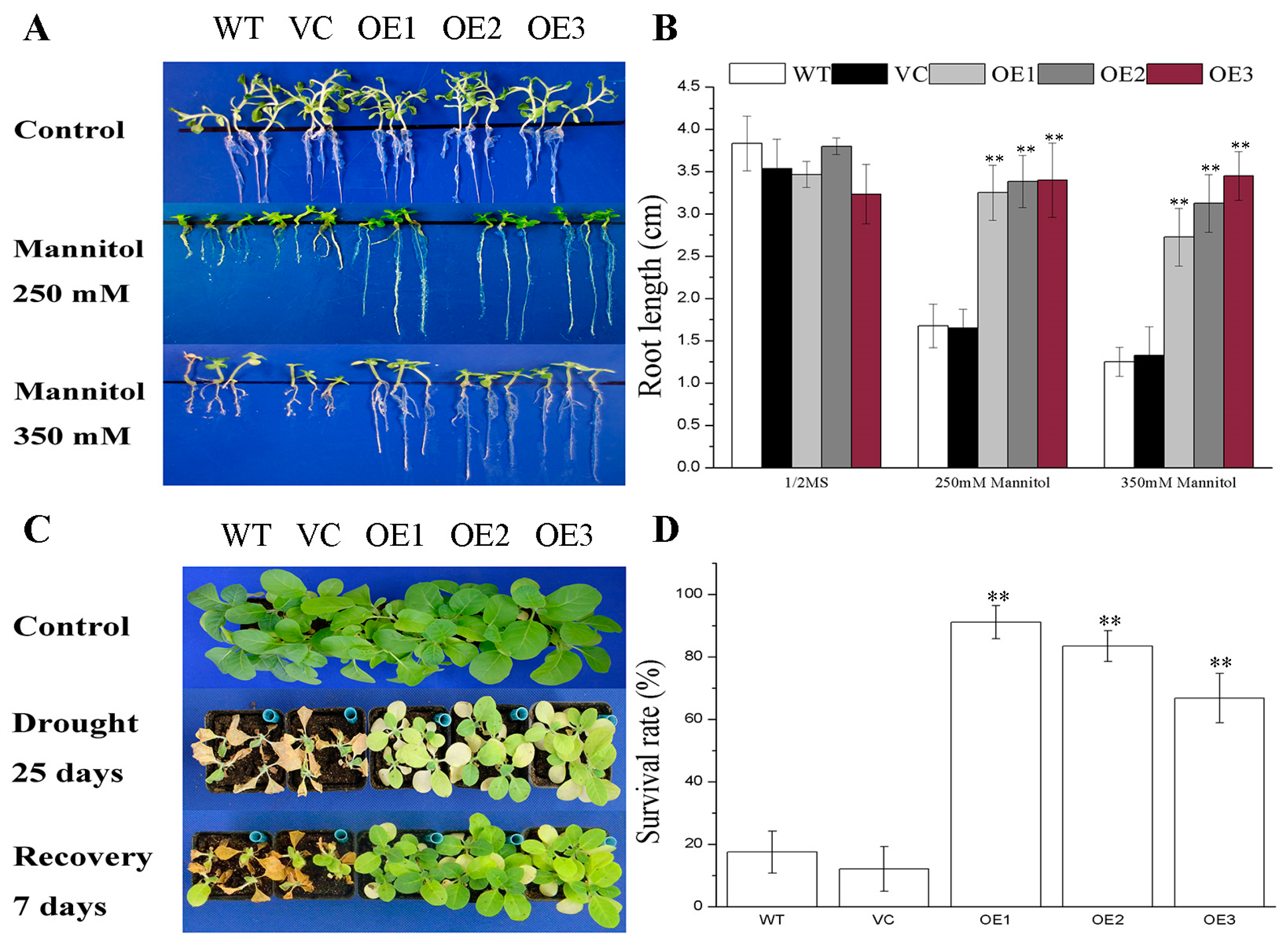
2.2. BdGF14g Improves Drought Tolerance at the Young Seedling Stage
2.3. BdGF14g Improves Drought Tolerance at the Mature Seedling Stage
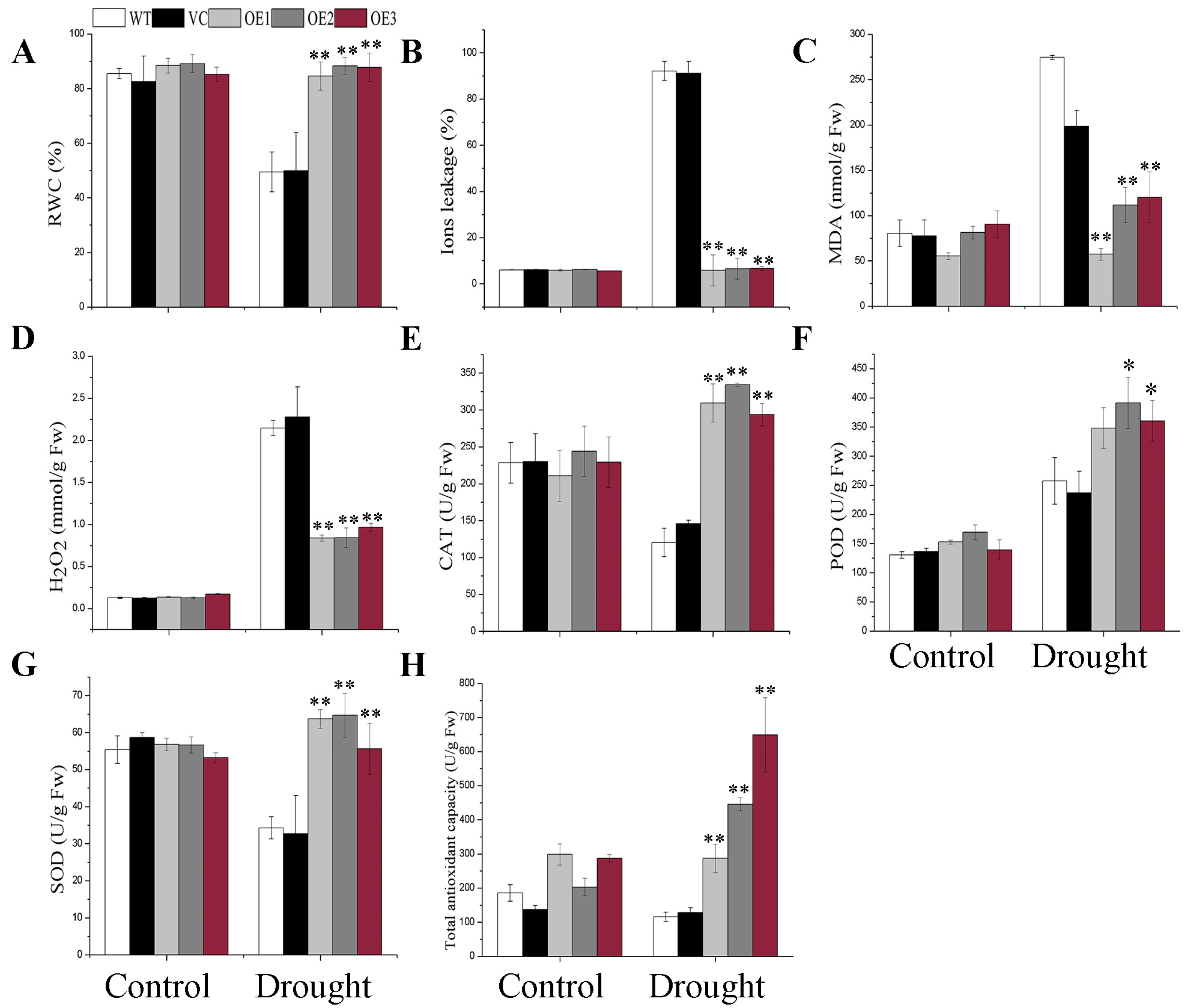
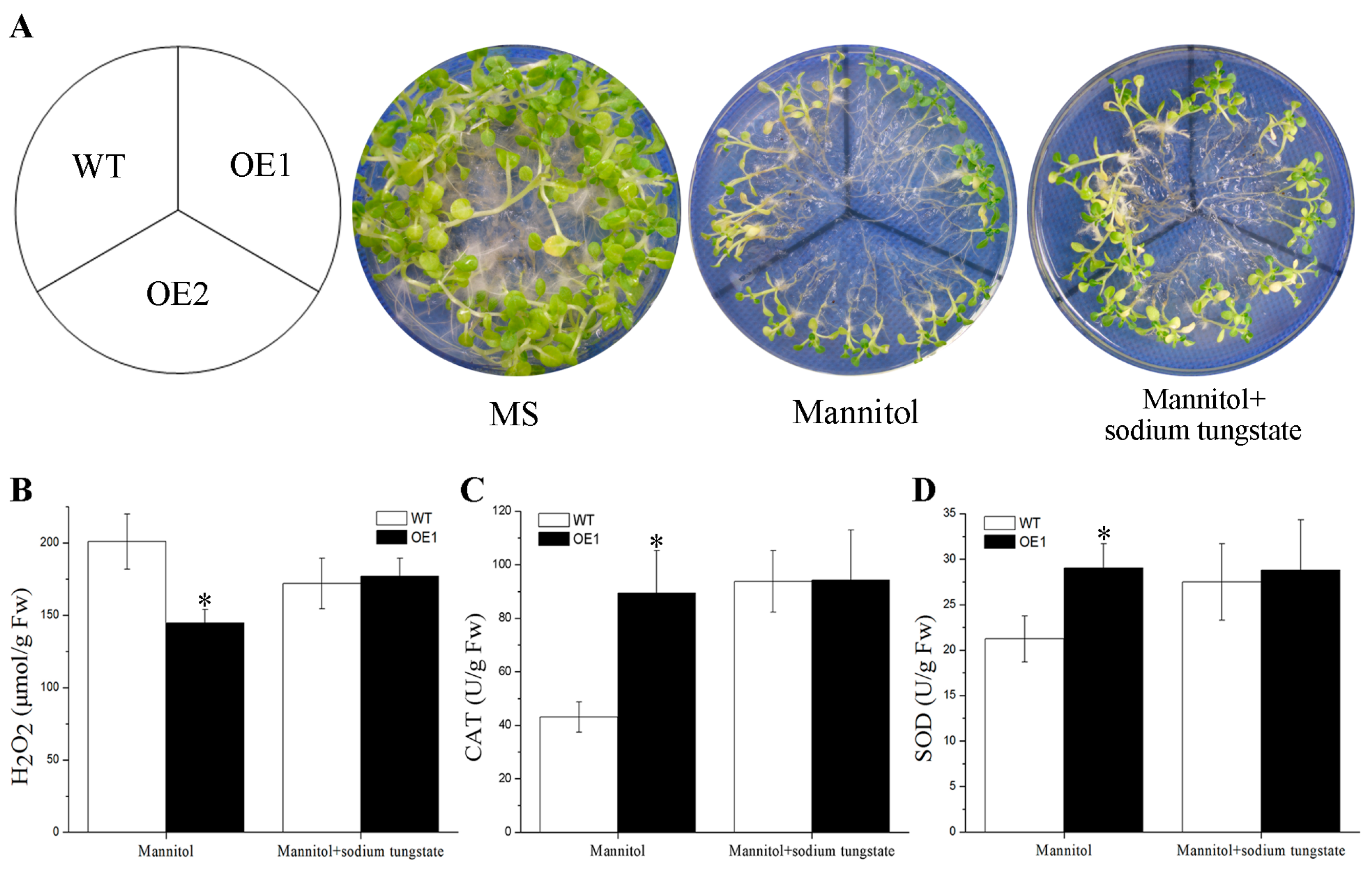
2.4. BdGF14g Increases ROS-Scavenging Ability under Water Deficiency
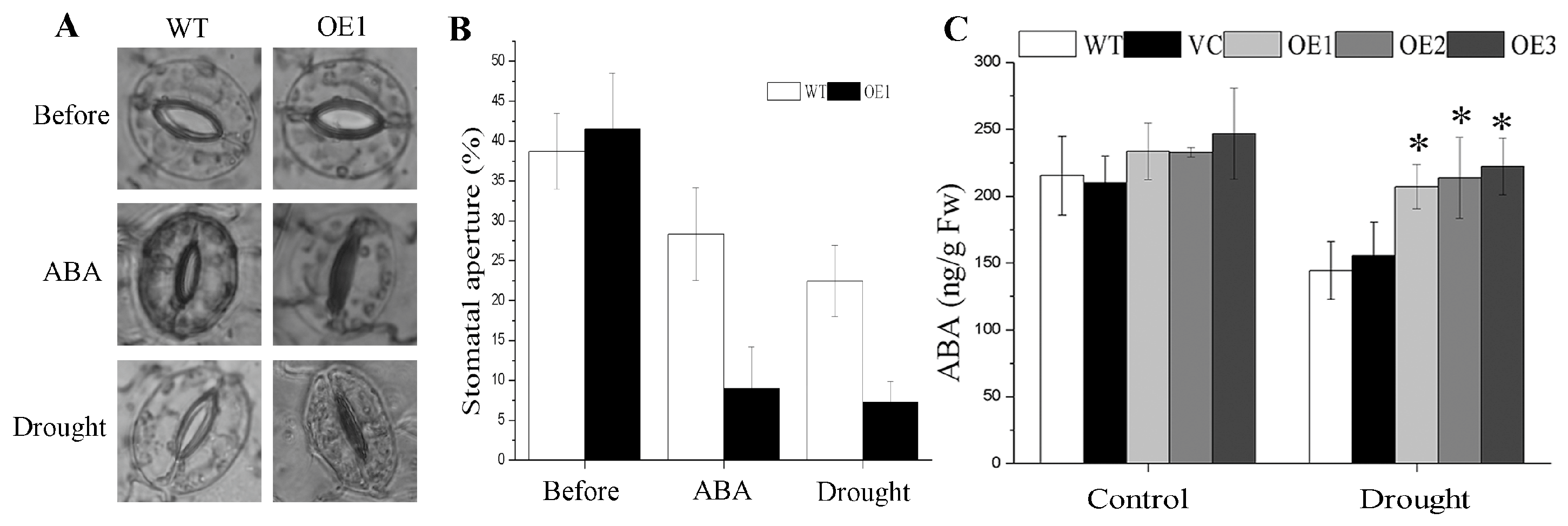
2.5. BdGF14g Overexpression Enhances Stomatal Closure under Water Deficiency and ABA Treatments
2.6. Drought Tolerance Conferred by BdGF14g Involving ABA in Transgenic Tobacco Plants
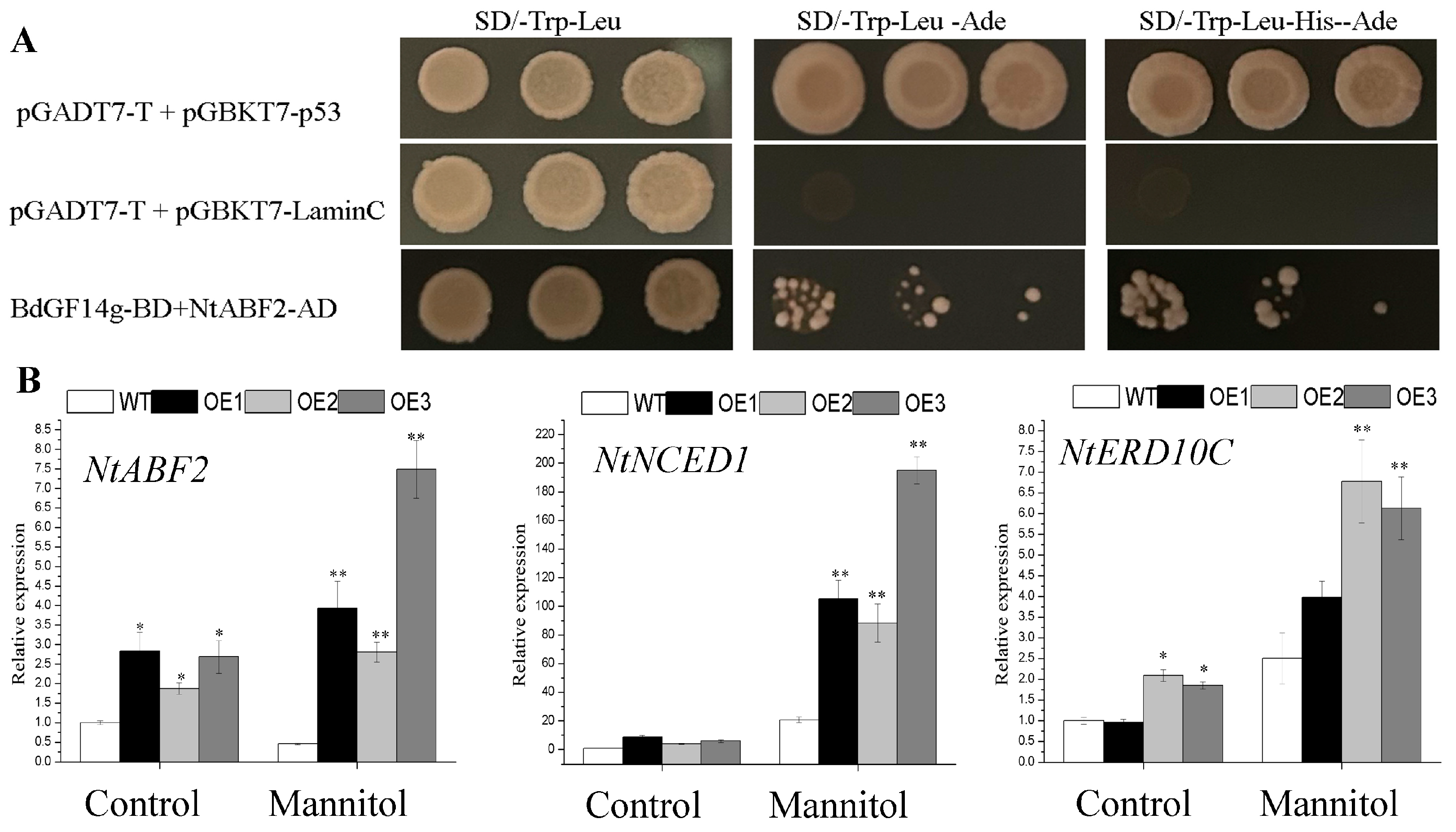
2.7. BdGF14g Interacts with NtABF2 and Increases the Expression of ABA-Related Gene in Transgenic Tobacco Plants
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Stress Treatments
4.2. Cloning and Subcellular Localization Analyses of BdGF14g
4.3. Tobacco Transformation and Stress Tolerance Analyses
4.4. Measurements of Aridity-Responsive Physiological Parameters
4.5. Stomatal Closure Assay
4.6. Y2H Assays
4.7. qRT-PCR Assays
4.8. Statistical Analyses
5. Conclusions
6. Patents
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Rosenquist, M.; Alsterfjord, M.; Larsson, C.; Sommarin, M. Data mining the Arabidopsis genome reveals fifteen 14-3-3 genes. Expression is demonstrated for two out of five novel genes. Plant Physiol. 2001, 127, 142–149. [Google Scholar] [CrossRef]
- Sehnke, P.C.; DeLille, J.M.; Ferl, R.J. Consummating signal transduction: The role of 14-3-3 proteins in the completion of signal-induced transitions in protein activity. Plant Cell 2002, 14S, S339–S354. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Tong, T.; Li, W.; Huang, Z.H.; Chen, G.; Zeng, F.R.; Riaz, A.; Amoanimaa-Dede, H.; Pan, R.; Zhang, W.Y.; et al. Molecular evolution of plant 14-3-3 proteins and function of Hv14-3-3A in stomatal regulation and drought tolerance. Plant Cell Physiol. 2023, 63, 1857–1872. [Google Scholar] [CrossRef] [PubMed]
- Mikhaylova, Y.V.; Puzanskiy, R.K.; Shishova, M.F. Evolution of 14-3-3 Proteins in Angiosperm Plants: Recurring Gene Duplication and Loss. Plants 2021, 10, 2724. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.J.; Sehnke, P.C.; Ferl, R.J. The 14-3-3 proteins: Cellular regulators of plant metabolism. Trends Plant Sci. 1999, 4, 367–371. [Google Scholar] [CrossRef] [PubMed]
- Chevalier, D.; Morris, E.R.; Walker, J.C. 14-3-3 and FHA domains mediate phosphoprotein interactions. Annu. Rev. Plant Biol. 2009, 60, 67–91. [Google Scholar] [CrossRef]
- Fiorillo, A.; Parmagnani, A.S.; Visconti, S.; Mannino, G.; Camoni, L.; Maffei, M.E. 14-3-3 proteins and the plasma membrane H+-ATPase are involved in maize (Zea mays) magnetic induction. Plants 2023, 12, 2887. [Google Scholar] [CrossRef] [PubMed]
- MacKintosh, C.; Meek, S.E. Regulation of plant NR activity by reversible phosphorylation, 14-3-3 proteins and proteolysis. Cell. Mol. Life Sci. 2001, 58, 205–214. [Google Scholar] [CrossRef]
- Kanczewska, J.; Marco, S.; Vandermeeren, C.; Maudoux, O.; Rigaud, J.L.; Boutry, M. Activation of the plant plasma membrane H+-ATPase by phosphorylation and binding of 14-3-3 proteins converts a dimer into a hexamer. Proc. Natl. Acad. Sci. USA 2005, 102, 11675–11680. [Google Scholar] [CrossRef]
- Cai, J.S.; Cai, W.W.; Huang, X.Y.; Yang, S.; Wen, J.Y.; Xia, X.Q.; Yang, F.; Shi, Y.Y.; Guan, D.Y.; He, S.L. Ca14-3-3 interacts with CaWRKY58 to positively modulate pepper response to low-phosphorus starvation. Front. Plant Sci. 2021, 11, 607878. [Google Scholar] [CrossRef]
- Wang, K.; Xu, F.Y.; Yuan, W.; Zhang, D.P.; Liu, J.P.; Sun, L.Y.; Cui, L.Y.; Zhang, J.H.; Xu, W.F. Rice G protein gamma subunit qPE9-1 modulates root elongation for phosphorus uptake by involving 14-3-3 protein OsGF14b and plasma membrane H+-ATPase. Plant J. 2021, 107, 1603–1615. [Google Scholar] [CrossRef] [PubMed]
- Taoka, K.; Ohki, I.; Tsuji, H.; Furuita, K.; Hayashi, K.; Yanase, T.; Yamaguchi, M.; Nakashima, C.; Purwestri, Y.A.; Tamaki, S.; et al. 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 2011, 476, 332–335. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, S.; Waksman, T.; Henderson, L.; Paliogianni, D.; Lutkemeyer, M.; Suetsugu, N.; Christie, J.M. Regulation of plant phototropic growth by NPH3/RPT2-like substrate phosphorylation and 14-3-3 binding. Nat. Commun. 2021, 12, 6129. [Google Scholar] [CrossRef] [PubMed]
- Song, P.Y.; Yang, Z.D.; Guo, C.; Han, R.; Wang, H.C.; Dong, J.; Kang, D.M.; Guo, Y.; Yang, S.H.; Li, J.G. 14-3-3 proteins regulate photomorphogenesis by facilitating light-induced degradation of PIF3. New Phytol. 2023, 237, 140–159. [Google Scholar] [CrossRef]
- Peng, Q.; Zhu, C.M.; Liu, T.; Zhang, S.; Feng, S.J.; Wu, C.Y. Phosphorylation of OsFD1 by OsCIPK3 promotes the formation of RFT1-containing florigen activation complex for long-day flowering in rice. Mol. Plant 2021, 14, 1135–1148. [Google Scholar] [CrossRef] [PubMed]
- Elena-Real, C.A.; Gonzalez-Arzola, K.; Perez-Mejias, G.; Diaz-Quintana, A.; Velazquez-Campoy, A.; Desvoyes, B.; Gutierrez, C.; De la Rosa, M.A.; Diaz-Moreno, I. Proposed mechanism for regulation of H2O2-induced programmed cell death in plants by binding of cytochrome c to 14-3-3 proteins. Plant J. 2021, 106, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Qi, H.; Lei, X.; Wang, Y.; Yu, S.; Liu, T.; Zhou, S.K.; Chen, J.Y.; Chen, Q.F.; Qiu, R.L.; Jiang, L.W.; et al. 14-3-3 proteins contribute to autophagy by modulating SINAT-mediated degradation of ATG13. Plant Cell 2022, 34, 4857–4876. [Google Scholar] [CrossRef]
- Chen, F.; Li, Q.; Sun, L.; He, Z. The rice 14-3-3 gene family and its involvement in responses to biotic and abiotic stress. DNA Res. 2006, 13, 53–63. [Google Scholar] [CrossRef]
- Deb, S.; Ghosh, P.; Patel, H.K.; Sonti, R.V. Interaction of the Xanthomonas effectors XopQ and XopX results in induction of rice immune responses. Plant J. 2020, 104, 332–350. [Google Scholar] [CrossRef]
- Ma, Y.M.; Yang, J.Y.; Dong, J.F.; Zhang, S.H.; Yang, W.; Zhao, J.L.; Yang, T.F.; Chen, L.; Zhou, L.; Wang, J.; et al. Overexpression of OsGF14f enhances quantitative leaf blast and bacterial blight resistance in rice. Int. J. Mol. Sci. 2022, 23, 7440. [Google Scholar] [CrossRef]
- Zhou, H.; Lin, H.; Chen, S.; Becker, K.; Yang, Y.; Zhao, J.; Kudla, J.; Schumaker, K.S.; Guo, Y. Inhibition of the Arabidopsis salt overly sensitive pathway by 14-3-3 proteins. Plant Cell 2014, 26, 1166–1182. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.Q.; He, C.X.; Wang, J.; Mao, Z.H.; Scott, A.; Randy, D.; Zhang, H. Overexpression of the Arabidopsis 14-3-3 protein GF14λ in cotton leads to a “stay-green” phenotype and improves stress tolerance under moderate drought conditions. Plant Cell Physiol. 2004, 45, 1007–1014. [Google Scholar] [CrossRef] [PubMed]
- Campo, S.; Peris-Peris, C.; Montesinos, L.; Penas, G.; Messeguer, J.; San, S.B. Expression of the maize ZmGF14-6 gene in rice confers tolerance to drought stress while enhancing susceptibility to pathogen infection. J. Exp. Bot. 2012, 63, 983–999. [Google Scholar] [CrossRef]
- He, Y.; Wu, J.; Lv, B.; Li, J.; Gao, Z.; Xu, W.; Baluska, F.; Shi, W.; Shaw, P.C.; Zhang, J. Involvement of 14-3-3 protein GRF9 in root growth and response under polyethylene glycol-induced water stress. J. Exp. Bot. 2015, 66, 2271–2281. [Google Scholar] [CrossRef]
- Ma, Y.; Wu, Z.; Dong, J.; Zhang, S.; Zhao, J.; Yang, T.; Yang, W.; Zhou, L.; Wang, J.; Chen, J.; et al. The 14-3-3 protein OsGF14f interacts with OsbZIP23 and enhances its activity to confer osmotic stress tolerance in rice. Plant Cell 2023, 35, 4173–4189. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Shi, Y.; Jiang, Q.; Li, H.; Fan, W.; Feng, Y.; Li, L.; Liu, B.; Lin, F.; Jing, W.; et al. A 14-3-3 protein positively regulates rice salt tolerance by stabilizing phospholipase C1. Plant Cell Environ. 2023, 46, 1232–1248. [Google Scholar] [CrossRef]
- Liu, J.P.; Sun, X.J.; Liao, W.C.; Zhang, J.H.; Liang, J.S.; Xu, W.F. Involvement of OsGF14b adaptation in the drought resistance of rice plants. Rice 2019, 12, 82. [Google Scholar] [CrossRef]
- Sharma, E.; Bhatnagar, A.; Bhaskar, A.; Majee, S.M.; Kieffer, M.; Kepinski, S.; Khurana, P.; Khurana, J.P. Stress-induced F-Box protein-coding gene OsFBX257 modulates drought stress adaptations and ABA responses in rice. Plant Cell Environ. 2023, 46, 1207–1231. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, H.; Zhou, S.; He, Y.; Luo, Q.; Zhang, F.; Qiu, D.; Feng, J.; Wei, Q.; Chen, L.; et al. Expression of TaGF14b, a 14-3-3 adaptor protein gene from wheat, enhances drought and salt tolerance in transgenic tobacco. Planta 2018, 248, 117–137. [Google Scholar] [CrossRef]
- He, Y.; Zhang, Y.; Chen, L.; Wu, C.; Luo, Q.; Zhang, F.; Wei, Q.; Li, K.; Chang, J.; Yang, G.; et al. A member of the 14-3-3 gene family in Brachypodium distachyon, BdGF14d, confers salt tolerance in transgenic tobacco plants. Front. Plant Sci. 2017, 8, 340. [Google Scholar] [CrossRef]
- Chang, H.C.; Tsai, M.C.; Wu, S.S.; Chang, I.F. Regulation of ABI5 expression by ABF3 during salt stress responses in Arabidopsis thaliana. Bot. Stud. 2019, 60, 16. [Google Scholar] [CrossRef] [PubMed]
- Schoonheim, P.J.; Sinnige, M.P.; Casaretto, J.A.; Veiga, H.; Bunney, T.D.; Quatrano, R.S.; de Boer, A.H. 14-3-3 adaptor proteins are intermediates in ABA signal transduction during barley seed germination. Plant J. 2007, 49, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Schoonheim Peter, J.; Costa Pereira Daniel, D.; De Boer Albertus, H. Dual role for 14-3-3 proteins and ABF transcription factors in gibberellic acid and abscisic acid signalling in barley Hordeum vulgare aleurone cells. Plant Cell Environ. 2009, 32, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Sun, M.; Jia, B.; Chen, C.; Qin, Z.; Yang, K.; Shen, Y.; Meiping, Z.; Mingyang, C.; Zhu, Y. A 14-3-3 family protein from wild soybean Glycine Soja regulates ABA sensitivity in Arabidopsis. PLoS ONE 2015, 10, e0146163. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jing, S.L.; Sun, X.M.; Yu, L.; Wang, E.S.; Cheng, Z.N.; Liu, H.M.; Jiang, P.; Qin, J.; Begum, S.; Song, B.T. Transcription factor StABI5-like 1 binding to the FLOWERING LOCUS T homologs promotes early maturity in potato. Plant Physiol. 2022, 189, 1677–1693. [Google Scholar] [CrossRef]
- Zhang, Q.; Yuan, W.; Wang, Q.; Cao, Y.; Xu, F.; Dodd, I.C.; Xu, W. ABA regulation of root growth during soil drying and recovery can involve auxin response. Plant Cell Environ. 2022, 45, 871–883. [Google Scholar] [CrossRef]
- Draper, J.; Mur, L.A.J.; Jenkins, G.; Ghosh-Biswas, G.C.; Bablak, P.; Hasterok, R.; Routledge, A.P.M. Brachypodium distachyon. A new model system for functional genomics in grasses. Plant Physiol. 2001, 127, 1539–1555. [Google Scholar] [CrossRef]
- Fujita, Y.; Fujita, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 2011, 124, 509–525. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, J.; Gong, Z.; Zhu, J.K. Abiotic stress responses in plants. Nat. Rev. Genet. 2022, 23, 104–119. [Google Scholar] [CrossRef]
- Waadt, R.; Seller, C.A.; Hsu, P.K.; Takahashi, Y.; Munemasa, S.; Schroeder, J.I. Plant hormone regulation of abiotic stress responses. Nat. Rev. Mol. Cell Biol. 2022, 23, 680–694. [Google Scholar] [CrossRef]
- Reuter, L.; Schmidt, T.; Manishankar, P.; Throm, C.; Keicher, J.; Bock, A.; Droste-Borel, I.; Oecking, C. Light-triggered and phosphorylation-dependent 14-3-3 association with NON-PHOTOTROPIC HYPOCOTYL 3 is required for hypocotyl phototropism. Nat. Commun. 2021, 12, 6128. [Google Scholar] [CrossRef]
- Fukayama, H.; Miyagawa, F.; Shibatani, N.; Koudou, A.; Sasayama, D.; Hatanaka, T.; Azuma, T.; Yamauchi, Y.; Matsuoka, D.; Morita, R. CO2-responsive CCT protein interacts with 14-3-3 proteins and controls the expression of starch synthesis-related genes. Plant Cell Environ. 2021, 44, 2480–2493. [Google Scholar] [CrossRef]
- Liu, S.; Chen, S.; Zhou, Y.; Shen, Y.; Qin, Z.; Wu, L. VERNALIZATION1 represses FLOWERING PROMOTING FACTOR1-LIKE1 in leaves for timely flowering in Brachypodium distachyon. Plant Cell 2023, 35, 3697–3711. [Google Scholar] [CrossRef]
- Liu, Z.; Jia, Y.; Ding, Y.; Shi, Y.; Li, Z.; Guo, Y.; Gong, Z.; Yang, S. Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to Fine-Tune CBF signaling during cold response. Mol. Cell 2017, 66, 117–128.e115. [Google Scholar] [CrossRef]
- Cao, H.; Xu, Y.; Yuan, L.; Bian, Y.; Wang, L.; Zhen, S.; Hu, Y.; Yan, Y. Molecular characterization of the 14-3-3 gene family in Brachypodium distachyon L. reveals high evolutionary conservation and diverse responses to abiotic stresses. Front. Plant Sci. 2016, 7, 1099. [Google Scholar] [CrossRef]
- Yang, L.; You, J.; Wang, Y.; Li, J.; Quan, W.; Yin, M.; Wang, Q.; Chan, Z. Systematic analysis of the G-box Factor 14-3-3 gene family and functional characterization of GF14a in Brachypodium distachyon. Plant Physiol. Bioch. 2017, 117, 1–11. [Google Scholar] [CrossRef]
- Ren, J.; Zhang, P.; Dai, Y.; Liu, X.; Lu, S.; Guo, L.; Gou, H.; Mao, J. Evolution of the 14-3-3 gene family in monocotyledons and dicotyledons and validation of MdGRF13 function in transgenic Arabidopsis thaliana. Plant Cell Rep. 2023, 42, 1345–1364. [Google Scholar] [CrossRef]
- Mittler, R.; Zandalinas, S.I.; Fichman, Y.; Van Breusegem, F. Reactive oxygen species signalling in plant stress responses. Nat. Rev. Mol. Cell Biol. 2022, 23, 663–679. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Lv, J.; Li, K.; Zhai, L.; Wu, Y.; Wu, T.; Zhang, X.; Han, Z.; Wang, Y. MdGRF11-MdARF19-2 module acts as a positive regulator of drought resistance in apple rootstock. Plant Sci. 2023, 335, 111782. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Lu, G.; Hao, Y.; Guo, H.; Guo, Y.; Zhao, J.; Cheng, H. ABP9, a maize bZIP transcription factor, enhances tolerance to salt and drought in transgenic cotton. Planta 2017, 246, 453–469. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Luo, X.; Sun, M.; Chen, C.; Ding, X.; Wang, X.; Yang, S.; Yu, Q.; Jia, B.; Ji, W.; et al. A Glycine soja 14-3-3 protein GsGF14o participates in stomatal and root hair development and drought tolerance in Arabidopsis thaliana. Plant Cell Physiol. 2014, 55, 99–118. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Li, G.J.; Bressan, R.A.; Song, C.P.; Zhu, J.K.; Zhao, Y. Abscisic acid dynamics, signaling, and functions in plants. J. Integr. Plant Biol. 2020, 62, 25–54. [Google Scholar] [CrossRef]
- Bi, C.; Ma, Y.; Wu, Z.; Yu, Y.; Liang, S.; Lu, K.; Wang, X.F. Arabidopsis ABI5 plays a role in regulating ROS homeostasis by activating CATALASE 1 transcription in seed germination. Plant Mol. Biol. 2017, 94, 197–213. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Zhang, J. Effect of abscisic acid on active oxygen species, antioxidative defence system and oxidative damage in leaves of maize seedlings. Plant Cell Physiol. 2001, 42, 1265–1273. [Google Scholar] [CrossRef] [PubMed]
- Zong, W.; Tang, N.; Yang, J.; Peng, L.; Ma, S.; Xu, Y.; Li, G.; Xiong, L. Feedback Regulation of ABA Signaling and Biosynthesis by a bZIP Transcription Factor Targets Drought-Resistance-Related Genes. Plant Physiol. 2016, 171, 2810–2825. [Google Scholar] [CrossRef]
- Shinozaki, K.; Yamaguchi-Shinozaki, K.; Seki, M. Regulatory network of gene expression in the drought and cold stress responses. Curr. Opin. Plant Biol. 2003, 6, 410–417. [Google Scholar] [CrossRef]
- Fujita, Y.; Fujita, M.; Satoh, R.; Maruyama, K.; Parvez, M.M.; Seki, M.; Hiratsu, K.; Ohme-Takagi, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AREB1 is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 2005, 17, 3470–3488. [Google Scholar] [CrossRef]
- Zhang, L.; Zhao, L.; Wang, L.; Liu, X.; Yu, Z.; Liu, J.; Wu, W.; Ding, L.; Xia, C.; Zhang, L.; et al. TabZIP60 is involved in the regulation of ABA synthesis-mediated salt tolerance through interacting with TaCDPK30 in wheat (Triticum aestivum L.). Planta 2023, 257, 107. [Google Scholar] [CrossRef]
- Song, J.; Sun, P.; Kong, W.; Xie, Z.; Li, C.; Liu, J.H. SnRK2.4-mediated phosphorylation of ABF2 regulates ARGININE DECARBOXYLASE expression and putrescine accumulation under drought stress. New Phytol. 2023, 238, 216–236. [Google Scholar] [CrossRef]
- Collin, A.; Daszkowska-Golec, A.; Szarejko, I. Updates on the Role of ABSCISIC ACID INSENSITIVE 5 (ABI5) and ABSCISIC ACID-RESPONSIVE ELEMENT BINDING FACTORs (ABFs) in ABA Signaling in Different Developmental Stages in Plants. Cells 2021, 10, 1996. [Google Scholar] [CrossRef]
- Shao, W.; Chen, W.; Zhu, X.; Zhou, X.; Jin, Y.; Zhan, C.; Liu, G.; Liu, X.; Ma, D.; Qiao, Y. Genome-wide identification and characterization of wheat 14-3-3 genes unravels the role of TaGRF6-A in salt stress tolerance by binding MYB transcription factor. Int. J. Mol. Sci. 2021, 22, 1904. [Google Scholar] [CrossRef] [PubMed]
- Horsch, R.B.; Fry, J.E.; Hoffmann, N.L.; Eichholtz, D.; Rogers, S.C.; Fraley, R.T. A simple and general method for transferring genes into plants. Science 1985, 227, 1229–1231. [Google Scholar] [CrossRef] [PubMed]
- Grondin, A.; Rodrigues, O.; Verdoucq, L.; Merlot, S.; Leonhardt, N. Aquaporins Contribute to ABA-Triggered Stomatal Closure through OST1-Mediated Phosphorylation. Plant Cell 2015, 27, 1945–1954. [Google Scholar] [CrossRef] [PubMed]
- Folter, S.; Immink, R.G. Yeast protein-protein interaction assays and screens. Methods Mol. Biol. 2011, 754, 145–165. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; He, Y.; Zhao, H.; Zhang, Y.; Yang, J.; Ou, X.; Zhang, J.; Zhu, Q. A 14-3-3 Protein-Encoding Gene, BdGF14g, Confers Better Drought Tolerance by Regulating ABA Biosynthesis and Signaling. Plants 2023, 12, 3975. https://doi.org/10.3390/plants12233975
Zhang Y, He Y, Zhao H, Zhang Y, Yang J, Ou X, Zhang J, Zhu Q. A 14-3-3 Protein-Encoding Gene, BdGF14g, Confers Better Drought Tolerance by Regulating ABA Biosynthesis and Signaling. Plants. 2023; 12(23):3975. https://doi.org/10.3390/plants12233975
Chicago/Turabian StyleZhang, Yang, Yuan He, Hongyan Zhao, Yan Zhang, Jing Yang, Xingqi Ou, Jinlong Zhang, and Qidi Zhu. 2023. "A 14-3-3 Protein-Encoding Gene, BdGF14g, Confers Better Drought Tolerance by Regulating ABA Biosynthesis and Signaling" Plants 12, no. 23: 3975. https://doi.org/10.3390/plants12233975
APA StyleZhang, Y., He, Y., Zhao, H., Zhang, Y., Yang, J., Ou, X., Zhang, J., & Zhu, Q. (2023). A 14-3-3 Protein-Encoding Gene, BdGF14g, Confers Better Drought Tolerance by Regulating ABA Biosynthesis and Signaling. Plants, 12(23), 3975. https://doi.org/10.3390/plants12233975




