Plastomes of Garcinia mangostana L. and Comparative Analysis with Other Garcinia Species
Abstract
:1. Introduction
2. Results
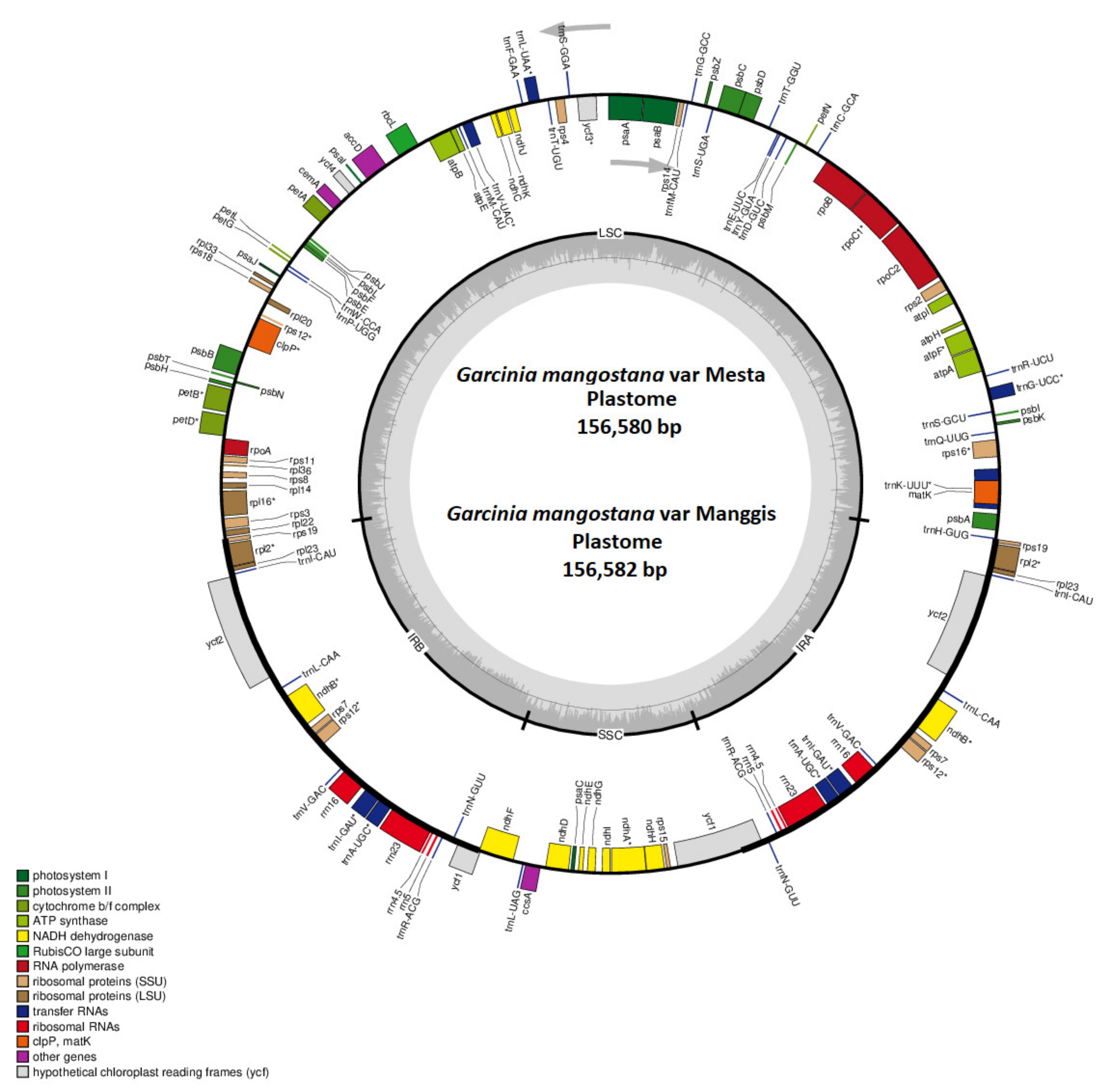
2.1. Characterization of the Mesta Plastome
2.2. Manggis Plastome Assembly
2.3. Plastome Feature Comparison
2.4. Codon Usage and Amino Acid Frequency
2.5. Simple Sequence Repeat (SSR) Analysis
2.6. Long Repeat Analysis
2.7. Contraction and Expansion of the Inverted Repeat Region
2.8. Comparative Plastome Analysis
2.9. Phylogenomic Analysis
3. Discussion
4. Materials and Methods
4.1. Mesta Plastome De Novo Genome Assembly
4.2. Manggis Plastome Assembly
4.3. Plastome Annotation
4.4. Open Reading Frame (ORF) Coordinate Adjustment
4.5. Identification of Simple-Sequence Repeats (SSRs)
4.6. Long Repeat Analysis
4.7. Codon Usage Analysis
4.8. Plastome Sequence Alignment and Comparative Analysis
4.9. Phylogenomic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mamat, S.F.; Azizan, K.A.; Baharum, S.N.; Noor, N.M.; Aizat, W.M. GC-MS and LC-MS analyses reveal the distribution of primary and secondary metabolites in mangosteen (Garcinia mangostana Linn.) fruit during ripening. Sci. Hortic. 2020, 262, 109004. [Google Scholar] [CrossRef]
- Aizat, W.M.; Jamil, I.N.; Ahmad-Hashim, F.H.; Noor, N.M. Recent updates on metabolite composition and medicinal benefits of mangosteen plant. PeerJ 2019, 7, e6324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sprecher, A. Étude sur la Semence et la Germination du Garcinia mangostana L.; Librairie Générale de l’Enseignement: Paris, France, 1919. [Google Scholar]
- Goh, H.-H.; Abu Bakar, S.; Kamal Azlan, N.D.; Zainal, Z.; Mohd Noor, N. Transcriptional reprogramming during Garcinia-type recalcitrant seed germination of Garcinia mangostana. Sci. Hortic. 2019, 257, 108727. [Google Scholar] [CrossRef]
- Chinawat, Y.; Subhadrabhanu, S. Phylogenetic relationship of Mangosteen and several wild relatives revealed by ITS Sequence data. J. Am. Soc. Hortic. Sci. 2004, 129, 368–373. [Google Scholar]
- Nazre, M.; Clyde, M.; Latiff, A. Phylogenetic relationships of locally cultivated Garcinia species with some wild relatives. Malays. Appl. Biol. 2007, 36, 31. [Google Scholar]
- Sweeney, P.W. Phylogeny and floral diversity in the genus Garcinia (Clusiaceae) and relatives. Int. J. Plant Sci. 2008, 169, 1288–1303. [Google Scholar] [CrossRef]
- Saleh, M. Taxonomic Revision and Molecular Studies of Garcinia Section Garcinia (Guttiferae); University of Edinburgh: Edinburgh, UK, 2006. [Google Scholar]
- Richards, A. Studies in Garcinia, dioecious tropical forest trees: The origin of the mangosteen (G. mangostana L.). Bot. J. Linn. Soc. 1990, 103, 301–308. [Google Scholar] [CrossRef]
- Nazre, M. New evidence on the origin of mangosteen (Garcinia mangostana L.) based on morphology and ITS sequence. Genet. Resour. Crop Evol. 2014, 61, 1147–1158. [Google Scholar] [CrossRef]
- Bi, Y.; Zhang, M.-F.; Xue, J.; Dong, R.; Du, Y.-P.; Zhang, X.-H. Chloroplast genomic resources for phylogeny and DNA barcoding: A case study on Fritillaria. Sci. Rep. 2018, 8, 1184. [Google Scholar] [CrossRef] [Green Version]
- Raspé, O. Inheritance of the Chloroplast Genome in Sorbus aucuparia L. (Rosaceae). J. Hered. 2001, 92, 507–509. [Google Scholar] [CrossRef] [Green Version]
- Cheon, K.-S.; Kim, K.-A.; Kwak, M.; Lee, B.; Yoo, K.-O. The complete chloroplast genome sequences of four Viola species (Violaceae) and comparative analyses with its congeneric species. PLoS ONE 2019, 14, e0214162. [Google Scholar] [CrossRef] [PubMed]
- Tonti-Filippini, J.; Nevill, P.G.; Dixon, K.; Small, I. What can we do with 1000 plastid genomes? Plant J. 2017, 90, 808–818. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jo, S.; Kim, H.-W.; Kim, Y.-K.; Sohn, J.-Y.; Cheon, S.-H.; Kim, K.-J. The complete plastome of tropical fruit Garcinia mangostana (Clusiaceae). Mitochondrial DNA Part B 2017, 2, 722–724. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abu Bakar, S.; Sampathrajan, S.; Loke, K.K.; Goh, H.H.; Mohd Noor, N. DNA-seq analysis of Garcinia mangostana. Genom. Data 2016, 7, 62–63. [Google Scholar] [CrossRef] [Green Version]
- Midin, M.R.; Loke, K.K.; Madon, M.; Nordin, M.S.; Goh, H.H.; Mohd Noor, N. SMRT sequencing data for Garcinia mangostana L. variety Mesta. Genom. Data 2017, 12, 134–135. [Google Scholar] [CrossRef]
- Abu Bakar, S.; Kumar, S.; Loke, K.K.; Goh, H.H.; Noor, N.M. DNA shotgun sequencing analysis of Garcinia mangostana L. variety Mesta. Genom. Data 2017, 12, 118–119. [Google Scholar] [CrossRef]
- Wee, C.-C.; Muhammad, N.A.N.; Subbiah, V.K.; Arita, M.; Nakamura, Y.; Goh, H.-H. Mitochondrial Genome of Garcinia mangostana L. variety Mesta. Sci. Rep. 2022, 12, 9480. [Google Scholar] [CrossRef]
- Wang, Y.; Zhao, B.; Lu, Z.; Shi, Y.; Li, J. The complete chloroplast genome provides insight into the polymorphism and adaptive evolution of Garcinia paucinervis. Biotechnol. Biotechnol. Equip. 2021, 35, 377–391. [Google Scholar] [CrossRef]
- Krumsiek, J.; Arnold, R.; Rattei, T. Gepard: A rapid and sensitive tool for creating dotplots on genome scale. Bioinformatics 2007, 23, 1026–1028. [Google Scholar] [CrossRef] [Green Version]
- Redwan, R.; Saidin, A.; Kumar, S. Complete chloroplast genome sequence of MD-2 pineapple and its comparative analysis among nine other plants from the subclass Commelinidae. BMC Plant Biol. 2015, 15, 196. [Google Scholar] [CrossRef] [Green Version]
- Gao, C.; Wu, C.; Zhang, Q.; Zhao, X.; Wu, M.; Chen, R.; Zhao, Y.; Li, Z. Characterization of chloroplast genomes from two Salvia medicinal plants and gene transfer among their mitochondrial and chloroplast genomes. Front. Genet. 2020, 11, 574962. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yang, Z.; Huang, S.; An, W.; Li, J.; Zheng, X. Comprehensive analysis of Rhodomyrtus tomentosa chloroplast genome. Plants 2019, 8, 89. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Wang, Z.; Wang, H.; Su, Y.; Wang, T. Patterns and rates of plastid rps12 gene evolution inferred in a phylogenetic context using plastomic data of ferns. Sci. Rep. 2020, 10, 9394. [Google Scholar] [CrossRef] [PubMed]
- Asaf, S.; Khan, A.L.; Khan, A.; Khan, G.; Lee, I.-J.; Al-Harrasi, A. Expanded inverted repeat region with large scale inversion in the first complete plastid genome sequence of Plantago ovata. Sci. Rep. 2020, 10, 3881. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, X.; Li, Q.; Li, Y.; Qian, J.; Han, J. Chloroplast genome of Aconitum barbatum var. puberulum (Ranunculaceae) derived from CCS reads using the PacBio RS platform. Front. Plant Sci. 2015, 6, 42. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, D.-M.; Zhao, C.-Y.; Liu, X.-F. Complete chloroplast genome sequences of Kaempferia galanga and Kaempferia elegans: Molecular structures and comparative analysis. Molecules 2019, 24, 474. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.; Xu, B.; Li, B.; Zhou, Q.; Wang, G.; Jiang, X.; Wang, C.; Xu, Z. Comparative analysis of codon usage patterns in chloroplast genomes of six Euphorbiaceae species. PeerJ 2020, 8, e8251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoch, B.; Maier, R.M.; Appel, K.; Igloi, G.L.; Kössel, H. Editing of a chloroplast mRNA by creation of an initiation codon. Nature 1991, 353, 178–180. [Google Scholar] [CrossRef]
- Kuroda, H.; Suzuki, H.; Kusumegi, T.; Hirose, T.; Yukawa, Y.; Sugiura, M. Translation of psbC mRNAs Starts from the Downstream GUG, not the Upstream AUG, and Requires the Extended Shine–Dalgarno Sequence in Tobacco Chloroplasts. Plant Cell Physiol. 2007, 48, 1374–1378. [Google Scholar] [CrossRef] [Green Version]
- Hirose, T.; Sugiura, M. Both RNA editing and RNA cleavage are required for translation of tobacco chloroplast ndhD mRNA: A possible regulatory mechanism for the expression of a chloroplast operon consisting of functionally unrelated genes. EMBO J. 1997, 16, 6804–6811. [Google Scholar] [CrossRef]
- Po, L.Q.; Zhong, X.Q. Codon usage in the chloroplast genome of rice (Oryza sativa L. ssp. japonica). Acta Agron. Sin. 2004, 30, 1220–1224. [Google Scholar]
- George, B.; Bhatt, B.S.; Awasthi, M.; George, B.; Singh, A.K. Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants. Curr. Genet. 2015, 61, 665–677. [Google Scholar] [CrossRef] [PubMed]
- Dong, F.; Lin, Z.; Lin, J.; Ming, R.; Zhang, W. Chloroplast genome of rambutan and comparative analyses in Sapindaceae. Plants 2021, 10, 283. [Google Scholar] [CrossRef] [PubMed]
- Xue, S.; Shi, T.; Luo, W.; Ni, X.; Iqbal, S.; Ni, Z.; Huang, X.; Yao, D.; Shen, Z.; Gao, Z. Comparative analysis of the complete chloroplast genome among Prunus mume, P. armeniaca, and P. salicina. Hortic. Res. 2019, 6, 89. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferrarini, M.; Moretto, M.; Ward, J.A.; Šurbanovski, N.; Stevanović, V.; Giongo, L.; Viola, R.; Cavalieri, D.; Velasco, R.; Cestaro, A. An evaluation of the PacBio RS platform for sequencing and de novo assembly of a chloroplast genome. BMC Genom. 2013, 14, 670. [Google Scholar] [CrossRef] [Green Version]
- Shearman, J.R.; Sonthirod, C.; Naktang, C.; Sangsrakru, D.; Yoocha, T.; Chatbanyong, R.; Vorakuldumrongchai, S.; Chusri, O.; Tangphatsornruang, S.; Pootakham, W. Assembly of the durian chloroplast genome using long PacBio reads. Sci. Rep. 2020, 10, 15980. [Google Scholar] [CrossRef]
- Fahrenkrog, A.M.; Matsumoto, G.; Toth, K.; Jokipii-Lukkari, S.; Salo, H.M.; Häggman, H.; Benevenuto, J.; Munoz, P. Chloroplast genome assemblies and comparative analyses of major Vaccinium berry crops. bioRxiv 2022, 481500. [Google Scholar] [CrossRef]
- Wu, Z.; Gui, S.; Quan, Z.; Pan, L.; Wang, S.; Ke, W.; Liang, D.; Ding, Y. A precise chloroplast genome of Nelumbo nucifera (Nelumbonaceae) evaluated with Sanger, Illumina MiSeq, and PacBio RS II sequencing platforms: Insight into the plastid evolution of basal eudicots. BMC Plant Biol. 2014, 14, 289. [Google Scholar] [CrossRef] [Green Version]
- Zhu, A.; Guo, W.; Gupta, S.; Fan, W.; Mower, J.P. Evolutionary dynamics of the plastid inverted repeat: The effects of expansion, contraction, and loss on substitution rates. New Phytol. 2016, 209, 1747–1756. [Google Scholar] [CrossRef] [Green Version]
- Wicke, S.; Schneeweiss, G.M.; de Pamphilis, C.W.; Müller, K.F.; Quandt, D. The evolution of the plastid chromosome in land plants: Gene content, gene order, gene function. Plant Mol. Biol. 2011, 76, 273–297. [Google Scholar] [CrossRef] [Green Version]
- Bock, R. Structure, function, and inheritance of plastid genomes. In Cell and Molecular Biology of Plastids; Springer: Berlin/Heidelberg, Germany, 2007; pp. 29–63. [Google Scholar]
- Li, Y.; Zhou, J.-G.; Chen, X.-L.; Cui, Y.-X.; Xu, Z.-C.; Li, Y.-H.; Song, J.-Y.; Duan, B.-Z.; Yao, H. Gene losses and partial deletion of small single-copy regions of the chloroplast genomes of two hemiparasitic Taxillus species. Sci. Rep. 2017, 7, 12834. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qi, F.; Zhao, Y.; Zhao, N.; Wang, K.; Li, Z.; Wang, Y. Structural variation and evolution of chloroplast tRNAs in green algae. PeerJ 2021, 9, e11524. [Google Scholar] [CrossRef] [PubMed]
- Shaul, O. How introns enhance gene expression. Int. J. Biochem. Cell Biol. 2017, 91, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Heyn, P.; Kalinka, A.T.; Tomancak, P.; Neugebauer, K.M. Introns and gene expression: Cellular constraints, transcriptional regulation, and evolutionary consequences. Bioessays 2015, 37, 148–154. [Google Scholar] [CrossRef]
- Liu, L.; Wang, Y.; He, P.; Li, P.; Lee, J.; Soltis, D.E.; Fu, C. Chloroplast genome analyses and genomic resource development for epilithic sister genera Oresitrophe and Mukdenia (Saxifragaceae), using genome skimming data. BMC Genom. 2018, 19, 235. [Google Scholar] [CrossRef] [Green Version]
- Downie, S.R.; Olmstead, R.G.; Zurawski, G.; Soltis, D.E.; Soltis, P.S.; Watson, J.C.; Palmer, J.D. Six independent losses of the chloroplast DNA rpl2 intron in dicotyledons: Molecular and phylogenetic implications. Evolution 1991, 45, 1245–1259. [Google Scholar] [CrossRef]
- Haberle, R.C.; Fourcade, H.M.; Boore, J.L.; Jansen, R.K. Extensive rearrangements in the chloroplast genome of Trachelium caeruleum are associated with repeats and tRNA genes. J. Mol. Evol. 2008, 66, 350–361. [Google Scholar] [CrossRef]
- Ogoma, C.A.; Liu, J.; Stull, G.W.; Wambulwa, M.C.; Oyebanji, O.; Milne, R.I.; Monro, A.K.; Zhao, Y.; Li, D.-Z.; Wu, Z.-Y. Deep insights into the plastome evolution and phylogenetic relationships of the tribe Urticeae (Family urticaceae). Front. Plant Sci. 2022, 13, 870949. [Google Scholar] [CrossRef]
- Yue, B.; Shi, J. The complete chloroplast genome sequence of Garcinia anomala (Clusiaceae) from Yunnan Province, China. Mitochondrial DNA Part B 2021, 6, 1899–1900. [Google Scholar] [CrossRef]
- Alqahtani, A.A.; Jansen, R.K. The evolutionary fate of rpl32 and rps16 losses in the Euphorbia schimperi (Euphorbiaceae) plastome. Sci. Rep. 2021, 11, 7466. [Google Scholar] [CrossRef]
- Yang, D.; Qiu, Q.; Xu, L.; Xu, Y.; Wang, Y. The complete chloroplast genome sequence of Garcinia pedunculata. Mitochondrial DNA Part B 2020, 5, 220–221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jin, D.-M.; Jin, J.-J.; Yi, T.-S. Plastome structural conservation and evolution in the clusioid clade of Malpighiales. Sci. Rep. 2020, 10, 9091. [Google Scholar] [CrossRef] [PubMed]
- Millen, R.S.; Olmstead, R.G.; Adams, K.L.; Palmer, J.D.; Lao, N.T.; Heggie, L.; Kavanagh, T.A.; Hibberd, J.M.; Gray, J.C.; Morden, C.W. Many parallel losses of infA from chloroplast DNA during angiosperm evolution with multiple independent transfers to the nucleus. Plant Cell 2001, 13, 645–658. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, S.; Jansen, R.K.; Park, S. Complete plastome sequence of Thalictrum coreanum (Ranunculaceae) and transfer of the rpl32 gene to the nucleus in the ancestor of the subfamily Thalictroideae. BMC Plant Biol. 2015, 15, 40. [Google Scholar] [CrossRef] [Green Version]
- Ramage, C.M.; Sando, L.; Peace, C.P.; Carroll, B.J.; Drew, R.A. Genetic diversity revealed in the apomictic fruit species Garcinia mangostana L.(mangosteen). Euphytica 2004, 136, 1–10. [Google Scholar] [CrossRef]
- Sando, L.; Peace, C.; Ramage, C.; Carrol, B.; Drew, R. Assessment of genetic diversity in Australian-grown mangosteen (Garcinia mangostana L.) and its wild relatives. In Proceedings of the II International Symposium on Biotechnology of Tropical and Subtropical Species; ISHS Acta Horticulturae 692; ISHS: Taipei, Taiwan, 2001; pp. 143–152. [Google Scholar]
- Sobir, S.; Poerwanto, R.; Santosa, E.; Sinaga, S.; Mansyah, E. Genetic variability in apomictic mangosteen (Garcinia mangostana) and its close relatives (Garcinia spp.) based on ISSR markers. Biodivers. J. Biol. Divers. 2011, 12, 59–63. [Google Scholar] [CrossRef]
- Zhu, M.; Feng, P.; Ping, J.; Li, J.; Su, Y.; Wang, T. Phylogenetic significance of the characteristics of simple sequence repeats at the genus level based on the complete chloroplast genome sequences of Cyatheaceae. Ecol. Evol. 2021, 11, 14327–14340. [Google Scholar] [CrossRef]
- Alzahrani, D.A.; Yaradua, S.S.; Albokhari, E.J.; Abba, A. Complete chloroplast genome sequence of Barleria prionitis, comparative chloroplast genomics and phylogenetic relationships among Acanthoideae. BMC Genom. 2020, 21, 393. [Google Scholar] [CrossRef]
- Asaf, S.; Ahmad, W.; Al-Harrasi, A.; Khan, A.L. Uncovering the first complete plastome genomics, comparative analyses, and phylogenetic dispositions of endemic medicinal plant Ziziphus hajarensis (Rhamnaceae). BMC Genom. 2022, 23, 83. [Google Scholar] [CrossRef]
- Zhan, X.; Zhang, Z.; Zhang, Y.; Gao, Y.; Jin, Y.; Shen, C.; Wang, H.; Feng, S. Complete Plastome of Physalis angulata var. villosa, Gene Organization, Comparative Genomics and Phylogenetic Relationships among Solanaceae. Genes 2022, 13, 2291. [Google Scholar] [CrossRef]
- Guo, Y.-Y.; Yang, J.-X.; Li, H.-K.; Zhao, H.-S. Chloroplast genomes of two species of Cypripedium: Expanded genome size and proliferation of AT-biased repeat sequences. Front. Plant Sci. 2021, 12, 609729. [Google Scholar] [CrossRef] [PubMed]
- Dunn, M.J.; Anderson, M.Z. To repeat or not to repeat: Repetitive sequences regulate genome stability in Candida albicans. Genes 2019, 10, 866. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, J.; Zhao, F.; Xu, Y.; Zhao, K.; Quan, H.; Su, Y.; Hao, P.; Liu, J.; Yu, B.; Yao, M. Complete chloroplast genome sequencing and phylogenetic analysis of two Dracocephalum plants. BioMed. Res. Int. 2020, 2020, 4374801. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Yang, X.; Zhang, C.; Yin, X.; Liu, S.; Li, X. Development of chloroplast microsatellite markers and analysis of chloroplast diversity in Chinese jujube (Ziziphus jujuba Mill.) and wild jujube (Ziziphus acidojujuba Mill.). PLoS ONE 2015, 10, e0134519. [Google Scholar] [CrossRef]
- Park, H.; Kim, C.; Lee, Y.M.; Kim, J.H. Development of chloroplast microsatellite markers for the endangered Maianthemum bicolor (Asparagaceae s.l.). Appl. Plant Sci. 2016, 4, 1600032. [Google Scholar] [CrossRef]
- Bock, R.; Knoop, V. Genomics of Chloroplasts and Mitochondria; Springer Science & Business Media: Berlin, Germany, 2012; Volume 35. [Google Scholar]
- Chen, H.; Shao, J.; Zhang, H.; Jiang, M.; Huang, L.; Zhang, Z.; Yang, D.; He, M.; Ronaghi, M.; Luo, X. Sequencing and analysis of Strobilanthes cusia (Nees) Kuntze chloroplast genome revealed the rare simultaneous contraction and expansion of the inverted repeat region in angiosperm. Front. Plant Sci. 2018, 9, 324. [Google Scholar] [CrossRef] [Green Version]
- Jo, I.-H.; Han, S.; Shim, D.; Ryu, H.; Hyun, T.K.; Lee, Y.; Kim, D.; So, Y.-S.; Chung, J.-W. Complete Chloroplast Genome of the Inverted Repeat-Lacking Species Vicia bungei and Development of Polymorphic Simple Sequence Repeat Markers. Front. Plant Sci. 2022, 13, 1571. [Google Scholar] [CrossRef]
- Kim, K.-J.; Lee, H.-L. Complete chloroplast genome sequences from Korean ginseng (Panax schinseng Nees) and comparative analysis of sequence evolution among 17 vascular plants. DNA Res. 2004, 11, 247–261. [Google Scholar] [CrossRef]
- Jo, S.; Kim, Y.-K.; Cheon, S.-H.; Fan, Q.; Kim, K.-J. Characterization of 20 complete plastomes from the tribe Laureae (Lauraceae) and distribution of small inversions. PLoS ONE 2019, 14, e0224622. [Google Scholar] [CrossRef]
- Catalano, S.A.; Saidman, B.O.; Vilardi, J.C. Evolution of small inversions in chloroplast genome: A case study from a recurrent inversion in angiosperms. Cladistics 2009, 25, 93–104. [Google Scholar] [CrossRef]
- Goulet, B.E.; Roda, F.; Hopkins, R. Hybridization in plants: Old ideas, new techniques. Plant Physiol. 2017, 173, 65–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.-J.; Yu, W.-B.; Yang, J.-B.; Song, Y.; de Pamphilis, C.W.; Yi, T.-S.; Li, D.-Z. GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 2020, 21, 241. [Google Scholar] [CrossRef] [PubMed]
- Kajitani, R.; Toshimoto, K.; Noguchi, H.; Toyoda, A.; Ogura, Y.; Okuno, M.; Yabana, M.; Harada, M.; Nagayasu, E.; Maruyama, H. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genome. Res. 2014, 24, 1384–1395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Tillich, M.; Lehwark, P.; Pellizzer, T.; Ulbricht-Jones, E.S.; Fischer, A.; Bock, R.; Greiner, S. GeSeq—Versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar] [CrossRef] [Green Version]
- Greiner, S.; Lehwark, P.; Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 2019, 47, W59–W64. [Google Scholar] [CrossRef] [Green Version]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [Green Version]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Harman, G.E.; Kubicek, C.P. Trichoderma and Gliocladium: Basic Biology, Taxonomy and Genetics; CRC Press: Boca Raton, FL, USA, 2002; Volume 1. [Google Scholar]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brudno, M.; Do, C.B.; Cooper, G.M.; Kim, M.F.; Davydov, E.; Green, E.D.; Sidow, A.; Batzoglou, S.; Program, N.C.S. LAGAN and Multi-LAGAN: Efficient tools for large-scale multiple alignment of genomic DNA. Genome. Res. 2003, 13, 721–731. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amiryousefi, A.; Hyvönen, J.; Poczai, P. IRscope: An online program to visualize the junction sites of chloroplast genomes. Bioinformatics 2018, 34, 3030–3031. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.E.; Mau, B.; Perna, N.T. progressiveMauve: Multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 2010, 5, e11147. [Google Scholar] [CrossRef] [Green Version]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef] [Green Version]
- Darriba, D.; Posada, D.; Kozlov, A.M.; Stamatakis, A.; Morel, B.; Flouri, T. ModelTest-NG: A new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol. 2020, 37, 291–294. [Google Scholar] [CrossRef] [Green Version]
- Kozlov, A.M.; Darriba, D.; Flouri, T.; Morel, B.; Stamatakis, A. RAxML-NG: A fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 2019, 35, 4453–4455. [Google Scholar] [CrossRef] [Green Version]






| Species | Plastome Size (bp) | Size (bp) | Number of Genes * | GC Content (%) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LSC | SSC | IR | All | Protein-Coding | rRNA | tRNA | All | LSC | SSC | IR | |||
| G. anomala | 156,774 | 85,586 | 17,082 | 27,053 | 128 (111) | 83 (77) | 8 (4) | 37 (30) | 36.1 | 33.5 | 30.3 | 42.1 | |
| G. gummi-gutta | 156,202 | 84,996 | 17,088 | 27,059 | 127 (110) | 83 (77) | 8 (4) | 36 (29) | 36.2 | 33.5 | 30.3 | 42.1 | |
| G. mangostana | Manggis | 156,582 | 85,385 | 17,137 | 27,030 | 128 (111) | 83 (77) | 8 (4) | 37 (30) | 36.2 | 33.6 | 30.2 | 42.2 |
| Mesta | 156,580 | 85,383 | 17,137 | 27,030 | 128 (111) | 83 (77) | 8 (4) | 37 (30) | 36.2 | 33.6 | 30.2 | 42.2 | |
| Thailand | 158,179 | 86,458 | 17,703 | 27,009 | 128 (111) | 83 (77) | 8 (4) | 37 (30) | 36.1 | 33.5 | 30.1 | 42.2 | |
| G. oblongifolia | 156,577 | 85,393 | 17,064 | 27,060 | 128 (111) | 83 (77) | 8 (4) | 37 (30) | 36.2 | 33.6 | 30.3 | 42.2 | |
| G. paucinervis | 157,702 | 85,989 | 17,737 | 26,988 | 128 (111) | 83 (77) | 8 (4) | 37 (30) | 36.2 | 33.6 | 30.3 | 42.2 | |
| G. pedunculata | 157,688 | 85,998 | 17,656 | 27,017 | 128 (111) | 83 (77) | 8 (4) | 37 (30) | 36.2 | 33.6 | 30.2 | 42.2 | |
| Function | Group | Gene Name |
|---|---|---|
| Protein synthesis and DNA replication | Transfer RNA | trnA-UGC (2×), trnC-GCA, trnD-GUC, trnE-UUC, trnF-GAA, trnfM-CAU, trnG-GCC, trnH-GUG, trnK-UUU, trnI-GAU (2×), trnL-CAA (2×), trnL-UAA, trnL-UAG, trnM-CAU, trnI-CAU (2×), trnN-GUU (2×), trnP-UGG, trnQ-UUG, trnR-ACG (2×), trnR-UCU, trnS-CGA, trnS-GCU, trnS-GGA, trnS-UGA, trnT-GGU, trnT-UGU, trnV-GAC (2×), trnV-UAC, trnW-CCA, trnY-GUA |
| Ribosomal RNA | rrn16 (2×), rrn23 (2×), rrn4.5 (2×), rrn5 (2×) | |
| Ribosomal protein small subunit | rps2, rps3, rps4, rps7 (2×), rps8, rps11, rps12 (2×), rps14, rps15, rps16, rps18, rps19 | |
| Ribosomal protein large subunit | rpl2 (2×), rpl14, rpl16, rpl20, rpl22, rpl23 (2×), rpl33, rpl36 | |
| Subunits of RNA polymerase | rpoA, rpoB, rpoC1, rpoC2 | |
| Photosynthesis | Photosystem I | psaA, psaB, psaC, psaI, psaJ |
| Photosystem II | psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbT, psbZ | |
| Cytochrome b6f complex | petA, petB, petD, petG, petL, petN | |
| ATP synthase | atpA, atpB, atpE, atpF, atpH, atpI | |
| NADH-dehydrogenase | ndhA, ndhB (2×), ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, ndhK | |
| Large subunit Rubisco | rbcL | |
| Miscellaneous group | Photosystem II protein N | psbN |
| Acetyl-CoA carboxylase | accD | |
| Cytochrome c biogenesis | ccsA | |
| Maturase | matK | |
| ATP-dependent protease | clpP | |
| Inner membrane protein | cemA | |
| Pseudogene unknown function | Conserved hypothetical chloroplast ORF | ycf1, ycf2 (2×), ycf3, ycf4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wee, C.-C.; Nor Muhammad, N.A.; Subbiah, V.K.; Arita, M.; Nakamura, Y.; Goh, H.-H. Plastomes of Garcinia mangostana L. and Comparative Analysis with Other Garcinia Species. Plants 2023, 12, 930. https://doi.org/10.3390/plants12040930
Wee C-C, Nor Muhammad NA, Subbiah VK, Arita M, Nakamura Y, Goh H-H. Plastomes of Garcinia mangostana L. and Comparative Analysis with Other Garcinia Species. Plants. 2023; 12(4):930. https://doi.org/10.3390/plants12040930
Chicago/Turabian StyleWee, Ching-Ching, Nor Azlan Nor Muhammad, Vijay Kumar Subbiah, Masanori Arita, Yasukazu Nakamura, and Hoe-Han Goh. 2023. "Plastomes of Garcinia mangostana L. and Comparative Analysis with Other Garcinia Species" Plants 12, no. 4: 930. https://doi.org/10.3390/plants12040930






