Proteome Analysis of Nicotiana tabacum Cells following Isonitrosoacetophenone Treatment Reveals Defence-Related Responses Associated with Priming
Abstract
:1. Introduction
2. Results
2.1. Two-Dimensional Electrophoresis
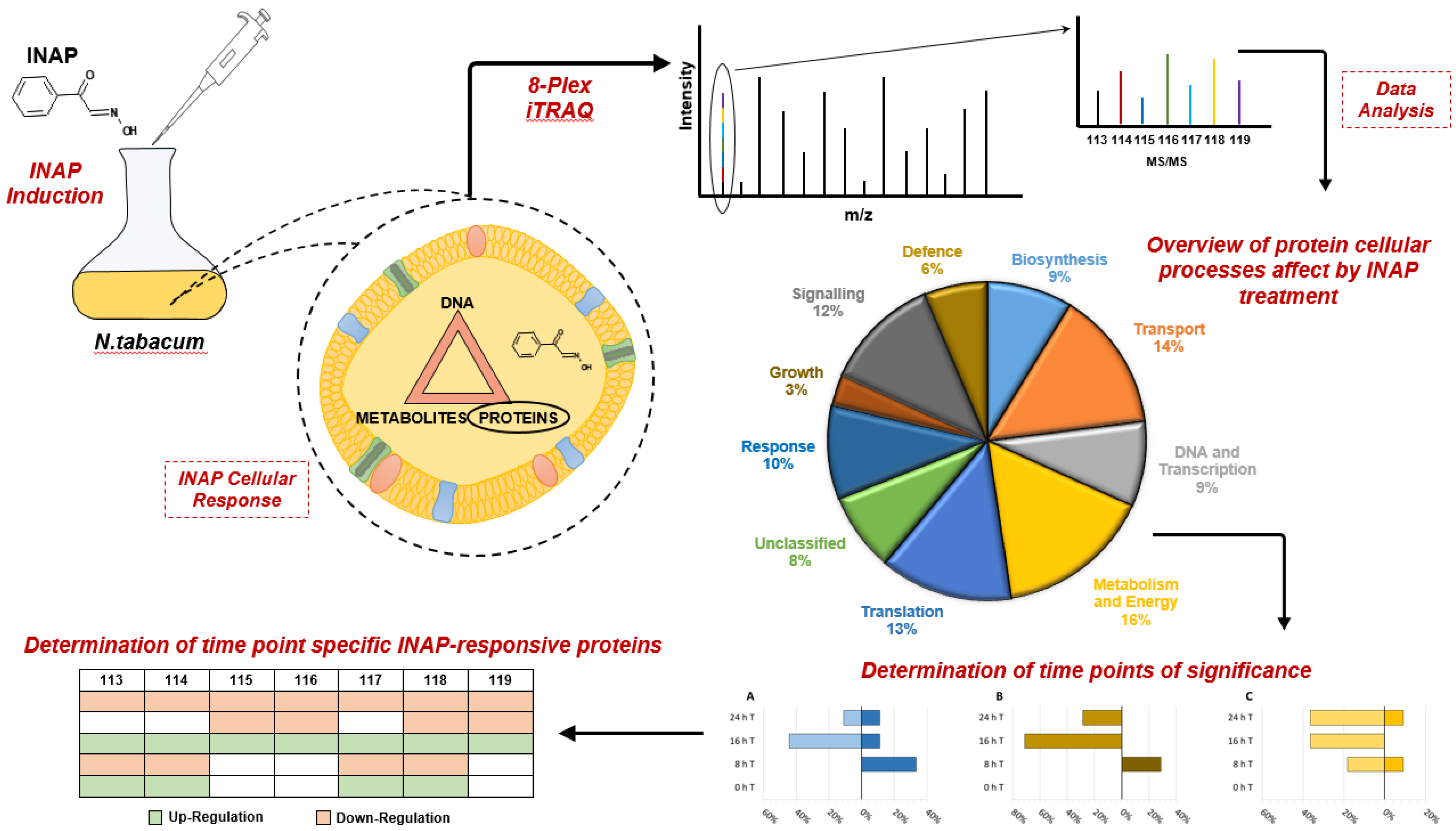
2.2. iTRAQ LC-MS Analysis
3. Discussion
3.1. Adaption-Related Proteins Responsive to INAP Treatment
3.1.1. Biosynthesis and Redirection of Resources
3.1.2. Defence
3.1.3. DNA and Transcription
3.1.4. Growth, Metabolism and Energy
3.1.5. Stress-Related Responses and Signalling Events
3.1.6. Translation
3.1.7. Transport
3.2. Basic Functionality Proteins
Mitochondrial Activity
4. Materials and Methods
4.1. Cell Suspension Cultures, INAP Treatment and Experimental Design
4.2. Protein Extraction and Two-Dimensional Gel Electrophoresis
4.3. Protein Extraction for iTRAQ
4.4. Sample Preparation and Labelling for 8-plex iTRAQ
4.5. OFFGEL Electrophoresis
4.6. LC MS/MS Analysis
4.7. Data Analysis and Database Searching
4.8. Data Analysis: Protein Identification Criteria and Bioinformatics Validation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Slaughter, A.; Daniel, X.; Flors, V.; Luna, E.; Hohn, B.; Mauch-Mani, B. Descendants of primed Arabidopsis plants exhibit resistance to biotic stress. Plant Physiol. 2012, 158, 835–843. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sanabria, N.M.-A.; Huang, J.-C.; Dubery, I.A. Self/non-self perception in plants in innate immunity and defense. Self/Non-Self 2010, 1, 40–45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van der Ent, S.; Koorneef, A.; Ton, J.; Pieterse, C.M.J. Induced resistance–orchestrating defence mechanisms through crosstalk and priming. Annu. Plant Rev. 2009, 34, 334–370. [Google Scholar] [CrossRef] [Green Version]
- Henry, G.; Thonart, P.; Ongena, M. PAMPs, MAMPs, DAMPs and others: An update on the diversity of plant immunity elicitors. Biotechnol. Agron. Soc. Environ. 2012, 16, 257–268. [Google Scholar]
- Muthamilarasan, M.; Prasad, M. Plant innate immunity: An updated insight into defence mechanism. J. Biosci. 2013, 38, 1–17. [Google Scholar] [CrossRef]
- Tugizimana, F.; Mhlongo, M.L.; Piater, L.A.; Dubery, I.A. Metabolomics in plant priming research: The way forward? International. J. Mol. Sci. 2018, 19, 1759. [Google Scholar] [CrossRef] [Green Version]
- Madala, N.E.; Steenkamp, P.A.; Piater, L.A.; Dubery, I.A. Metabolomic analysis of isonitrosoacetophenone-induced perturbations in phenolic metabolism of Nicotiana tabacum Cells. Phytochemistry 2013, 94, 82–90. [Google Scholar] [CrossRef]
- Djami-Tchatchou, A.T.; Maake, M.P.; Piater, L.A.; Dubery, I.A. Isonitrosoacetophenone drives transcriptional reprogramming in Nicotiana tabacum cells in support of innate immunity and defense. PLoS ONE 2015, 10, e0117377. [Google Scholar] [CrossRef] [Green Version]
- Segura, A.; Godoy, P.; van Dillewijn, P.; Hurtado, A.; Arroyo, N.; Sanatacruz, S.; Ramos, J.L. Proteomic analysis reveals the participation of energy- and stress-related proteins in the response of Pseudomonas putida DOT-T1E to Toluene. J. Bacteriol. 2005, 187, 5937–5945. [Google Scholar] [CrossRef] [Green Version]
- Berthelot, K.; Estevez, Y.; Deffieux, A.; Peruch, F. Isopentenyl disphosphate isomerase: A checkpoint to isoprenoid biosynthesis. Biochimie 2012, 94, 1621–1634. [Google Scholar] [CrossRef]
- Tugizimana, F.; Steenkamp, P.A.; Piater, L.A.; Dubery, I.A. Ergosterol-induced sesquiterpenoid synthesis in tobacco cells. Molecules 2012, 17, 1698–1715. [Google Scholar] [CrossRef]
- Lim, E.K.; Higgins, G.S.; Li, Y.; Bowles, D.J. Regioselectivity of glucosylation of caffeic acid by a UDP-glucose:glucosyltransferase is maintained in planta. Biochem. J. 2003, 373, 987–992. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wheatley, E.R.; Davies, D.R.; Bolwen, G.P. Characterization and immunolocation of an 87 kDa polypeptide associated with UDP-glucuronic acid decarboxylase activity from differing tobacco cells (Nicotiana tabacum L.). Phytochemistry 2002, 61, 771–780. [Google Scholar] [CrossRef] [PubMed]
- Mehta, A.; Brasileiro, A.C.M.; Souza, D.S.L.; Romano, E.; Campos, M.A.; Grossi-de-Sá, M.F.; Silva, M.S.; Franco, O.L.; Fragoso, R.R.; Bevitori, R.; et al. Plant-pathogen interactions: What is proteomics telling us? FEBS J. 2008, 275, 3731–3746. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Farooq, M.; Aziz, T.; Basra, M.A.; Cheema, M.A.; Rehman, H. Chilling tolerance in hybrid maize induced by seed priming with salicylic acid. J. Agron. Crop Sci. 2008, 194, 161–168. [Google Scholar] [CrossRef]
- Kellokumpu, S.; Hassinen, A.; Glumoff, T. Glycosyltransferase complexes in eukaryotes: Long-known, prevalent but still unrecognized. Cell. Mol. Life Sci. 2016, 73, 305–325. [Google Scholar] [CrossRef]
- Souza, C.D.A.; Kim, S.S.; Kienow, L.; Schneider, K.; McKim, S.M.; Haughn, G.W.; Kombrink, E.; Douglas, C.J. A novel fatty acyl-CoA synthetase is required for pollen development and sporopollenin biosynthesis in Arabidopsis. Plant Cell 2009, 21, 507–525. [Google Scholar] [CrossRef] [Green Version]
- Shockey, J.M.; Fulda, M.S.; Browse, J.A. Arabidopsis contains nine long-chain acyl-coenzyme A synthetase genes that participate in fatty acid and glycerolipid metabolism. Plant Physiol. 2002, 129, 1710–1722. [Google Scholar] [CrossRef] [Green Version]
- Zhou, S.; Wei, S.; Boone, B.; Levy, S. Microarray analysis of genes affected by salt stress in tomato. Afr. J. Environ. Sci. Technol. 2007, 1, 14–26. [Google Scholar]
- Pritchard, L.; Birch, P. A systems biology perspective on plant-microbe interactions: Biochemical and structural targets of pathogen effectors. Plant Sci. 2011, 180, 584–603. [Google Scholar] [CrossRef]
- Nuhse, T.S. Cell wall integrity signaling and innate immunity in plants. Front. Plant Sci. 2012, 3, 280. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carmo, L.S.T.; Resende, R.O.; Silva, L.P.; Ribeiro, S.G.; Mehta, A. Identification of host proteins modulated by the virulence factor AC2 of tomato chlorotic mottle virus in Nicotiana benthamiana. Proteomics 2013, 13, 1947–1960. [Google Scholar] [CrossRef] [PubMed]
- Jami, S.K.; Clark, G.B.; Turlapati, S.A.; Handley, C.; Roux, S.J.; Kirti, P.B. Ectopic expression of an annexin from Brassica juncea confers tolerance to abiotic and biotic stress treatments in transgenic tobacco. Plant Physiol. Biochem. 2008, 46, 1019–1030. [Google Scholar] [CrossRef] [PubMed]
- Shadle, G.; Chen, F.; Reddy, M.S.S.; Jackson, L.; Nakashima, J.; Dixon, R.A. Down-regulation of hydroxycinnamoyl CoA: Shikimate hydroxycinnamoyl transferase in transgenic alfalfa affects lignification, development and forage quality. Phytochemistry 2007, 68, 1521–1529. [Google Scholar] [CrossRef]
- Negrel, J.; Lotfy, S.; Javelle, F. Modulation of the activity of two hydroxycinnamoyl transerfases in wound-healing potato tuber discs in response to pectinase or abscisic acid. J. Plant Physiol. 1995, 146, 318–322. [Google Scholar] [CrossRef]
- Federici, L.; Di Matteo, A.; Fernandez-Recio, J.; Tsernoglou, D.; Cervone, F. Polygalacturonase inhibiting proteins: Players in plant innate immunity? Trends Plant Sci. 2006, 11, 65–70. [Google Scholar] [CrossRef]
- Ton, J.; Mauch-Mani, B. β-Amino-butyric acid-induced resistance against necrotrophic pathogens is based on ABA-dependent priming for callose. Plant J. 2004, 38, 119–130. [Google Scholar] [CrossRef]
- Mauch-Mani, B.; Mauch, F. The role of abscisic acid in plant-pathogen interactions. Curr. Opin. Plant Biol. 2005, 8, 409–414. [Google Scholar] [CrossRef]
- Deepak, S.A.; Ishii, H.; Park, P. Acibenzolar-S-methyl primes cell wall strengthening genes and reactive oxygen species forming/scavenging enzymes in cucumber after fungal pathogen attack. Physiol. Molec. Plant Pathol. 2006, 69, 52–61. [Google Scholar] [CrossRef]
- Kimura, S.; Suzuki, T.; Yanagawa, Y.; Yamamoto, T.; Nakagawa, H.; Tanaka, I.; Hashimoto, J.; Sakaguchi, K. Characterization of plant proliferating cell nuclear antigen (PCNA) and flap endonuclease-1 (FEN-1), and their distribution in mitotic and meiotic cell cycles. Plant J. 2001, 28, 643–653. [Google Scholar] [CrossRef] [Green Version]
- Hua, J.; Grisafi, P.; Cheng, S.H.; Fink, G.R. Plant growth homeostasis in controlled by the Arabidopsis BON1 and BAP1 genes. Genes Dev. 2001, 15, 2263–2272. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ando, S.; Takumi, S.; Ueda, Y.; Ueda, T.; Mori, N.; Nakamura, C. Nicotiana tabacum cDNAs encoding α and β subunits of a heterotrimeric GTP-binding protein isolated from hairy root tissues. Genes Genet. Syst. 2000, 75, 211–221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Collado-Romero, M.; Alós, E.; Prieto, P. Unravelling the proteomic profile of rice meiocytes during early meiosis. Front. Plant Sci. 2014, 5, 356. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barabasz, A.; Klimecka, M.; Kendziorek, M.; Weremczuk, A.; Ruszczyńska, A.; Bulska, E.; Antosiewicz, D.M. The ratio of Zn to Cd supply as a determinant of metal-homeostasis gene expression in tobacco and its modulation by overexpressing the metal exporter AtHMA4. J. Exp. Bot. 2016, 67, 6201–6214. [Google Scholar] [CrossRef] [Green Version]
- Bosch, M.; Hepler, P.K. Pectin methylesterases and pectin dynamics in pollen tubes. Plant Cell 2005, 17, 3219–3226. [Google Scholar] [CrossRef] [Green Version]
- Conrath, U.; Thulke, O.; Katz, V.; Schwindling, S.; Kohler, A. Priming as a mechanism in induced systemic resistance in plants. Eur. J. Plant Pathol. 2001, 107, 113–119. [Google Scholar] [CrossRef]
- Mazari, K.; Landa, P.; Přerostova, S.; Műller, K.; Vaňkova, R.; Soudek, P.; Vaňek, T. Thorium impact on tobacco root transcriptome. J. Hazard. Mater. 2017, 325, 163–169. [Google Scholar] [CrossRef]
- Petriccione, M.; Di Cecco, I.; Arena, S.; Scaloni, A.; Scortichini, M. Proteomics changes in Actinidia chinensis shoot during systemic infection with a pandemic Pseudomonas syringae pv. actinidiae strain. J. Proteom. 2013, 78, 461–476. [Google Scholar] [CrossRef]
- Showalter, A.M. Structure and function of plant cell wall proteins. The Plant Cell 1993, 5, 9–23. [Google Scholar] [CrossRef]
- Gerber, I.B.; Laukens, K.; De Vijlder, T.; Witters, E.; Dubery, I.A. Proteomic profiling of cellular targets of lipopolysaccharide-induced signalling in Nicotiana tabacum BY-2 cells. Biochim. Et Biophys. Acta 2008, 1784, 1750–1762. [Google Scholar] [CrossRef]
- Zheng, Z.L.; Yang, Z. The Rop GTPase: An emerging signalling switch in plants. Plant Mol. Biol. 2000, 44, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Mazel, A.; Leshem, Y.; Tiwari, B.S.; Levine, A. Induction of salt and osmotic stress tolerance by overexpression of an intracellular vesicle trafficking protein AtRab7 (AtRabG3e). Plant Physiol. 2004, 134, 118–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ludwig, A.A.; Saitoh, H.; Felix, G.; Freymark, G.; Miersch, O.; Wasternack, C.; Boller, T.; Jones, J.D.G.; Romeis, T. Ethylene-mediated cross-talk between calcium-dependent protein kinase and MAPK signalling controls stress responses in plants. Proc. Natl. Acad. Sci. USA 2005, 102, 10736–10741. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, X.; Wang, Z.Q.; Xiao, R.; Wang, Y.; Xie, Y.; Zhou, X. iTRAQ analysis of the tobacco leaf proteome reveals that RNA-directed DNA methylation (RdDM) has important roles in defense against Geminivirus-Betasatellite infection. J. Proteom. 2017, 152, 88–101. [Google Scholar] [CrossRef] [PubMed]
- Devos, S.; Laukens, K.; Deckers, P.; Van der Straaten, D.; Beekman, T.; Inzé, D.; Van Onckelen, H.; Witters, E.; Prinsen, E. A hormone and proteome approach to picturing the initial metabolic events during Plasmodiophora brassica infection on Arabidopsis. MPMI 2006, 12, 1693–1707. [Google Scholar] [CrossRef] [Green Version]
- Hulsmans, S.; Rodriguez, M.; De Coninck, B.; Rolland, R. The SnRK1 energy sensor in plant biotic interactions. Trends Plant Sci. 2016, 21, 648–661. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J. Phospholipase D and phosphatidic acid in plant defence response: From protein-protein and lipid-protein interactions to hormone signalling. J. Exp. Bot. 2015, 66, 1721–1736. [Google Scholar] [CrossRef] [Green Version]
- Wang, X. Regulatory functions of phospholipase D and phosphatidic acid in plant growth, development, and stress response. Plant Physiol. 2005, 139, 566–573. [Google Scholar] [CrossRef] [Green Version]
- Liscovitch, M.; Czarny, M.; Fiucci, G.; Tang, X. Phospholipase D: Molecular and cell biology of a novel gene family. Biochem. J. 2000, 345, 401–415. [Google Scholar] [CrossRef]
- van Loon, L.C.; Rep, M.; Pieterse, C.M.J. Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef] [Green Version]
- Dubery, I.A. An elicitor- and pathogen-induced cDNA from potato encodes a stress-responsive cyclophilin. Biol. Plant. 2007, 51, 327–332. [Google Scholar] [CrossRef]
- Crouzet, J.; Roland, J.; Peeters, E.; Trombik, T.; Ducos, E.; Nader, J.; Boutry, M. NtPDR1, a plasma membrane ABC transporter from Nicotiana tabacum, is involved in diterpene transport. Plant Mol. Biol. 2013, 82, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Tahara, H.; Yokota, E.; Igarashi, H.; Orii, H.; Yao, M.; Sonobe, S.; Hashimoto, T.; Hussey, P.J.; Shimmen, T. Clathrin is involved in organization of mitotic spindle and phragmoplast as well as in endocytosis in tobacco cell cultures. Protoplasma 2007, 230, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Satoh-Nagasawa, N.; Mori, M.; Nakazawa, N.; Kawamoto, T.; Nagato, Y.; Sakurai, K.; Takahashi, H.; Watanabe, A.; Akagi, H. Mutations in rice (Oryza sativa) heavy metal ATPase 2 (OsHMA2) restrict the translocation of zinc and cadmium. Plant Cell Physiol. 2012, 53, 213–224. [Google Scholar] [CrossRef] [Green Version]
- Morsomme, P.; Boutry, M. The plant plasma membrane H+-ATPase: Structure, function and regulation. Biochim. Et Biophys. Acta 2000, 1465, 1–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marti, L.; Fornaciari, S.; Renna, L.; Stefano, G.; Brandizzi, F. COPII-mediated traffic in plants. Trends Plant Sci. 2010, 15, 522–528. [Google Scholar] [CrossRef]
- Nziengui, H.; Schoefs, B. Functions of reticulons in plants: What we can learn from animal and yeasts. Cell. Mol. Life Sci. 2009, 66, 584–595. [Google Scholar] [CrossRef]
- Wasternack, C. Jasmonates: An update on biosynthesis, signal transduction and action in plant stress response, growth and development. Ann. Bot. 2007, 100, 681–697. [Google Scholar] [CrossRef] [Green Version]
- Azevedo, R.A.; Lancien, M.; Lea, P.J. The aspartic acid metabolic pathway, an exciting and essential pathway in plants. Amino Acids 2006, 30, 143–162. [Google Scholar] [CrossRef]
- Forizs, L.; Lestrade, S.; Mol, A.; Dierick, J.F.; Gerbaux, C.; Diallo, B.; Jaziri, M.E.; Baucher, M.; Vandeputte, O.M. Metabolic shift in the phytopathogen Rhodococcus fasciana in response to cell-free extract of infected tobacco plant tissues. Curr. Microbiol. 2009, 58, 483–487. [Google Scholar] [CrossRef]
- Balmer, A.; Pastor, V.; Glauser, G.; Mauch-Mani, B. Tricarboxylates induce defense priming against bacteria in Arabidopsis thaliana. Front. Plant Sci. 2018, 9, 1221. [Google Scholar] [CrossRef]
- Downs, C.A.; Heckathorn, S.A. The mitochondrial small heat-shock protein protects NADH:ubiquinone oxidoreductase of the electron transport chain during heat stress in plants. FEBS Lett. 1998, 430, 246–250. [Google Scholar] [CrossRef]
- Van Aken, O. Mitochondrial redox systems as central hubs in plant metabolism and signaling. Plant Physiol. 2021, 186, 36–52. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Vignani, R.; Scali, M.; Cresti, M. A universal and rapid protocol for protein extraction from recalcitrant plant tissues for proteomic analysis. Electrophoresis 2006, 27, 2782–2786. [Google Scholar] [CrossRef] [PubMed]
- Khoza, T.; Dubery, I.A.; Piater, I.A. Identification of candidate ergosterol-responsive proteins associated with the plasma membrane of Arabidopsis thaliana. Int. J. Mol. Sci. 2019, 20, 1302. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fairbanks, G.; Steck, T.L.; Wallach, D.F. Electrophoretic analysis of the major polypeptides of the human erythrocyte membrane. Biochemistry 1971, 10, 2606–2617. [Google Scholar] [CrossRef] [PubMed]
- Nesvizhskii, A.I.; Keller, A.; Kolker, E.; Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 2003, 75, 4646–4658. [Google Scholar] [CrossRef]
- Koskinen, V.R.; Emery, P.A.; Creasy, D.M.; Cottrell, J.S. Hierarchical clustering of shotgun proteomics data. Mol. Cell. Proteom. 2011, 10, M110.003822. [Google Scholar] [CrossRef] [Green Version]
- Kosová, K.; Vítámvás, P.; Prášil, I.T.; Renaut, J. Plant proteome changes under abiotic stress–Contribution of proteomics studies to understanding plant stress response. J. Proteom. 2011, 74, 1301–1322. [Google Scholar] [CrossRef]
- Kosová, K.; Vítamvás, P.; Urban, M.O.; Prášil, I.T.; Renaut, J. Plant abiotic stress prpteomics: The major facors determining alterations in cellular proteome. Front. Plant Sci. 2018, 9, 122. [Google Scholar] [CrossRef] [Green Version]



| Protein Name | Accession Number | 0 h (T) | 8 h (T) | 16 h (T) | 24 h (T) | Gene Accession Number | Gene Name |
|---|---|---|---|---|---|---|---|
| Biosynthesis | |||||||
| UDP-glucose:protein transglucosylase-like protein SlUPTG1 1 | Q6IV07_SOLLC | 0.0 | 0.2 | −0.3 | 0.0 | XM 016639837.1 a | Alpha-1,4-glucan-protein synthase |
| Uncharacterised protein 2 | M1C547_SOLTU | 0.0 | 0.7 | 0.3 | 0.5 | XM 016633077.1 a | Isopentenyl-diphosphate delta-isomerase I |
| Uncharacterised protein 2 | M1C5Y4_SOLTU | 0.0 | 1.2 | −0.3 | 0.4 | NM 001326080.1 | UDP-glucuronic acid decarboxylase 6-like |
| Uncharacterised protein 1 | K4BVZ4_SOLLC | 0.0 | −0.2 | −0.6 | −0.3 | XM 016647169.1 b | Dolichyl-diphosphooligosaccharide-protein glycosyltrasnferase |
| Uncharacterised protein 1 | K4CET6_SOLLC (+1) | 0.0 | 0.2 | −0.8 | −0.3 | XM 016657513.1 b | Long chain acyl-CoA synthetase 8-like |
| Cystathionine gamma-synthase 3 | Q9ZPL5_TOBAC | 0.0 | 0.0 | −0.6 | −0.4 | AF097180.1 | Alpha-1,4-glucan-protein synthase |
| Defence | |||||||
| Annexin 3 | Q9XEN8_TOBAC | 0.0 | 0.6 | 0.4 | 0.4 | NM 001325957.1 | |
| Putative hydroxycinnamoyl transferase 5 | B5LAV0_CAPAN (+5) | 0.0 | 0.5 | −0.1 | 0.3 | NM 001325623.1 | Shikimate O-hydroxy-cinnamoyltransferase |
| Superoxide dismutase (Cu-Zn) 8 | SODC_NICPL | 0.0 | −0.4 | −0.8 | −0.4 | XM 016594235.1 a | |
| Polygalacturonase inhibiting protein 4 | C4PG28_9SOLN | 0.0 | −0.3 | −0.6 | −0.3 | XM 016650938.1 a | Polygalacturonase inhibitor-like |
| Uncharacterised protein 1 | K4ASK1_SOLLC | 0.0 | 0.1 | −0.9 | −0.3 | XM 016578137.1 a | Callose synthase 9-like |
| DNA and Transcription | |||||||
| Uncharacterised protein 1 | K4CMV1_SOLLC | 0.0 | −0.6 | −0.7 | −0.9 | XR 001654730.1 b | Replication protein A 70kDa DNA-binding |
| XM 016640989.1 b | |||||||
| Endonuclease 1 1 | G3XKQ7_SOLLC (+2) | 0.0 | −0.2 | −0.6 | −0.5 | XM 016636178.1 a | |
| Histone H2B 6 | A2IBL2_NICBE | 0.0 | 0.1 | −0.6 | −0.2 | ||
| Histone H2B 3 | H2B_TOBAC | 0.0 | −0.4 | −0.7 | −0.2 | ||
| Growth | |||||||
| Uncharacterised protein 1 | K4BLX1_SOLLC | 0.0 | 0.0 | −0.7 | −0.4 | XM 016647000.1 b | BONZAI 1-like |
| XM 016649340.1 b | |||||||
| DWARF1/DIMINUTO 1 | Q66YT8_SOLLC | 0.0 | −0.2 | −0.7 | −0.4 | XM 016612143.1 a | Delta(24)-sterol reductase-like |
| Uncharacterised protein 1 | K4BGV0_SOLLC | 0.0 | 0.1 | 0.7 | 0.0 | XM 016614569.1 b | CDK5RAP3-like protein |
| Metabolism and Energy | |||||||
| Uncharacterised protein 1 | K4BDG0_SOLLC (+2) | 0.0 | −0.1 | −1.3 | −0.7 | XM 016594393.1 b | Very-long-chain 3-oxoacyl-CoA reductase 1-like |
| XM 016626206.1 b | |||||||
| Uncharacterised protein 2 | M1A251_SOLTU | 0.0 | −0.4 | −0.8 | −0.5 | XM 016617189.1 b | Suberization-associated anionic peroxidase 1-like |
| Probable pectate lyase P18 1 | K4BDF4_SOLLC | 0.0 | 0.1 | −0.2 | 0.0 | XM 016621815.1 b | |
| Uncharacterised protein 1 | K4AT35_SOLLC (+2) | 0.0 | −0.2 | 0.0 | 0.6 | XM 0166417471 a | 26S Proteasome non-ATPase |
| XM 016629739.1 a | |||||||
| Glycylpeptide N-tetradecanoyltransferase 2 | M0ZRQ4_SOLTU | 0.0 | 0.5 | 0.1 | 0.4 | XM 016628510.1 a | |
| Methylenetetrahydro-folate reductase 1 | K4C2P8_SOLLC (+2) | 0.0 | 0.7 | 0.4 | 0.5 | XM 016660349.1 a | |
| XM 016660348.1 a XM 016660347.1 a | |||||||
| Response | |||||||
| Glycine-rich RNA-binding protein 3 | B2YKT9_TOBAC | 0.0 | 0.1 | 0.4 | 0.6 | ||
| Putative methyltransferase 2 | A4UV20_SOLTU (+3) | 0.0 | −0.3 | −0.8 | −0.4 | NM 001324795.1 | |
| LC052785.1 | |||||||
| Uncharacterised protein 2 | M1C4X5_SOLTU | 0.0 | −0.3 | −0.9 | −0.9 | XM 016650151.1 b XM 016650150.1 b | Probable methyltransferase |
| Signalling | |||||||
| Ras-related GTP-binding protein 3 | Q40569_TOBAC | 0.0 | 0.1 | −0.7 | −0.4 | NM 001325922.1 X72212.1 | |
| Uncharacterised protein 2 | M1B9Y5_SOLTU | 0.0 | 0.0 | −0.7 | −0.3 | XM 016620909.1 b | Putative receptor kinase |
| Calcium-dependent protein kinase 3 | O81390_TOBAC | 0.0 | −0.5 | −0.7 | −0.5 | NM 001324640.1 AF072908.1 | |
| Beta-tubulin 9 | Q676U1_NICAT | 0.0 | 0.6 | 0.2 | 0.2 | NM 001325510.1 | |
| KP316400.1 | |||||||
| Alpha subunit of SnRK1 10 | M1LH79_9SOLN | 0.0 | 0.8 | 0.1 | 0.2 | XM 016633060.1 a | |
| Phospholipase D 1 | K4BAK2_SOLLC | 0.0 | 0.6 | −1.1 | −0.6 | XM 016601192.1 b | Ras-related protein RAB8-1 |
| Translation | |||||||
| Peptidyl-prolyl cis-trans isomerase 5 | B1PDK0_CAPAN | 0.0 | 0.4 | 0.6 | 0.2 | XM 016614927.1 b | |
| Transport | |||||||
| Clathrin heavy chain 1 | K4C1T2_SOLLC | 0.0 | −0.2 | −0.6 | −0.5 | XM 016641028.1 a | Clathrin heavy chain 1 |
| Plasma membrane ATPase 4 8 | PMA4_NICPL | 0.0 | 0.0 | −0.7 | −0.4 | XM 016619330.1 a | |
| Uncharacterised protein 2 | M1D6E0_SOLTU | 0.0 | −0.1 | −0.7 | −0.4 | XM 016602228.1 a | ATPase 2 |
| Pleiotropic drug resistance protein 1 8 | PDR1_NICPL | 0.0 | −0.2 | −1.1 | −0.5 | XM 016622655.1 a | |
| Clathrin heavy chain 1 | K4C5S4_SOLLC | 0.0 | −0.3 | −0.8 | −0.8 | XM 016613710.1 a | |
| Reticulon-like protein 2 | M0ZGB8_SOLTU | 0.0 | 0.1 | −0.8 | 0.0 | XM 016611959.1 b | |
| Uncharacterised protein 1 | K4CIM3_SOLLC (+2) | 0.0 | −0.4 | −0.9 | −1.0 | XM 016622951.1 b | Protein transport protein SEC16B |
| Basic Functionality Proteins | |||||||
| Glycine dehydrogenase (decarboxylating) 2 | GCSP_SOLTU (+2) | 0.0 | 0.4 | 0.5 | 0.5 | XM 016601634.1 a | |
| Aspartate aminotransferase 1 | K4CG60_SOLLC | 0.0 | 0.5 | 0.5 | 0.6 | XM 016642299.1 a | |
| Uncharacterised protein 1 | K4C412_SOLLC | 0.0 | −0.2 | −0.4 | −0.6 | XM 016617152.1 a | Fumarate hydratase 1 |
| Tobacco pre-pro-cysteine proteinase3 | Q43579_TOBAC (+2) | 0.0 | 0.1 | 0.0 | 0.3 | Z13959.1 | |
| Cytochrome c oxidase subunit 2 3 | Q5MA02_TOBAC | 0.0 | 0.1 | −0.7 | −0.5 | ||
| Mitochondrial small heat shock protein 5 | D9IAX1_CAPAN (+1) | 0.0 | −0.1 | −0.6 | 0.0 | ||
| Uncharacterised protein 2 | M0ZUV5_SOLTU | 0.0 | −0.2 | −0.8 | −0.4 | XM 016634781.1 b | Cytochrome P450 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
da Camara, N.; Dubery, I.A.; Piater, L.A. Proteome Analysis of Nicotiana tabacum Cells following Isonitrosoacetophenone Treatment Reveals Defence-Related Responses Associated with Priming. Plants 2023, 12, 1137. https://doi.org/10.3390/plants12051137
da Camara N, Dubery IA, Piater LA. Proteome Analysis of Nicotiana tabacum Cells following Isonitrosoacetophenone Treatment Reveals Defence-Related Responses Associated with Priming. Plants. 2023; 12(5):1137. https://doi.org/10.3390/plants12051137
Chicago/Turabian Styleda Camara, Nikita, Ian A. Dubery, and Lizelle A. Piater. 2023. "Proteome Analysis of Nicotiana tabacum Cells following Isonitrosoacetophenone Treatment Reveals Defence-Related Responses Associated with Priming" Plants 12, no. 5: 1137. https://doi.org/10.3390/plants12051137





