Abstract
The pharmacological properties of plants lie in the content of secondary metabolites that are classified into different categories based on their biosynthesis, structures, and functions. MicroRNAs (miRNAs) are small non-coding RNA molecules that play crucial post-transcriptional regulatory roles in plants, including development and stress-response signaling; however, information about their involvement in secondary metabolism is still limited. Cumin is one of the most popular seeds from the plant Cuminum cyminum, with extensive applications in herbal medicine and cooking; nevertheless, no previous studies focus on the miRNA profile of cumin. In this study, the miRNA profile of C. cyminum and its association with the biosynthesis of secondary metabolites were determined using NGS technology. The sequencing data yielded 10,956,054 distinct reads with lengths ranging from 16 to 40 nt, of which 349 miRNAs were found to be conserved and 39 to be novel miRNAs. Moreover, this work identified 1959 potential target genes for C. cyminum miRNAs. It is interesting to note that several conserved and novel miRNAs have been found to specifically target important terpenoid backbone, flavonoid biosynthesis, and lipid/fatty acid pathways enzymes. We believe this investigation will aid in elucidating the implications of miRNAs in plant secondary metabolism.
1. Introduction
Cuminum cyminum L. (Apiaceae), commonly known as cumin, is an annual, aromatic, herbaceous plant [1,2,3,4] native to Egypt, the Mediterranean, and the countries of South Asia. This plant has been widely grown in India, China, Turkey, Pakistan, Morocco, Egypt, Syria, Mexico, Iran, and Chile [5]. Cumin is a commercially important seed spice known for its distinct aromatic impact in a variety of culinary cultures around the world [4,5,6,7,8], pharmacological attributes in both traditional and current medicine [1,3,5,6], as well as in beverage, perfumery, alimentary [4,5], and beauty industries [2]. Asia, the Middle East, and North Africa have been identified as the largest producers and exporters of cumin seeds and products [4]. The world production of cumin in 2012 was approximately 300,000 tons, with approximately 70% of the production coming from India. The main import markets for cumin seeds are the United States (15.8%), Egypt (13%), Brazil (6%), the United Kingdom (5.6%), and Spain (5.5%) [5]. Dietary cumin provides high amounts of fat, protein, dietary fiber, vitamins B, C, E, and minerals, especially iron [5,6]. Moreover, C. cyminum contains a variety of bioactive compounds, the most prominent of which are alkaloids, flavonoids, and terpenoids [1,4]. Cuminaldehyde, cymene, cumin alcohol, and other terpenoids are the main volatile biomolecules found in C. cyminum essential oil [6,8]. Furthermore, cumin seeds are traditionally used to treat stomach aches, dyspepsia, indigestion, flatulence, toothache, hypertension, scorpion stings, obesity, jaundice, vomiting, chronic diarrhea, fever, ulcers, and cold in different countries [1,3,5,6].
MicroRNAs (miRNAs) are short non-coding RNA sequences (20–24 nucleotides) [9,10,11,12] found in animals, plants, and some viruses [12,13,14] that regulate gene expression at the post-transcriptional level [12,14] through mRNA inhibition. In plants, miRNAs are synthesized by RNA polymerase II (Pol II) as a primary transcript (pri-miRNA) and then processed by a DICER-LIKE endonuclease complex 1 (DCL1) to form the mature miRNA [9,10,11,13,14]. Interestingly, these tiny molecules are found in all major plant lineages and can be grouped into two classes: (a) highly expressed and evolutionarily conserved miRNAs and (b) non-conserved/species specific/novel miRNAs that expressed at low levels or induced under specific conditions [9]. Plant miRNAs largely control transcription factors and play an important role in plant growth and development, as well as abiotic and biotic stress response signaling [9,11,14,15]. Intriguingly, some of the previous research has also shown that a number of microRNAs are involved in secondary plant metabolism [9,14]. For instance, Barozai and colleagues (2013) identified 17 novel miRNAs in carrots, out of which five were involved in plant metabolism [16], while microRNAs miR5140, miR159, miR172, miR5303, miR5658, miR477, miR1426, miR530, and miR5079 from Withania somnifera (Indian ginseng) were found to be associated with the biosynthesis of steroid, stilbenoid, diarylheptanoid, gingerol, phenylpropanoid, isoquinoline, glycosyltransferases, ubiquinone, carotenoid, and other diterpenoids [17]. Similarly, miR4995, miR5021, and miR6300 from the medicinal plant Cinnamomum camphora are linked to the regulation of the biosynthesis of highly valuable terpenoids [18]. MicroRNAs from tree peony (Paeonia sp.) seeds such as miR414, miR156b, miR2673b, miR7826, and novel-m0027-5p were also reported to be associated with the fatty acids and lipid metabolism through post-transcriptional regulation [19]. In using high-throughput sequencing technology, six miRNAs (miRNAs miR164d, miR168b, novelmiRNA-108, novelmiRNA-23, novelmiRNA-58, and novelmiRNA-191) potentially involved in lipid biosynthesis were identified in medicinal plant Sea buckthorn (Hippophae L.). Recently, Gutiérrez-García et al. (2022) identified several miRNAs in the leaves of the medicinal plant Curry (Murraya koenigii), such as mko-miR5082, mko-miR396g-5p, mko-miRN7-3p, mko-miR827b, mko-mir156, mko-mir396c, mko-miRN4-5p, mko-miR168b, mko-miR858, mko-miR8610.1, and mko-miR5082. These are involved in the biosynthesis of terpenoids and flavonoids [20].
It has been suggested that artificial miRNAs (amiRNAs) can be used to manipulate target transcripts in order to increase secondary metabolite production in planta. However, a deeper comprehension of their mechanisms of action is necessary for utilizing them in the metabolic engineering process. Therefore, the current study aims to generate a secondary metabolism biosynthesis pathway-associated miRNA profile from C. cyminum using NGS technology.
2. Results
2.1. Sequence Analysis of C. cyminum Small RNAs
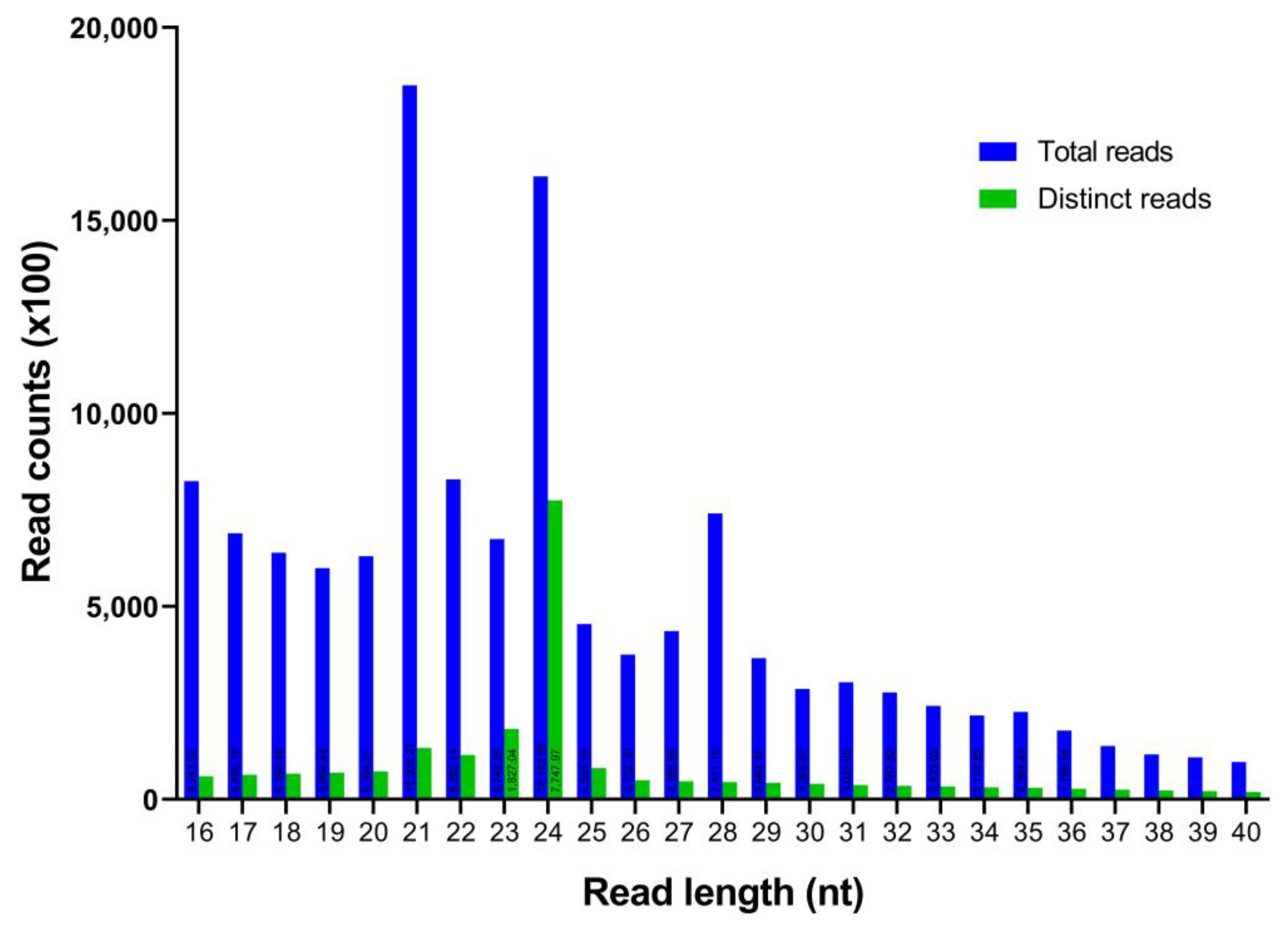
In this work, a total of 17,058,681 raw reads from the C. cyminum seedling samples were obtained (NCBI SRA accession number: SRP376517). Subsequently, the adaptors, low-quality reads, and other short RNAs like rRNA, snoRNA, snRNA, and tRNA were removed (Table 1), acquiring a total of 2,126,409 distinct reads with lengths ranging from 16 to 40 nt. According to the size distribution of unique C. cyminum reads, 24 nt was found to be the most prevalent one (36.44%), followed by 23 nt (8.59%), 21 nt (6.25%), and 22 nt (5.40%) (Figure 1).

Table 1.
Categorization of sequencing reads (srna-workbench, Bowtie, Mireap_0.22b, and UNAFold bioinformatic tools were used during the analysis).
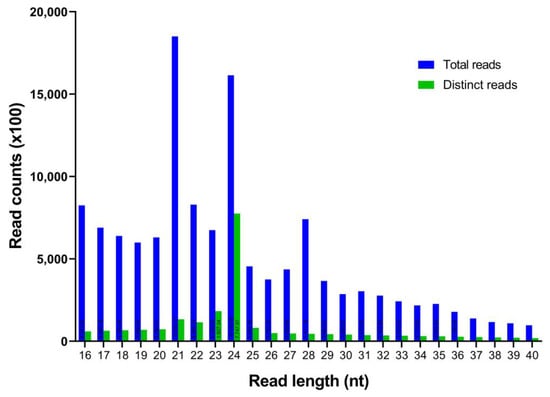
Figure 1.
Length distribution and abundance of the cumin small RNA sequences obtained by Illumina sequencing.
2.2. Identification of Conserved and Novel miRNAs in C. cyminum
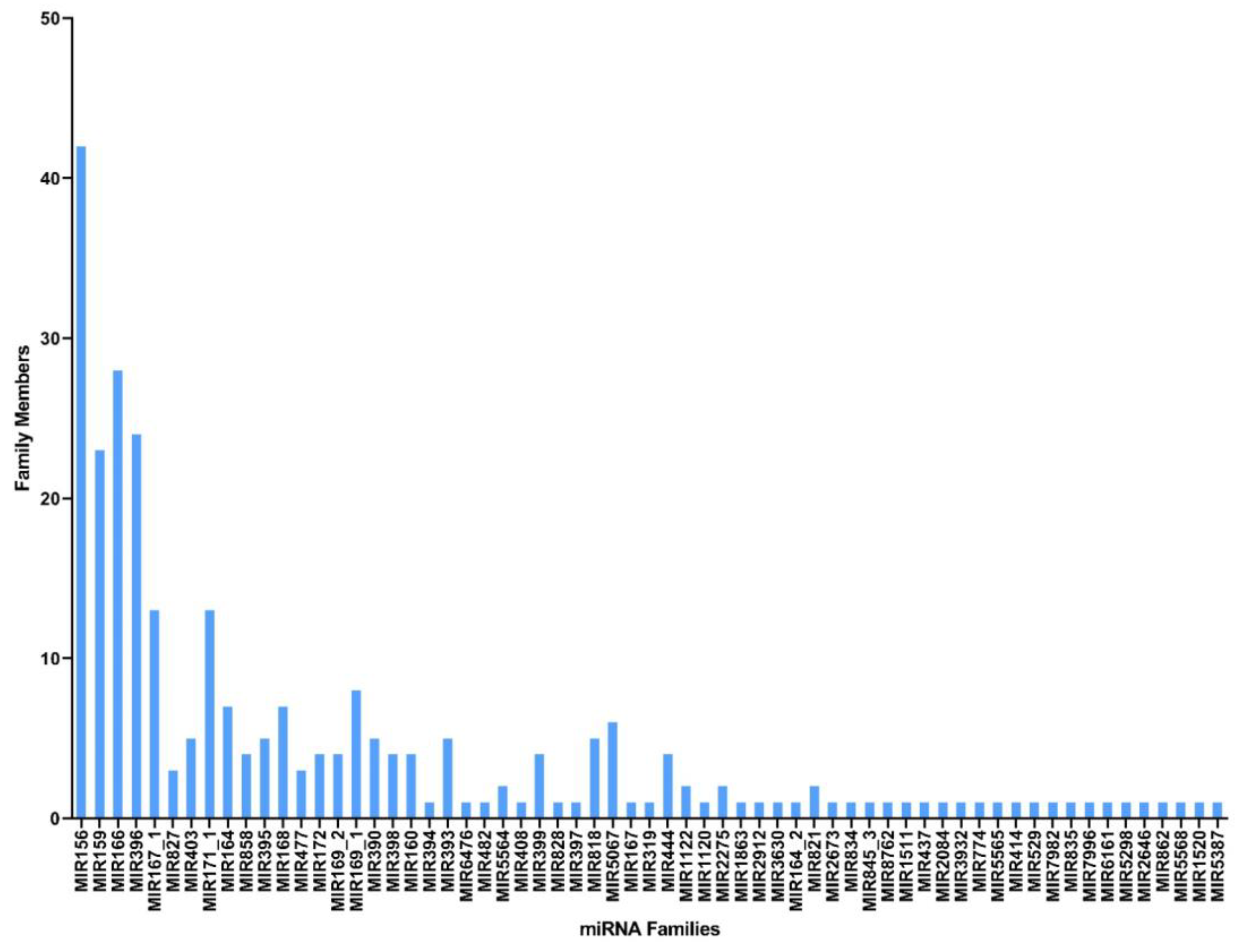
Unique reads obtained in this study were first matched against miRbase-22 (https://www.mirbase.org/ accessed on 10 January 2022) using the BLASTn program, and a total of 142 conserved cumin miRNAs spanning 63 miRNA families were identified (Table 2 shows the conserved miRNAs with the highest read counts for each family; for further info, please refer to Supplementary Materials). The frequency of the conserved miRNAs varied greatly between families, and all showed substantial homology with their respective homologs (maximum of single mismatch). Conserved miRNA data showed that the most abundant miRNA families were miR156, miR166, miR396, and miR159, with 42, 28, 24, and 23 members, respectively (Figure 2); moreover, the read counts of the miRNA families varied from 1 to 400,512, where the miR156 family had the highest number of reads (400,512), followed by the miR159 (184,400) and miR166 (144,536) families (Supplementary Table S1).

Table 2.
Summary of conserved miRNAs in C. cyminum.
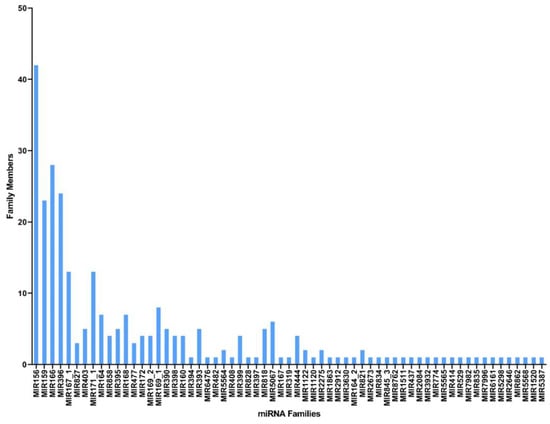
Figure 2.
miRNA families with the corresponding number of members found in C. cyminum.
Following the identification of conserved miRNAs, the remaining unaligned 1,057,102 reads were subjected to predict novel C. cyminum miRNA candidates by applying strict filtering criteria. A total of 39 novel cumin miRNAs were identified in this study (Table 3), and their secondary hairpin structures were deduced (Figure 3). The read counts of these novel miRNA candidates varied from 2 to 1382. The cci-miRN10-3p displayed the highest number of reads (1382), followed by cci-miRN2-3p (975) and cci-miRN20-5p (262). The precursor sequences of these novel miRNA candidates had high Minimum Folding Free Energy Index (MFEI) values ranging from 0.70 to 1.41 with an average of 0.98 ± 0.14, distinguishing them from other types of RNAs such as tRNAs (0.64), rRNAs (0.59), and mRNAs (0.62–0.66) [21].

Table 3.
Potential novel miRNA candidates identified from C. cyminum.

Figure 3.
Secondary structures (stem-loops) of C. cyminum novel miRNA precursors. Mature miRNAs are highlighted in red font.
2.3. Target Prediction of Conserved and Novel C. cyminum miRNAs and Their Functional Analysis
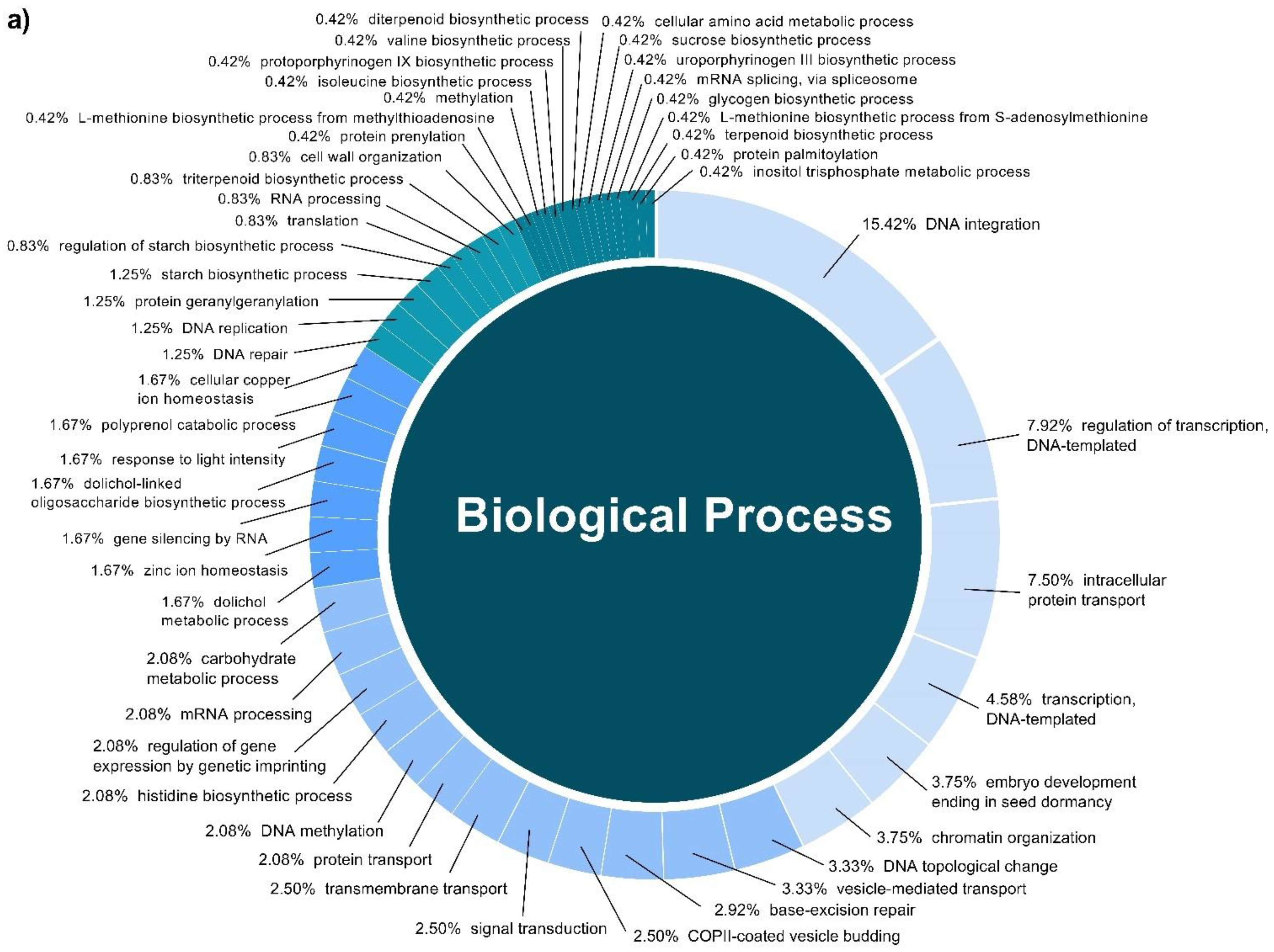
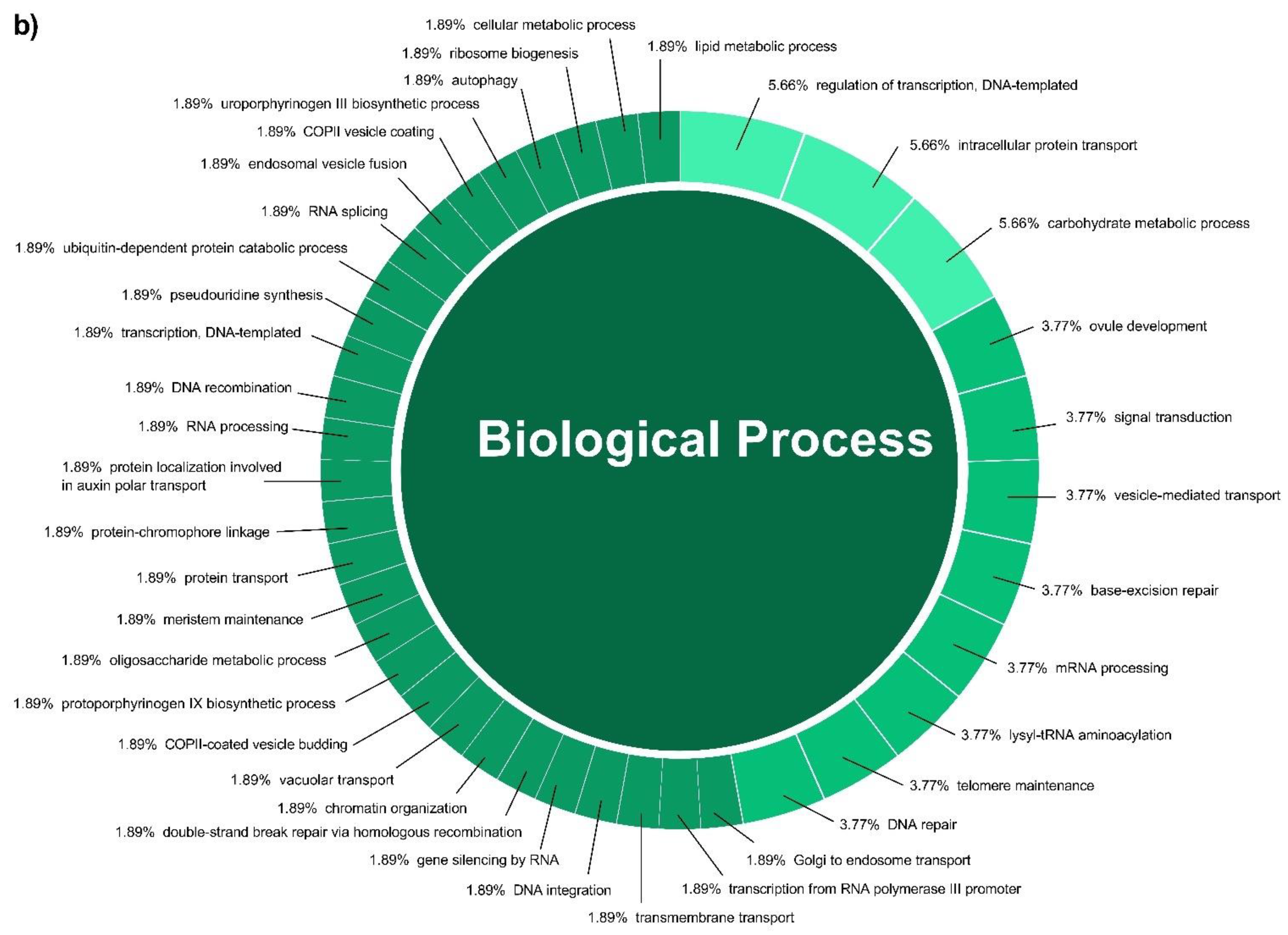
In this study, a total of 3037 putative target genes for C. cyminum miRNAs (1959 target genes for conserved miRNAs and 1078 target genes for novel miRNAs) were identified, among which cci-miR156b-3p (240) had the highest possible number of targets among conserved miRNAs, followed by cci-miR156d-3p (239) and cci-miR156f-3p (218), whereas cci-miRN10-3p (274) targeted the most number of transcripts among novel miRNAs. The Gene Ontology (GO) analysis showed that the top terms in the Biological Process (BP) category for the targets of conserved miRNAs were “DNA integration” (15.42%), “regulation of transcription, DNA-templated” (7.92%), and “intracellular protein transport” (7.50%); moreover, “histidine biosynthetic process” (2.08%), “dolichol metabolic process” (1.67%), “dolichol-linked oligosaccharides biosynthetic process” (1.67%), and “triterpenoid biosynthetic process” (0.83%) were key terms related to secondary metabolism biosynthesis. While “regulation of transcription, DNA-templated” (5.66%), “intracellular protein transport”, and “carbohydrate metabolic process” (5.66%) were the three main terms for the targets of novel miRNAs in the BP category and, in particular, “protoporphyrinogen IX biosynthetic process” (0.66%) is involved in the biosynthesis of secondary metabolites (Figure 4). Furthermore, gene network analysis revealed the coregulation of numerous target genes (Figure 5).
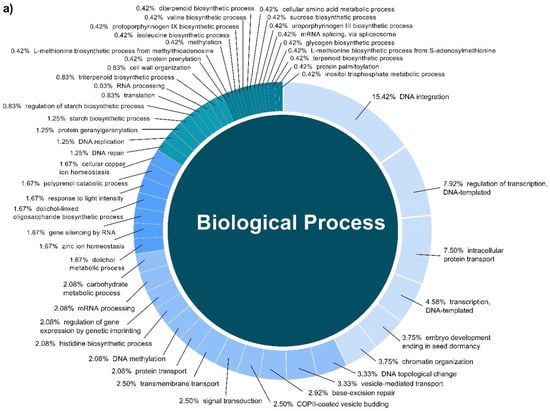
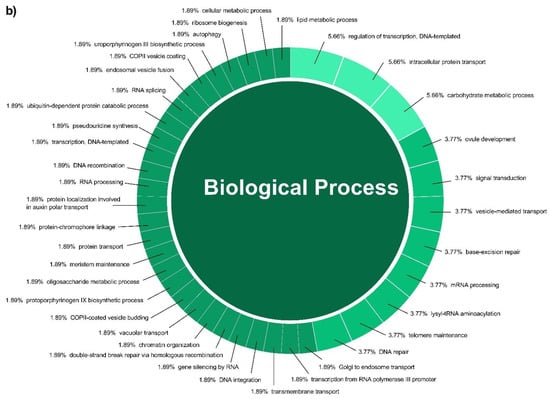
Figure 4.
Biological Process categories of the GO analysis of both conserved (a) and novel (b) C. cyminum miRNAs’ putative target genes.
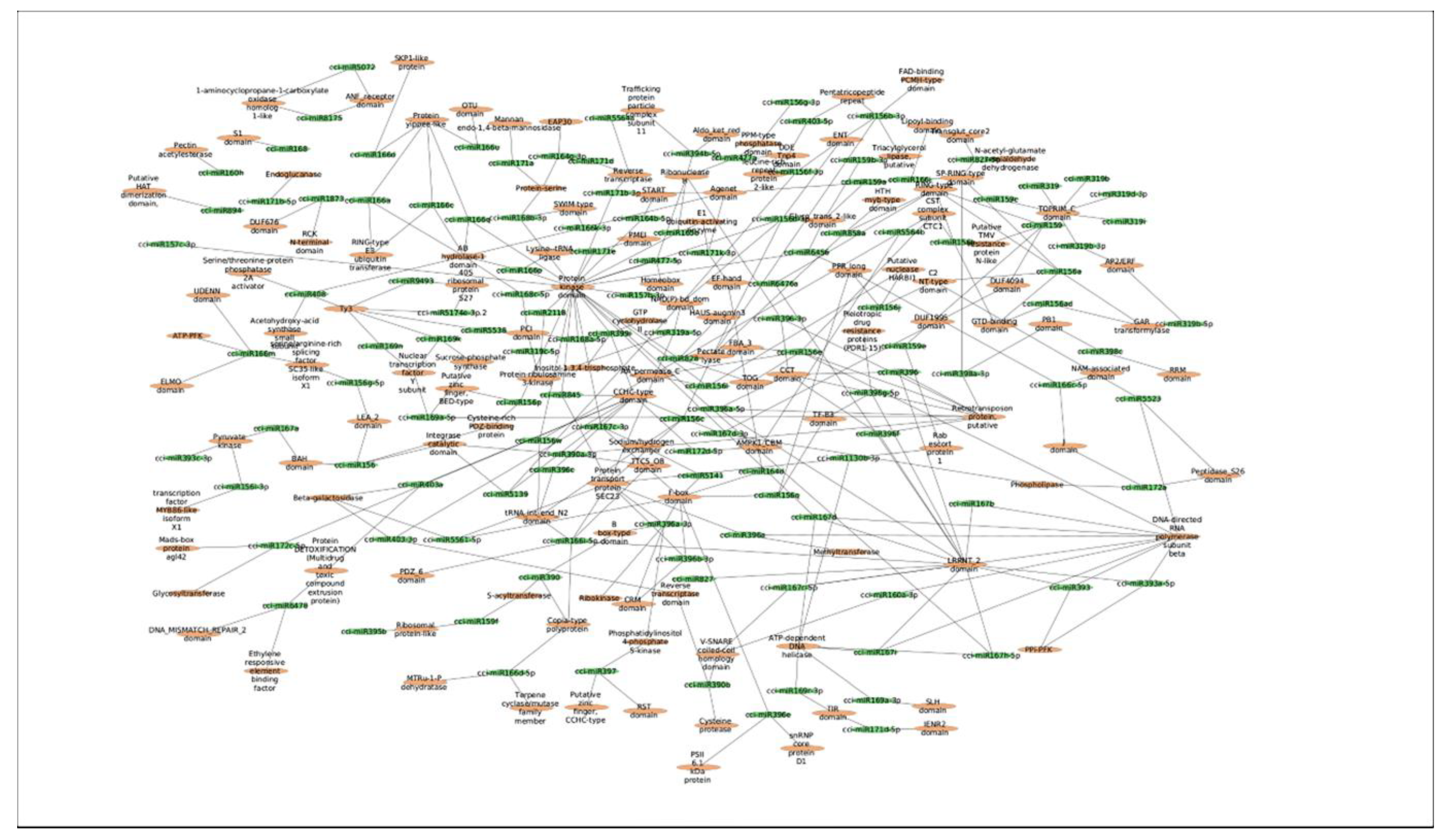
Figure 5.
MFE-based network interaction of C. cyminum miRNAs and their corresponding potential targets.
2.4. Identification of C. cyminum miRNA Targets Involved in Plant Secondary Metabolite Biosynthesis
Secondary metabolites produced by particular enzymes are thought to be responsible for C. cyminum’s pharmacological activities. Thus, as stated above, the main objective of the study is the identification of C. cyminum miRNAs that are involved in the biosynthesis of secondary metabolites as they are associated with target genes that encode the enzymes involved in different metabolic and biosynthesis pathways. Data analysis revealed that a total of 70 C. cyminum miRNA target genes are associated with the biosynthesis of steroids, diterpenoids, sesquiterpenoids, triterpenoids, terpenoids, ubiquinones and other terpenoid-quinones; of which fourteen conserved miRNAs (cci-miR156q, cci-miR157b-3p, cci-miR159, cci-miR159c, cci-miR166d-5p, cci-miR172c-5p, cci-miR172d-5p, cci-miR319d-3p, cci-miR394b-5p, cci-miR396a-3p, cci-miR397, cci-miR2118, cci-miR5564b, and cci-miR6476a) denoted control over key terpene-regulating enzymes (Cycloartenol synthase, homogentisate phytyltransferase, homogentisate geranylgeranyltransferase, and ent-kaurene synthase) within the plant metabolism, leading to the biosynthesis of different terpenoids shown in Figure 6, Figure 7 and Figure 8.
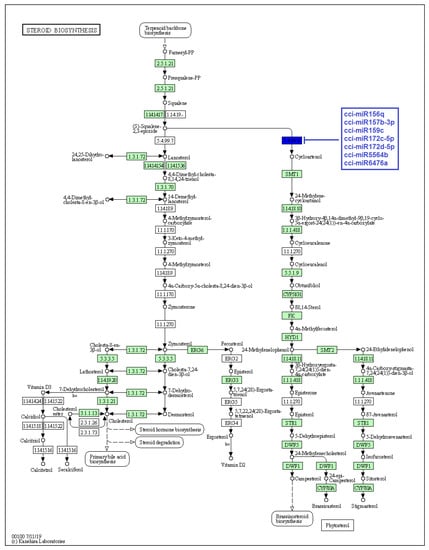
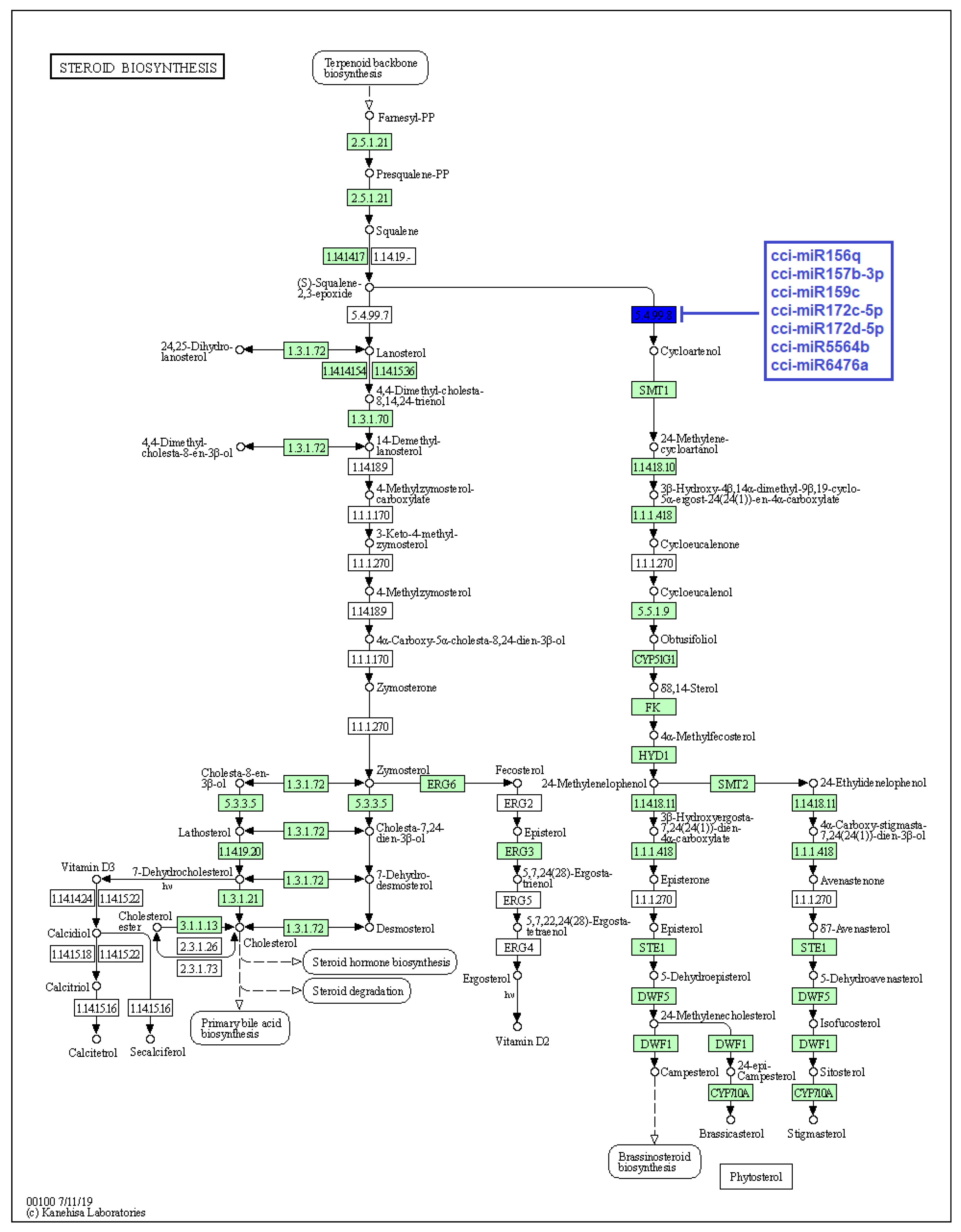
Figure 6.
Target enzymes of conserved miRNAs of C. cyminum in the steroid biosynthesis via terpenoid backbone biosynthesis pathway. EC:5.4.99.8—Cycloartenol synthase. The blue box represents the targeted enzyme of the corresponding known miRNAs.
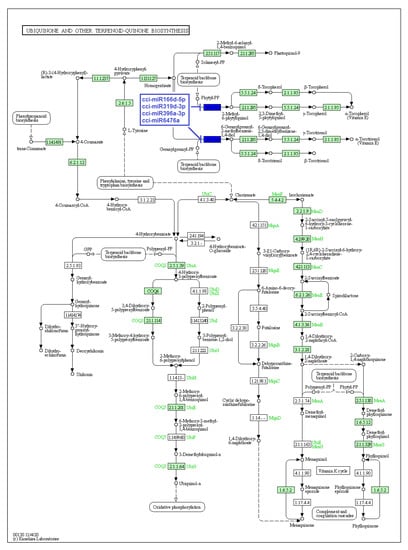
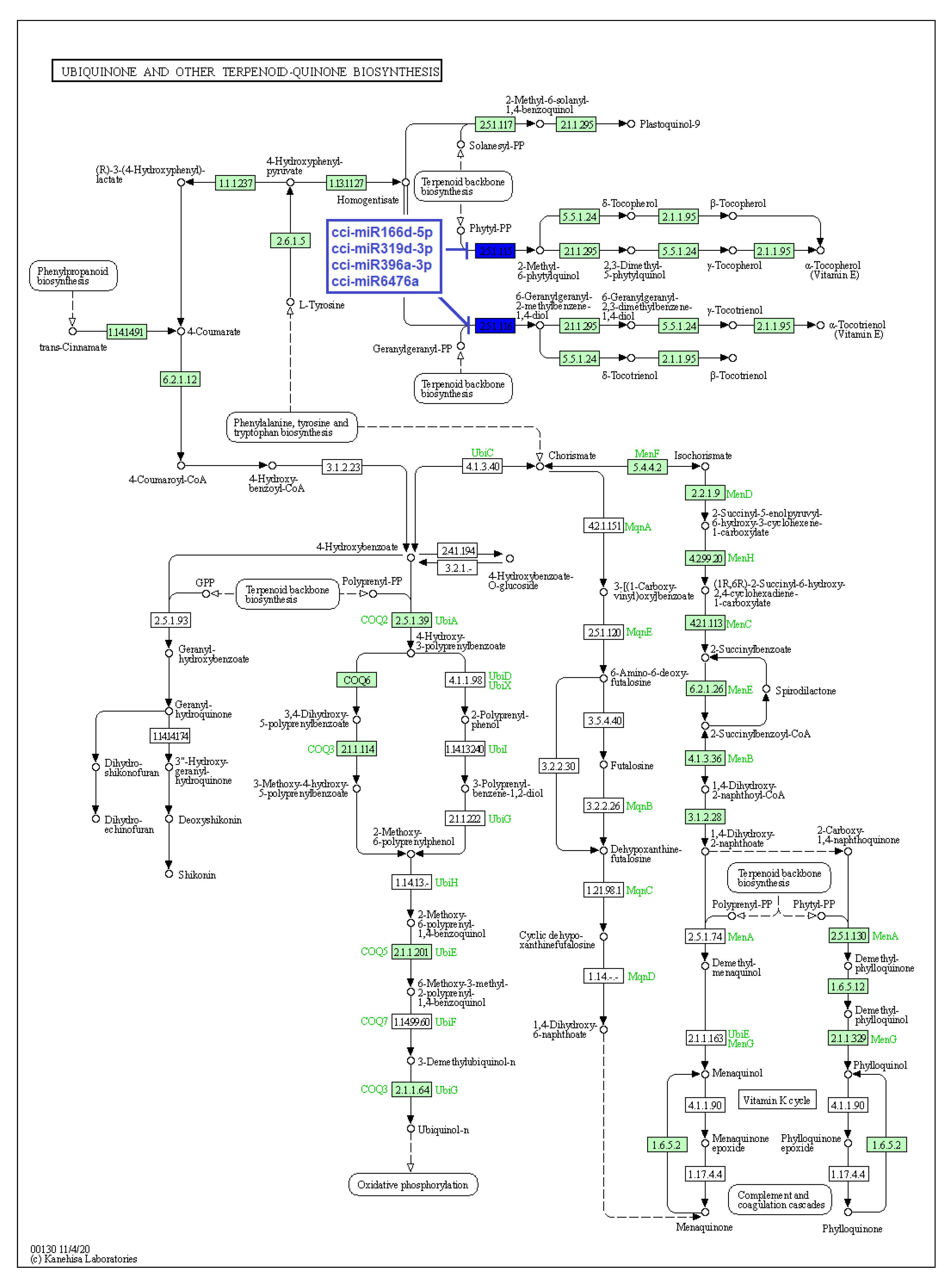
Figure 7.
Target enzymes of conserved miRNAs of C. cyminum in the ubiquinone and other terpenoid-quinone biosynthesis pathways. EC:2.5.1.115—Homogentisate phytyltransferase; EC: 2.5.1.116—Homogentisate genarylgeranyltransferase. The blue boxes represent the targeted enzymes of the corresponding known miRNAs.
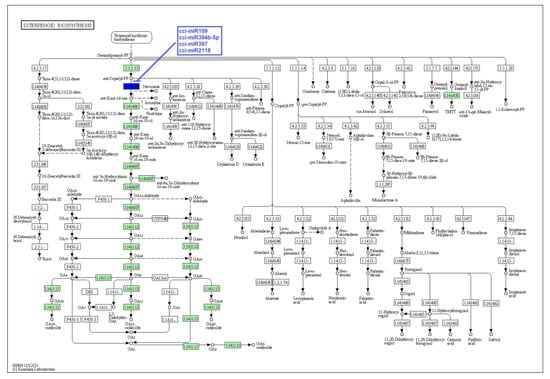
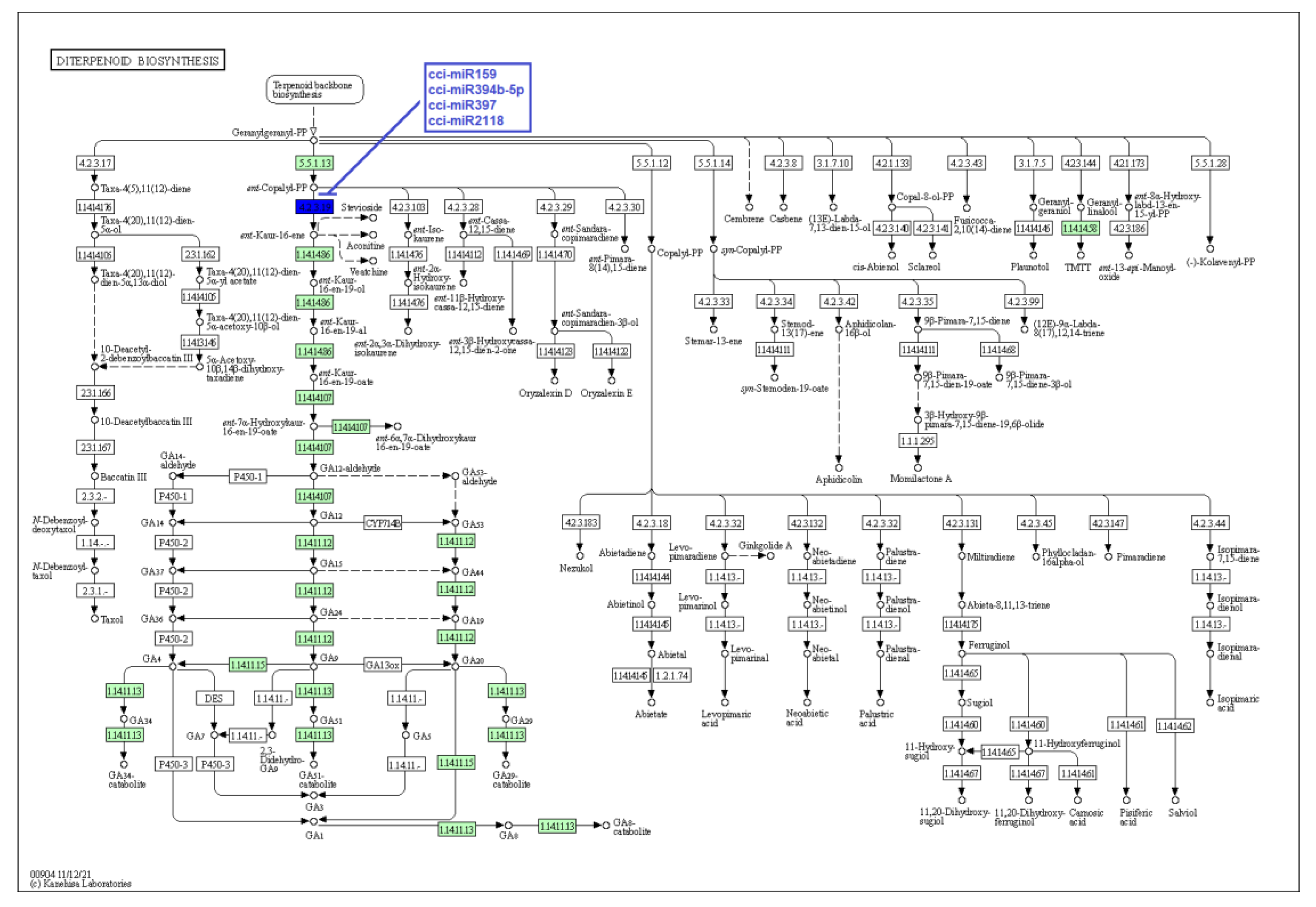
Figure 8.
Target enzymes of conserved miRNAs of C. cyminum in the diterpenoid biosynthesis pathway. EC:4.2.3.19—Ent-kaurene synthase. The blue box represents the targeted enzyme of the corresponding known miRNAs.
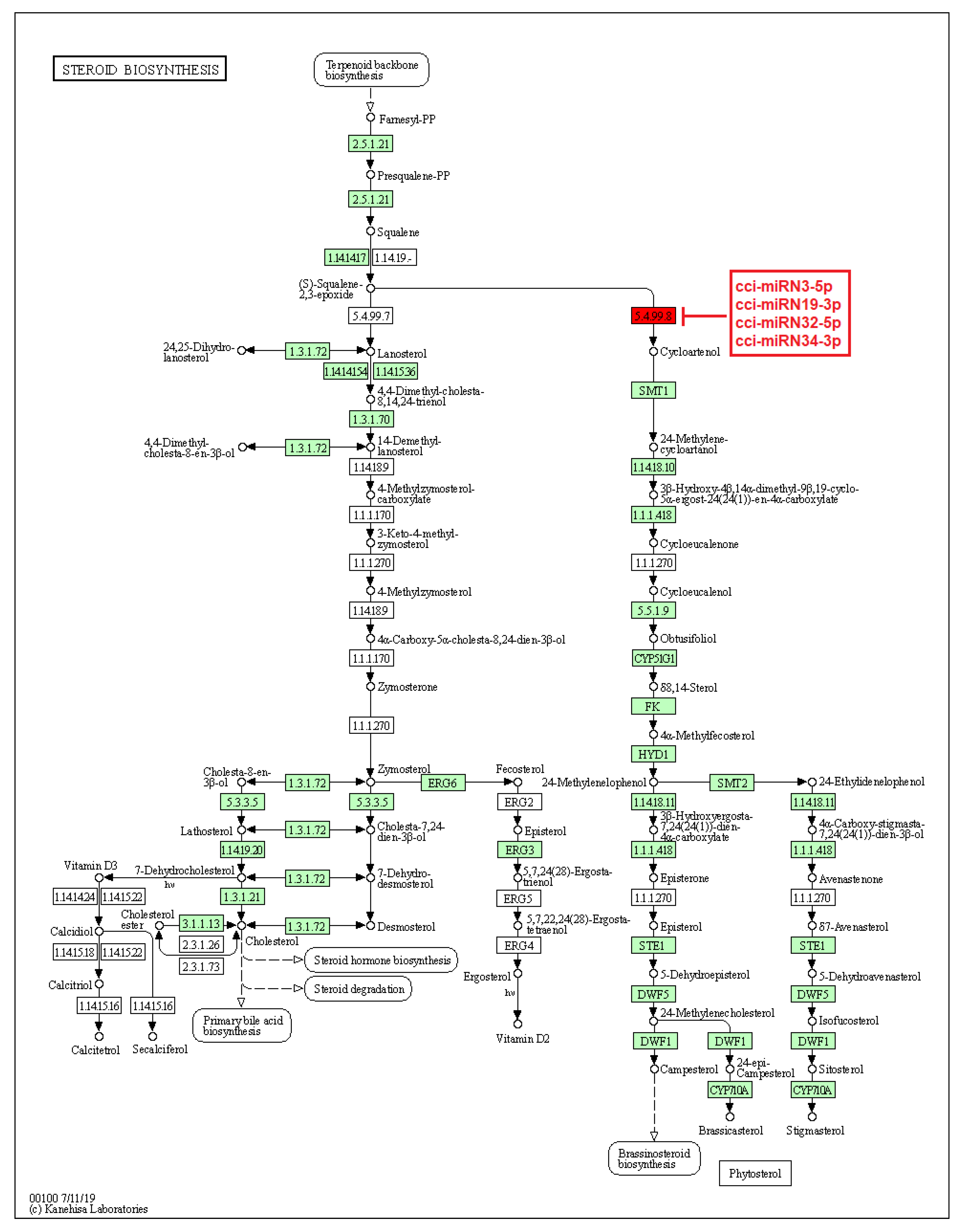
On the other hand, a few identified novel C. cyminum miRNAs were also found to be associated with secondary metabolite biosynthetic pathways, including the biosynthesis of stilbenoid, diarylheptanoid, gingerol, phenylpropanoid, flavonoids, diterpenoids, terpenoids, sesquiterpenoids, and triterpenoids. Four of the novel miRNAs (cci-miRN3-5p, cci-miRN19-3p, cci-miRN32-5p, and cci-miRN34-3p) were considerably linked with the enzyme cycloartenol synthase within the terpenoid backbone biosynthesis pathway (Figure 9), while novel miRNA cci-miRN2-3p was revealed to be correlated with the enzyme beta-carotene 3-hydroxylase (Figure 10) associated with the production of B-end group carotenoids.
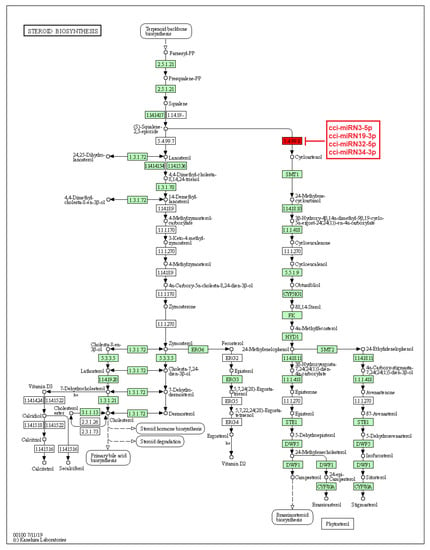
Figure 9.
Target enzymes of novel miRNAs of C. cyminum in the steroid biosynthesis via terpenoid backbone biosynthesis pathway. EC:5.4.99.8—Cycloartenol synthase. The red box represents the targeted enzyme of the corresponding novel miRNAs.
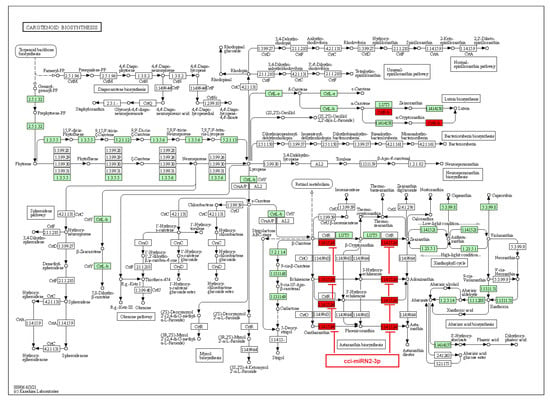
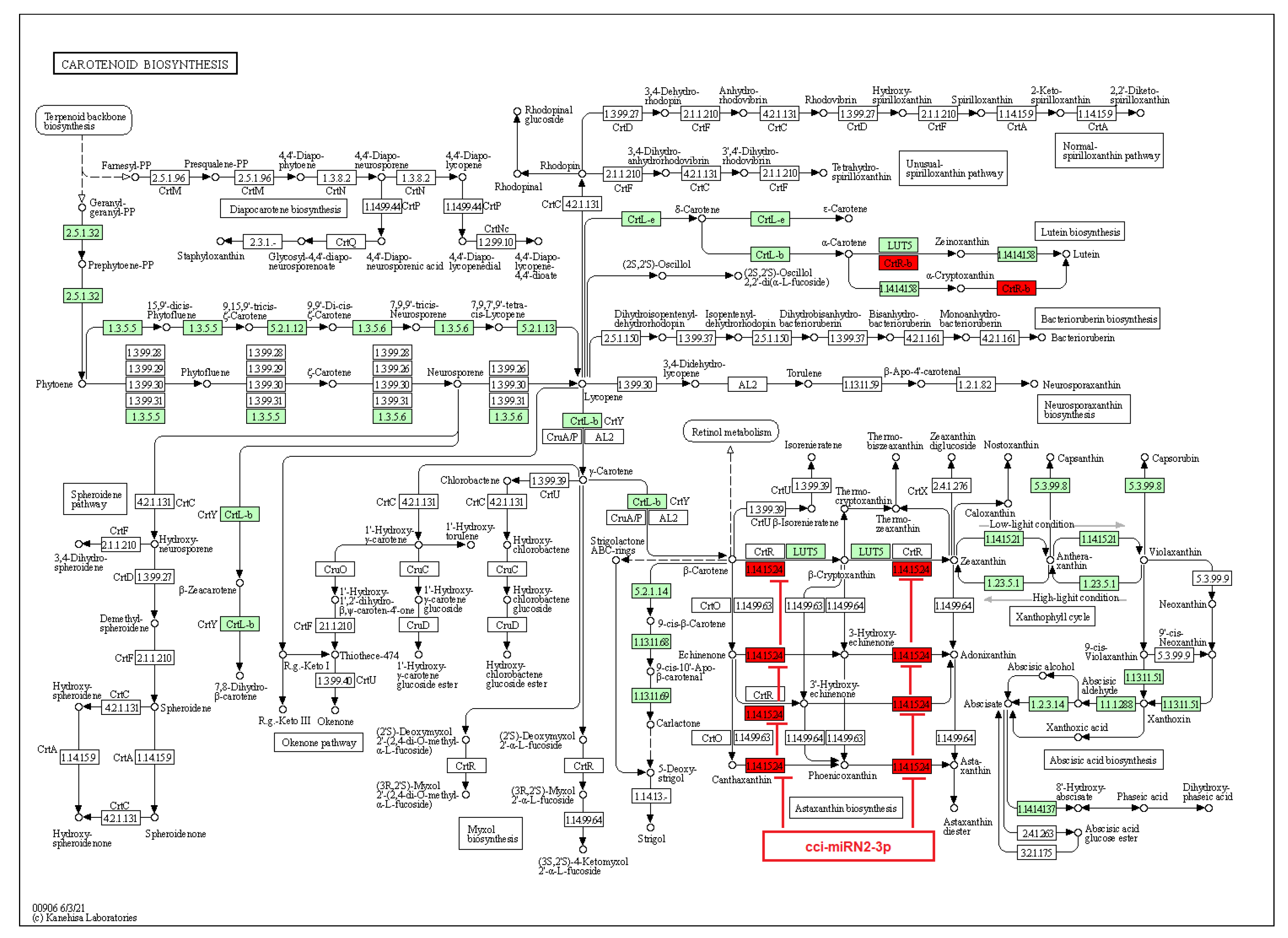
Figure 10.
Target enzymes of novel miRNAs of C. cyminum in the carotenoid biosynthesis pathway. EC:1.14.15.24—Beta-carotene 3-hydroxylase. The red boxes represent the targeted enzymes of the corresponding novel miRNAs.
2.5. Experimental Validation of C. cyminum miRNAs by qPCR
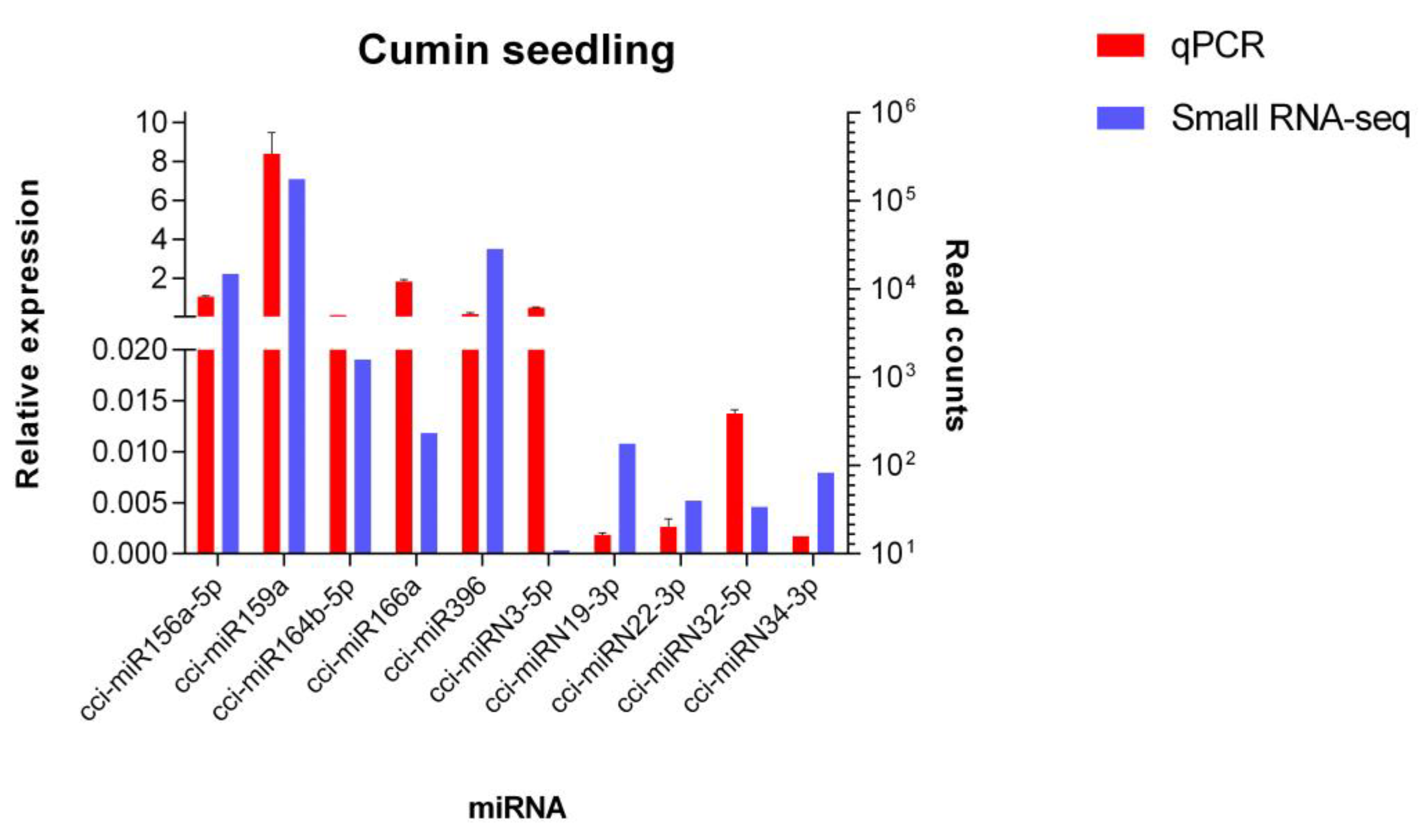
In order to validate the data obtained by high-throughput sequencing, a qPCR analysis was performed. Five conserved and five novel C. cyminum miRNAs were randomly selected for the experiments. The results showed similar relative expression patterns with respect to the read counts for all miRNAs analyzed (Figure 11).
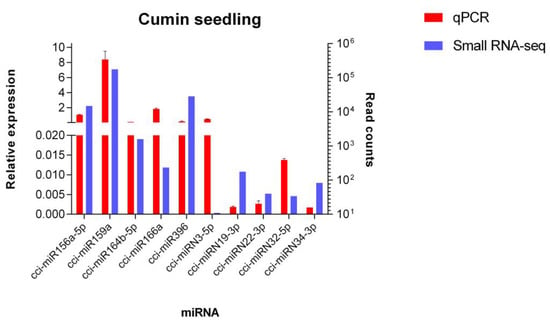
Figure 11.
Quantitative PCR analysis of novel and conserved miRNAs from C. cyminum. The relative expression of five conserved miRNAs (cci-miR156a-5p, cci-miR159a, cci-miR164b-5p, cci-miR166a, and cci-miR396) and five novel miRNAs (cci-miRN3-5p, cci-miRN19-3p, cci-miRN22-3p, cci-miRN32-5p, and cci-miRN34-3p) was detected by qPCR. U6 was used as an endogenous control. The analysis was performed in triplicate, and the error bars indicate the standard deviations.
3. Discussion
Secondary metabolites are synthesized by plants to adapt and respond to the environment’s biotic and abiotic stressors [22]. Moreover, these secondary metabolites have multiple applications in a number of industries that aroused commercial interest and a field of study to explore overproduction alternatives [22]. MiRNAs are short non-coding RNAs that control the plant’s secondary metabolism by targeting the mRNA of crucial enzymes involved in the biosynthetic pathways [11]. Therefore, it becomes imperative to identify the miRNAs and their targets in unexplored medicinal plant species. The present study investigated the miRNA profile of cumin (C. cyminum) by Illumina small RNA sequencing technology to better understand their involvement in secondary metabolism.
It is well established that most of the plant miRNA families are highly conserved in the plant kingdom, while some others are specific to certain species (novel miRNAs). The degree of conservation of a miRNA indicates its evolutionary status and is closely related to its expression patterns and functions [23]. In this study, the conserved C. cyminum miRNAs from young cumin seedlings (15-day-old seedlings) were distributed in 63 families, from which miR156 was the most persistent family with 42 members and 400,512 read counts. Liu et al. (2018) reported that miR156 controls the phase transition in plant development and accumulates during the juvenile stage, decreasing when the plant ages, which is consistent with our results of cumin miRNAs from the seedling stage [23].
The pharmacological potential of C. cyminum is attributed to the presence of bioactive compounds such as terpenoids, phenols, and flavonoids [24]. Several studies reported that cuminaldehyde is the most prominent chemical compound in C. cyminum seeds, followed by cuminic alcohol, γ-terpinene, p-cymene, and β-pinene [25,26]. Therefore, terpenoids play a vital role in cumin’s secondary metabolism. Terpenoids backbone is produced through successive condensations of isopentenyl diphosphate (IPP) and its isomer, dimethylallyl diphosphate (DMAPP). After different enzymatic steps, the backbone is arranged and modified to give rise to a morphological and functional diversity of terpenoids [27,28]. Specifically, terpenoids can be classified depending on the number of carbons (or isoprenoid units, C5H8) in C10 monoterpenes, C15 sesquiterpenes, C20 diterpenes, C30 triterpenes, and C40 tetraterpenes. Particularly, C30 triterpenoids contain a carbon skeleton of four or five rings of varied configurations, while steroids are tetracyclic compounds produced from terpenoid precursors and contain a perhydro-1,2-cyclopentane-phenanthrene moiety [29]. For both triterpenoids and steroids, a common step is needed in their biosynthesis pathway, which consists of the cyclization of squalene (C30 compound) into cycloartenol by the enzyme cycloartenol synthase [27]. This enzyme belongs to the oxidosqualene cyclase gene family [30]. Interestingly, the present study revealed that cycloartenol synthase is targeted by the conserved cumin miRNAs cci-miR156q, cci-miR157b-3p, cci-miR159c, cci-miR172c-5p, cci-miR172d-5p, cci-miR5564b, and cci-miR6476a and the novel cumin miRNAs cci-miRN3-5p, cci-miRN19-3p, cci-miRN32-5p, and cci-miRN34-5p. It suggests these cumin miRNAs can target a key regulator enzyme in both sterol and triterpenoid production due to squalene cyclization being a crucial branch point in their biosynthesis pathway [30]. Our results coincide with the findings of Srivastava et al. (2018), who reported that miR5140 and miR159 families were involved in the gene regulation of cycloartenol synthase and sterol delta-7 reductase 1, which are directly involved in the biosynthesis of withanolides, a group of steroids produced by the medicinal plant Withania somnifera [17]. Similarly, Jeena et al. (2021) identified the cleavage site of cycloartenol synthase by Bm-miR172c-5p from the medicinal plant Bacopa monnieri and suggested that this miRNA could be a key regulator in the plant response to abiotic stress [31].
Additionally, our study showed that the conserved cumin miRNAs cci-miR166d-5p, cci-miR319d-3p, cci-miR396a-3p, and cci-miR6476a target both enzymes homogentisate phytyltransferase and homogentisate genarylgeranyltransferase. These enzymes are responsible for condensing homogentisic acid to phytyl-pyrophosphate (Phytyl-PP) and geranylgeranyl pyrophosphate (GGPP) for later production of tocopherols and tocotrienols in the Ubiquinone and Other Terpenoid-Quinone Biosynthetic Pathway. Importantly, these enzymes perform the first committed reaction in their biosynthesis due to an irreversible enzymatic reaction in which the product must continue through the pathway [32]. Our data are consistent with the report of Zheng et al. (2019), in which several miRNAs of soybean (Glycine max) targeted genes of the same biosynthetic pathway under biotic stress [33].
Moreover, in this investigation, we showed that the conserved cumin miRNAs cci-miR159, cci-miR394b-5p, cci-miR397, and cci-miR2118 target the enzyme ent-kaurene synthase, responsible for producing ent-Kaur-16-ene from ent-copalyl diphosphate, playing a key bifunctional role in the biosynthesis of phytohormones and terpenoids [34,35]. Additionally, this enzyme has been conserved in all land plants and reported in higher concentrations in rapidly dividing plant tissues [36]. Likewise, Barozai et al. (2011) found in a computational study that the miRNA families miR158 and miR393 from Picea glauca and Picea sitchensis targeted ent-kaurene synthase [37].
Regarding carotenoids, they are C40 tetraterpenoids that play an important role in the physical condition of plants and are biomolecules of great commercial interest due to their different applications in fragrances, food, cosmetics, and biofuels [38]. In addition, they have antioxidant, anti-inflammatory, and anti-allergic properties [39]. Carotenoids are made by two C20 moieties of GGPP, linked together at their tails to give up a C40 linear hydrocarbon skeleton adaptable to different structural changes [39]. Each GGPP moiety is produced by the linear sequential addition of three IPP molecules to one DMAPP molecule [39,40,41]. We noticed that the novel miRNA cci-miRN2-3p targets the enzyme beta-carotene 3-hydroxylase, which plays a vital role in several sequential steps for the synthesis of astaxanthins. Astaxanthins are carotenoids with high pharmaceutical importance synthesized by ketolase and beta-carotene 3-hydroxylase enzymes from β-carotene [42]. Importantly, beta-carotene 3-hydroxylase and other related enzymes are responsible for rate-limiting steps in β-carotene catalysis [43]. Therefore, the cumin novel miRNA cci-miRN2-3p could represent a key regulator in astaxanthins biosynthesis. Similarly, another study of potato seedling miRNAs revealed that the novel miRNA stu-novel-miR5125 targeted beta-carotene 3-hydroxylase, which could be upregulated by low-temperature stress [44]. Furthermore, a miRNA regulatory network from Cryptomeria fortunei, a medicinal plant from China, indicated that several miRNAs, including pde-miR159, mdm-miR396a, and hbr-miR6173 were related to the carotenoid biosynthesis [45].
Finally, the qPCR experiment to validate the sequencing analysis was done with five conserved and five novel C. cyminum miRNAs. The results demonstrated that both the Illumina sequencing and the qPCR assays displayed similar expression patterns. This behavior was also observed in the leaves of the medicinal plant C. camphora [18], 15-day-old seedlings of wild-type A. thaliana [46], and 15-day-old seedlings of Tibetan barley (Hordeum vulgare L. var. nudum) [47].
4. Materials and Methods
4.1. Plant Materials, RNA Extraction and Quality Control
C. cyminum seeds were collected from a local supermarket and washed thoroughly with 2% sodium hypochlorite and colloidal silver (10 drops of Microdacyn® in 100 mL of distilled water). For seed germination, Petri dishes were prepared with sterile filter paper, 2 mL of MS 1/2 medium, and a thin layer of cumin seeds, which were left incubating at 25 °C for 15 days with a 12-h light/dark photoperiod. Seedling samples of healthy 15-days-old-C. cyminum were collected and instantly frozen in liquid nitrogen. Total RNA (including small RNA) was isolated from 100 mg of the seedling samples using the Direct-zol RNA micro kit (Zymo Research, Irvine, CA, USA) following the manufacturer’s instructions. RNA concentration and purity were quantified utilizing a Nanodrop2000 Spectrophotometer (Thermo Scientific, Wilmington, DE, USA). The integrity of the samples was assessed on Tapestation (Agilent, Santa Clara, CA, USA).
4.2. Small RNA Library Construction and Sequencing
A single small RNA sequencing library from pooled RNA samples (3 biological replicates) was prepared using QIAseq® miRNA Library Kit (Qiagen, Germantown, MD, USA). Briefly, 63 ng of total RNA was used as starting material, and 3′ adapters were ligated to the specific 3′OH group of miRNAs, followed by ligation of 5′ adapter. Subsequently, the adapter-ligated fragments were reverse transcribed with Unique Molecular Index (UMI) assignment, and the cDNA was enriched and barcoded by PCR amplification. The resultant cDNA library was then quantified by Qubit fluorometer (Thermo Fisher Scientific, Waltham, MA, USA), and the fragment size distribution of the library was evaluated on Agilent 2200 TapeStation system. Finally, high-throughput single-ended sequencing was performed using an Illumina NovaSeq 6000 for 75 cycles, following the manufacturer’s instructions.
4.3. Small RNA Sequencing Data Analysis
Illumina GA raw data was processed by srna-workbench (V3.0_ALPHA). For this step, 3′ adaptors and low-quality reads (<Q30) were removed, and reads matching other ncRNAs such as rRNAs, tRNAs, snRNAs, and snoRNAa, as well as sequences smaller than 16 bp and larger than 40 bp were eliminated. The remaining small RNA sequences were aligned against miRbase22 (http://www.mirbase.org, accessed on 10 January 2022) in order to identify conserved C. cyminum miRNAs. Those sequences that did not show homology were considered for the prediction of novel miRNAs using bowtie (https://bowtie-bio.sourceforge.net/index.shtml, accessed on 10 January 2022) and Mireap_0.22b [48]. Conserved miRNAs were identified by the homology approach against mature Viridiplantae miRNA sequences due to the unavailability of the C. cyminum genome. In this study, only novel miRNAs with suitable precursor secondary structures and with MFEI values ≥ 0.70 were considered. The secondary structures of the precursors were predicted using the UNAFold web server (http://www.unafold.org, evaluated on 16–17 June 2022), and the MFEI values were calculated using the following formula:
4.4. Prediction of C. cyminum miRNA Targets, Their Functional Annotation, and Pathway Analysis
The psRNATarget tool (https://www.zhaolab.org/psRNATarget/, accessed on 10 January 2022) was used to predict the target genes of both novel and conserved miRNAs with copy numbers equal to or greater than five. Potential Cuminum cyminum miRNA targets were annotated using the Gene Ontology (GO) in the bioprocesses (BP) domain. Additionally, a biological network of the miRNAs and their targets was built using the MFE values of the miRNA-target interaction and visualized using Cytoscape 3.2 (https://cytoscape.org/releasenotes320.html, accessed on 10 January 2022) in order to identify the coregulation of possible targets. Finally, a thorough analysis of miRNA targets linked to pathways for the biosynthesis of secondary metabolites was performed.
4.5. Extraction of Small RNA and Experimental Validation of C. cyminum miRNAs by qPCR
In order to validate the identified conserved and novel miRNAs, small RNA was extracted from frozen C. cyminum seedlings using a mini miRNeasy kit (Qiagen, Maryland, USA) following the manufacturer’s instructions. Quality and quantity of small RNA samples were measured using NanoDrop One UV–Vis microvolume spectrophotometer (Thermo Scientific, Wilmington, DE, USA). Resulting small RNAs were then polyadenylated and reverse transcribed using the Mir-X miRNA First-Strand Synthesis kit (Clontech, Mountain View, CA, USA), and finally, the qPCR was performed using the TB Green® Advantage® qPCR Premix (Takara Bio USA, Inc., San José, CA, USA) in a StepOne™ Real-Time PCR System (Applied Biosystems, Carlsbad, CA, USA). The primers for cumin novel miRNAs were listed in Supplementary Table S2. The reactions were performed in a 48-well optical plate with the following conditions: initial polymerase activation step for 10 s at 95 °C, denaturation by 45 cycles of 5 s at 95 °C, annealing and extension for 20 s at 55 °C. Following the amplification cycle, a melting curve analysis with temperature ranges of 56–95 °C and increments of 0.5 °C every 10 s was performed. For each sample, the reactions were run with three technical and six biological replicates, and the relative expression of the miRNAs was normalized to U6, the endogenous control.
5. Conclusions
In this study, a total of 353 conserved and 39 novel cumin miRNAs were identified using Illumina high-throughput sequencing technology. Among the identified miRNAs, fourteen conserved miRNAs (cci-miR156q, cci-miR157b-3p, cci-miR159, cci-miR159c, cci-miR166d-5p, cci-miR172c-5p, cci-miR172d-5p, cci-miR319d-3p, cci-miR394b-5p, cci-miR396a-3p, cci-miR397, cci-miR2118, cci-miR5564b, and cci-miR6476a) and four novel miRNAs (cci-miRN3-5p, cci-miRN19-3p, cci-miRN32-5p, and cci-miRN34-3p) were found to target enzymes that were significantly involved in the terpenoid backbone biosynthesis pathway, while one novel miRNA was discovered to target the beta-carotene 3-hydroxylase enzyme connected to the carotenoid biosynthesis pathway. A total of ten random miRNAs (cci-miR156a-5p, cci-miR159a, cci-miR164b-5p, cci-miR166a, cci-miR396, cci-miRN3-5p, cci-miRN19-3p, cci-miRN22-3p, cci-miRN32-5p, and cci-miRN34-3p) were experimentally verified using qPCR. To the best of our knowledge, this is the first report of microRNA profile from the culinary and medicinal plant C. cyminum and its relationship with the biosynthesis of secondary metabolites. This study could support miRNA-mediated transgenic research for the overproduction of valuable plant secondary metabolites for both medical and commercial purposes.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/plants12091756/s1, Table S1: Conserved miRNAs in Cuminum cyminum, Table S2: Sequence of primers of C. cyminum novel miRNAs.
Author Contributions
Conceptualization, A.S. (Ashutosh Sharma) and S.P.; methodology, A.R.-C., C.G.-G. and A.G.U.-P.; software, A.R.-C., S.P., A.S. (Aashish Srivastava) and A.S. (Ashutosh Sharma); validation, S.P., A.S. (Ashutosh Sharma) and A.R.-C.; formal analysis, A.R.-C., S.P. and A.S. (Ashutosh Sharma); investigation, A.R.-C.; resources, A.S. (Ashutosh Sharma); data curation, L.A.-M.; writing—original draft preparation, A.R.-C., C.G.-G. and H.M.N.I.; writing—review and editing, A.S. (Ashutosh Sharma), S.P. and S.S.S.J.A.; visualization, A.R.-C.; supervision, A.S. (Ashutosh Sharma); project administration, A.S. (Ashutosh Sharma); funding acquisition, A.S. (Ashutosh Sharma). All authors have read and agreed to the published version of the manuscript.
Funding
The authors would like to thank the Regional Department of Bioengineering, South-Central Region of Tecnologico de Monterrey, for their support regarding the equipment and required infrastructure for this project.
Data Availability Statement
Publicly available datasets were analyzed in this study. This data can be found here: https://www.ncbi.nlm.nih.gov/search/all/?term=SRP376517.
Acknowledgments
We thank CONACYT, Mexico, for the scholarship support for the completion of the Ph.D. thesis of Almendra Reyes Calderón (s).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Singh, N.; Yadav, S.S.; Kumar, S.; Narashiman, B. A review on traditional uses, phytochemistry, pharmacology, and clinical research of dietary spice Cuminum cyminum L. Phyther. Res. 2021, 35, 5007–5030. [Google Scholar] [CrossRef] [PubMed]
- Minooeianhaghighi, M.H.; Sepehrian, L.; Shokri, H. Effets antifongiques des huiles essentielles de Lavandula binaludensis et Cuminum cyminum contre Candida albicans isolés de patientes atteintes de candidose vulvo-vaginale récidivante. J. Mycol. Med. 2017, 27, 65–71. [Google Scholar] [CrossRef]
- Balaram, P.K.; Vidhya, N.; Rojamathi, K.; Revathi Satheesh, E.; Sowmiya, P.; Sabarish, T.; Mahalingam, P. Antibacterial and Phytochemical Analysis of Cuminum Cyminum (Cumin) and Illicium Verum (Star Anise) against Clinical Pathogens. WJPR 2021, 10, 938–947. [Google Scholar]
- Archangia, A.; Bahram, H.; Mohammadi-Nejad, G. Association between seed yield-related traits and cDNA-AFLP markers in cumin (Cuminum cyminum) under drought and irrigation regimes. Ind. Crop. Prod. 2019, 133, 276–283. [Google Scholar] [CrossRef]
- Chaudhry, Z.; Khera, R.A.; Hanif, M.A.; Ayub, M.A.; Sumrra, S.H. Medicinal Plants of South Asia; Elsevier: Amsterdam, The Netherlands, 2020; pp. 165–178. ISBN 9780081026595. [Google Scholar]
- Keerthiga, N.; Anitha, R.; Rajeshkumar, S.; Lakshmi, T.; Keerthiga, N.; Rajeshkumar, S. Antioxidant Activity of Cumin Oil Mediated Silver Nanoparticles. Pharmacogn. J. 2019, 11, 787–789. [Google Scholar] [CrossRef]
- Karimirad, R.; Behnamian, M.; Dezhsetan, S. LWT—Food Science and Technology Application of chitosan nanoparticles containing Cuminum cyminum oil as a delivery system for shelf life extension of Agaricus bisporus. LWT Food Sci. Technol. 2019, 106, 218–228. [Google Scholar] [CrossRef]
- Dinparvar, S.; Bagirova, M.; Allahverdiyev, A.M.; Abamor, E.S.; Safarov, T.; Aydogdu, M.; Aktas, D. A nanotechnology-based new approach in the treatment of breast cancer: Biosynthesized silver nanoparticles using Cuminum cyminum L. seed extract. J. Photochem. Photobiol. B Biol. 2020, 208, 111902. [Google Scholar] [CrossRef] [PubMed]
- Armenta-Medina, A.; Gillmor, C.S. An introduction to methods for discovery and functional analysis of MicroRNAs in plants. Methods Mol. Biol. 2019, 1932, 1–14. [Google Scholar] [CrossRef]
- Ding, Y.; Ding, L.; Xia, Y.; Wang, F.; Zhu, C. Emerging Roles of microRNAs in Plant Heavy Metal Tolerance and Homeostasis. J. Agric. Food Chem. 2020, 68, 1958–1965. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Li, Y.; Cao, X.; Qi, Y. MicroRNAs and Their Regulatory Roles in Plant-Environment Interactions. Annu. Rev. Plant Biol. 2019, 70, 489–525. [Google Scholar] [CrossRef]
- Griffiths-Jones, S. MiRBase: MicroRNA sequences and annotation. Curr. Protoc. Bioinforma. 2010, 29, 1291–12910. [Google Scholar] [CrossRef]
- Saliminejad, K.; Khorram Khorshid, H.R.; Soleymani Fard, S.; Ghaffari, S.H. An overview of microRNAs: Biology, functions, therapeutics, and analysis methods. J. Cell. Physiol. 2019, 234, 5451–5465. [Google Scholar] [CrossRef] [PubMed]
- Hossain, R.; Quispe, C.; Saikat, A.S.M.; Jain, D.; Habib, A.; Janmeda, P.; Islam, M.T.; Radha; Daştan, S.D.; Kumar, M.; et al. Biosynthesis of Secondary Metabolites Based on the Regulation of MicroRNAs. Biomed Res. Int. 2022, 2022, 1–20. [Google Scholar] [CrossRef]
- Que, F.; Hou, X.L.; Wang, G.L.; Xu, Z.S.; Tan, G.F.; Li, T.; Wang, Y.H.; Khadr, A.; Xiong, A.S. Advances in research on the carrot, an important root vegetable in the Apiaceae family. Hortic. Res. 2019, 6, 69. [Google Scholar] [CrossRef]
- Barozai, M.Y.K.; Kakar, S.U.R.; Sarangzai, A.M. Profiling the carrot (Daucus carota L.) MicroRNAs and their targets. Pak. J. Bot. 2013, 45, 353–358. [Google Scholar]
- Srivastava, S.; Sanchita; Singh, R.; Srivastava, G.; Sharma, A. Comparative Study of Withanolide Biosynthesis-Related miRNAs in Root and Leaf Tissues of Withania somnifera. Appl. Biochem. Biotechnol. 2018, 185, 1145–1159. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhong, Y.; Yu, F.; Xu, M. Deep sequencing identifies miRNAs and their target genes involved in the biosynthesis of terpenoids in Cinnamomum camphora. Ind. Crops Prod. 2020, 145, 111853. [Google Scholar] [CrossRef]
- Yin, D.D.; Li, S.S.; Shu, Q.Y.; Gu, Z.Y.; Wu, Q.; Feng, C.Y.; Xu, W.Z.; Wang, L.S. Identification of microRNAs and long non-coding RNAs involved in fatty acid biosynthesis in tree peony seeds. Gene 2018, 666, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-García, C.; Ahmed, S.S.S.J.; Ramalingam, S.; Selvaraj, D.; Srivastava, A.; Paul, S.; Sharma, A. Identification of microRNAs from medicinal plant Murraya Koenigii by high-throughput sequencing and their functional implications in secondary metabolite biosynthesis. Plants 2022, 11, 46. [Google Scholar] [CrossRef]
- Zhang, B.H.; Pan, X.P.; Cox, S.B.; Cobb, G.P.; Anderson, T.A. Evidence that miRNAs are different from other RNAs. Cell. Mol. Life Sci. 2006, 63, 246–254. [Google Scholar] [CrossRef]
- Vargas-Hernández, M.; Vázquez-Marrufo, G.; Aguilar-Ruiz, C.A.; González-Márquez, M.A.; Rocha, O.; Cerna-Pantoja, D.; Cruz-Hernández, A. MicroRNAs Associated with secondary metabolites production. In Plant Physiological Aspects of Phenolic Compounds; Soto-Hernández, M., García-Mateos, R., Palma-Tenango, M., Eds.; IntechOpen: London, UK, 2019; ISBN 978-1-78984-034-6. [Google Scholar]
- Liu, H.; Yu, H.; Tang, G.; Huang, T. Small but powerful: Function of microRNAs in plant development. Plant Cell Rep. 2018, 37, 515–528. [Google Scholar] [CrossRef] [PubMed]
- Srinivasan, K. Cumin (Cuminum cyminum) and black cumin (Nigella sativa) seeds: Traditional uses, chemical constituents, and nutraceutical effects. Food Qual. Saf. 2018, 2, 1–16. [Google Scholar] [CrossRef]
- Bettaieb, I.; Bourgou, S.; Wannes, W.A.; Hamrouni, I.; Limam, F.; Marzouk, B. Essential oils, phenolics, and antioxidant activities of different parts of cumin (Cuminum cyminum L.). J. Agric. Food Chem. 2010, 58, 10410–10418. [Google Scholar] [CrossRef]
- Rana, A.; Kumar, V.; Mirza, A.; Panghal, A. Efficacy of cumin (Cuminum cyminum L.) as a bionutrient and its management. Ann. Biol. 2018, 34, 218–222. [Google Scholar]
- Rogowska, A.; Szakiel, A. Enhancement of phytosterol and triterpenoid production in plant hairy root cultures—Simultaneous stimulation or competition? Plants 2021, 10, 2028. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, J.; Wang, C.; Chen, H.; Liu, Y.; Wang, Y.; Gao, W. Comprehensive Identification and Profiling of miRNAs Involved in Terpenoid Synthesis of Gleditsia sinensis Lam. Forests 2022, 13, 108. [Google Scholar] [CrossRef]
- Bergman, M.E.; Davis, B.; Phillips, M.A. Medically useful plant terpenoids: Biosynthesis, occurrence, and mechanism of action. Molecules 2019, 24, 3961. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Tang, Q.; Mo, C.; Bai, L.; Tu, D.; Ma, X. Cloning and characterization of squalene synthase and cycloartenol synthase from Siraitia grosvenorii. Acta Pharm. Sin. B 2017, 7, 215–222. [Google Scholar] [CrossRef] [PubMed]
- Jeena, G.S.; Joshi, A.; Shukla, R.K. Bm-miR172c-5p Regulates Lignin Biosynthesis and Secondary Xylem Thickness by Altering the Ferulate 5 Hydroxylase Gene in Bacopa monnieri. Plant Cell Physiol. 2021, 62, 894–912. [Google Scholar] [CrossRef]
- Venkatesh, T.V.; Karunanandaa, B.; Free, D.L.; Rottnek, J.M.; Baszis, S.R.; Valentin, H.E. Identification and characterization of an Arabidopsis homogentisate phytyltransferase paralog. Planta 2006, 223, 1134–1144. [Google Scholar] [CrossRef]
- Zeng, W.; Sun, Z.; Lai, Z.; Yang, S.; Chen, H.; Yang, X.; Tao, J.; Tang, X. Determination of the MiRNAs related to bean pyralid larvae resistance in soybean using small RNA and transcriptome sequencing. Int. J. Mol. Sci. 2019, 20, 2966. [Google Scholar] [CrossRef] [PubMed]
- Kawaide, H.; Hayashi, K.I.; Kawanabe, R.; Sakigi, Y.; Matsuo, A.; Natsume, M.; Nozaki, H. Identification of the single amino acid involved in quenching the ent-kauranyl cation by a water molecule in ent-kaurene synthase of Physcomitrella patens. FEBS J. 2011, 278, 123–133. [Google Scholar] [CrossRef]
- Reid, B.; Meirong, J.; Reuben, J.P. A pair of threonines mark ent-kaurene synthases for phytohormone biosynthesis. Phytochemistry 2021, 184, 112672. [Google Scholar] [CrossRef]
- Aach, H.; Bode, H.; Robinson, D.G.; Graebe, J.E. ent-Kaurene synthase is located in proplastids of meristematic shoot tissues. Planta 1997, 202, 211–219. [Google Scholar] [CrossRef]
- Barozai, M.Y.K.; Baloch, I.A.; Din, M. Computational identification of microRNAs and their targets in two species of evergreen spruce tree (Picea). World Acad. Sci. Eng. Technol. 2011, 5, 1290–1295. [Google Scholar]
- Nagegowda, D.A.; Gupta, P. Advances in biosynthesis, regulation, and metabolic engineering of plant specialized terpenoids. Plant Sci. 2020, 294, 110457. [Google Scholar] [CrossRef] [PubMed]
- Stephen, N.M.; Gayathri, R.; Niranjana, R.; Prasad, Y.; Das, A.K.; Baskaran, V.; Ganesan, P. Carotenoids: Types, sources, ans biosynthesis. In Plant Secondary Metabolites; Mohammed Wasim, S., Vasudha, B., Kamlesh, P., Eds.; Apple Academic Press: New York, NY, USA, 2017; Volume 2, pp. 103–132. ISBN 9781771883566. [Google Scholar]
- Kumari, S.; Goyal, A.; Garg, M. Phytochemistry and Pharmacological Update on Tetraterpenoids. Nat. Prod. J. 2020, 11, 617–628. [Google Scholar] [CrossRef]
- Zhao, X.; Zhang, Y.; Long, T.; Wang, S.; Yang, J. Regulation Mechanism of Plant Pigments Biosynthesis: Anthocyanins, Carotenoids, and Betalains. Metabolites 2022, 12, 871. [Google Scholar] [CrossRef]
- Du, H.; Liao, X.; Gao, Z.; Li, Y.; Lei, Y.; Chen, W.; Chen, L.; Fan, X.; Zhang, K.; Chen, S.; et al. Effects of methanol on carotenoids as well as biomass and fatty acid biosynthesis in Schizochytrium limacinum B4D1. Appl. Environ. Microbiol. 2019, 85, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Guo, J.; Chen, Q.; Wang, B.; He, X.; Zhuge, Q.; Wang, P. Different color regulation mechanism in willow barks determined using integrated metabolomics and transcriptomics analyses. BMC Plant Biol. 2022, 22, 1–15. [Google Scholar] [CrossRef]
- Yan, C.; Zhang, N.; Wang, Q.; Fu, Y.; Wang, F.; Su, Y.; Xue, B.; Zhou, L.; Liao, H. The Effect of Low Temperature Stress on the Leaves and MicroRNA Expression of Potato Seedlings. Front. Ecol. Evol. 2021, 9, 1–16. [Google Scholar] [CrossRef]
- Zhang, Y.; Cui, J.; Hu, H.; Xue, J.; Yang, J.; Xu, J. Integrated Four Comparative-Omics Reveals the Mechanism of the Terpenoid Biosynthesis in Two Different Overwintering Cryptomeria fortunei Phenotypes. Front. Plant Sci. 2021, 12, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Pegler, J.L.; Oultram, J.M.J.; Grof, C.P.L.; Eamens, A.L. Profiling the abiotic stress responsive microRNA landscape of arabidopsis thaliana. Plants 2019, 8, 58. [Google Scholar] [CrossRef] [PubMed]
- Dou, X.; Zhou, Z.; Zhao, L. Identification and expression analysis of miRNAs in germination and seedling growth of Tibetan hulless barley. Genomics 2021, 113, 3735–3749. [Google Scholar] [CrossRef] [PubMed]
- Qibin, L.; Jiang, W. MIREAP: MicroRNA Discovery By Deep Sequencing (Software, Version 0.2) 2008, Sourceforge. Available online: https://sourceforge.net/projects/mireap/ (accessed on 1 June 2019).
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).