Transcriptome Analysis of Early Lateral Root Formation in Tomato
Abstract
1. Introduction
2. Results
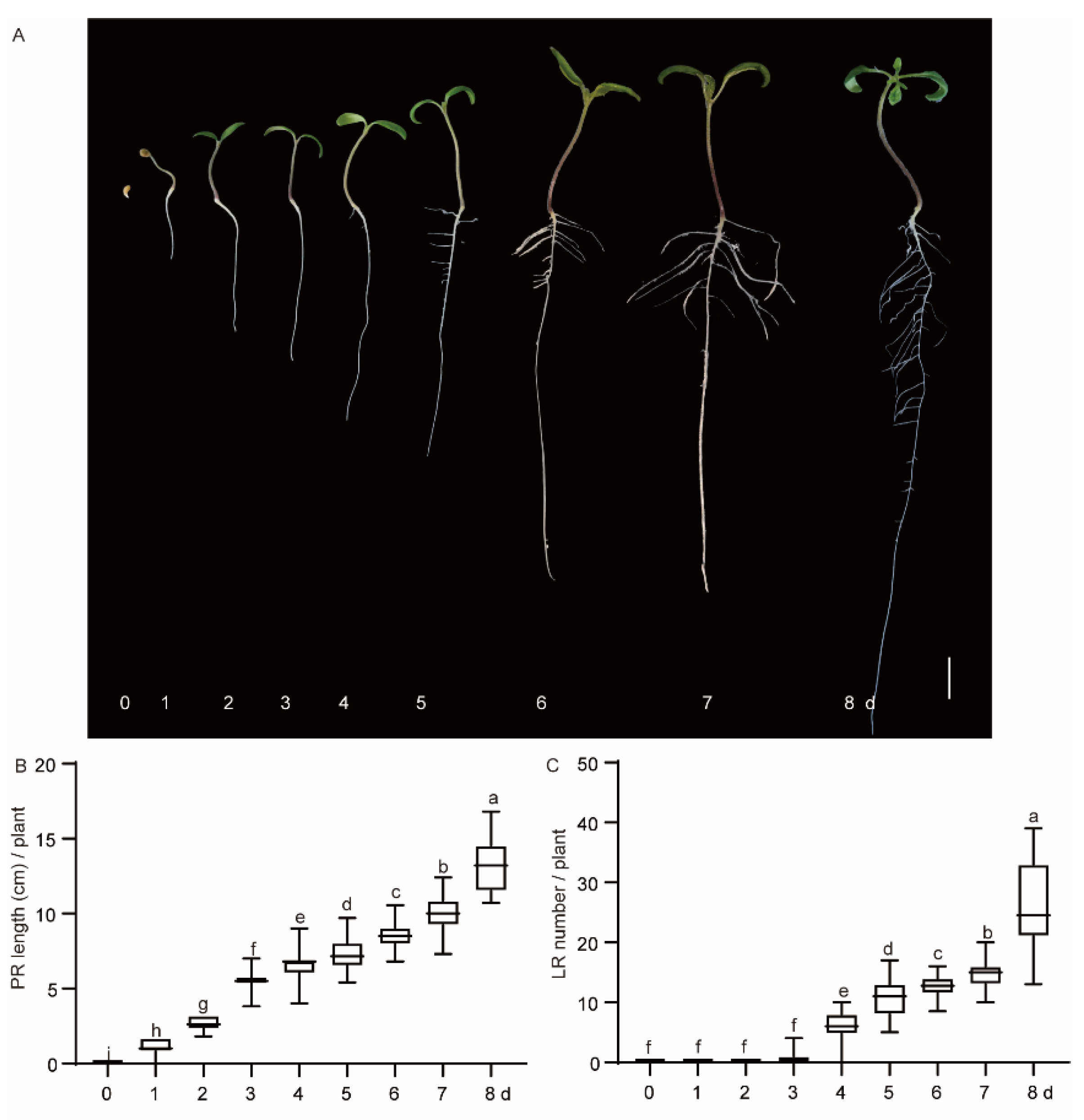
2.1. Temporal Dynamics of Lateral Root (LR) Development in Tomato Seedlings
2.2. Cytological Analysis of Tomato LRs
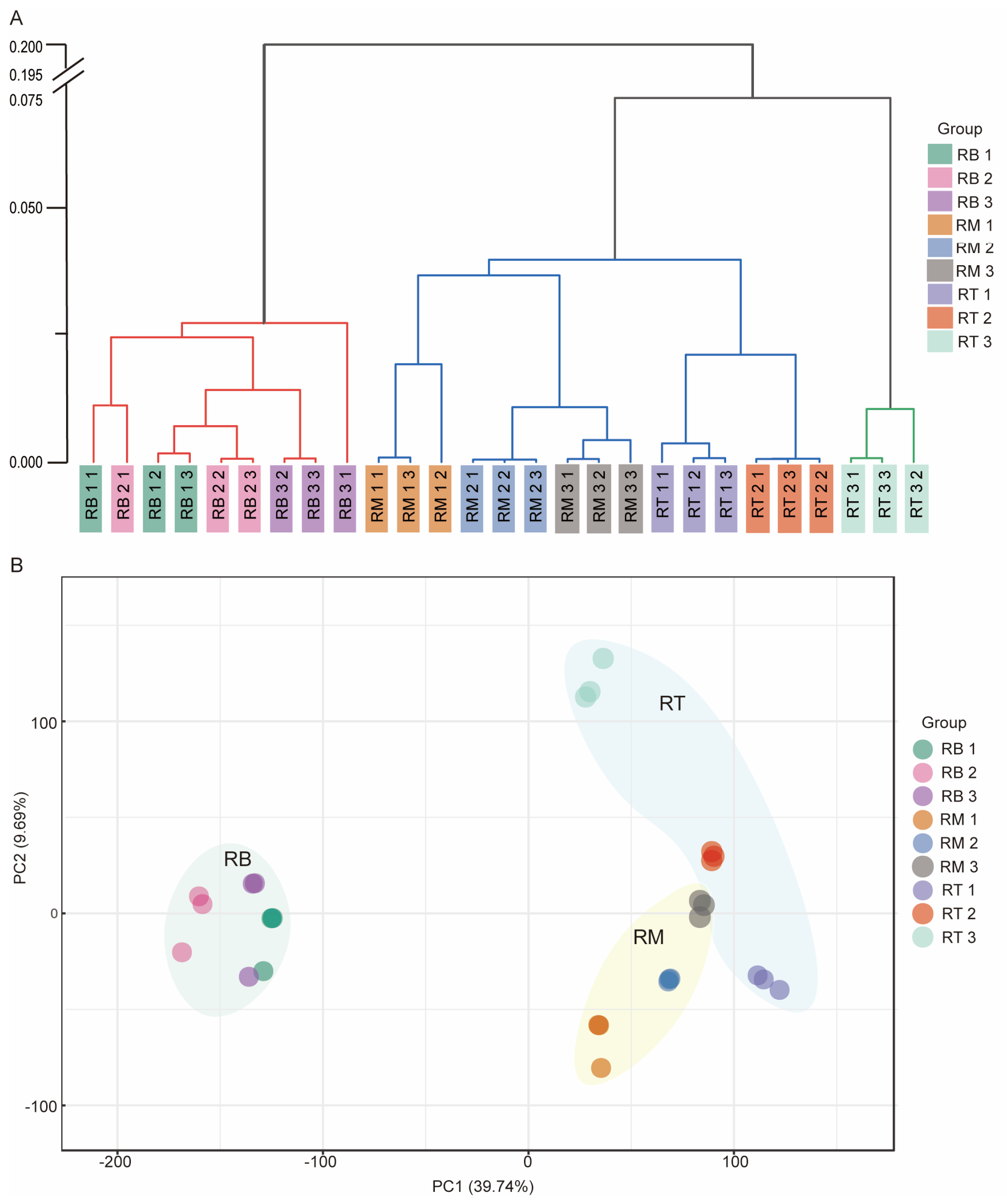
2.3. Cluster Analysis of Transcriptome Data from Different Stages of LR Development
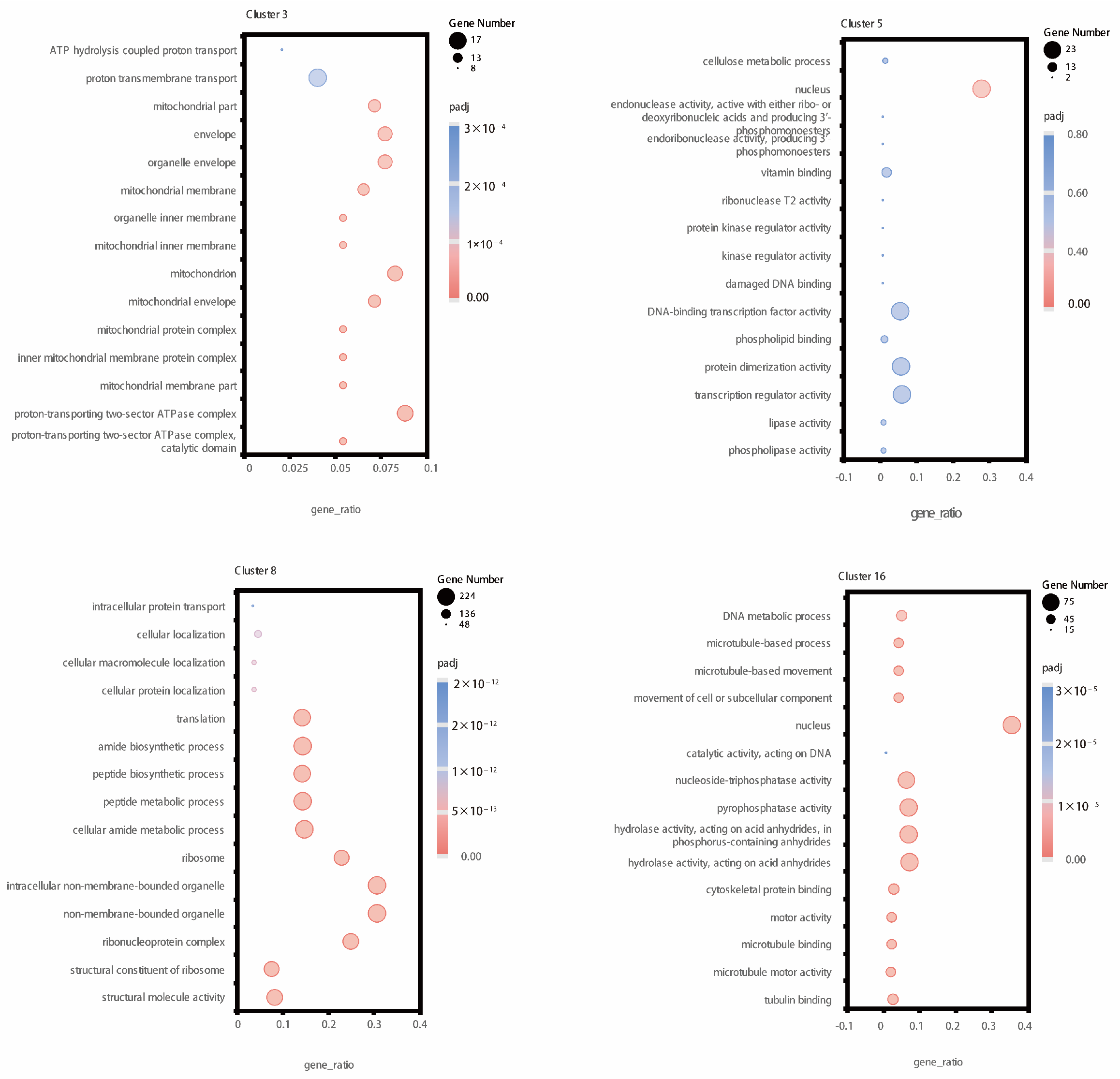
2.4. Co-Expression Analysis of Genes in 16 Clusters of the Tomato Root Transcriptome
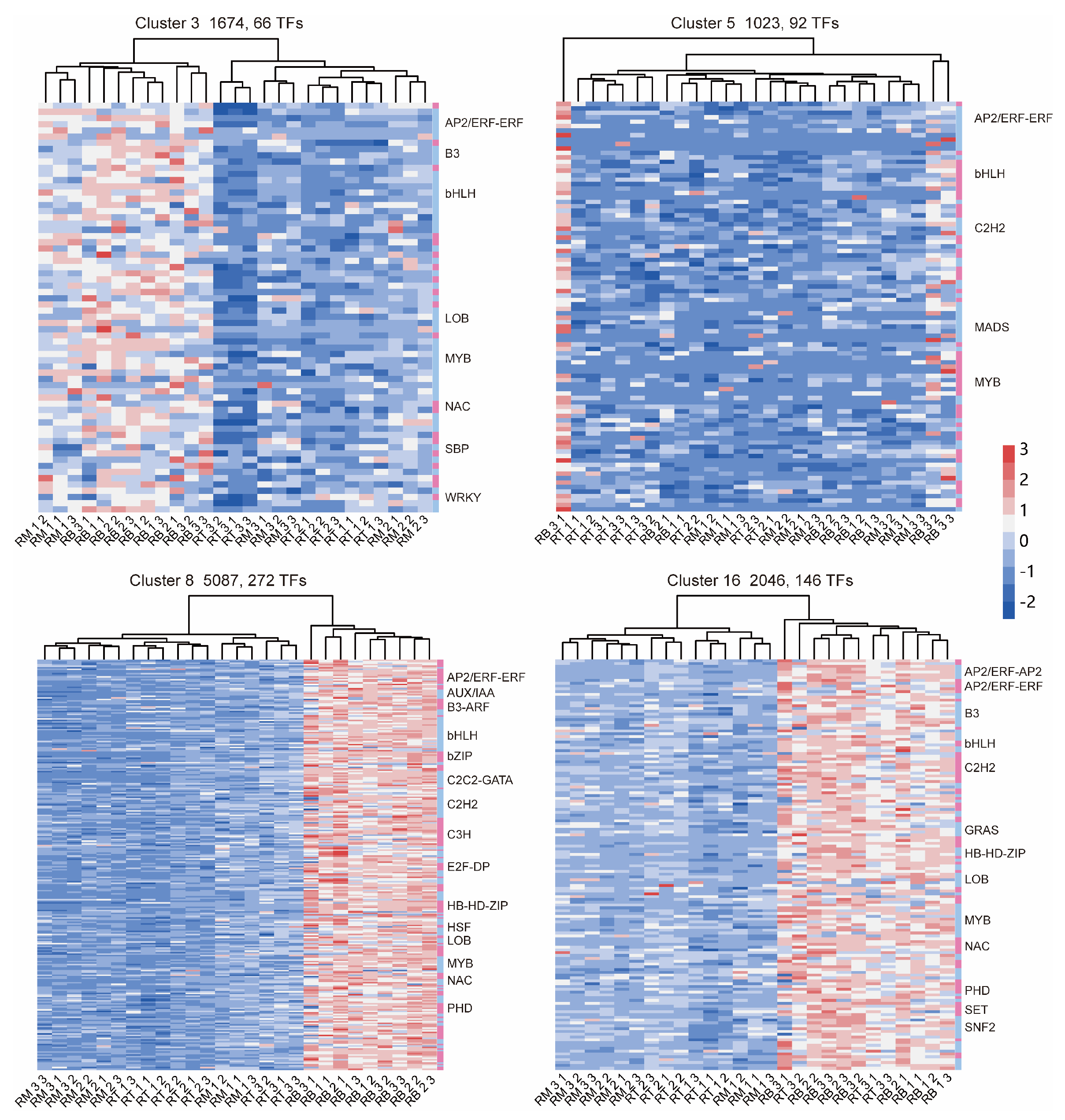
2.5. Expression of TFs Related to LR Formation
2.6. Marker Gene Expression in LR Development and by Quantitative Real-Time PCR (qPCR) Validation
2.7. Determination of Endogenous IAA Content in Tomato
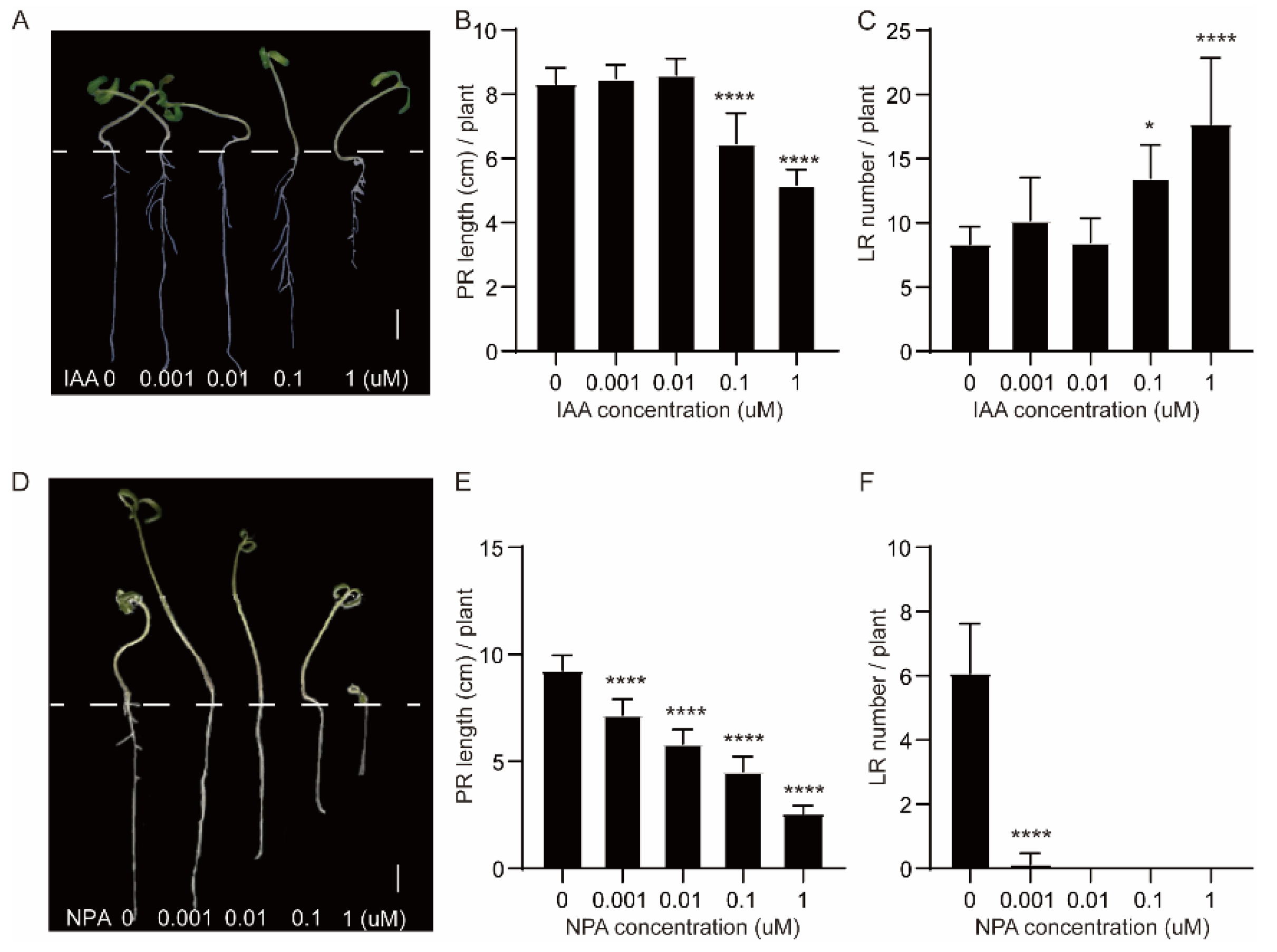
2.8. Tomato Seedlings Treated with Different Concentrations of IAA and NPA
3. Discussion
4. Materials and Methods
4.1. Plant Material and Growth Conditions
4.2. Paraffin Sectioning of Tomato Seedling Roots
4.3. RNA Extraction and Transcriptome Sequencing
4.4. Sample Correlation
4.5. Gene Expression Analysis and Identification of Differentially Expressed Genes (DEGs)
4.6. K-Means Analysis and GO Enrichment Analysis
4.7. Validation of RNA-Seq Data by qPCR
4.8. Measuring Endogenous Levels of IAA, IAA Precursors, and IAA Metabolites
4.9. IAA and NPA Treatments at Different Concentrations
4.10. Statistical Analysis Methods
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Singh, H.; Singh, Z.; Kashyap, R.; Yadav, S.R. Lateral root branching: Evolutionary innovations and mechanistic divergence in land plants. New Phytol. 2023, 238, 1379–1385. [Google Scholar] [CrossRef] [PubMed]
- Javed, S.; Chai, X.; Wang, X.; Xu, S. The phytohormones underlying the plant lateral root development in fluctuated soil environments. Plant Soil 2023, 1–14. [Google Scholar] [CrossRef]
- Zhang, H.; San, M.L.; Jang, S.G.; Lee, J.H.; Kim, N.E.; Lee, A.R.; Park, S.Y.; Cao, F.Y.; Chin, J.H.; Kwon, S.W. Genome-Wide Association Study of Root System Development at Seedling Stage in Rice. Genes 2020, 12, 1395. [Google Scholar] [CrossRef] [PubMed]
- Malamy, J.E. Intrinsic and environmental response pathways that regulate root system architecture. Plant Cell Environ. 2005, 28, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Mitsuda, N.; Yoshizumi, T.; Horii, Y.; Oshima, Y.; Ohme-Takagi, M.; Matsui, M.; Kakimoto, T. Two types of bHLH transcription factor determine the competence of the pericycle for lateral root initiation. Nat. Plants 2021, 7, 633–643. [Google Scholar] [CrossRef] [PubMed]
- Vilches-Barro, A.; Maizel, A. Talking through walls: Mechanisms of lateral root emergence in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2015, 23, 31–38. [Google Scholar] [CrossRef]
- Swarup, K.; Benkova, E.; Swarup, R.; Casimiro, I.; Peret, B.; Yang, Y.; Parry, G.; Nielsen, E.; De Smet, I.; Vanneste, S.; et al. The auxin influx carrier LAX3 promotes lateral root emergence. Nat. Cell Biol. 2008, 10, 946–954. [Google Scholar] [CrossRef] [PubMed]
- Vermeer, J.E.M.; von Wangenheim, D.; Barberon, M.; Lee, Y.; Stelzer, E.H.K.; Maizel, A.; Geldner, N. A Spatial Accommodation by Neighboring Cells Is Required for Organ Initiation in Arabidopsis. Science 2014, 343, 178–183. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.; Vu, L.D.; Van den Broeck, L.; Lin, Z.F.; Ramakrishna, P.; van de Cotte, B.; Gaudinier, A.; Goh, T.; Slane, D.; Beeckman, T.; et al. RALFL34 regulates formative cell divisions in Arabidopsis pericycle during lateral root initiation. J. Exp. Bot. 2016, 67, 4863–4875. [Google Scholar] [CrossRef]
- Li, S.N.; Li, Q.Q.; Tian, X.; Mu, L.J.; Ji, M.L.; Wang, X.P.; Li, N.; Liu, F.; Shu, J.; Crawford, N.M.; et al. PHB3 regulates lateral root primordia formation via NO-mediated degradation of AUXIN/INDOLE-3-ACETIC ACID proteins. J. Exp. Bot. 2022, 73, 4034–4045. [Google Scholar] [CrossRef]
- Cavallari, N.; Artner, C.; Benkova, E. Auxin-Regulated Lateral Root Organogenesis. CSH Perspect. Biol. 2021, 13, a039941. [Google Scholar] [CrossRef] [PubMed]
- Hu, F.; Fang, D.; Zhang, W.; Dong, K.; Ye, Z.; Cao, J. Lateral root primordium: Formation, influencing factors and regulation. Plant Physiol. Biochem. 2024, 207, 108429. [Google Scholar] [CrossRef] [PubMed]
- De Smet, S.; Cuypers, A.; Vangronsveld, J.; Remans, T. Gene Networks Involved in Hormonal Control of Root Development in Arabidopsis thaliana: A Framework for Studying Its Disturbance by Metal Stress. Int. J. Mol. Sci. 2015, 16, 19195–19224. [Google Scholar] [CrossRef] [PubMed]
- Ditengou, F.A.; Teale, W.D.; Kochersperger, P.; Flittner, K.A.; Kneuper, I.; van der Graaff, E.; Nziengui, H.; Pinosa, F.; Li, X.; Nitschke, R.; et al. Mechanical induction of lateral root initiationin Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2008, 105, 18818–18823. [Google Scholar] [CrossRef] [PubMed]
- Teale, W.D.; Ditengou, F.A.; Dovzhenko, A.D.; Li, X.; Molendijk, A.M.; Ruperti, B.; Paponov, I.; Palme, K. Auxin as a model for the integration of hormonal signal processing and transduction. Mol. Plant 2008, 1, 229–237. [Google Scholar] [CrossRef] [PubMed]
- De Smet, I.; Lau, S.; Voss, U.; Vanneste, S.; Benjamins, R.; Rademacher, E.H.; Schlereth, A.; De Rybel, B.; Vassileva, V.; Grunewald, W.; et al. Bimodular auxin response controls organogenesis in Arabidopsis. Proc. Natl. Acad. Sci. USA 2010, 107, 2705–2710. [Google Scholar] [CrossRef]
- De Smet, I.; Vassileva, V.; De Rybel, B.; Levesque, M.P.; Grunewald, W.; Van Damme, D.; Van Noorden, G.; Naudts, M.; Van Isterdael, G.; De Clercq, R.; et al. Receptor-like kinase ACR4 restricts formative cell divisions in the Arabidopsis root. Science 2008, 322, 594–597. [Google Scholar] [CrossRef] [PubMed]
- Banda, J.; Bellande, K.; von Wangenheim, D.; Goh, T.; Guyomarc’h, S.; Laplaze, L.; Bennett, M.J. Lateral Root Formation in Arabidopsis: A Well-Ordered LRexit. Trends Plant Sci. 2019, 24, 826–839. [Google Scholar] [CrossRef] [PubMed]
- Perotti, M.F.; Ribone, P.A.; Cabello, J.V.; Ariel, F.D.; Chan, R.L. AtHB23 participates in the gene regulatory network controlling root branching, and reveals differences between secondary and tertiary roots. Plant J. 2019, 100, 1224–1236. [Google Scholar] [CrossRef]
- Yuan, T.T.; Xu, H.H.; Li, J.; Lu, Y.T. Auxin abolishes SHI-RELATED SEQUENCE5-mediated inhibition of lateral root development in Arabidopsis. New Phytol. 2020, 225, 297–309. [Google Scholar] [CrossRef]
- Péret, B.; Middleton, A.M.; French, A.P.; Larrieu, A.; Bishopp, A.; Njo, M.; Wells, D.M.; Porco, S.; Mellor, N.; Band, L.R.; et al. Sequential induction of auxin efflux and influx carriers regulates lateral root emergence. Mol. Syst. Biol. 2013, 9, 699. [Google Scholar] [CrossRef] [PubMed]
- Porco, S.; Larrieu, A.; Du, Y.J.; Gaudinier, A.; Goh, T.; Swarup, K.; Swarup, R.; Kuempers, B.; Bishopp, A.; Lavenus, J.; et al. Lateral root emergence in Arabidopsis is dependent on transcription factor LBD29 regulation of auxin influx carrier LAX3. Development 2016, 143, 3340–3349. [Google Scholar] [CrossRef] [PubMed]
- Kumpf, R.P.; Shi, C.L.; Larrieu, A.; Sto, I.M.; Butenko, M.A.; Péret, B.; Riiser, E.S.; Bennett, M.J.; Aalen, R.B. Floral organ abscission peptide IDA and its HAE/HSL2 receptors control cell separation during lateral root emergence. Proc. Natl. Acad. Sci. USA 2013, 110, 5235–5240. [Google Scholar] [CrossRef] [PubMed]
- Howe, E.; Holton, K.; Nair, S.; Schlauch, D.; Sinha, R.; Quackenbush, J. MeV: MultiExperiment Viewer. In Biomedical Informatics for Cancer Research; Springer: Boston, MA, USA, 2010; pp. 267–277. [Google Scholar] [CrossRef]
- Bellande, K.; Trinh, D.C.; Gonzalez, A.A.; Dubois, E.; Petitot, A.S.; Lucas, M.; Champion, A.; Gantet, P.; Laplaze, L.; Guyomarc’h, S. PUCHI represses early meristem formation in developing lateral roots of Arabidopsis thaliana. J. Exp. Bot. 2023, 73, 3496. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, T.; Tanaka, A.; Inahashi, H.; Nishizawa, N.K.; Tsutsumi, N.; Inukai, Y.; Nakazono, M. Fine control of aerenchyma and lateral root development through AUX/IAA- and ARF-dependent auxin signaling. Proc. Natl. Acad. Sci. USA 2019, 116, 20770–20775. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.W.; Li, Z.W.; Ma, B.; Hou, Q.C.; Wan, X.Y. Phylogeny and Functions of LOB Domain Proteins in Plants. Int. J. Mol. Sci. 2020, 21, 2278. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.R.; Wang, R.; Zheng, M.; Liu, X.B.; Meng, F.; Wu, H.L.; Yao, Y.Y.; Xin, M.M.; Peng, H.R.; Ni, Z.F.; et al. TaWRKY51 promotes lateral root formation through negative regulation of ethylene biosynthesis in wheat (Triticum aestivum L.). Plant J. 2018, 96, 372–388. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Li, K.N.; Liu, X.Y.; Ruan, H.; Zheng, M.M.; Yu, Z.J.; Gai, J.Y.; Yang, S.P. Transcription Factor GmWRKY46 Enhanced Phosphate Starvation Tolerance and Root Development in Transgenic Plants. Front. Plant Sci. 2021, 12, 700651. [Google Scholar] [CrossRef] [PubMed]
- Gate, T.; Hill, L.; Miller, A.J.; Sanders, D. AtIAR1 is a Zn transporter that regulates auxin metabolism in Arabidopsis thaliana. J. Exp. Bot. 2023, 75, 1437–1450. [Google Scholar] [CrossRef]
- Cao, J.; Tan, X.; Cheng, X. Over-expression of the BnVIT-L2 gene improves the lateral root development and biofortification under iron stress. Plant Physiol. Biochem. 2024, 208, 108501. [Google Scholar] [CrossRef]
- He, W.; Truong, H.A.; Zhang, L.; Cao, M.; Arakawa, N.; Xiao, Y.; Zhong, K.; Hou, Y.; Busch, W. Identification of mebendazole as an ethylene signaling activator reveals a role of ethylene signaling in the regulation of lateral root angles. Cell Rep. 2024, 43, 113763. [Google Scholar] [CrossRef]
- Wei, S.J.; Tanaka, R.; Kawakatsu, T.; Teramoto, S.; Tanaka, N.; Shenton, M.; Uga, Y.; Yabe, S. Genome- and Transcriptome-wide Association Studies to Discover Candidate Genes for Diverse Root Phenotypes in Cultivated Rice. Rice 2023, 16, 55. [Google Scholar] [CrossRef]
- Yamauchi, T.; Tanaka, A.; Nakazono, M.; Inukai, Y. Age-dependent analysis dissects the stepwise control of auxin-mediated lateral root development in rice. Plant Physiol. 2024, 194, 819–831. [Google Scholar] [CrossRef] [PubMed]
- Ranjan, A.; Sinha, R.; Singla-Pareek, S.L.; Pareek, A.; Singh, A.K. Shaping the root system architecture in plants for adaptation to drought stress. Physiol. Plant. 2022, 174, e13651. [Google Scholar] [CrossRef] [PubMed]
- De Smet, I.; Signora, L.; Beeckman, T.; Inzé, D.; Foyer, C.H.; Zhang, H.M. An abscisic acid-sensitive checkpoint in lateral root development of Arabidopsis. Plant J. 2003, 33, 543–555. [Google Scholar] [CrossRef] [PubMed]
- Stoeckle, D.; Thellmann, M.; Vermeer, J.E.M. Breakout—Lateral root emergence in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2018, 41, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Singh, A. GIGANTEA regulates lateral root formation by modulating auxin signaling in Arabidopsis thaliana. Plant Signal Behav. 2022, 17, 2096780. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.W.; Kim, J. EXPANSINA17 Up-Regulated by LBD18/ASL20 Promotes Lateral Root Formation During the Auxin Response. Plant Cell Physiol. 2013, 54, 1600–1611. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Chen, B.X.; Chen, T.; Chen, J.H.; Lin, X.Y.; Yue, X.L.; An, L.Z.; Zhang, H. BYPASS1-LIKE regulates lateral root initiation via exocytic vesicular trafficking-mediated PIN recycling in Arabidopsis. J. Integr. Plant Biol. 2022, 64, 965–978. [Google Scholar] [CrossRef]
- Swarup, R.; Péret, B. AUX/LAX family of auxin influx carriers-an overview. Front. Plant Sci. 2012, 3, 225. [Google Scholar] [CrossRef]
- Huang, S.B.; Taylor, N.L.; Ströher, E.; Fenske, R.; Millar, A.H. Succinate dehydrogenase assembly factor 2 is needed for assembly and activity of mitochondrial complex II and for normal root elongation in Arabidopsis. Plant J. 2013, 73, 429–441. [Google Scholar] [CrossRef]
- Purushotham, P.; Ho, R.Y.; Zimmer, J. Architecture of a catalytically active homotrimeric plant cellulose synthase complex. Science 2020, 369, 1089–1094. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Wen, C.K. Arabidopsis aux1rcr1 mutation alters AUXIN RESISTANT1 targeting and prevents expression of the auxin reporter DR5:GUS in the root apex. J. Exp. Bot. 2013, 64, 921–933. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.L.; Li, Q.; Zhu, Y.Y.; Afzal, M.S.; Li, L.G. An auxin-induced β-type endo-1,4-β-glucanase in poplar is involved in cell expansion and lateral root formation. Planta 2018, 247, 1149–1161. [Google Scholar] [CrossRef] [PubMed]
- Stöckle, D.; Reyes-Hernández, B.J.; Barro, A.V.; Nenadic, M.; Winter, Z.; Marc-Martin, S.; Bald, L.; Ursache, R.; Fujita, S.; Maizel, A.; et al. Microtubule-based perception of mechanical conflicts controls plant organ morphogenesis. Sci. Adv. 2022, 8, eabm4974. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, C.G.; Bellinger, M. An overview of plant division-plane orientation. New Phytol. 2018, 219, 505–512. [Google Scholar] [CrossRef] [PubMed]
- Qian, Y.; Wang, X.; Liu, Y.; Wang, X.; Mao, T. HY5 inhibits lateral root initiation in Arabidopsis through negative regulation of the microtubule-stabilizing protein TPXL5. Plant Cell 2023, 35, 1092–1109. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.; Niu, Q.W.; Ng, K.H.; Chua, N.H. The role of miR156/SPLs modules in Arabidopsis lateral root development. Plant J. 2015, 83, 673–685. [Google Scholar] [CrossRef] [PubMed]
- Gasch, A.P.; Eisen, M.B. Exploring the conditional coregulation of yeast gene expression through fuzzy k-means Clustering. Genome Biol. 2002, 3, research0059.1. [Google Scholar] [CrossRef]
- Cui, Q.; Xie, L.; Dong, C.; Gao, L.; Shang, Q. Stage-specific events in tomato graft formation and the regulatory effects of auxin and cytokinin. Plant Sci. 2021, 304, 110803. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, A.; Shang, Q. Transcriptome Analysis of Early Lateral Root Formation in Tomato. Plants 2024, 13, 1620. https://doi.org/10.3390/plants13121620
Zhang A, Shang Q. Transcriptome Analysis of Early Lateral Root Formation in Tomato. Plants. 2024; 13(12):1620. https://doi.org/10.3390/plants13121620
Chicago/Turabian StyleZhang, Aiai, and Qingmao Shang. 2024. "Transcriptome Analysis of Early Lateral Root Formation in Tomato" Plants 13, no. 12: 1620. https://doi.org/10.3390/plants13121620
APA StyleZhang, A., & Shang, Q. (2024). Transcriptome Analysis of Early Lateral Root Formation in Tomato. Plants, 13(12), 1620. https://doi.org/10.3390/plants13121620





