Heterologous Expression of Chrysanthemum TCP Transcription Factor CmTCP13 Enhances Salinity Tolerance in Arabidopsis
Abstract
:1. Introduction
2. Results
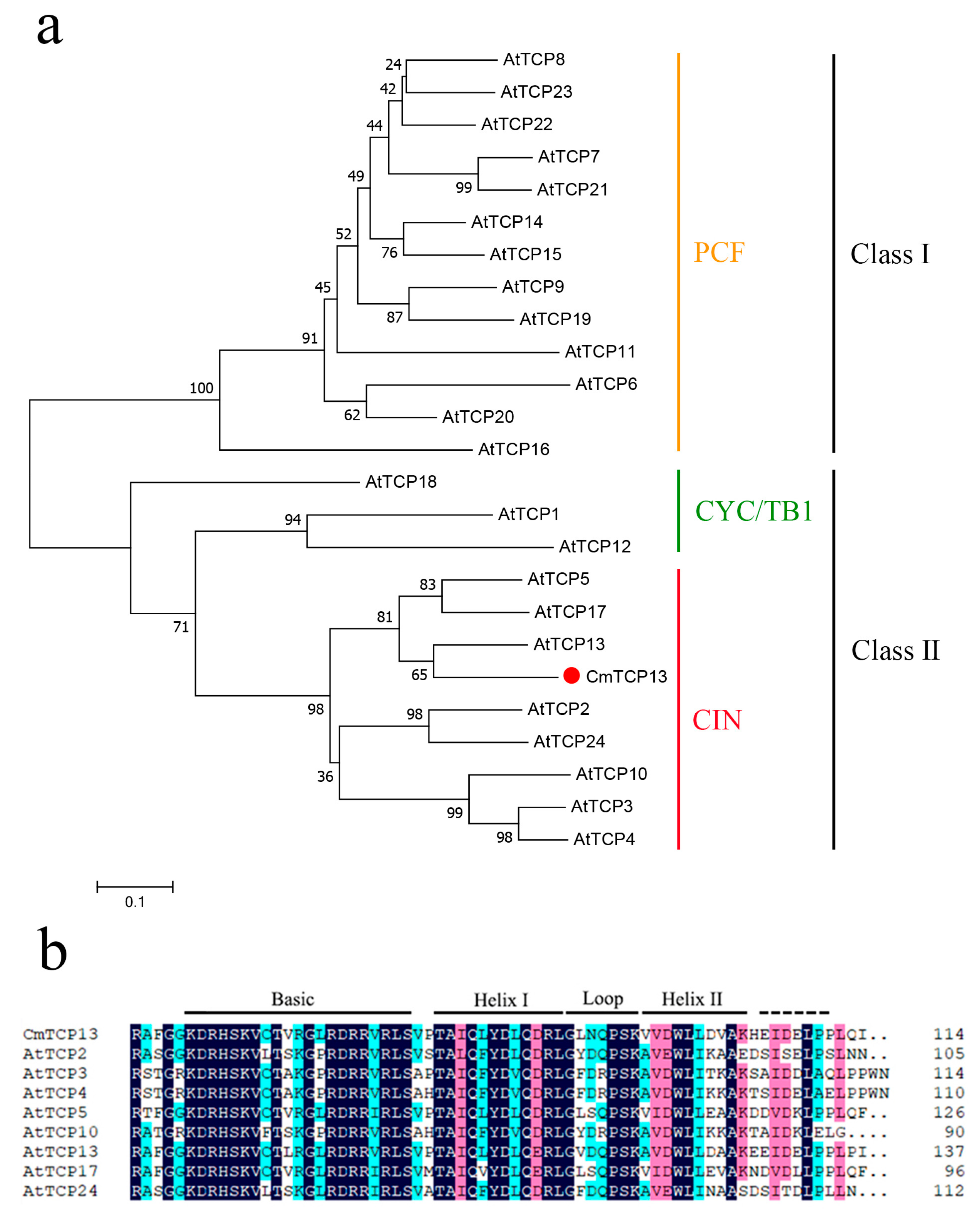
2.1. Cloning and Sequence Analysis of CmTCP13
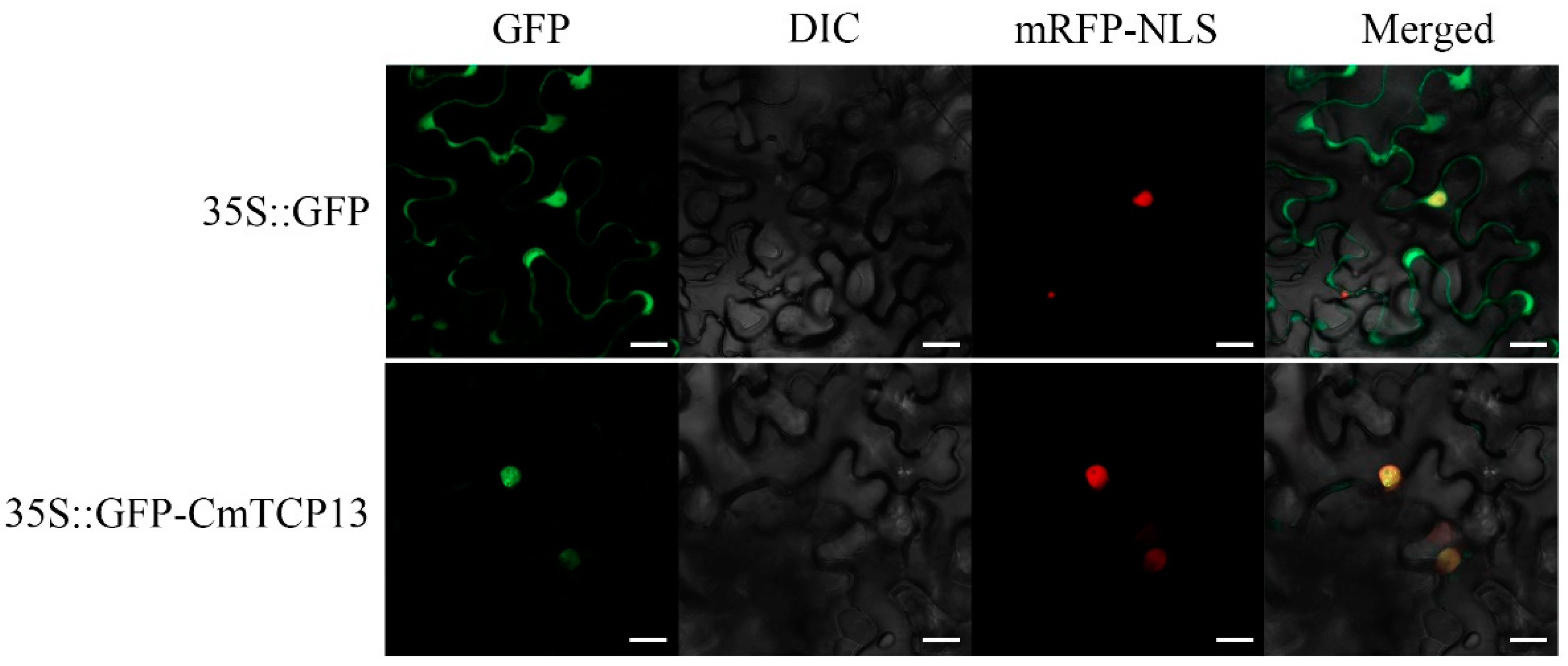
2.2. Subcellular Localization of CmTCP13
2.3. Transcript Profile of CmTCP13 in Response to Salinity
2.4. Cis-Acting Regulatory Element Analysis of CmTCP13 Promoter Sequence
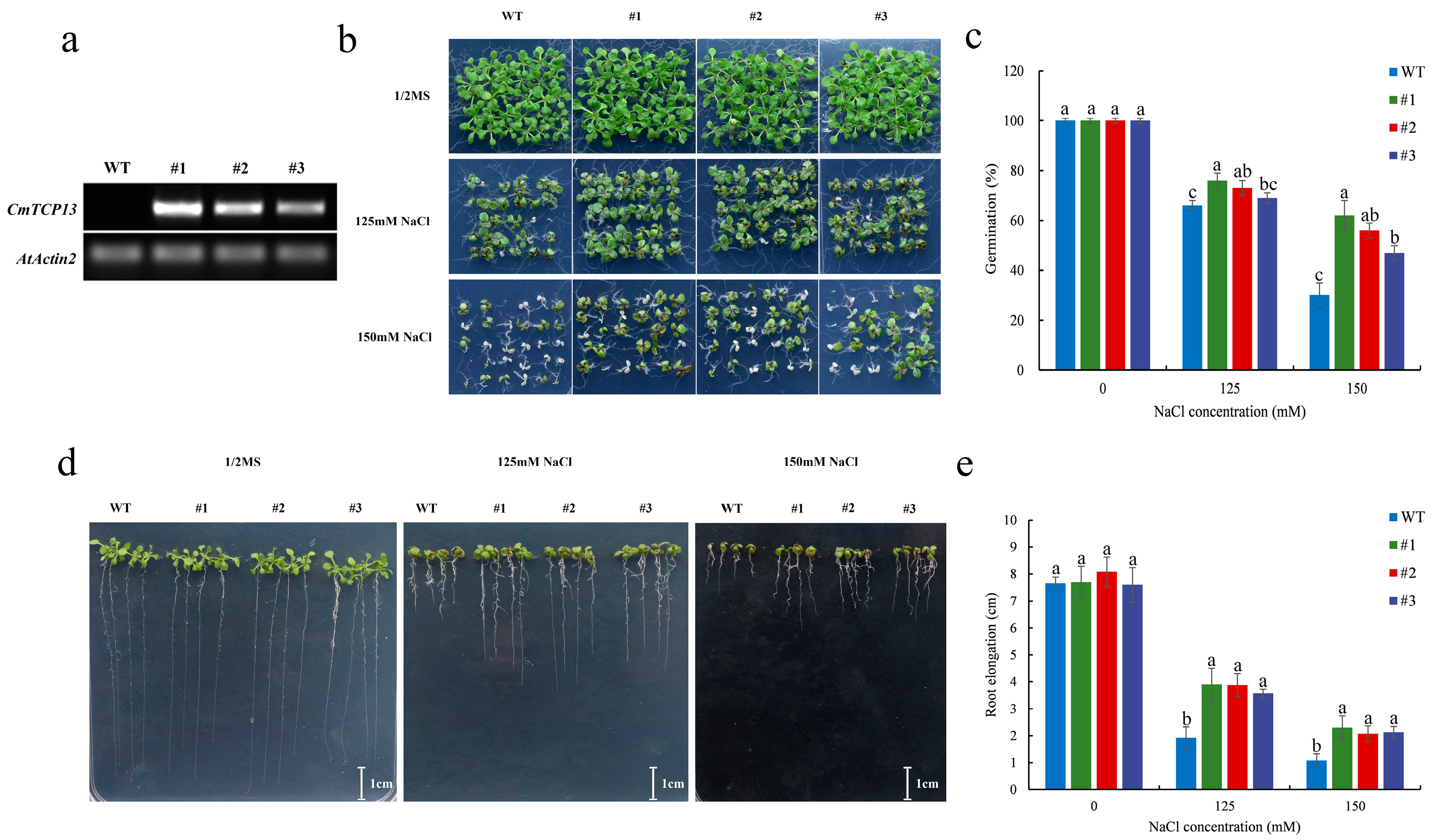
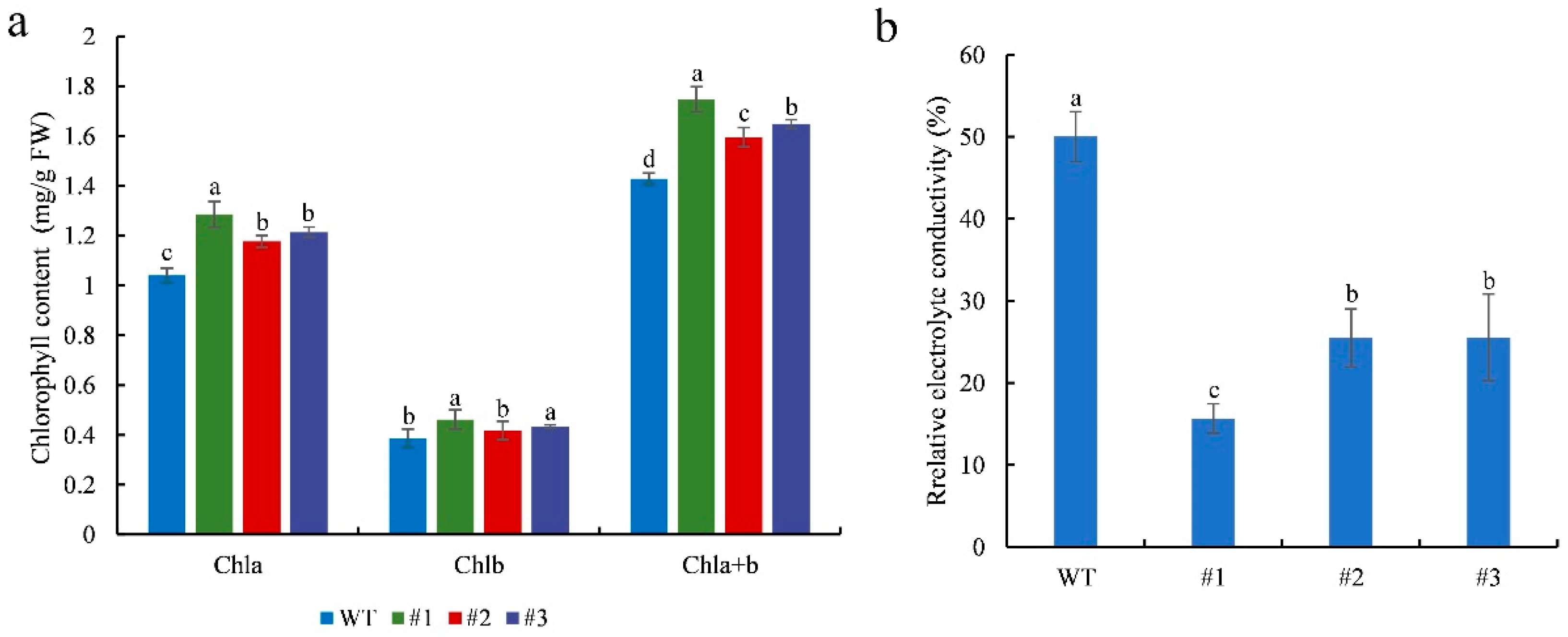
2.5. CmTCP13 Overexpression in Arabidopsis Enhanced Salinity Tolerance
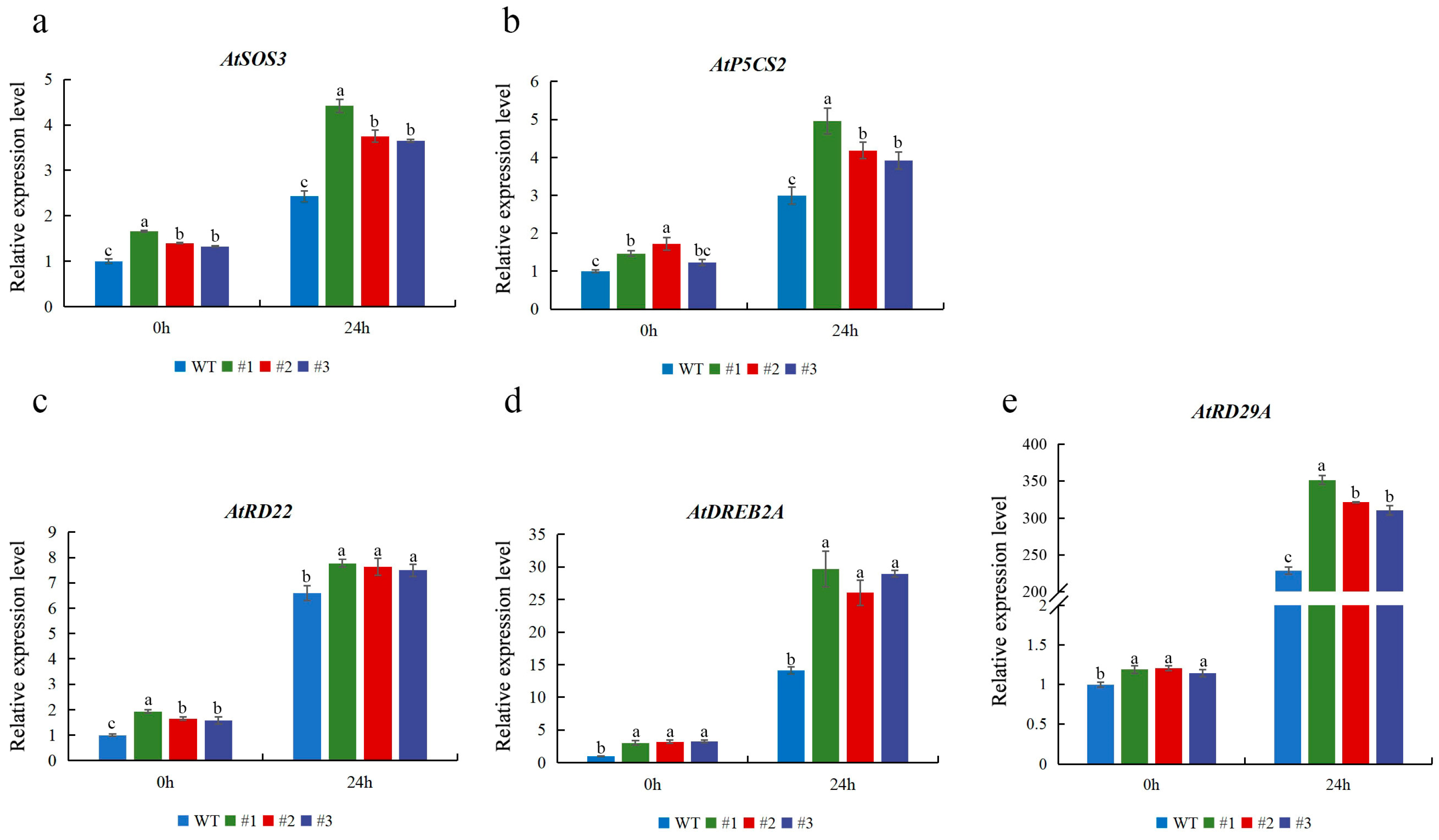
2.6. CmTCP13 Overexpression Altered the Expression of Stress-Responsive Genes
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Salinity Stress Treatments
4.2. Isolation and Sequence Analysis of CmTCP13
4.3. Subcellular Localization of CmTCP13
4.4. Ectopic Expression of CmTCP13 in Arabidopsis
4.5. Cis-Acting Element Analysis in the Promoter Region
4.6. Salinity Tolerance Assay of Transgenic Arabidopsis Plants
4.7. Measurements of Electrolyte Leakage and Chlorophyll Content
4.8. qRT-PCR Analyses
4.9. Statistical Analyses
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Deinlein, U.; Stephan, A.B.; Horie, T.; Luo, W.; Xu, G.; Schroeder, J.I. Plant salt-tolerance mechanisms. Trends Plant Sci. 2014, 19, 371–379. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Koussevitzky, S.; Mittler, R.; Miller, G. ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ. 2012, 35, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Xiong, L.; Schumaker, K.S.; Zhu, J.-K. Cell signaling during cold, drought, and salt stress. Plant Cell 2002, 14, 165–183. [Google Scholar] [CrossRef] [PubMed]
- Vinocur, B.; Altman, A. Recent advances in engineering plant tolerance to abiotic stress: Achievements and limitations. Curr. Opin. Biotechnol. 2005, 16, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed]
- Sakuma, Y.; Maruyama, K.; Osakabe, Y.; Qin, F.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 2006, 18, 1292–1309. [Google Scholar] [CrossRef]
- Yao, W.; Wang, S.; Zhou, B.; Jiang, T. Transgenic poplar overexpressing the endogenous transcription factor ERF76 gene improves salinity tolerance. Tree Physiol. 2016, 36, 896–908. [Google Scholar] [CrossRef] [PubMed]
- Martín-Trillo, M.; Cubas, P. TCP genes: A family snapshot ten years later. Trends Plant Sci. 2010, 15, 31–39. [Google Scholar] [CrossRef]
- Nicolas, M.; Cubas, P. TCP factors: New kids on the signaling block. Curr. Opin. Plant Biol. 2016, 33, 33–41. [Google Scholar] [CrossRef]
- Yao, X.; Ma, H.; Wang, J.; Zhang, D. Genome-wide comparative analysis and expression pattern of TCP gene families in Arabidopsis thaliana and Oryza sativa. J. Integr. Plant Biol. 2007, 49, 885–897. [Google Scholar] [CrossRef]
- Ma, X.; Ma, J.; Fan, D.; Li, C.; Jiang, Y.; Luo, K. Genome-wide identification of TCP family transcription factors from Populus euphratica and their involvement in leaf shape regulation. Sci. Rep. 2016, 6, 32795. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhai, Z.; Li, Y.; Geng, S.; Song, G.; Guan, J.; Jia, M.; Wang, F.; Sun, G.; Feng, N. Genome-wide identification and expression profiling of the TCP family genes in spike and grain development of wheat (Triticum aestivum L.). Front. Plant Sci. 2018, 9, 1282. [Google Scholar] [CrossRef]
- Navaud, O.; Dabos, P.; Carnus, E.; Tremousaygue, D.; Hervé, C. TCP transcription factors predate the emergence of land plants. J. Mol. Evol. 2007, 65, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Li, S. The Arabidopsis thaliana TCP transcription factors: A broadening horizon beyond development. Plant Signal. Behav. 2015, 10, e1044192. [Google Scholar] [CrossRef] [PubMed]
- Resentini, F.; Felipo-Benavent, A.; Colombo, L.; Blázquez, M.A.; Masiero, S. TCP14 and TCP15 mediate the promotion of seed germination by gibberellins in Arabidopsis thaliana. Mol. Plant 2014, 8, 482–485. [Google Scholar] [CrossRef]
- Yu, H.; Zhang, L.; Wang, W.; Tian, P.; Wang, W.; Wang, K.; Gao, Z.; Liu, S.; Zhang, Y.; F, I.V. TCP5 controls leaf margin development by regulating KNOX and BEL-like transcription factors in Arabidopsis. J. Exp. Bot. 2021, 72, 1809–1821. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, J.; Li, C.; Deng, X.; Wang, T.; Dong, L. Genome-wide analysis of TCP transcription factor family in sunflower and identification of HaTCP1 involved in the regulation of shoot branching. BMC Plant Biol. 2023, 23, 222. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Sun, M.; Pan, H.; Cheng, T.; Zhang, Q. Two Cyc2CL transcripts (Cyc2CL-1 and Cyc2CL-2) may play key roles in the petal and stamen development of ray florets in chrysanthemum. BMC Plant Biol. 2021, 21, 105. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Irish, V.F. Temporal control of plant organ growth by TCP transcription factors. Curr. Biol. 2015, 25, 1765–1770. [Google Scholar] [CrossRef] [PubMed]
- Giraud, E.; Ng, S.; Carrie, C.; Duncan, O.; Low, J.; Lee, C.P.; Van Aken, O.; Millar, A.H.; Murcha, M.; Whelan, J. TCP transcription factors link the regulation of genes encoding mitochondrial proteins with the circadian clock in Arabidopsis thaliana. Plant Cell 2010, 22, 3921–3934. [Google Scholar] [CrossRef]
- Yao, Y.; Dong, L.; Fu, X.; Zhao, L.; Wei, J.; Cao, J.; Sun, Y.; Liu, J. HrTCP20 dramatically enhance drought tolerance of sea buckthorn (Hippophae rhamnoides L.) by mediating the JA signaling pathway. Plant Physiol. Biochem. 2022, 174, 51–62. [Google Scholar] [CrossRef] [PubMed]
- Almeida, D.M.; Gregorio, G.B.; Oliveira, M.M.; Saibo, N.J. Five novel transcription factors as potential regulators of OsNHX1 gene expression in a salt tolerant rice genotype. Plant Mol. Biol. 2017, 93, 61–77. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, P.; Tyagi, A.K. OsTCP19 influences developmental and abiotic stress signaling by modulating ABI4-mediated pathways. Sci. Rep. 2015, 5, 9998. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.; Cai, Z.; Du, H.; Wang, H. Genome-wide analysis of TCP family genes in Zea mays L. identified a role for ZmTCP42 in drought tolerance. Int. J. Mol. Sci. 2019, 20, 2762. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Liu, H.; Gao, Y.; Xiong, R.; Wu, M.; Zhang, K.; Xiang, Y. The TCP transcription factor PeTCP10 modulates salt tolerance in transgenic Arabidopsis. Plant Cell Rep. 2021, 40, 1971–1987. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Shi, X.; Wang, Z.; Qu, M.; Gao, C.; Wang, C.; Wang, Y. Acetylation of transcription factor BpTCP20 by acetyltransferase BpPDCE23 modulates salt tolerance in birch. Plant Physiol. 2024, 195, kiae168. [Google Scholar] [CrossRef] [PubMed]
- Song, A.; Su, J.; Wang, H.; Zhang, Z.; Zhang, X.; Van de Peer, Y.; Chen, F.; Fang, W.; Guan, Z.; Zhang, F. Analyses of a chromosome-scale genome assembly reveal the origin and evolution of cultivated chrysanthemum. Nat. Commun. 2023, 14, 2021. [Google Scholar] [CrossRef]
- Manassero, N.G.U.; Viola, I.L.; Welchen, E.; Gonzalez, D.H. TCP transcription factors: Architectures of plant form. Biomol. Concepts 2013, 4, 111–127. [Google Scholar] [CrossRef] [PubMed]
- Parapunova, V.; Busscher, M.; Busscher-Lange, J.; Lammers, M.; Karlova, R.; Bovy, A.G.; Angenent, G.C.; de Maagd, R.A. Identification, cloning and characterization of the tomato TCP transcription factor family. BMC Plant Biol. 2014, 14, 157. [Google Scholar] [CrossRef]
- Francis, A.; Dhaka, N.; Bakshi, M.; Jung, K.-H.; Sharma, M.K.; Sharma, R. Comparative phylogenomic analysis provides insights into TCP gene functions in Sorghum. Sci. Rep. 2016, 6, 38488. [Google Scholar] [CrossRef]
- Wang, J.; Guan, Y.; Ding, L.; Li, P.; Zhao, W.; Jiang, J.; Chen, S.; Chen, F. The CmTCP20 gene regulates petal elongation growth in Chrysanthemum morifolium. Plant Sci. 2019, 280, 248–257. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.-M.; Liu, B.-W.; Ma, F.-F.; Sun, X.; Zheng, C.-S. Proteomic profiling of root system development proteins in chrysanthemum overexpressing the CmTCP20 gene. Plant Sci. 2019, 287, 110175. [Google Scholar] [CrossRef]
- Zhang, T.; Qu, Y.; Wang, H.; Wang, J.; Song, A.; Hu, Y.; Chen, S.; Jiang, J.; Chen, F. The heterologous expression of a chrysanthemum TCP-P transcription factor CmTCP14 suppresses organ size and delays senescence in Arabidopsis thaliana. Plant Physiol. Biochem. 2017, 115, 239–248. [Google Scholar] [CrossRef] [PubMed]
- Danisman, S.; van Dijk, A.D.; Bimbo, A.; van der Wal, F.; Hennig, L.; de Folter, S.; Angenent, G.C.; Immink, R.G. Analysis of functional redundancies within the Arabidopsis TCP transcription factor family. J. Exp. Bot. 2013, 64, 5673–5685. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, P.; Das Gupta, M.; Joseph, A.P.; Chatterjee, N.; Srinivasan, N.; Nath, U. Identification of specific DNA binding residues in the TCP family of transcription factors in Arabidopsis. Plant Cell 2010, 22, 1174–1189. [Google Scholar] [CrossRef] [PubMed]
- Urano, K.; Maruyama, K.; Koyama, T.; Gonzalez, N.; Inzé, D.; Yamaguchi-Shinozaki, K.; Shinozaki, K. CIN-like TCP13 is essential for plant growth regulation under dehydration stress. Plant Mol. Biol. 2022, 108, 257–275. [Google Scholar] [CrossRef]
- Wang, Y.; Yu, Y.; Wan, H.; Tang, J.; Ni, Z. The sea-island cotton GbTCP4 transcription factor positively regulates drought and salt stress responses. Plant Sci. 2022, 322, 111329. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yang, Y.-N.; Xue, L.-J.; Zou, M.-J.; Liu, J.-Y.; Chen, F.; Xue, H.-W. Rice ABI5-Like1 regulates abscisic acid and auxin responses by affecting the expression of ABRE-containing genes. Plant Physiol. 2011, 156, 1397–1409. [Google Scholar] [CrossRef]
- Zhang, J.; Jia, W.; Yang, J.; Ismail, A.M. Role of ABA in integrating plant responses to drought and salt stresses. Field Crops Res. 2006, 97, 111–119. [Google Scholar] [CrossRef]
- Wang, Y.; Mostafa, S.; Zeng, W.; Jin, B. Function and mechanism of jasmonic acid in plant responses to abiotic and biotic stresses. Int. J. Mol. Sci. 2021, 22, 8568. [Google Scholar] [CrossRef]
- Wang, J.; Song, L.; Gong, X.; Xu, J.; Li, M. Functions of jasmonic acid in plant regulation and response to abiotic stress. Int. J. Mol. Sci. 2020, 21, 1446. [Google Scholar] [CrossRef]
- Yang, J.; Duan, G.; Li, C.; Liu, L.; Han, G.; Zhang, Y.; Wang, C. The crosstalks between jasmonic acid and other plant hormone signaling highlight the involvement of jasmonic acid as a core component in plant response to biotic and abiotic stresses. Front. Plant Sci. 2019, 10, 1349. [Google Scholar] [CrossRef]
- Song, W.; Shao, H.; Zheng, A.; Zhao, L.; Xu, Y. Advances in roles of salicylic acid in plant tolerance responses to biotic and abiotic stresses. Plants 2023, 12, 3475. [Google Scholar] [CrossRef]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wu, Y.; Dong, G.; Zhu, G.; Zhou, G. Progress of research on the physiology and molecular regulation of sorghum growth under salt stress by gibberellin. Int. J. Mol. Sci. 2023, 24, 6777. [Google Scholar] [CrossRef] [PubMed]
- Walker, J.C.; Howard, E.A.; Dennis, E.S.; Peacock, W.J. DNA sequences required for anaerobic expression of the maize alcohol dehydrogenase 1 gene. Proc. Natl. Acad. Sci. USA 1987, 84, 6624–6628. [Google Scholar] [CrossRef]
- Park, H.C.; Kim, M.L.; Kang, Y.H.; Jeon, J.M.; Yoo, J.H.; Kim, M.C.; Park, C.Y.; Jeong, J.C.; Moon, B.C.; Lee, J.H.; et al. Pathogen- and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol. 2004, 135, 2150–2161. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Cui, M.-Y.; Hu, Y.; Gao, K.; Xie, Y.-G.; Jiang, Y.; Feng, J.-Y. Ectopic expression of FvWRKY42, a WRKY transcription factor from the diploid woodland strawberry (Fragaria vesca), enhances resistance to powdery mildew, improves osmotic stress resistance, and increases abscisic acid sensitivity in Arabidopsis. Plant Sci. 2018, 275, 60–74. [Google Scholar] [CrossRef]
- Santos, C.V. Regulation of chlorophyll biosynthesis and degradation by salt stress in sunflower leaves. Sci. Hortic. 2004, 103, 93–99. [Google Scholar] [CrossRef]
- Eckhardt, U.; Grimm, B.; Hörtensteiner, S. Recent advances in chlorophyll biosynthesis and breakdown in higher plants. Plant Mol. Biol. 2004, 56, 1–14. [Google Scholar] [CrossRef]
- Sanders, D. Plant biology: The salty tale of Arabidopsis. Curr. Biol. 2000, 10, 486–488. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Chen, Z.-Z.; Zhou, X.-F.; Yin, H.-B.; Li, X.; Xin, X.-F.; Hong, X.-H.; Zhu, J.-K.; Gong, Z. Overexpression of SOS (Salt Overly Sensitive) genes increases salt tolerance in transgenic Arabidopsis. Mol. Plant 2009, 2, 22–31. [Google Scholar] [CrossRef]
- Halfter, U.; Ishitani, M.; Zhu, J.-K. The Arabidopsis SOS2 protein kinase physically interacts with and is activated by the calcium-binding protein SOS3. Proc. Natl. Acad. Sci. USA 2000, 97, 3735–3740. [Google Scholar] [CrossRef]
- Qiu, Q.-S.; Guo, Y.; Dietrich, M.A.; Schumaker, K.S.; Zhu, J.-K. Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc. Natl. Acad. Sci. USA 2002, 99, 8436–8441. [Google Scholar] [CrossRef] [PubMed]
- Strizhov, N.; Ábrahám, E.; Ökrész, L.; Blickling, S.; Zilberstein, A.; Schell, J.; Koncz, C.; Szabados, L. Differential expression of two P5CS genes controlling proline accumulation during salt-stress requires ABA and is regulated by ABA1, ABI1 and AXR2 in Arabidopsis. Plant J. 1997, 12, 557–569. [Google Scholar] [CrossRef] [PubMed]
- Amini, S.; Ghobadi, C.; Yamchi, A. Proline accumulation and osmotic stress: An overview of P5CS gene in plants. J. Plant Mol. Breed. 2015, 3, 44–55. [Google Scholar]
- Zhu, J.-K. Plant salt tolerance. Trends Plant Sci. 2001, 6, 66–71. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, P.K.; Agarwal, P.; Reddy, M.; Sopory, S.K. Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep. 2006, 25, 1263–1274. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, H.; Ding, L.; Song, A.; Shen, F.; Jiang, J.; Chen, S.; Chen, F. Transcriptomic and hormone analyses reveal mechanisms underlying petal elongation in Chrysanthemum morifolium ‘Jinba’. Plant Mol. Biol. 2017, 93, 593–606. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Wang, F.-Q.; Zhang, Z.-Y. Interactions among mediator subunits of tobacco by bimolecular fluorescence complementation (BiFC) method. J. Agric. Biotechnol. 2012, 20, 38–47. [Google Scholar]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Murray, M.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4325. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Song, A.; Gao, C.; Wang, L.; Wang, Y.; Sun, J.; Jiang, J.; Chen, F.; Chen, S. Chrysanthemum WRKY gene CmWRKY17 negatively regulates salt stress tolerance in transgenic chrysanthemum and Arabidopsis plants. Plant Cell Rep. 2015, 34, 1365–1378. [Google Scholar] [CrossRef]
- Yadav, N.S.; Shukla, P.S.; Jha, A.; Agarwal, P.K.; Jha, B. The SbSOS1 gene from the extreme halophyte Salicornia brachiata enhances Na+ loading in xylem and confers salt tolerance in transgenic tobacco. BMC Plant Biol. 2012, 12, 188. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, H.K. Chlorophylls and carotenoids: Pigments of photosynthetic biomembranes. Method. Enzymol. 1987, 148, 350–382. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


| Motif Name | Motif Sequence | Function | No. | Position |
|---|---|---|---|---|
| ABRE | ACGTG/CACGTG/GCAACGTGTC | Abscisic acid response element | 5 | 1704, −2123, −1780, −1778, 1781 |
| ARE | AAACCA | Anaerobic-responsive element | 6 | 195, −1655, −1191, 1903, −871, 1536 |
| ATC-motif | AGTAATCT | Light-responsive element | 1 | 1722 |
| AuxRR-core | GGTCCAT | Auxin response element | 1 | 555 |
| Box 4 | ATTAAT | Light-responsive element | 1 | 905 |
| Box III | atCATTTTCACt | Protein binding site | 1 | 291 |
| CGTCA-motif | CGTCA | MeJA-responsive element | 2 | 75, −1715 |
| ERE | ATTTTAAA | Ethylene-responsive element | 1 | 636 |
| G-box | CACGTT/CACGTG/ACACGTGGC/TACGTG | Light-responsive element | 4 | −1703, −1780, 1799, −2123 |
| GA-motif | ATAGATAA | Light-responsive element | 1 | 606 |
| GATA-motif | AAGATAAGATT/AAGGATAAGG | Light-responsive element | 3 | −211, 2302, −2164 |
| GT-1 motif | GAAAAA | Salinity stress-inducible element | 7 | 95, 425, 1385, 2048, −294, −1145, −2342 |
| HD-Zip 3 | GTAAT(G/C)ATTAC | Protein binding site | 1 | 701 |
| I-box | AGATAAGG/cCATATCCAAT | Part of a light-responsive element | 2 | 239, −1832 |
| LTR | CCGAAA | Low-temperature response element | 1 | 423 |
| MBS | CAACTG | MYB binding site involved in drought-inducibility | 1 | 1267 |
| MYB | TAACCA/CAACCA/CAACAG | MYB/YB recognition site | 5 | −1027, −2065, −1844, −1840, −2003 |
| MYC | CATGTG | MYC/YC recognition site | 1 | 1283 |
| P-box | CCTTTTG | Gibberellin-responsive element | 1 | 695 |
| TCA-element | CCATCTTTTT | Salicylic acid-responsive element | 1 | 289 |
| TGA-element | AACGAC | Auxin-responsive element | 1 | −44 |
| TGACG-motif | TGACG | MeJA-responsive element | 2 | 5, −5 |
| WUN-motif | AAATTTCCT | Wound-responsive element | 1 | −111 |
| Circadian | CAAAGATATC | Circadian regulation element | 1 | −214 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chong, X.; Liu, Y.; Li, P.; Wang, Y.; Zhou, T.; Chen, H.; Wang, H. Heterologous Expression of Chrysanthemum TCP Transcription Factor CmTCP13 Enhances Salinity Tolerance in Arabidopsis. Plants 2024, 13, 2118. https://doi.org/10.3390/plants13152118
Chong X, Liu Y, Li P, Wang Y, Zhou T, Chen H, Wang H. Heterologous Expression of Chrysanthemum TCP Transcription Factor CmTCP13 Enhances Salinity Tolerance in Arabidopsis. Plants. 2024; 13(15):2118. https://doi.org/10.3390/plants13152118
Chicago/Turabian StyleChong, Xinran, Yanan Liu, Peiling Li, Yue Wang, Ting Zhou, Hong Chen, and Haibin Wang. 2024. "Heterologous Expression of Chrysanthemum TCP Transcription Factor CmTCP13 Enhances Salinity Tolerance in Arabidopsis" Plants 13, no. 15: 2118. https://doi.org/10.3390/plants13152118







