Association Mapping of Seed Coat Color Characteristics for Near-Isogenic Lines of Colored Waxy Maize Using Simple Sequence Repeat Markers
Abstract
:1. Introduction
2. Results
2.1. Anthocyanin Content and Seed Coat Color Analysis and Correlation Analysis
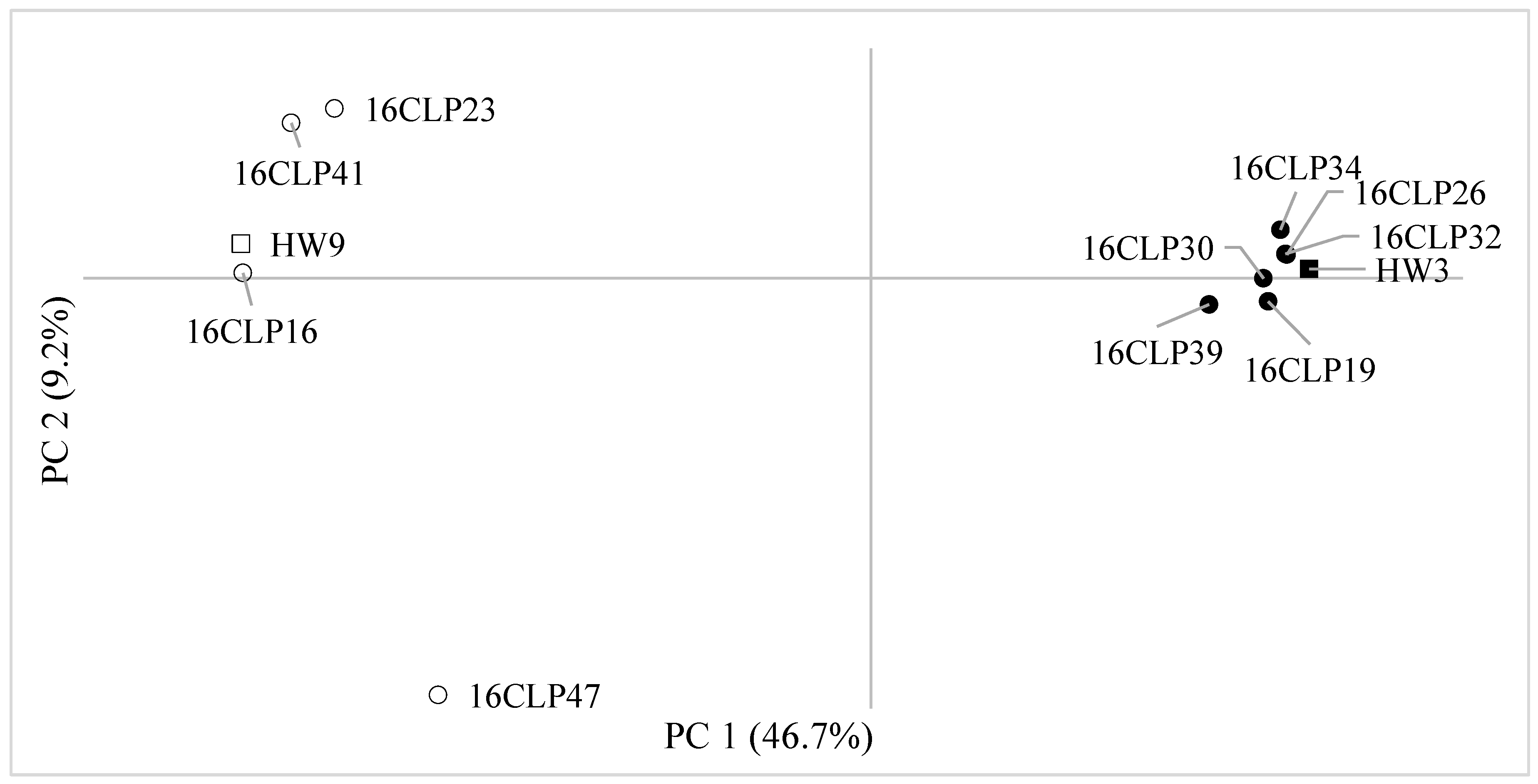
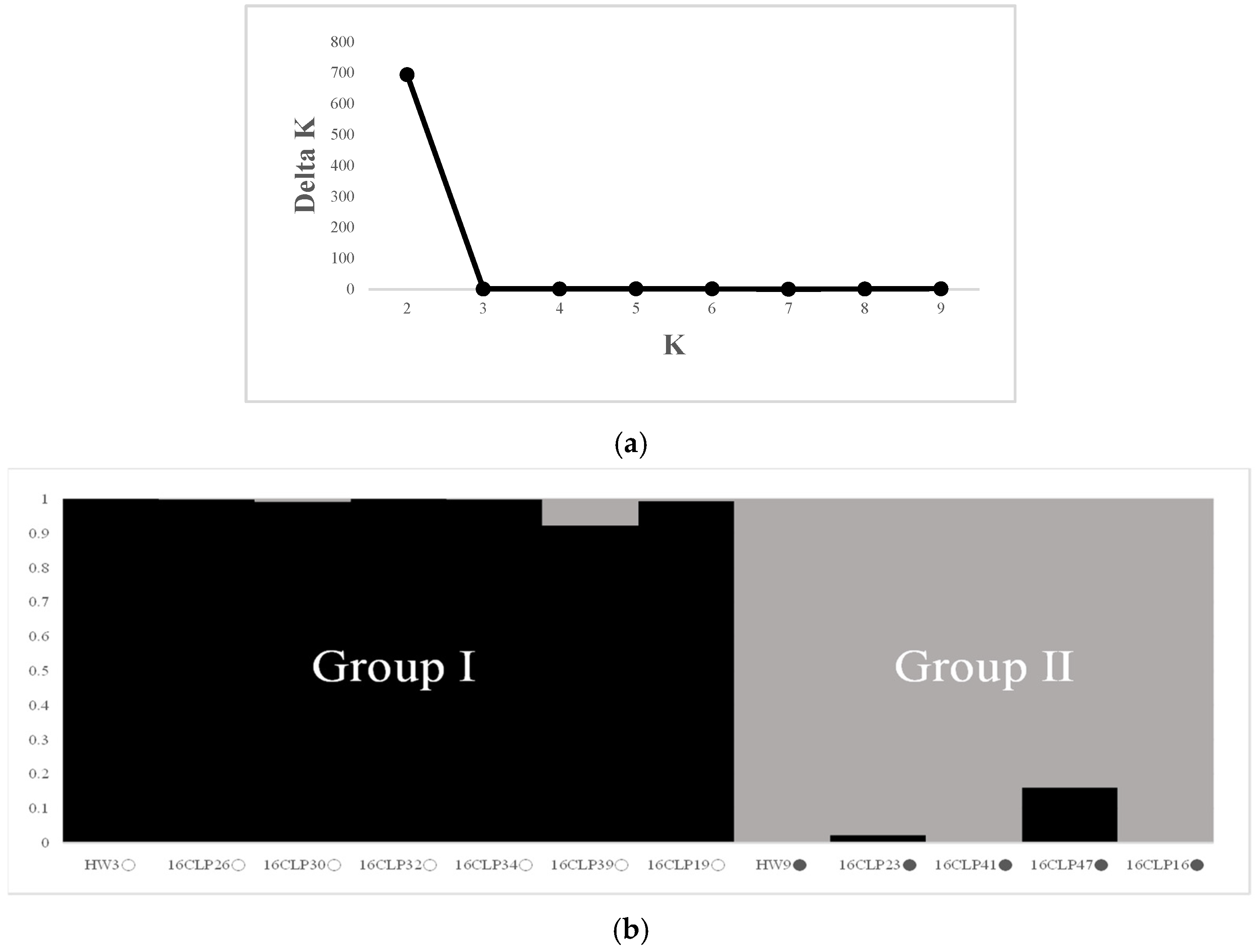
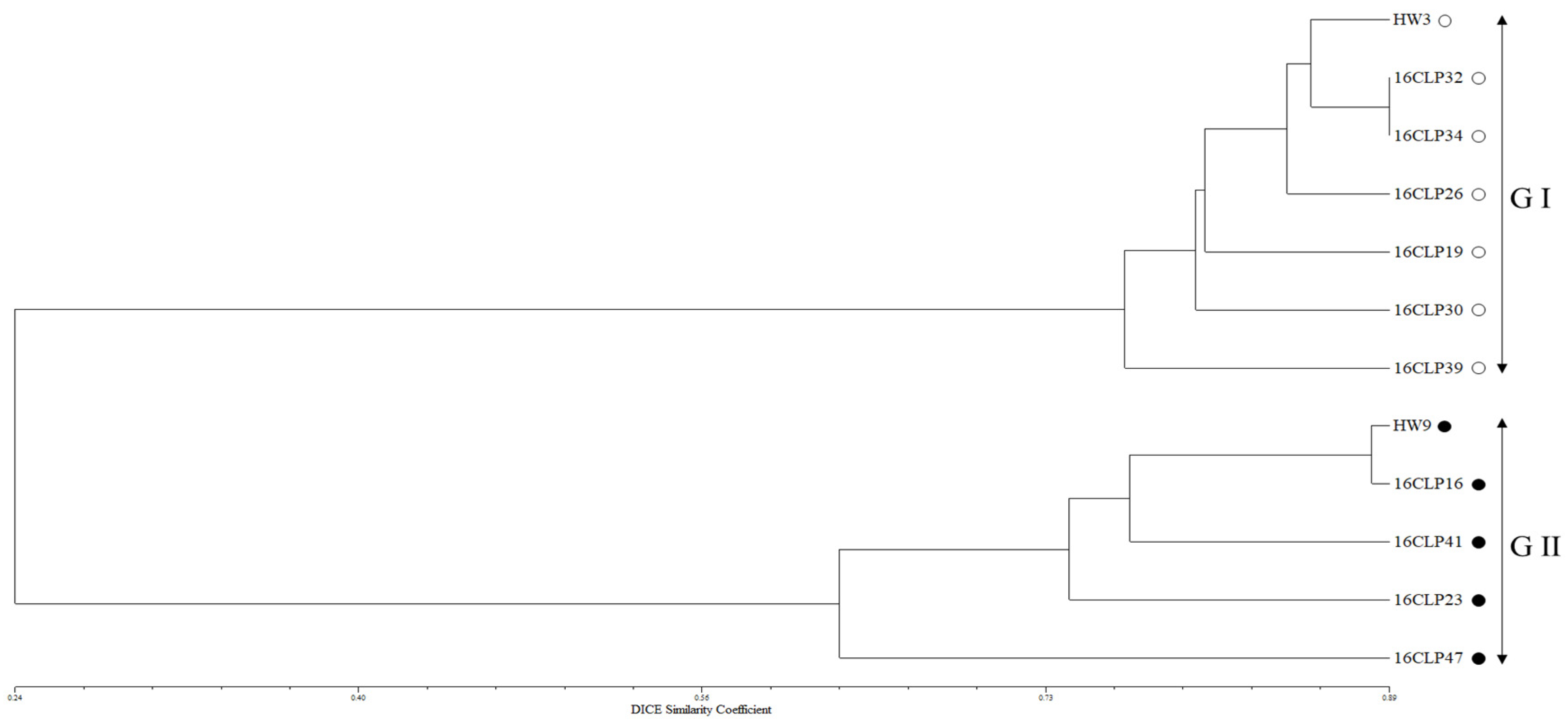
2.2. Genetic Diversity and Relationships and Population Structure among 10 NILs of Colored Waxy Maize and Two Parental Lines (HW3, HW9) of “Mibaek 2ho” Variety Waxy Maize
2.3. AMA among 10 NILs Using SSR Markers and Anthocyanin Content and Seed Coat Color Characteristics
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Seed Trait Analysis
4.2. DNA Extraction and SSR Amplification Analysis
4.3. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Erenstein, O.; Jaleta, M.; Sonder, K.; Mottaleb, K.; Prasanna, B.M. Global maize production, consumption and trade: Trends and R&D implications. Food Secur. 2022, 14, 1295–1319. [Google Scholar]
- Sprague, G.F.; Brimhall, B.; Hixon, R.M. Some effects of the waxy gene in corn on properties of the endosperm starch. J. Am. Soc. Agron. 1943, 35, 817–822. [Google Scholar] [CrossRef]
- Nelson, O.E.; Rines, H.W. The enzymatic deficiency in the waxy mutant of maize. Biochem. Biophys. Res. Commun. 1962, 9, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Sa, K.J.; Park, J.Y.; Woo, S.Y.; Ramekar, R.V.; Jang, C.S.; Lee, J.K. Mapping of QTL traits in corn using a RIL population derived from a cross of dent corn x waxy corn. Genes Genom. 2015, 37, 1–14. [Google Scholar] [CrossRef]
- Wu, X.; Chen, D.; Lu, Y.; Liu, W.; Yang, X.; Li, X.; Du, J.; Li, L. Molecular characteristics of two new waxy mutations in China waxy maize. Mol. Breed. 2017, 37, 27. [Google Scholar]
- Talukder, Z.A.; Muthusamy, V.; Chhabra, R.; Gain, N.; Reddappa, S.B.; Mishra, S.J.; Kasana, R.; Bhatt, V.; Chand, G.; Katral, A.; et al. Combining higher accumulation of amylopectin, lysine and tryptophan in maize hybrids through genomics-assisted stacking of waxy1 and opaque2 genes. Sci. Rep. 2022, 12, 706. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.R.; Sa, K.J.; Nam-Gung, M.; Park, K.J.; Ryu, S.H.; Mo, C.Y.; Lee, J.K. Genetic characterization and association mapping in near-isogenic lines of waxy maize using seed characteristics and SSR markers. Genes Genom. 2021, 43, 79–90. [Google Scholar] [CrossRef]
- Chatham, L.A.; Paulsmeyer, M.; Juvik, J.A. Prospects for economical natural colorants: Insights from maize. Theor. Appl. Genet. 2019, 132, 2927–2946. [Google Scholar] [CrossRef] [PubMed]
- Brewer, M.S. Natural antioxidants: Sources, compounds, mechanisms of action, and potential applications. Compr. Rev. Food Sci. Food Saf. 2011, 10, 221–247. [Google Scholar] [CrossRef]
- Dong, Y.; Wu, X.; Han, L.; Bian, J.; He, C.; El-Omar, E.; Gong, L.; Wang, M. The potential roles of dietary anthocyanins in inhibiting vascular endothelial cell senescence and preventing cardiovascular diseases. Nutrients 2022, 14, 2836. [Google Scholar] [CrossRef]
- Shi, N.; Chen, X.; Chen, T. Anthocyanins in colorectal cancer prevention review. Antioxidants 2021, 10, 1600. [Google Scholar] [CrossRef] [PubMed]
- Park, K.J.; Park, J.Y.; Ryu, S.H.; Goh, B.D.; Seo, J.S.; Min, H.K.; Jung, T.W.; Huh, C.S.; Ryu, I.M. A new waxy corn hybrid cultivar, “Mibaek 2” with good eating quality and lodging resistance. Korean J. Breed. Sci. 2007, 39, 108–109. [Google Scholar] [CrossRef] [PubMed]
- Leon, K.; Mery, D.; Pedreschi, F.; Leon, J. Color measurement in L* a* b* units from RGB digital images. Food Res. Int. 2006, 39, 1084–1091. [Google Scholar] [CrossRef]
- Dana, W.; Ivo, W. Computer image analysis of seed shape and seed color for flax cultivar description. Comp. Elect. Agric. 2008, 61, 126–135. [Google Scholar] [CrossRef]
- Quevedo, R.; Aguilera, J.; Pedreschi, F. Color of salmon fillets by computer vision and sensory panel. Food Bioprocess Technol. 2010, 3, 637–643. [Google Scholar] [CrossRef]
- Park, Y.J.; Lee, J.K.; Kim, N.S. Simple Sequence Repeat Polymorphisms (SSRPs) for Evaluation of Molecular Diversity and Germplasm Classification of Minor Crops. Molecules 2009, 14, 4546–4569. [Google Scholar] [CrossRef]
- Park, K.J.; Sa, K.J.; Koh, H.J.; Lee, J.K. QTL analysis for eating quality-related traits in an F2:3 population derived from waxy corn x sweet corn cross. Breed. Sci. 2013, 63, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.K.; Sa, K.J.; Park, D.H.; Lim, S.E.; Ryu, S.H.; Park, J.Y.; Park, K.J.; Rhee, H.I.; Lee, M.; Lee, J.K. Construction of genetic linkage map and identification of QTLs related to agronomic traits in DH population of maize (Zea mays L.) using SSR markers. Genes Genom. 2019, 41, 667–678. [Google Scholar] [CrossRef] [PubMed]
- Pejic, I.; Ajmone-Marsan, P.; Morgante, M.; Kozumplick, V.; Castiglioni, P.; Taramino, G.; Motto, M. Comparative analysis of genetic similarity among maize inbred lines detected by RFLPs, RAPDs, SSR, and AFLPs. Theor. Appl. Genet. 1998, 97, 1248–1255. [Google Scholar] [CrossRef]
- Stich, B.; Melchinger, A.E.; Frisch, M.; Maurer, H.P.; Hecknberger, M.; Reif, J.C. Linkage disequilibrium in European elite maize germplasm investigated with SSRs. Theor. Appl. Genet. 2005, 111, 723–730. [Google Scholar] [CrossRef]
- Szalma, S.J.; Hostert, B.M.; LeDeaux, J.R.; Stuber, C.W.; Holland, J.B. QTL mapping with near-isogenic lines in maize. Theor. Appl. Genet. 2007, 114, 1211–1228. [Google Scholar] [CrossRef]
- Kim, O.G.; Sa, K.J.; Lee, J.R.; Lee, J.K. Genetic analysis of maize germplasm in the Korean Genebank and association with agronomic traits and simple sequence repeat markers. Genes Genom. 2017, 39, 843–853. [Google Scholar] [CrossRef]
- Andorf, C.; Beavis, W.D.; Hufford, M.; Smith, S.; Suza, W.P.; Wang, K.; Woodhouse, M.; Yu, J.; Lübberstedt, T. Technological advances in maize breeding: Past, present and future. Theor. Appl. Genet. 2019, 132, 817–849. [Google Scholar] [CrossRef]
- Tanksley, S.D.; Nelson, J.C. Advanced backcross QTL analysis: A method for the simultaneous discovery and transfer of valuable QTLs from unadapted germplasm into elite breeding lines. Theor. Appl. Genet. 1996, 92, 191–203. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Rosenberg, N.A.; Donnelly, P. Association Mapping in Structured Populations. Am. J. Hum. Genet. 2000, 67, 170–181. [Google Scholar] [CrossRef]
- Skot, L.; Humphreys, M.O.; Armstead, I. An association mapping approach to identify flowering time genes in natural populations of Loliumperenne (L.). Mol. Breed. 2005, 15, 233–245. [Google Scholar] [CrossRef]
- Lim, S.E.; Sa, K.J.; Lee, J.K. Bulk segreant analysis identifies SSR markers associated with leaf- and seed-related traits in Perilla crop (Perilla frutescens L.). Gens Genome 2021, 43, 323–332. [Google Scholar]
- Sivakumar, S.; Dhasarathan, M.; Karthikeyan, A.; Bharathi, P.; Kumari vinodhana, N.; Ganesamurthy, K.; Senthil, N. Population structure and association mapping studies for yield-related traits in Maize (Zea mays L.). Cur. Plant Biol. 2019, 18, 100103. [Google Scholar] [CrossRef]
- Ha, Y.J.; Sa, K.J.; Lee, J.K. Identifying SSR markers associated with seed characteristics in Perilla (Perilla frutescens L.). Physiol. Mol. Biol. Plants 2021, 27, 93–105. [Google Scholar] [CrossRef]
- Pasam, R.K.; Sharma, R.; Malosetti, M.; van Eeuwijk, F.A.; Haseneyer, G.; Kilian, B.; Graner, A. Genome-wide association studies for agronomical traits in a worldwide spring barley collection. BMC Plant Biol. 2012, 27, 12–16. [Google Scholar]
- Raboin, L.M.; Ballini, E.; Tharreau, D.; Ramanantsoanirina, A.; Frouin, J.; Courtois, B.; Ahmadi, N. Association mapping of resistance to rice blast in upland field conditions. Rice 2016, 9, 59. [Google Scholar] [CrossRef]
- Qaseem, M.F.; Qureshi, R.; Muqaddasi, Q.H.; Shaheen, H.; Kousar, R.; Röder, M.S. Genome-wide association mapping in bread wheat subjected to independent and combined high temperature and drought stress. PLoS ONE 2018, 13, e0199121. [Google Scholar] [CrossRef]
- Sa, K.J.; Park, H.; Jang, S.J.; Lee, J.K. Association Mapping of Amylose Content in Maize RIL Population Using SSR and SNP Markers. Plants 2023, 12, 239. [Google Scholar] [CrossRef] [PubMed]
- Fujita, D.; Tagle, A.G.; Ebron, L.A.; Fukuta, Y.; Kobayashi, N. Characterization of near-isogenic lines carrying QTL for high spikelet number with the genetic background of an indica rice variety IR64 (Oryza sativa L.). Breed. Sci. 2012, 62, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Mia, M.S.; Liu, H.; Wang, X.; Yan, G. Multiple Near-Isogenic Lines Targeting a QTL Hotspot of Drought Tolerance Showed Contrasting Performance Under Post-anthesis Water Stress. Front. Plant Sci. 2019, 10, 271. [Google Scholar] [CrossRef] [PubMed]
- Pea, G.; Aung, H.H.; Frascaroli, F.; Landi, P.; Pè, M.E. Extensive genomic characterization of a set of near-isogenic lines for heterotic QTL in maize (Zea mays L.). BMC Genom. 2013, 14, 61. [Google Scholar] [CrossRef]
- Farré, A.; Sayers, L.; Leverington-Waite, M.; Goram, R.; Orford, S.; Wingen, L.; Mumford, C.; Griffiths, S. Application of a library of near isogenic lines to understand context dependent expression of QTL for grain yield and adaptive traits in bread wheat. BMC Plant Biol. 2016, 16, 161. [Google Scholar] [CrossRef]
- Borba, T.C.O.; Brondani, R.P.V.; Breseghello, F.; Coelho, A.S.G.; Mendonça, J.A.; Rangel, P.H.N.; Brondani, C. Association mapping for yield and grain quality traits in rice (Oryza sativa L.). Genet. Mol. Biol. 2010, 33, 515–524. [Google Scholar] [CrossRef]
- Falke, K.C.; Frisch, M. Power and false-positive rate in QTL detection with near-isogenic line libraries. Heredity 2011, 106, 576–584. [Google Scholar] [CrossRef]
- Stuber, C.W.; Polacco, M.; Senior, M.L. Synergy of empirical breeding, marker-assisted selection, genomics, and genetic engineering to increase crop yield potential. Crop Sci. 1999, 39, 1571–1583. [Google Scholar] [CrossRef]
- Zhang, Q.; Wu, C.; Ren, F.; Li, Y.; Zhang, C. Association analysis of important agronomical traits of maize inbred lines with SSRs. Aust. J. Crop Sci. 2012, 6, 1131–1138. [Google Scholar]
- Kandianis, C.B.; Stevens, R.; Liu, W.; Palacios, N.; Montgomery, K.; Pixley, K.; White, W.S.; Rocheford, T. Genetic architecture controlling variation in grain carotenoid composition and concentrations in two maize populations. Theor. Appl. Genet. 2013, 126, 2879–2895. [Google Scholar] [CrossRef] [PubMed]
- Ashokkumar, K.; Govindaraj, M.; Karthikeyan, A.; Shobhana, V.G.; Warkentin, T.D. Genomics-Integrated Breeding for Carotenoids and Folates in Staple Cereal Grains to Reduce Malnutrition. Front. Genet. 2020, 11, 414. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Tang, Y.; Jin, W.; Liu, Y.; Li, G.; Zhong, W.; Huang, J.; Wang, W. QTL Mapping of Zeaxanthin Content in Sweet Corn Using Recombinant Inbred Line Population across Different Environments. Plants 2023, 12, 3506. [Google Scholar] [CrossRef] [PubMed]
- Oh, Y.S.; Lee, J.H.; Yoon, S.H.; Oh, C.H.; Choi, D.S.; Choe, E.; Jung, M.Y. Characterization and Quantification of Anthocyanins in Grape Juices Obtained from the Grapes Cultivated in Korea by HPLC/DAD, HPLC/MS, and HPLC/MS/MS. J. Food Sci. 2008, 73, C378–C389. [Google Scholar] [CrossRef] [PubMed]
- Dellaporta, S.L.; Wood, J.; Hicks, J.B. A simple and rapid method for plant DNA preparation. Version II. Plant Mol. Biol. Rep. 1983, 1, 19–21. [Google Scholar] [CrossRef]
- Liu, K.; Muse, S.V. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef] [PubMed]
- Dice, L.R. Measures of the amount of ecologic association between species. Ecology 1945, 26, 297–302. [Google Scholar] [CrossRef]
- Rohlf, F.J. NTSYS- pc: Numerical Taxonomy and Multivariate Analysis System; Version 2.02; Exeter Software: Setauket, NY, USA, 1998. [Google Scholar]
- Pritchard, J.K.; Wen, W. Documentation for STRUCTURE Software; Version 2; 2003. Available online: http://pritc.h.bsd.uchicago.edu/structure.html (accessed on 1 January 2023).
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics 2007, 23, 2633–2635. [Google Scholar] [CrossRef]

| NILs | Kuromanin | Peonidin | R | V | L* | a* | b* | 750 nm |
|---|---|---|---|---|---|---|---|---|
| 16CLP26 | 1.5 | 0.895 | 59.1 | 59.1 | 26.3 | 151.2 | 139.2 | 0.394 |
| 16CLP30 | 0.187 | 0.164 | 120.8 | 120.9 | 86.2 | 151.9 | 143.6 | 0.479 |
| 16CLP32 | 0.305 | 0.28 | 88.3 | 88.3 | 46.4 | 159.1 | 145.7 | 0.467 |
| 16CLP34 | 1.42 | 0.355 | 124.7 | 124.7 | 80.0 | 160.2 | 146.3 | 0.498 |
| 16CLP39 | 19.5 | 7.34 | 20.8 | 20. 9 | 7.43 | 134.2 | 129.7 | 0.278 |
| 16CLP19 | 0.707 | 0.495 | 68.7 | 68.7 | 32.7 | 153.9 | 141.3 | 0.416 |
| 16CLP23 | 1.09 | 0.579 | 66.7 | 66.7 | 31.4 | 153.8 | 141.4 | 0.413 |
| 16CLP41 | 0.175 | 0.016 | 115.9 | 115.9 | 67.2 | 163.7 | 152.8 | 0.467 |
| 16CLP47 | 5.73 | 1.29 | 17.9 | 17.9 | 6.91 | 132.6 | 129.2 | 0.269 |
| 16CLP16 | 1.20 | 0.683 | 45.6 | 45.6 | 20.9 | 143.7 | 134.9 | 0.381 |
| Mean | 3.18 | 1.21 | 72.9 | 72.9 | 40.5 | 150.4 | 140.4 | 0.406 |
| Min | 0.175 | 0.016 | 17.9 | 17.9 | 6.91 | 132.6 | 129.2 | 0.269 |
| Max | 19.5 | 7.34 | 124.7 | 124.7 | 86.2 | 163.7 | 152.8 | 0.498 |
| SD | 5.66 | 2.07 | 37.2 | 37.1 | 27.1 | 10.01 | 7.07 | 0.076 |
| Peonidin | R | V | L* | a* | b* | 750 nm | |
|---|---|---|---|---|---|---|---|
| Kuromanin | 0.988 ** | −0.618 | −0.618 | −0.54 | −0.708 * | −0.669 * | −0.737 * |
| Peonidin | −0.595 | −0.595 | −0.531 | −0.657 * | −0.633 * | −0.687 * | |
| R | 1.000 ** | 0.980 ** | 0.874 ** | 0.915 ** | 0.953 ** | ||
| V | 0.980 ** | 0.874 ** | 0.915 ** | 0.953 ** | |||
| L* | 0.760 * | 0.824 ** | 0.893 ** | ||||
| a* | 0.974 ** | 0.936 ** | |||||
| b* | 0.920 ** |
| Chromosome | No. of SSR Loci | Total Alleles | Mean of Alleles | MAF | GD | PIC |
|---|---|---|---|---|---|---|
| Ch.1 | 26 | 76 | 2.92 | 0.551 | 0.545 | 0.463 |
| Ch.2 | 27 | 71 | 2.63 | 0.583 | 0.496 | 0.410 |
| Ch.3 | 27 | 74 | 2.74 | 0.599 | 0.512 | 0.427 |
| Ch.4 | 35 | 101 | 2.89 | 0.574 | 0.533 | 0.450 |
| Ch.5 | 24 | 65 | 2.71 | 0.601 | 0.513 | 0.433 |
| Ch.6 | 25 | 75 | 3.00 | 0.597 | 0.514 | 0.445 |
| Ch.7 | 41 | 111 | 2.71 | 0.583 | 0.511 | 0.426 |
| Ch.8 | 28 | 74 | 2.64 | 0.607 | 0.499 | 0.414 |
| Ch.9 | 25 | 67 | 2.68 | 0.580 | 0.518 | 0.428 |
| Ch.10 | 27 | 87 | 3.22 | 0.642 | 0.496 | 0.442 |
| Total | 285 | 801 | - | - | - | - |
| Mean | 29 | 80.1 | 2.81 | 0.592 | 0.514 | 0.434 |
| Min | 24 | 65 | 2.63 | 0.551 | 0.496 | 0.410 |
| Max | 41 | 111 | 3.22 | 0.642 | 0.545 | 0.463 |
| Trait | Marker | Chr. | GLM | Marker R2 | Trait | Marker | Chr. | GLM | Marker R2 |
|---|---|---|---|---|---|---|---|---|---|
| Kuromanin | bnlg1017 | 2 | ** | 0.963 | L* | umc1030 | 3 | * | 0.506 |
| bnlg238 | 6 | ** | 0.946 | umc2215 | 1 | * | 0.765 | ||
| umc1131 | 9 | ** | 0.948 | a* | bnlg1017 | 2 | * | 0.870 | |
| umc1447 | 5 | ** | 0.937 | umc1024 | 2 | * | 0.489 | ||
| umc1490 | 6 | ** | 0.993 | umc1030 | 3 | * | 0.612 | ||
| umc1798 | 2 | ** | 0.953 | umc1063 | 6 | * | 0.593 | ||
| umc1935 | 5 | ** | 0.930 | umc1490 | 6 | * | 0.727 | ||
| umc1946 | 2 | ** | 0.937 | umc2122 | 10 | * | 0.617 | ||
| umc2172 | 10 | * | 0.949 | umc2196 | 6 | ** | 0.804 | ||
| umc2196 | 6 | ** | 0.962 | umc2255 | 3 | ** | 0.723 | ||
| umc2255 | 3 | * | 0.457 | b* | bnlg1017 | 2 | * | 0.896 | |
| umc2320 | 6 | * | 0.415 | umc1012 | 3 | * | 0.582 | ||
| Peonidin | bnlg1017 | 2 | ** | 0.988 | umc1030 | 3 | * | 0.575 | |
| bnlg238 | 6 | ** | 0.973 | umc1061 | 10 | * | 0.590 | ||
| umc1131 | 9 | ** | 0.974 | umc1063 | 6 | * | 0.583 | ||
| umc1447 | 5 | ** | 0.973 | umc1490 | 6 | * | 0.601 | ||
| umc1490 | 6 | ** | 0.987 | umc2122 | 10 | * | 0.608 | ||
| umc1798 | 2 | ** | 0.988 | umc2196 | 6 | * | 0.721 | ||
| umc1935 | 5 | ** | 0.973 | umc2255 | 3 | * | 0.621 | ||
| umc1946 | 2 | ** | 0.973 | 750 nm | umc1030 | 3 | * | 0.553 | |
| umc2172 | 10 | ** | 0.987 | umc1063 | 6 | * | 0.591 | ||
| umc2196 | 6 | ** | 0.984 | umc1365 | 5 | * | 0.409 | ||
| umc2255 | 3 | * | 0.473 | umc1490 | 6 | ** | 0.769 | ||
| umc2320 | 6 | * | 0.405 | umc2122 | 10 | * | 0.582 | ||
| R | umc1030 | 3 | * | 0.568 | umc2196 | 6 | * | 0.727 | |
| umc1063 | 6 | * | 0.598 | umc2255 | 3 | * | 0.597 | ||
| umc2122 | 10 | * | 0.591 | ||||||
| umc2196 | 6 | * | 0.628 | ||||||
| umc2215 | 1 | * | 0.732 | ||||||
| umc2255 | 3 | * | 0.516 | ||||||
| V | umc1030 | 3 | * | 0.568 | |||||
| umc1063 | 6 | * | 0.598 | ||||||
| umc2122 | 10 | * | 0.591 | ||||||
| umc2196 | 6 | * | 0.628 | ||||||
| umc2215 | 1 | * | 0.732 | ||||||
| umc2255 | 3 | * | 0.516 |
| Abbreviation | Character | When/How Measured | Category |
|---|---|---|---|
| Kuromanin | Cyanidin 3-O-glucoside chloride | after harvest | ppm (μg/mL) |
| Peonidin | Peonidin-3-glucoside | after harvest | ppm (μg/mL) |
| R | Red color value of RGB color space | after harvest | Red color degree (0~255) |
| V | Value of HSV color space | after harvest | Darkness–lightness (0~255) |
| L* | L* value of CIELAB color space | after harvest | Darkness–lightness (0~255) |
| a* | a* value of CIELAB color space | after harvest | Greenness–redness (0~255) |
| b* | b* value of CIELAB color space | after harvest | Blueness–yellowness (0~255) |
| 750 nm | Hyperspectral image measurement | after harvest | Visible light (400~1000 nm region) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Heo, T.H.; Park, H.; Kim, N.-W.; Cho, J.; Mo, C.; Ryu, S.-H.; Choi, J.-K.; Park, K.J.; Sa, K.J.; Lee, J.K. Association Mapping of Seed Coat Color Characteristics for Near-Isogenic Lines of Colored Waxy Maize Using Simple Sequence Repeat Markers. Plants 2024, 13, 2126. https://doi.org/10.3390/plants13152126
Heo TH, Park H, Kim N-W, Cho J, Mo C, Ryu S-H, Choi J-K, Park KJ, Sa KJ, Lee JK. Association Mapping of Seed Coat Color Characteristics for Near-Isogenic Lines of Colored Waxy Maize Using Simple Sequence Repeat Markers. Plants. 2024; 13(15):2126. https://doi.org/10.3390/plants13152126
Chicago/Turabian StyleHeo, Tae Hyeon, Hyeon Park, Nam-Wook Kim, Jungeun Cho, Changyeun Mo, Si-Hwan Ryu, Jae-Keun Choi, Ki Jin Park, Kyu Jin Sa, and Ju Kyong Lee. 2024. "Association Mapping of Seed Coat Color Characteristics for Near-Isogenic Lines of Colored Waxy Maize Using Simple Sequence Repeat Markers" Plants 13, no. 15: 2126. https://doi.org/10.3390/plants13152126





