LzSCL9, a Novel GRAS Transcription Factor in Lanzhou Lily (Lilium davidii var. unicolor), Participates in Regulation of Trichokonins-Primed Heat Stress Tolerance
Abstract
1. Introduction
2. Results
2.1. GRAS Gene Expression Was Significantly Regulated by TK Treatment under HS
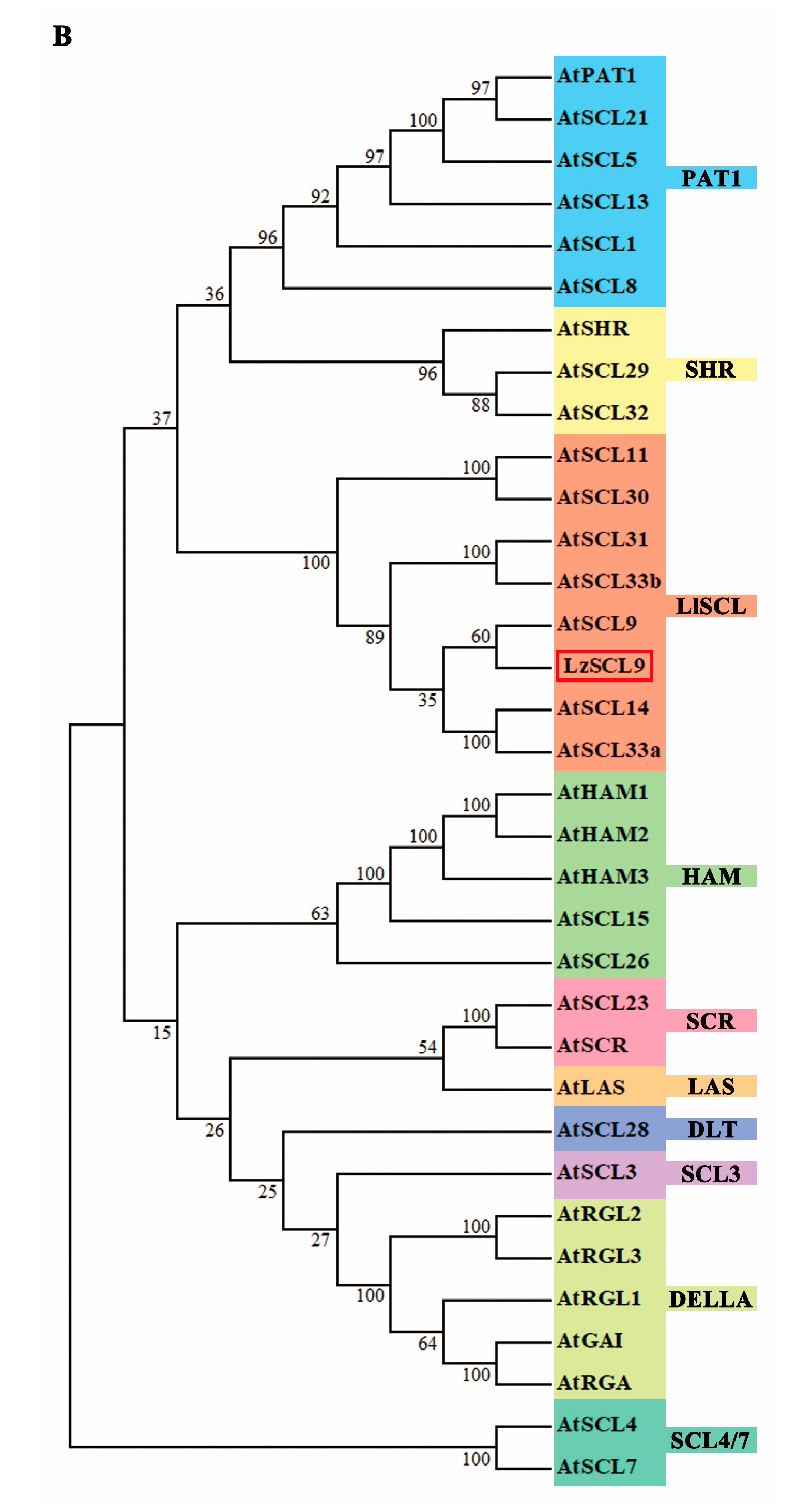
2.2. Cloning and Sequence Analysis of LzSCL9
2.3. LzSCL9 Transcript Was Induced by HS and HS + TKs Treatment
2.4. LzSCL9 Overexpression Enhanced the Thermotolerance of Lily
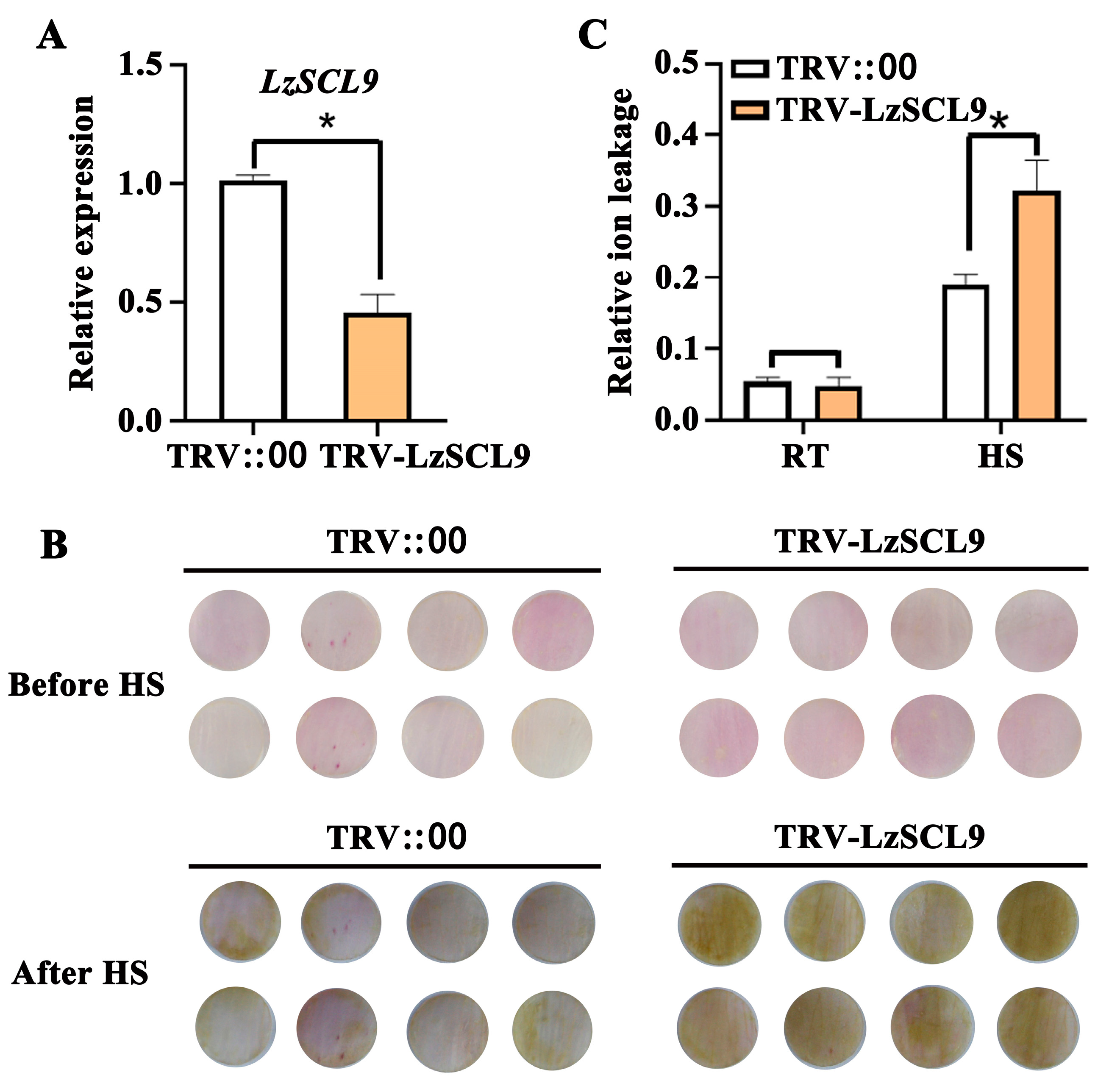
2.5. LzSCL9 Silencing Reduced the Thermotolerance of Lily
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. De Novo Transcriptome Sequencing and Analysis
4.3. Cloning and Sequence Analysis of LzSCL9
4.4. Gene Expression Analysis Using RT-qPCR
4.5. Transient Overexpression of LzSCL9 in Lily Petals
4.6. Silencing of LzSCL9 in Lily Petals Using VIGS
4.7. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Challinor, A.J.; Watson, J.; Lobell, D.B.; Howden, S.M.; Smith, D.R.; Chhetri, N. A meta-analysis of crop yield under climate change and adaptation. Nat. Clim. Chang. 2014, 4, 287–291. [Google Scholar] [CrossRef]
- Jung, J.H.; Barbosa, A.D.; Hutin, S.; Kumita, J.R.; Gao, M.; Derwort, D.; Silva, C.S.; Lai, X.; Pierre, E.; Geng, F.; et al. A prion-like domain in ELF3 functions as a thermosensor in Arabidopsis. Nature 2020, 585, 256–260. [Google Scholar] [CrossRef]
- Bohn, L.; Huang, J.; Weidig, S.; Yang, Z.; Heidersberger, C.; Genty, B.; Falter-Braun, P.; Christmann, A.; Grill, E. The temperature sensor TWA1 is required for thermotolerance in Arabidopsis. Nature 2024, 629, 1126–1132. [Google Scholar] [CrossRef]
- Li, Z.; Li, Z.R.; Ji, Y.L.; Wang, C.Y.; Wang, S.F.; Shi, Y.T.; Le, J.; Zhang, M. The heat shock factor 20-HSF4-cellulose synthase A2 module regulates heat stress tolerance in maize. Plant Cell. 2024, 36, 2652–2667. [Google Scholar] [CrossRef]
- Guo, M.; Liu, J.H.; Ma, X.; Luo, D.X.; Gong, Z.H.; Lu, M.H. The plant heat stress transcription factors (HSFs): Structure, regulation, and function in response to abiotic stresses. Front. Plant Sci. 2016, 7, 114. [Google Scholar] [CrossRef]
- Schramm, F.; Larkindale, J.; Kiehlmann, E.; Ganguli, A.; Englich, G.; Vierling, E.; Koskull-Döring, P.V. A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of arabidopsis. Plant J. 2008, 53, 264–274. [Google Scholar] [CrossRef] [PubMed]
- Li, S.J.; Fu, Q.T.; Chen, L.G.; Huang, W.D.; Yu, D.Q. Arabidopsis thaliana WRKY25, WRKY26, and WRKY33 coordinate induction of plant thermotolerance. Planta 2011, 233, 1237–1252. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Tian, X.J.; Wang, F.; Zhang, L.Y.; Xin, M.M.; Hu, Z.R.; Yao, Y.Y.; Ni, Z.F.; Sun, Q.X.; Peng, H.R. Characterization of wheat MYB genes responsive to high temperatures. BMC Plant Biol. 2017, 17, 208. [Google Scholar] [CrossRef]
- Cai, W.W.; Yang, S.; Wu, R.J.; Cao, J.S.; Shen, L.; Guan, D.Y.; He, S.L. Pepper NAC-type transcription factor NAC2c balances the trade-off between growth and defense responses. Plant Physiol. 2021, 186, 2169–2189. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Wang, Q.; Li, W.Q.; Hu, T.X.; Wang, Q.Q.; Yin, Y.; Liu, X.H.; He, S.; Zhang, M.K.; Liang, Y.; et al. Overexpression of SlBBX17 affects plant growth and enhances heat tolerance in tomato. Int. J. Biol. Macromol. 2022, 206, 799–811. [Google Scholar] [CrossRef]
- Li, Z.X.; Tang, J.; Srivastava, R.; Bassham, D.C.; Howell, S.H. The transcription factor bZIP60 links the unfolded protein response to the heat stress response in maize. Plant Cell. 2020, 32, 3559–3575. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Li, T.; Zhang, D.H.; Teng, N.J. Lily HD-Zip I Transcription Factor LlHB16 Promotes Thermotolerance by Activating LlHSFA2 and LlMBF1c. Plant Cell Physiol. 2022, 63, 1729–1744. [Google Scholar] [CrossRef] [PubMed]
- Koini, M.A.; Alvey, L.; Allen, T.; Tilley, C.A.; Harberd, N.P.; Whitelam, G.C.; Franklin, K.A. High temperature-mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Curr. Biol. 2009, 19, 408–413. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.H.; Jiang, F.L.; Wen, J.Q.; Shi, X.P.; Wu, C.Y.; Liu, M.; Wu, Z. Identification and heat resistance analysis of SlGRAS4 gene in tomato. Act. Bot. Boreal -Occident. Sin. 2021, 41, 539–548. (In Chinese) [Google Scholar]
- Cenci, A.; Rouard, M. Evolutionary analyses of GRAS transcription factors in angiosperms. Front. Plant Sci. 2017, 8, 273. [Google Scholar] [CrossRef]
- Pysh, L.D.; Wysocka-Diller, J.W.; Camilleri, C.; Bouchez, D.; Benfey, P.N. The GRAS gene family in Arabidopsis: Sequence characterization and basic expression analysis of the SCARECROW-LIKE genes. Plant J. 1999, 18, 111–119. [Google Scholar] [CrossRef]
- Day, R.B.; Tanabe, S.; Koshioka, M.; Mitsui, T.; Itoh, H.; Ueguchi-Tanaka, M.; Matsuoka, M.; Kaku, H.; Shibuya, N.; Minami, E. Two rice GRAS family genes responsive to N-acetylchitooligosaccharide elicitor are induced by phytoactive gibberellins: Evidence for cross-talk between elicitor and gibberellin signaling in rice cells. Plant Mol. Biol. 2004, 54, 261–272. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Zhang, N.; Zhu, X.; Yang, J.W.; Li, S.G.; Che, Y.Z.; Liu, W.G.; Si, H.J. Identification and expression analysis of StGRAS gene family in potato (Solanum tuberosum L.). Comput. Biol. Chem. 2019, 80, 195–205. [Google Scholar] [CrossRef] [PubMed]
- Neves, C.; Ribeiro, B.; Amaro, R.; Expósito, J.; Grimplet, J.; Fortes, A.M. Network of GRAS transcription factors in plant development, fruit ripening and stress responses. Hortic. Res. 2023, 10, uhad220. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.L.; Xue, B.; Jones, W.T.; Rikkerink, E.; Dunker, A.K.; Uversky, V.N. A functionally required unfoldome from the plant kingdom: Intrinsically disordered N-terminal domains of GRAS proteins are involved in molecular recognition during plant development. Plant Mol. Biol. 2011, 77, 205–223. [Google Scholar] [CrossRef]
- Khan, Y.; Xiong, Z.; Zhang, H.; Liu, S.; Yaseen, T.; Hui, T. Expression and roles of GRAS gene family in plant growth, signal transduction, biotic and abiotic stress resistance and symbiosis formation—A review. Plant Biol. 2022, 24, 404–416. [Google Scholar] [CrossRef] [PubMed]
- Kumari, P.; Gahlaut, V.; Kaur, E.; Singh, S.; Kumar, S.; Jaiswal, V. Genome-Wide Identification of GRAS Transcription Factors and Their Potential Roles in Growth and Development of Rose (Rosa chinensis). J. Plant Growth Regul. 2023, 42, 1505–1521. [Google Scholar] [CrossRef]
- Shen, J.J.; Jiang, Y.X.; Pan, J.; Sun, L.H.; Li, Q.Q.; He, W.J.; Sun, P.Y.; Zhao, B.S.; Zhao, H.J.; Ke, X.B.; et al. The GRAS transcription factor CsTL regulates tendril formation in cucumber. Plant Cell 2024, 36, 2818–2833. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.J.; Li, X.; Huang, J.; Gropp, G.M.; Gjetvaj, B.; Lindsay, D.L.; Wei, S.; Coutu, C.; Chen, Z.X.; Wan, X.C.; et al. SCARECROW-LIKE15 interacts with HISTONE DEACETYLASE19 and is essential for repressing the seed maturation programme. Nat. Commun. 2015, 6, 7243. [Google Scholar] [CrossRef]
- Stuurman, J.; Jaggi, F.; Kuhlemeier, C. Shoot meristem maintenance is controlled by a GRAS-gene mediated signal from differentiating cells. Genes Dev. 2002, 16, 2213–2218. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Zhu, Y.; Chang, H.; Wang, C.H.; Yang, J.; Shi, J.C.; Gao, J.P.; Yang, W.B.; Lan, L.Y.; Wang, Y.R.; et al. An SHR–SCR module specifies legume cortical cell fate to enable nodulation. Nature 2021, 589, 586–590. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Qin, L.; Lee, S.; Fu, X.; Richards, D.E.; Cao, D.; Luo, D.; Harberd, N.P.; Peng, J. Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 2004, 131, 1055–1064. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Li, X.W.; Fan, S.D.; Zhou, L.J.; Wang, Y. Overexpression of HcSCL13, a Halostachys caspica GRAS transcription factor, enhances plant growth and salt stress tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2020, 151, 243–254. [Google Scholar] [CrossRef]
- Wang, Y.T.; Feng, C.; Zhai, Z.F.; Peng, X.; Wang, Y.Y.; Sun, Y.T.; Li, J.; Shen, X.S.; Xiao, Y.Q.; Zhu, S.J.; et al. The apple microR171i-SCARECROW-LIKE PROTEINS26.1 module enhances drought stress tolerance by integrating ascorbic acid metabolism. J. Plant Physiol. 2020, 184, 194–211. [Google Scholar] [CrossRef]
- Wang, Z.; Wong, D.C.J.; Wang, Y.; Xu, G.; Ren, C.; Liu, Y.; Kuang, Y.; Fan, P.; Li, S.; Xin, H.; et al. GRAS-domain transcription factor PAT1 regulates jasmonic acid biosynthesis in grape cold stress response. Plant Physiol. 2021, 186, 1660–1678. [Google Scholar] [CrossRef]
- Jha, D.K.; Chanwala, J.; Sandeep, I.S.; Dey, N. Comprehensive identification and expression analysis of GRAS gene family under abiotic stress and phytohormone treatments in Pearl millet. Funct. Plant Biol. 2021, 48, 1039–1052. [Google Scholar] [CrossRef] [PubMed]
- Park, H.J.; Jung, W.Y.; Lee, S.S.; Song, J.H.; Kwon, S.Y.; Kim, H.; Ahn, J.C.; Cho, H.S. Use of heat stress responsive gene expression levels for early selection of heat tolerant Cabbage (Brassica oleracea L.). Int. J. Mol. Sci. 2013, 14, 11871–11894. [Google Scholar] [CrossRef] [PubMed]
- Li, L.Y.; Li, X.Z.; Chen, S.W.; Zhang, F.F.; Zhao, T.T.; Yang, G.Y. Expression and Function Analysis of Walnut JrGRAS2 Gene under Heat Stress. Bull. Bot. Res. 2018, 38, 125–131. (In Chinese) [Google Scholar]
- Li, W.M.; Wang, Y.J.; Ren, H.; Guo, Z.H.; Li, N.; Zhao, C.Z.; Xie, Z.K. Transcriptomic and physiological analyses identifying Lanzhou lily (Lilium davidii var. unicolor) drought adaptation strategies. Hortic. Plant J. 2023, 9, 145–157. [Google Scholar] [CrossRef]
- Feng, X.L.; Yue, L.; Zhao, X.H. A New Lilium Cultivar ‘Sunflower’. Acta Hortic. Sin. 2020, 47, 2069–2070. (In Chinese) [Google Scholar] [CrossRef]
- Gong, B.H.; Yi, J.; Wu, J.; Sui, J.J.; Khan, M.A.; Wu, Z.; Zhong, X.H.; Seng, S.S.; He, J.N.; Yi, M.F. LlHSFA1, a novel heat stress transcription factor in lily (Lilium longiflorum), can interact with LlHSFA2 and enhance the thermotolerance of transgenic Arabidopsis thaliana. Plant Cell Rep. 2014, 33, 1519–1533. [Google Scholar] [CrossRef]
- Wu, Z.; Li, T.; Cao, X.; Zhang, D.H.; Teng, N.J. Lily WRKY factor LlWRKY22 promotes thermotolerance through autoactivation and activation of LlDREB2B. Hortic. Res. 2022, 9, uhac186. [Google Scholar] [CrossRef]
- Wu, Z.; Liang, J.H.; Zhang, S.; Zhang, B.; Zhao, Q.C.; Li, G.Q.; Yang, X.; Wang, C.P.; He, J.N.; Yi, M.F. A canonical DREB2-type transcription factor in lily is posttranslationally regulated and mediates heat stress response. Front. Plant Sci. 2018, 9, 243. [Google Scholar] [CrossRef]
- Wu, Z.; Li, T.; Xiang, J.; Teng, R.D.; Zhang, D.H.; Teng, N.J. A lily membrane-associated NAC transcription factor LlNAC014 is involved in thermotolerance via activation of the DREB2-HSFA3 module. J. Exp. Bot. 2023, 74, 945–963. [Google Scholar] [CrossRef]
- Wu, Z.; Li, T.; Liu, X.Y.; Yuan, G.Z.; Hou, H.Z.; Teng, N.J. A novel R2R3-MYB transcription factor LlMYB305 from Lilium longiflorum plays a positive role in thermotolerance via activating heat-protective genes. Environ. Exp. Bot. 2021, 184, 104399. [Google Scholar] [CrossRef]
- Li, T.; Wu, Z.; Zhang, Y.Y.; Xu, S.J.; Xiang, J.; Ding, L.P.; Teng, N.J. An AP2/ERF member LlERF012 confers thermotolerance via activation of HSF pathway in lily. Plant Cell Environ. 2024; online ahead of print. [Google Scholar] [CrossRef]
- Cao, X.; Yi, J.; Wu, Z.; Luo, X.; Zhong, X.H.; Wu, J.; Ali, K.M.; Zhao, Y.; Yi, M.F. Involvement of Ca2+ and CaM3 in regulation of thermotolerance in lily (Lilium longiflorum). Plant Mol. Biol. Rep. 2013, 31, 1293–1304. [Google Scholar] [CrossRef]
- Chen, Q.M.; Yin, H.; Li, X.Y.; Yi, M.F. Effects of salicylic acid on the activities of antioxidant systems in lily plants under high temperature stress. J. China Agric. Univ. 2008, 13, 44–48. (In Chinese) [Google Scholar]
- Cao, X.; Sui, J.J.; Li, H.Y.; Yue, W.X.; Liu, T.; Hou, D.; Liang, J.H.; Wu, Z. Enhancing heat stress tolerance in Lanzhou lily (Lilium davidii var. unicolor) with Trichokonins isolated from Trichoderma longibrachiatum SMF2. Front. Plant Sci. 2023, 14, 1182977. [Google Scholar] [CrossRef] [PubMed]
- Morohashi, K.; Minami, M.; Takase, H.; Hotta, Y.; Hiratsuka, K. Isolation and characterization of a novel GRAS gene that regulates meiosis-associated gene expression. J. Biol. Chem. 2003, 278, 20865–20873. [Google Scholar] [CrossRef]
- Clarke, S.M.; Mur, L.A.J.; Wood, J.E.; Scott, I.M. Salicylic acid dependent signaling promotes basal thermotolerance but is not essential for acquired thermotolerance in Arabidopsis thaliana. Plant J. 2004, 38, 432–447. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Zhang, R.; Zong, X.F.; Wang, S.G.; He, G.H. Effect of salicylic acid on heat resistance of rice seedling under heat stress. Chin. J. Eco-Agric. 2009, 17, 1168–1171. [Google Scholar] [CrossRef]
- Pan, Q.H.; Zhan, J.C.; Liu, H.T.; Zhang, J.H.; Chen, J.Y.; Wen, P.F. Salicylic acid synthesized by benzoic acid 2-hydroxylase participates in the development of thermotolerance in pea plants. Plant Sci. 2006, 171, 226–233. [Google Scholar] [CrossRef]
- Wang, L.J.; Fan, L.; Loescher, W.; Duan, W.; Liu, G.J.; Cheng, J.S.; Luo, H.B.; Li, S.H. Salicylic acid alleviates decreases in photosynthesis under heat stress and accelerates recovery in grapevine leaves. BMC Plant Biol. 2010, 10, 34. [Google Scholar] [CrossRef]
- Zhang, Y.P.; Zhu, X.H.; Ding, H.D.; Yang, S.J.; Chen, Y.Y. Foliar application of 24-epibrassinolide alleviates high-temperature-induced inhibition of photosynthesis in seedlings of two melon cultivars. Photosynthetica 2013, 51, 341–349. [Google Scholar] [CrossRef]
- Mazorra, L.M.; Núnez, M.; Hechavarria, M.; Coll, F.; Sánchez-Blanco, M.J. Influence of brassinosteroids on antioxidant enzymes activity in tomato under different temperatures. Biol. Plantarum. 2002, 45, 593–596. [Google Scholar] [CrossRef]
- Li, Z.X.; Li, X.; Han, W.Y. Physiology mechanism of exogenous 24-epibrassinolide-induced heat resistance in tea plants (Camellia sinensis L.). Acta Agric. Zhejiangensis. 2016, 28, 959–965. [Google Scholar]
- Alam, M.N.; Zhang, L.H.; Yang, L.; Islam, M.R.; Liu, Y.; Luo, H.; Yang, P.F.; Wang, Q.F.; Chan, Z.L. Transcriptomic profiling of tall fescue in response to heat stress and improved thermotolerance by melatonin and 24-epibrassinolide. BMC Genom. 2018, 19, 224. [Google Scholar] [CrossRef]
- Zhang, J.; Shi, Y.; Zhang, X.Z.; Du, H.M.; Xu, B.; Huang, B.R. Melatonin suppression of heat-induced leaf senescence involves changes in abscisic acid and cytokinin biosynthesis and signaling pathways in perennial ryegrass (Lolium perenne L.). Environ. Exp. Bot. 2017, 138, 36–45. [Google Scholar] [CrossRef]
- Sehar, Z.; Fatma, M.; Khan, S.; Mir, I.R.; Abdi, G.; Khan, N.A. Melatonin influences methyl jasmonate-induced protection of photosynthetic activity in wheat plants against heat stress by regulating ethylene-synthesis genes and antioxidant metabolism. Sci. Rep. 2023, 13, 7468. [Google Scholar] [CrossRef]
- Teng, Z.Y.; Chen, C.Z.; He, Y.R.; Pan, S.H.; Liu, D.D.; Zhu, L.Y.; Liang, K.X.; Li, Y.F.; Li, H. Melatonin confers thermotolerance and antioxidant capacity in Chinese cabbage. Plant Physiol. Bioch. 2024, 212, 108736. [Google Scholar] [CrossRef]
- Shekhawat, K.; Almeida-Trapp, M.; García-Ramírez, G.X.; Hirt, H. Beat the heat: Plant- and microbe-mediated strategies for crop thermotolerance. Trends Plant Sci. 2022, 27, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Dastogeer, K.M.G.; Zahan, M.I.; Rhaman, M.S.; Sarker, M.S.A.; Chakraborty, A. Microbe-Mediated Thermotolerance in Plants and Pertinent Mechanisms—A Meta-Analysis and Review. Front. Microbiol. 2022, 13, 833566. [Google Scholar] [CrossRef]
- Reva, M.; Cano, C.; Herrera, M.; Bago, A. Arbuscular Mycorrhizal Inoculation Enhances Endurance to Severe Heat Stress in Three Horticultural Crops. HortScience 2021, 56, 396–406. [Google Scholar] [CrossRef]
- Tuang, Z.K.; Wu, Z.J.; Jin, Y.; Wang, Y.Z.; Oo, P.P.Z.; Zuo, G.X.; Shi, H.Z.; Yang, W.N. Pst DC3000 infection alleviates subsequent freezing and heat injury to host plants via a salicylic acid-dependent pathway in Arabidopsis. Plant Cell Environ. 2020, 43, 801–817. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Guo, Y.J.; Li, J.J.; Wang, L.; Wang, H.; Liu, G.D.; Fang, W.P.; Qiang, S.; Strasser, R.J.; Chen, S.G. Natural plant inducer 2-Amino-3-Methylhexanoic acid protects physiological activity against high-temperature damage to tea (Camellia sinensis). Sci. Hortic. 2023, 312, 111836. [Google Scholar] [CrossRef]
- Song, X.Y.; Shen, Q.T.; Xie, S.T.; Chen, X.L.; Sun, C.Y.; Zhang, Y.Z. Broad-spectrum antimicrobial activity and high stability of trichokonins from Trichoderma koningii SMF2 against plant pathogens. FEMS Microbiol. Lett. 2006, 260, 119–125. [Google Scholar] [CrossRef]
- Luo, Y.; Zhang, D.D.; Dong, X.W.; Zhao, P.B.; Chen, L.L.; Song, X.Y.; Wang, X.J.; Chen, X.L.; Shi, M.; Zhang, Y.Z. Antimicrobial peptaibols induce defense responses and systemic resistance in tobacco against tobacco mosaic virus. FEMS Microbiol. Lett. 2010, 313, 120–126. [Google Scholar] [CrossRef]
- Liu, J.Z.; Feng, L.L.; Gu, X.T.; Deng, X.; Qiu, Q.; Li, Q.; Zhang, Y.Y.; Wang, M.Y.; Deng, Y.W.; Wang, E.; et al. An H3K27me3 demethylase-HSFA2 regulatory loop orchestrates transgenerational thermomemory in Arabidopsis. Cell Res. 2019, 29, 379–390. [Google Scholar] [CrossRef]
- Friedrich, T.; Oberkofler, V.; Trindade, I.; Altmann, S.; Brzezinka, K.; Lämke, J.; Gorka, M.; Kappel, C.; Sokolowska, E.; Skirycz, A.; et al. Heteromeric HSFA2/HSFA3 complexes drive transcriptional memory after heat stress in Arabidopsis. Nat. Commun. 2021, 12, 3426. [Google Scholar] [CrossRef] [PubMed]
- Xue, G.P.; Sadat, S.; Drenth, J.; McIntyre, C.L. The heat shock factor family from Triticum aestivum in response to heat and other major abiotic stresses and their role in regulation of heat shock protein genes. J. Exp. Bot. 2014, 65, 539–557. [Google Scholar] [CrossRef]
- Liu, X.N.; Chen, H.Y.; Li, S.C.; Lecourieux, D.; Duan, W.; Fan, P.G.; Liang, Z.H.; Wang, L.J. Natural variations of HSFA2 enhance thermotolerance in grapevine. Hortic. Res. 2023, 10, uhac250. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Hwang, J.E.; Chan, J.L.; Kim, D.Y.; Sang, Y.L.; Lim, C.O. Arabidopsis DREB2C functions as a transcriptional activator of HsfA3 during the heat stress response. Biochem. Bioph. Res. Commun. 2010, 401, 238–244. [Google Scholar] [CrossRef]
- Huang, J.Y.; Zhao, X.B.; Bürger, M.; Wang, Y.R.; Chory, J. Two interacting ethylene response factors regulate heat stress response. Plant Cell 2021, 33, 338–357. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.P.; Wu, Z.; Teng, R.D.; Xu, S.J.; Cao, X.; Yuan, G.Z.; Zhang, D.H.; Teng, N.J. LlWRKY39 is involved in thermotolerance by activating LlMBF1c and interacting with LlCaM3 in lily (Lilium longiflorum). Hortic. Res. 2021, 8, 36. [Google Scholar] [CrossRef] [PubMed]
- Albertos, P.; Dündar, G.; Schenk, P.; Carrera, S.; Cavelius, P.; Sieberer, T.; Poppenberger, B. Transcription factor BES1 interacts with HSFA1 to promote heat stress resistance of plants. EMBO J. 2022, 41, e108664. [Google Scholar] [CrossRef]
- Zhu, X.L.; Wang, B.Q.; Wei, X.H. Genome wide identification and expression pattern analysis of the GRAS family in quinoa. Funct. Plant Biol. 2021, 48, 948–962. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.Y.; Gao, X.Q.; Ma, K.H.; Li, D.P.; Jia, C.X.; Zhai, M.Z.; Xu, Z.G. The walnut transcription factor JrGRAS2 contributes to high temperature stress tolerance involving in Dof transcriptional regulation and HSP protein expression. BMC Plant Biol. 2018, 18, 367. [Google Scholar] [CrossRef]
- Liu, X.Y.; Widmer, A. Genome-wide comparative analysis of the GRAS gene family in Populus, Arabidopsis and Rice. Plant Mol. Biol. Rep. 2014, 32, 1129–1145. [Google Scholar] [CrossRef]
- Weng, Y.H.; Chen, X.Y.; Hao, Z.D.; Lu, L.; Wu, X.R.; Zhang, J.Q.; Wu, J.X.; Shi, J.S.; Chen, J.H. Genome-wide analysis of the GRAS gene family in Liriodendron chinense reveals the putative function in abiotic stress and plant development. Front. Plant Sci. 2023, 14, 1211853. [Google Scholar] [CrossRef]
- Li, C.; Wang, K.; Chen, S.; Zhang, X.L.; Zhang, X.Y.; Fan, L.X.; Dong, J.H.; Xu, L.; Wang, Y.; Li, Y.; et al. Genome-wide identification of RsGRAS gene family reveals positive role of RsSHRc gene in chilling stress response in radish (Raphanus sativus L.). Plant Physiol. Bioch. 2022, 192, 285–297. [Google Scholar] [CrossRef]
- Zeng, X.; Ling, H.; Chen, X.M.; Guo, S. Genome-wide identification, phylogeny and function analysis of GRAS gene family in Dendrobium catenatum (Orchidaceae). Gene 2019, 705, 5–15. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Levesque, M.P.; Vernoux, T.; Jung, J.W.; Paquette, A.J.; Gallagher, L.; Wang, J.Y.; Blilou, I.; Scheres, B.; Benfey, P.N. An evolutionarily conserved mechanism delimiting SHR movement defines a single layer of endodermis in plants. Science 2007, 316, 421–425. [Google Scholar] [CrossRef] [PubMed]
- Curaba, J.; Talbot, M.; Li, Z.Y.; Helliwell, C. Over-expression of microRNA171 affects phase transitions and floral meristem determinancy in barley. BMC Plant Biol. 2013, 13, 6. [Google Scholar] [CrossRef]
- Bakery, A.; Vraggalas, S.; Shalha, B.; Chauchan, H.; Benhamed, M.; Fragkostefanakis, S. Heat stress transcription factors as the central molecular rheostat to optimize plant survival and recovery from heat stress. New Phytol. 2024; early view. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Hall, T. BioEdit: An important software for molecular biology. Gerf Bull. Biosci. 2011, 2, 60–61. [Google Scholar]




| Sequences(5′-3′) | |
|---|---|
| Primers for gene cloning | |
| LzSCL9-F | ATGGTGAACGAGCTAAAAGTTGACGATTT |
| LzSCL9-R | TCATCGACTATTGGGTTTCCACGCAGAGA |
| Primers for RT-qPCR | |
| qPCR-LzSCL9-F | CTCCATCTGACCAGCCATC |
| qPCR-LzSCL9-R | TAGCGCCAAAGGCCCAATTA |
| 18S rRNA-F | AGTTGGTGGAGCGATTTGTCT |
| 18S rRNA-R | CCTGTTATTGCCTCAAACTTCC |
| Primers for transient overexpression | |
| SK-LzSCL9-XbaI-F | ctccaccgcggtggcggccgctctagaATGGTGAACGAGCTAAAAGTT |
| SK-LzSCL9-BamHI-R | atatcgaattcctgcagcccgggggatccTCATCGACTATTGGGTTTCCACGC |
| Primers for VIGS | |
| TRV2-LzSCL9-XbaI-F | ctgtgagtaaggttaccgaattctctagaTGCGAGGGAACAGA |
| TRV2-LzSCL9-BamHI-R | gcctcgagacgcgtgagctcggtaccTCATCGACTATTGGGTTTCCACGC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, X.; Ding, L.; Liang, J.; Zhou, Y.; Chen, X.; Li, H.; Liu, T.; Yue, W.; Sui, J.; Jiang, L.; et al. LzSCL9, a Novel GRAS Transcription Factor in Lanzhou Lily (Lilium davidii var. unicolor), Participates in Regulation of Trichokonins-Primed Heat Stress Tolerance. Plants 2024, 13, 2330. https://doi.org/10.3390/plants13162330
Cao X, Ding L, Liang J, Zhou Y, Chen X, Li H, Liu T, Yue W, Sui J, Jiang L, et al. LzSCL9, a Novel GRAS Transcription Factor in Lanzhou Lily (Lilium davidii var. unicolor), Participates in Regulation of Trichokonins-Primed Heat Stress Tolerance. Plants. 2024; 13(16):2330. https://doi.org/10.3390/plants13162330
Chicago/Turabian StyleCao, Xing, Liping Ding, Jiahui Liang, Yanrong Zhou, Xiulan Chen, Haiyan Li, Tao Liu, Wenxiu Yue, Juanjuan Sui, Liangbao Jiang, and et al. 2024. "LzSCL9, a Novel GRAS Transcription Factor in Lanzhou Lily (Lilium davidii var. unicolor), Participates in Regulation of Trichokonins-Primed Heat Stress Tolerance" Plants 13, no. 16: 2330. https://doi.org/10.3390/plants13162330
APA StyleCao, X., Ding, L., Liang, J., Zhou, Y., Chen, X., Li, H., Liu, T., Yue, W., Sui, J., Jiang, L., Qian, Y., Yang, D., Wang, B., Zhang, H., Wu, Z., & Song, X. (2024). LzSCL9, a Novel GRAS Transcription Factor in Lanzhou Lily (Lilium davidii var. unicolor), Participates in Regulation of Trichokonins-Primed Heat Stress Tolerance. Plants, 13(16), 2330. https://doi.org/10.3390/plants13162330







