Genome-Wide Identification of GH17s Family Genes and Biological Function Analysis of SlA6 in Tomato
Abstract
:1. Introduction
2. Results
2.1. Identification of GH17 Family Members and Analysis of Their Physicochemical Properties
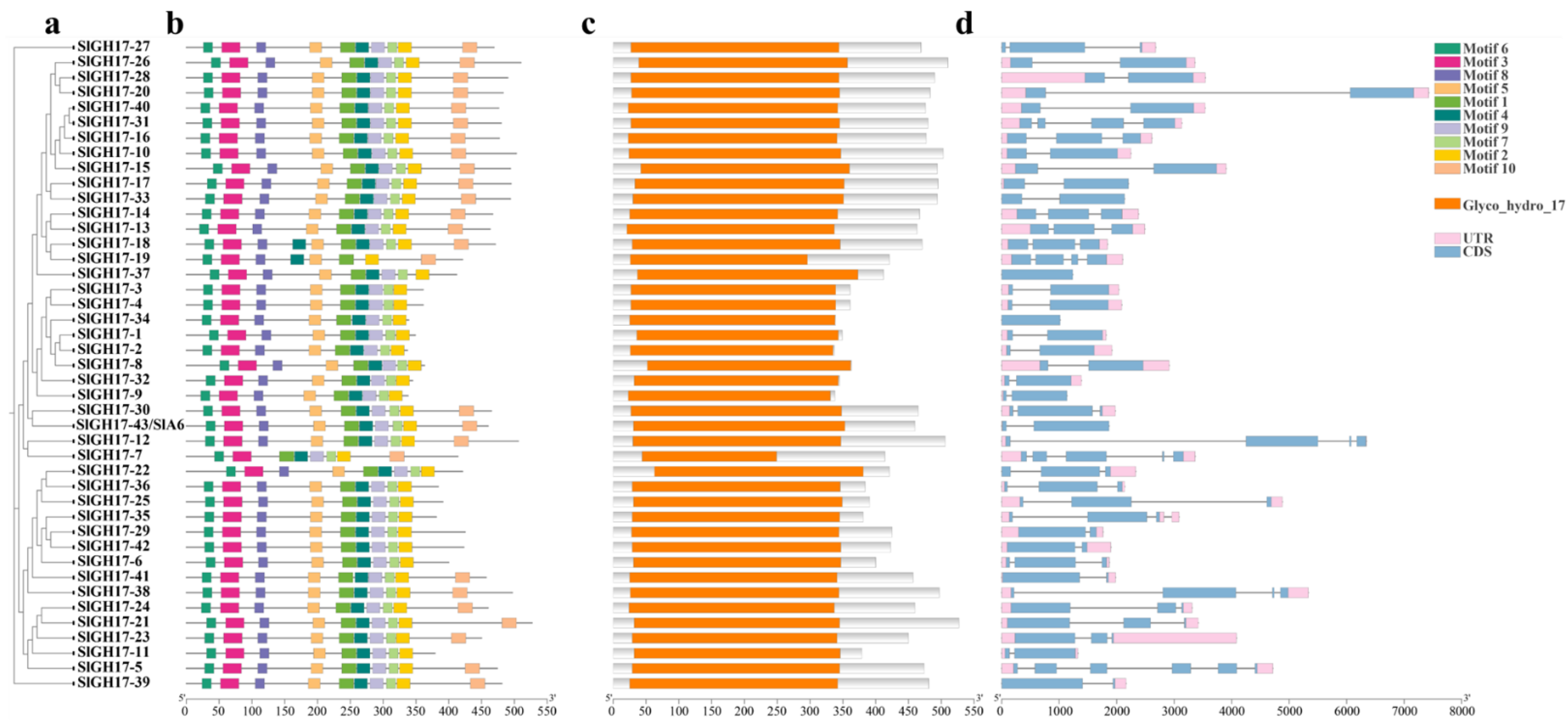
2.2. Protein Motifs, Conserved Domains, and Gene Structure Analysis of GH17 Family Members
2.3. Evolutionary and Covariance Analyses of GH17 Family Members
2.4. Cis-Acting Elements Analysis of the Promoter Region of GH17 Family Members
2.5. Expression Pattern Analysis of the SlGH17s Family Members in Tomato Tissues
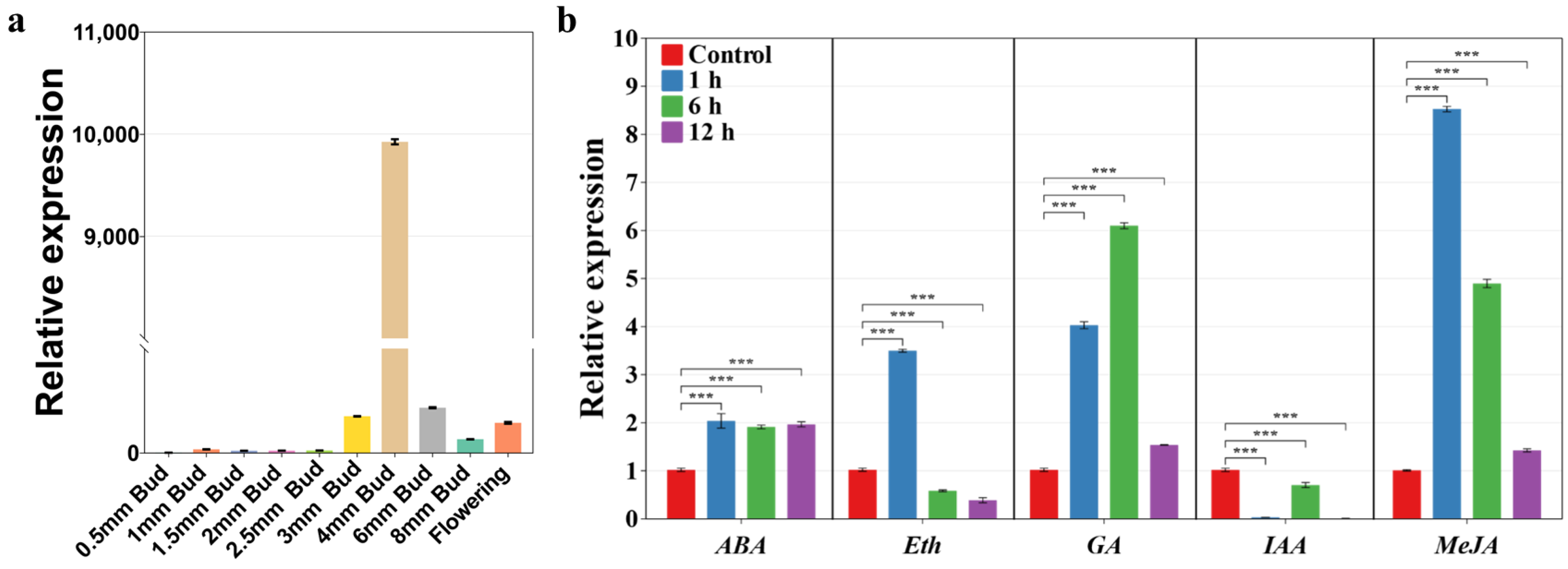
2.6. Expression Pattern Analysis of SlA6 during Tomato Flower Development and Response to Phytohormones
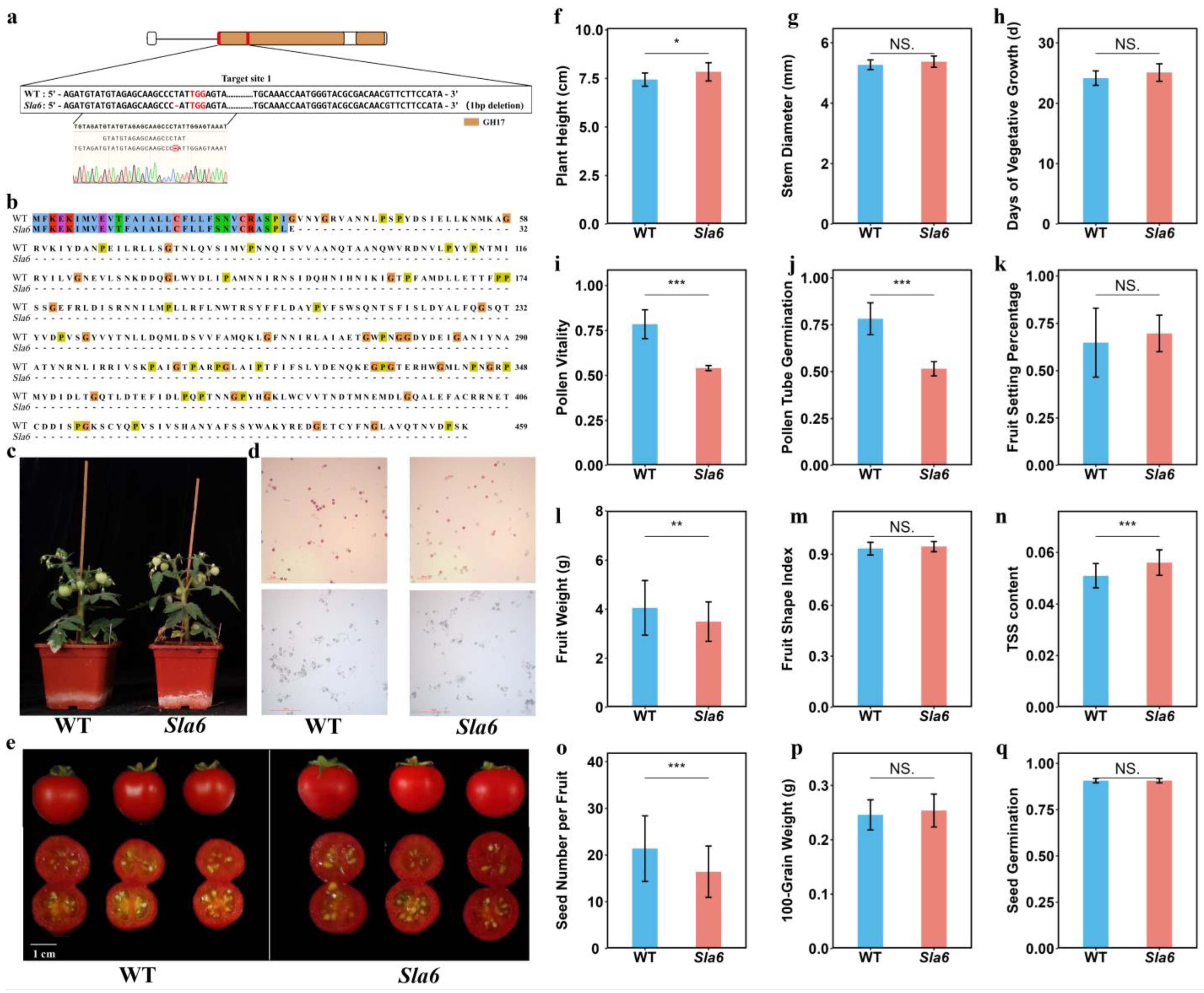
2.7. Knocking Out of SlA6 Affects Pollen and Fruit Development in Tomato
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Identification of GH17 Family Members
4.3. Structural and Evolutionary Analyses of GH17 Family Members
4.4. Expression Pattern Analysis of GH17 Family Members
4.5. Expression Pattern Analysis of SlA6
4.6. Quantitative Real-Time PCR Analysis
4.7. Generating SlA6 Mutants Using CRISPR/Cas9 Technology
4.8. Identification of Transgenic Plant Phenotypes
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Simmons, C.R. The Physiology and Molecular Biology of Plant 1,3-β-D-Glucanases and 1,3;1,4-β-D-The Physiology and Molecular Biology of Plant 1,3-β-D-Glucanases and 1,3;1,4-β-D-Glucanases. Crit. Rev. Plant Sci. 1994, 13, 325–387. [Google Scholar] [CrossRef]
- Shrivastava, S. Industrial Applications of Glycoside Hydrolases; Springer: Singapore, 2020. [Google Scholar]
- Zhang, Q.; Zhang, X.; Pettolino, F.; Zhou, G.; Li, C. Changes in cell wall polysaccharide composition, gene transcription and alternative splicing in germinating barley embryos. J. Plant Physiol. 2016, 191, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Kuge, T.; Nagoya, H.; Tryfona, T.; Kurokawa, T.; Yoshimi, Y.; Dohmae, N.; Tsubaki, K.; Dupree, P.; Tsumuraya, Y.; Kotake, T. Action of an endo-beta-1,3(4)-glucanase on cellobiosyl unit structure in barley beta-1,3:1,4-glucan. Biosci. Biotechnol. Biochem. 2015, 79, 1810–1817. [Google Scholar] [CrossRef] [PubMed]
- Benitez-Alfonso, Y.; Faulkner, C.; Pendle, A.; Miyashima, S.; Helariutta, Y.; Maule, A. Symplastic intercellular connectivity regulates lateral root patterning. Dev. Cell 2013, 26, 136–147. [Google Scholar] [CrossRef]
- Levy, A.; Guenoune-Gelbart, D.; Epel, B.L. beta-1,3-Glucanases: Plasmodesmal gate keepers for intercellular communication. Plant Signal. Behav. 2007, 2, 404–407. [Google Scholar] [CrossRef]
- Rinne, P.L.; Paul, L.K.; Vahala, J.; Kangasjarvi, J.; van der Schoot, C. Axillary buds are dwarfed shoots that tightly regulate GA pathway and GA-inducible 1,3-beta-glucanase genes during branching in hybrid aspen. J. Exp. Bot. 2016, 67, 5975–5991. [Google Scholar] [CrossRef]
- Li, Y.; Wang, W.; Feng, Y.; Tu, M.; Wittich, P.E.; Bate, N.J.; Messing, J. Transcriptome and metabolome reveal distinct carbon allocation patterns during internode sugar accumulation in different sorghum genotypes. Plant Biotechnol. J. 2019, 17, 472–487. [Google Scholar] [CrossRef]
- Pervaiz, T.; Liu, T.; Fang, X.; Ren, Y.; Li, X.; Liu, Z.; Fiaz, M.; Fang, J.; Shangguan, L. Identification of GH17 gene family in Vitis vinifera and expression analysis of GH17 under various adversities. Physiol. Mol. Biol. Plants 2021, 27, 1423–1436. [Google Scholar] [CrossRef]
- Shah, T.; Qin, Z.; Pu, L.; Li, X.; Liu, D.; Cui, X. A β-1,3-glucanase gene from Panax notoginseng confers resistance in tobacco to Fusarium solani. Ind. Crops Prod. 2020, 143, 111947. [Google Scholar] [CrossRef]
- Jung, J.Y.; Min, C.W.; Jang, J.W.; Gupta, R.; Kim, J.H.; Kim, Y.H.; Cho, S.W.; Song, Y.H.; Jo, I.H.; Rakwal, R.; et al. Proteomic analysis reveals a critical role of the glycosyl hydrolase 17 protein in Panax ginseng leaves under salt stress. Int. J. Mol. Sci. 2023, 24, 3693. [Google Scholar] [CrossRef]
- Akiyama, T.; Pillai, M.A. Molecular cloning, characterization and in vitro expression of a novel endo-1,3-b-glucanase up-regulated by ABA and drought stress in rice (Oryza sativa L.). Plant Sci. 2001, 161, 1089–1098. [Google Scholar] [CrossRef]
- Pu, Y.; Hou, L.; Guo, Y.; Ullah, I.; Yong, Y.; Yue, Y. Genome-wide analysis of the callose enzyme families of fertile and sterile flower buds of the Chinese cabbage (Brassica rapa L. ssp. Pekinensis). FEBS Open Bio 2019, 9, 1432–1449. [Google Scholar] [CrossRef] [PubMed]
- Tsuchiya, T.; Toriyama, K.; Yoshikawa, M.; Ejiri, S.; Hinata, K. Tapetum-specific expression of the gene for an endo-beta-1,3-glucanase causes male sterility in transgenic tobacco. Plant Cell Physiol. 1995, 36, 487–494. [Google Scholar] [CrossRef] [PubMed]
- Stieglitz, H.; Stern, H. Regulation of beta-1,3-glucanase activity in developing anthers of Lilium. Dev. Biol. 1973, 34, 169–173. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Han, X.; Lu, T.G. Callose synthesis during reproductive development in monocotyledonous and dicotyledonous plants. Plant Signal. Behav. 2016, 11, e1062196. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.Z.; Zhang, G.S.; Zhu, W.W.; Ba, Q.S.; Niu, N.; Wang, J.W.; Ma, S.C.; Wang, J.S. Relationship between male sterility and beta-1,3-glucanase activity and callose deposition-related gene expression in wheat (Triticum aestivum L.). Genet. Mol. Res. 2015, 14, 574–584. [Google Scholar] [CrossRef]
- Nishikawa, S.I.; Zinkl, G.M.; Swanson, R.J.; Maruyama, D.; Preuss, D. Callose (β-1,3 glucan) is essential for Arabidopsis pollen wall patterning, but not tube growth. BMC Plant Biol. 2005, 5, 22. [Google Scholar] [CrossRef] [PubMed]
- Hird, D.L.; Worrall, D.; Hodge, R.; Smartt, S.; Paul, W.; Scott, R. The anther-specific protein encoded by the Brassica napus and Arabidopsis thaliana A6 gene displays similarity to beta-1,3-glucanases. Plant J. 1993, 4, 1023–1033. [Google Scholar] [CrossRef]
- Zhang, Z.B.; Zhu, J.; Gao, J.F.; Wang, C.; Li, H.; Li, H.; Zhang, H.Q.; Zhang, S.; Wang, D.M.; Wang, Q.X.; et al. Transcription factor AtMYB103 is required for anther development by regulating tapetum development, callose dissolution and exine formation in Arabidopsis. Plant J. 2007, 52, 528–538. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Yang, Y.; Su, D.; Yu, C.; Xian, Z.; Pan, Z.; Guan, H.; Hu, G.; Chen, D.; Li, Z.; et al. The SlHB8 acts as a negative regulator in tapetum development and pollen wall formation in Tomato. Hortic. Res. 2022, 9, c185. [Google Scholar] [CrossRef]
- Lu, P.; Chai, M.; Yang, J.; Ning, G.; Wang, G.; Ma, H. The Arabidopsis CALLOSE DEFECTIVE MICROSPORE1 gene is required for male fertility through regulating callose metabolism during microsporogenesis. Plant Physiol. 2014, 164, 1893–1904. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.; Xu, L.; Liu, Y.; Dong, H.; Zhou, D.; Zhang, Y.; Lin, S.; Cao, J.; Huang, L. Comparative transcriptome analysis and ChIP-sequencing reveals stage-specific gene expression and regulation profiles associated with pollen wall formation in Brassica rapa. BMC Genom. 2019, 20, 264. [Google Scholar] [CrossRef]
- Wan, L.; Zha, W.; Cheng, X.; Liu, C.; Lv, L.; Liu, C.; Wang, Z.; Du, B.; Chen, R.; Zhu, L.; et al. A rice beta-1,3-glucanase gene Osg1 is required for callose degradation in pollen development. Planta 2011, 233, 309–323. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Khedkar, S.; Bork, P. SMART: Recent updates, new developments and status in 2020. Nucleic Acids Res. 2021, 49, D458–D460. [Google Scholar] [CrossRef]
- Chow, C.; Yang, C.; Wu, N.; Wang, H.; Tseng, K.; Chiu, Y.; Lee, T.; Chang, W. PlantPAN 4.0: Updated database for identifying conserved non-coding sequences and exploring dynamic transcriptional regulation in plant promoters. Nucleic Acids Res. 2024, 52, D1569–D1578. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Li, Y.; Chen, Y.; Zhou, L.; You, S.; Deng, H.; Chen, Y.; Alseekh, S.; Yuan, Y.; Fu, R.; Zhang, Z.; et al. MicroTom Metabolic Network: Rewiring Tomato Metabolic Regulatory Network throughout the Growth Cycle. Mol. Plant 2020, 13, 1203–1218. [Google Scholar] [CrossRef]
- Bokszczanin, K.L.; Krezdorn, N.; Fragkostefanakis, S.; Muller, S.; Rycak, L.; Chen, Y.; Hoffmeier, K.; Kreutz, J.; Paupiere, M.J.; Chaturvedi, P.; et al. Identification of novel small ncRNAs in pollen of tomato. BMC Genom. 2015, 16, 714. [Google Scholar] [CrossRef]
- Song, S.; Huang, B.; Pan, Z.; Zhong, Q.; Yang, Y.; Chen, D.; Zhu, L.; Hu, G.; He, M.; Wu, C.; et al. The SlTPL3-SlWUS module regulates multi-locule formation in tomato by modulating auxin and gibberellin levels in the shoot apical meristem. J. Integr. Plant Biol. 2022, 64, 2150–2167. [Google Scholar] [CrossRef]




| Name | ID | Protein Length/bp | Molecular Weight/Da | Theoretical pI | Instability Coefficient | Aliphatic Amino Acid Index | Hydrophilicity Coefficient |
|---|---|---|---|---|---|---|---|
| SlGH17-1 | Solyc01g008610 | 348 | 39.23 | 5.78 | 34.29 | 93.3 | −0.108 |
| SlGH17-2 | Solyc01g008620 | 336 | 37.45 | 6.09 | 38.35 | 87.41 | −0.289 |
| SlGH17-3 | Solyc01g059965 | 360 | 39.72 | 7.84 | 46.03 | 89.08 | −0.156 |
| SlGH17-4 | Solyc01g060020 | 360 | 39.72 | 7.84 | 46.03 | 89.08 | −0.156 |
| SlGH17-5 | Solyc01g109570 | 473 | 50.30 | 4.95 | 38.16 | 81.71 | −0.079 |
| SlGH17-6 | Solyc02g070450 | 399 | 44.69 | 6.12 | 41.81 | 91.45 | 0.017 |
| SlGH17-7 | Solyc02g080660 | 413 | 45.27 | 5.46 | 41.71 | 83.56 | −0.105 |
| SlGH17-8 | Solyc02g086700 | 362 | 40.82 | 5.9 | 32.19 | 90.19 | −0.087 |
| SlGH17-9 | Solyc03g025650 | 337 | 36.56 | 5 | 29.84 | 103.83 | 0.159 |
| SlGH17-10 | Solyc03g058450 | 502 | 54.68 | 5.68 | 35.14 | 78.25 | −0.217 |
| SlGH17-11 | Solyc03g082900 | 378 | 41.68 | 5.15 | 31.76 | 107.28 | 0.226 |
| SlGH17-12 | Solyc04g007910 | 505 | 54.64 | 5.33 | 40.92 | 90.2 | 0.055 |
| SlGH17-13 | Solyc04g011720 | 462 | 52.12 | 7.06 | 30.75 | 85.82 | −0.187 |
| SlGH17-14 | Solyc04g011730 | 466 | 52.98 | 8.73 | 32.12 | 83.03 | −0.248 |
| SlGH17-15 | Solyc04g015190 | 493 | 53.91 | 5.9 | 40.06 | 94.89 | 0.126 |
| SlGH17-16 | Solyc04g051590 | 476 | 52.85 | 6.66 | 31.22 | 83.57 | −0.097 |
| SlGH17-17 | Solyc05g006210 | 494 | 55.02 | 5.69 | 39.82 | 89.03 | −0.016 |
| SlGH17-18 | Solyc05g015160 | 470 | 53.50 | 6.72 | 29.86 | 82.13 | −0.233 |
| SlGH17-19 | Solyc05g015170 | 420 | 47.84 | 6.46 | 26.5 | 81.64 | −0.199 |
| SlGH17-20 | Solyc05g025500 | 482 | 52.71 | 6.32 | 31.91 | 83.36 | −0.1 |
| SlGH17-21 | Solyc05g054440 | 526 | 57.62 | 9.13 | 39.78 | 76.16 | −0.227 |
| SlGH17-22 | Solyc06g073710 | 420 | 46.91 | 8.67 | 38.68 | 97.79 | −0.027 |
| SlGH17-23 | Solyc06g076170 | 449 | 49.37 | 4.93 | 29.93 | 82.09 | −0.041 |
| SlGH17-24 | Solyc07g005330 | 459 | 50.92 | 9.05 | 37.72 | 77.52 | −0.25 |
| SlGH17-25 | Solyc07g008150 | 390 | 42.83 | 8.73 | 26.21 | 93.1 | −0.051 |
| SlGH17-26 | Solyc07g017730 | 509 | 56.33 | 7.19 | 33.18 | 88.82 | −0.092 |
| SlGH17-27 | Solyc07g049370 | 468 | 51.32 | 6.72 | 33.84 | 81.71 | −0.162 |
| SlGH17-28 | Solyc08g074390 | 489 | 52.95 | 5.76 | 32.4 | 85.36 | −0.088 |
| SlGH17-29 | Solyc08g083310 | 424 | 45.63 | 8.05 | 30.06 | 91.32 | 0.03 |
| SlGH17-30 | Solyc09g057630 | 464 | 51.04 | 9.28 | 31.17 | 85.93 | −0.136 |
| SlGH17-31 | Solyc10g078510 | 479 | 53.38 | 5.43 | 30.27 | 84.49 | −0.073 |
| SlGH17-32 | Solyc10g079860 | 344 | 37.80 | 9.42 | 46.67 | 93.02 | −0.193 |
| SlGH17-33 | Solyc11g012030 | 493 | 55.20 | 4.87 | 38.07 | 90.77 | −0.057 |
| SlGH17-34 | Solyc11g065280 | 338 | 37.39 | 4.84 | 45.4 | 88.85 | −0.212 |
| SlGH17-35 | Solyc11g068440 | 380 | 42.56 | 6.06 | 27.47 | 90.11 | −0.056 |
| SlGH17-36 | Solyc11g071520 | 383 | 42.56 | 9.15 | 38.18 | 93.94 | 0.025 |
| SlGH17-37 | Solyc11g072230 | 411 | 45.19 | 5.38 | 37.27 | 98.69 | 0.209 |
| SlGH17-38 | Solyc12g008580 | 496 | 53.95 | 5.91 | 34.82 | 79.31 | −0.073 |
| SlGH17-39 | Solyc12g014420 | 480 | 51.46 | 5.59 | 39.02 | 82.9 | −0.149 |
| SlGH17-40 | Solyc12g019890 | 475 | 52.35 | 4.89 | 32.02 | 81.96 | −0.052 |
| SlGH17-41 | Solyc12g040860 | 456 | 50.39 | 6 | 29.34 | 84.87 | −0.134 |
| SlGH17-42 | Solyc12g055840 | 422 | 45.54 | 6.52 | 32.48 | 88.48 | 0.006 |
| SlGH17-43/SlA6 | Solyc12g098560 | 459 | 51.89 | 5.09 | 39.87 | 85.62 | −0.22 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, D.; Zeng, Z.; Yu, C.; Hu, H.; Lin, Y.; Wu, C.; Yang, Y.; Zhong, Q.; Zhang, X.; Huang, C.; et al. Genome-Wide Identification of GH17s Family Genes and Biological Function Analysis of SlA6 in Tomato. Plants 2024, 13, 2443. https://doi.org/10.3390/plants13172443
Chen D, Zeng Z, Yu C, Hu H, Lin Y, Wu C, Yang Y, Zhong Q, Zhang X, Huang C, et al. Genome-Wide Identification of GH17s Family Genes and Biological Function Analysis of SlA6 in Tomato. Plants. 2024; 13(17):2443. https://doi.org/10.3390/plants13172443
Chicago/Turabian StyleChen, Da, Zaohai Zeng, Canye Yu, Huimin Hu, Yuxiang Lin, Caiyu Wu, Yinghua Yang, Qiuxiang Zhong, Xinyue Zhang, Caihong Huang, and et al. 2024. "Genome-Wide Identification of GH17s Family Genes and Biological Function Analysis of SlA6 in Tomato" Plants 13, no. 17: 2443. https://doi.org/10.3390/plants13172443






