New Aspects of HECT-E3 Ligases in Cell Senescence and Cell Death of Plants
Abstract
:1. Introduction
2. The HECT E3s Family in Plants and Human Being
3. The Patterns of HECT E3s’ Substrate Recruitment and Catalytic Activity Regulation
4. The Roles of HECT E3s in Cell Senescence/Aging
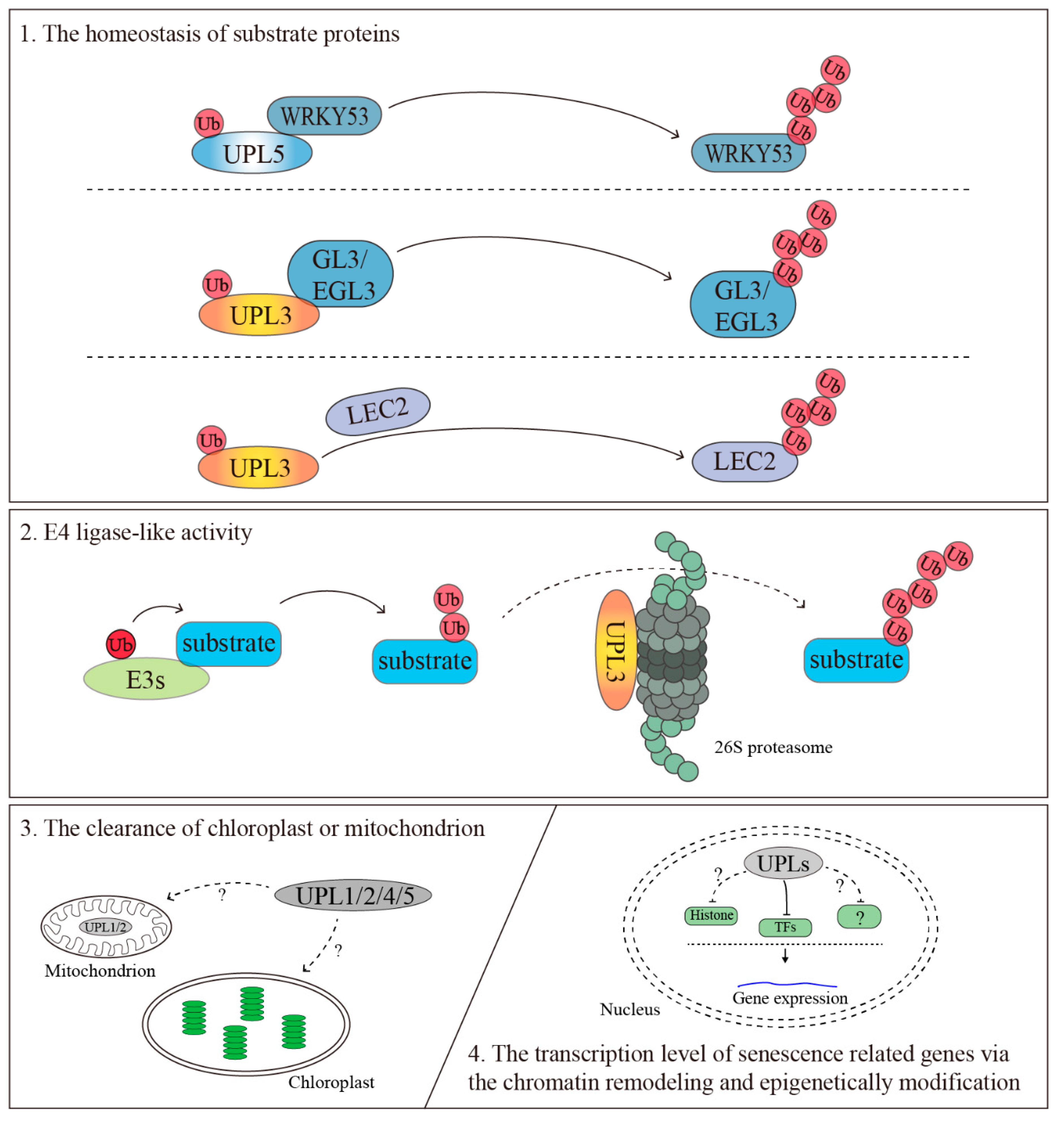
4.1. The Homeostasis of Substrate Proteins
4.2. The Clearance of the Chloroplast or Mitochondrion
4.3. The Transcriptional Regulation of Senescence-Related Genes via the Chromatin Remodeling and Epigenetic Modification
4.4. E4 Ligase-Like Activity
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Multani, D.S.; Meeley, R.B.; Paterson, A.H.; Gray, J.; Briggs, S.P.; Johal, G.S. Plant-pathogen microevolution: Molecular basis for the origin of a fungal disease in maize. Proc. Natl. Acad. Sci. USA 1998, 95, 1686–1691. [Google Scholar] [CrossRef] [PubMed]
- Page, T.; Harrison, E.; Breeze, E.; Lim, P.O.; Gil Nam, H.; Lin, J.F.; Wu, S.H.; Swidzinski, J.; Ishizaki, K.; Leaver, C.J.; et al. Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. Plant J. 2005, 42, 567–585. [Google Scholar]
- Buchanan-Wollaston, V.; Ainsworth, C. Leaf senescence in Brassica napus: Cloning of senescence related genes by subtractive hybridisation. Plant Mol. Boil. 1997, 33, 821–834. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Ma, Z.; Wang, H.; Niu, K.; Cao, Y.; Sun, L.; Geng, Y.; Yang, B.; Gao, F.; Chen, Z.; et al. Ubiquitylome study identifies increased histone 2A ubiquitylation as an evolutionarily conserved aging biomarker. Nat. Commun. 2019, 10, 2191. [Google Scholar] [CrossRef] [PubMed]
- Rotin, D.; Kumar, S. Physiological functions of the HECT family of ubiquitin ligases. Nat. Rev. Mol. Cell Boil. 2009, 10, 398–409. [Google Scholar] [CrossRef] [PubMed]
- Scheffner, M.; Kumar, S. Mammalian HECT ubiquitin-protein ligases: Biological and pathophysiological aspects. Biochim. Biophys. Acta 2014, 1843, 61–74. [Google Scholar] [CrossRef] [PubMed]
- Huibregtse, J.M.; Scheffner, M.; Beaudenon, S.; Howley, P.M. A family of proteins structurally and functionally related to the E6-AP ubiquitin-protein ligase. Proc. Natl. Acad. Sci. USA 1995, 92, 2563–2567. [Google Scholar] [CrossRef]
- Downes, B.P.; Stupar, R.M.; Gingerich, D.J.; Vierstra, R.D. The HECT ubiquitin-protein ligase (UPL) family in Arabidopsis: UPL3 has a specific role in trichome development. Plant J. 2003, 35, 729–742. [Google Scholar] [CrossRef]
- Marín, I. Evolution of Plant HECT Ubiquitin Ligases. PLoS ONE 2013, 8, e68536. [Google Scholar] [CrossRef]
- Marín, I. Animal HECT ubiquitin ligases: Evolution and functional implications. BMC Evol. Boil. 2010, 10, 56. [Google Scholar] [CrossRef]
- Scheffner, M.; Nuber, U.; Huibregtse, J.M. Protein ubiquitination involving an E1–E2–E3 enzyme ubiquitin thioester cascade. Nature 1995, 373, 81–83. [Google Scholar] [CrossRef] [PubMed]
- Metzger, M.B.; Pruneda, J.N.; Klevit, R.E.; Weissman, A.M. RING-type E3 ligases: Master manipulators of E2 ubiquitin-conjugating enzymes and ubiquitination. Biochim. Biophys. Acta 2014, 1843, 47–60. [Google Scholar] [CrossRef] [PubMed]
- Huang, L. Structure of an E6AP-UbcH7 Complex: Insights into Ubiquitination by the E2-E3 Enzyme Cascade. Science 1999, 286, 1321–1326. [Google Scholar] [CrossRef] [PubMed]
- Berndsen, C.E.; Wolberger, C. New insights into ubiquitin E3 ligase mechanism. Nat. Struct. Mol. Boil. 2014, 21, 301–307. [Google Scholar] [CrossRef] [PubMed]
- Sluimer, J.; Distel, B. Regulating the human HECT E3 ligases. Cell. Mol. Life Sci. 2018, 75, 3121–3141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huibregtse, J.M.; Scheffner, M.; Howley, P.M. Localization of the E6-AP regions that direct human papillomavirus E6 binding, association with p53, and ubiquitination of associated proteins. Mol. Cell. Boil. 1993, 13, 4918–4927. [Google Scholar] [CrossRef] [PubMed]
- Zanier, K.; Charbonnier, S.; Sidi, A.O.M.O.; McEwen, A.G.; Ferrario, M.G.; Poussin-Courmontagne, P.; Cura, V.; Brimer, N.; Babah, K.O.; Ansari, T.; et al. Structural basis for hijacking of cellular LxxLL motifs by papillomavirus E6 oncoproteins. Science 2013, 339, 694–698. [Google Scholar] [CrossRef]
- Kühnle, S.; Kogel, U.; Glockzin, S.; Marquardt, A.; Ciechanover, A.; Matentzoglu, K.; Scheffner, M. Physical and Functional Interaction of the HECT Ubiquitin-protein Ligases E6AP and HERC2. J. Boil. Chem. 2011, 286, 19410–19416. [Google Scholar] [CrossRef] [Green Version]
- Izzi, L.; Attisano, L. Ubiquitin-dependent regulation of TGFbeta signaling in cancer. Neoplasia 2006, 8, 677–688. [Google Scholar] [CrossRef]
- Kavsak, P.; Rasmussen, R.K.; Causing, C.G.; Bonni, S.; Zhu, H.; Thomsen, G.H.; Wrana, J.L. Smad7 binds to Smurf2 to form an E3 ubiquitin ligase that targets the TGF beta receptor for degradation. Mol. Cell 2000, 6, 1365–1375. [Google Scholar] [CrossRef]
- Ogunjimi, A.A.; Briant, D.J.; Pece-Barbara, N.; Le Roy, C.; Di Guglielmo, G.M.; Kavsak, P.; Rasmussen, R.K.; Seet, B.T.; Sicheri, F.; Wrana, J.L. Regulation of Smurf2 Ubiquitin Ligase Activity by Anchoring the E2 to the HECT Domain. Mol. Cell 2005, 19, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Malakhova, O.A.; Zhang, D.-E. ISG15 inhibits Nedd4 ubiquitin E3 activity and enhances the innate antiviral response. J. Boil. Chem. 2008, 283, 8783–8787. [Google Scholar] [CrossRef] [PubMed]
- Okumura, A.; Pitha, P.M.; Harty, R.N. ISG15 inhibits Ebola VP40 VLP budding in an L-domain-dependent manner by blocking Nedd4 ligase activity. Proc. Natl. Acad. Sci. USA 2008, 105, 3974–3979. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Timmins, J.; Schoehn, G.; Ricard-Blum, S.; Scianimanico, S.; Vernet, T.; Ruigrok, R.W.H.; Weissenhorn, W. Ebola virus matrix protein VP40 interaction with human cellular factors Tsg101 and Nedd4. J. Mol. Boil. 2003, 326, 493–502. [Google Scholar] [CrossRef]
- Yasuda, J.; Nakao, M.; Kawaoka, Y.; Shida, H. Nedd4 Regulates Egress of Ebola Virus-Like Particles from Host Cells. J. Virol. 2003, 77, 9987–9992. [Google Scholar] [CrossRef] [Green Version]
- Han, Z.; Lu, J.; Liu, Y.; Davis, B.; Lee, M.S.; Olson, M.A.; Ruthel, G.; Freedman, B.D.; Schnell, M.J.; Wrobel, J.E.; et al. Small-Molecule Probes Targeting the Viral PPxY-Host Nedd4 Interface Block Egress of a Broad Range of RNA Viruses. J. Virol. 2014, 88, 7294–7306. [Google Scholar] [CrossRef] [Green Version]
- Gallagher, E.; Gao, M.; Liu, Y.-C.; Karin, M. Activation of the E3 ubiquitin ligase Itch through a phosphorylation-induced conformational change. Proc. Natl. Acad. Sci. USA 2006, 103, 1717–1722. [Google Scholar] [CrossRef] [Green Version]
- Zhu, K.; Shan, Z.; Chen, X.; Cai, Y.; Cui, L.; Yao, W.; Wang, Z.; Shi, P.; Tian, C.; Lou, J.; et al. Allosteric auto-inhibition and activation of the Nedd4 family E3 ligase Itch. EMBO Rep. 2017, 18, 1618–1630. [Google Scholar] [CrossRef]
- Wan, L.; Zou, W.; Gao, D.; Inuzuka, H.; Fukushima, H.; Berg, A.H.; Drapp, R.; Shaik, S.; Hu, D.; Lester, C.; et al. Cdh1 regulates osteoblast function through an APC/C-independent modulation of Smurf1. Mol. Cell 2011, 44, 721–733. [Google Scholar] [CrossRef]
- Sander, B.; Xu, W.; Eilers, M.; Popov, N.; Lorenz, S. A conformational switch regulates the ubiquitin ligase HUWE1. eLife 2017, 6, 36. [Google Scholar] [CrossRef]
- Vierstra, R.D. Proteolysis in plants: Mechanisms and functions. Plant Mol. Boil. 1996, 32, 275–302. [Google Scholar] [CrossRef] [PubMed]
- Vierstra, R.D. Protein degradation in plants. Ann. Rev. Plant Physiol. Plant Biol. 1993, 44, 385–410. [Google Scholar] [CrossRef]
- Bernassola, F.; Karin, M.; Ciechanover, A.; Melino, G. The HECT Family of E3 Ubiquitin Ligases: Multiple Players in Cancer Development. Cancer Cell 2008, 14, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Schmid, M.; Davison, T.S.; Henz, S.R.; Pape, U.J.; Demar, M.; Vingron, M.; Schölkopf, B.; Weigel, D.; Lohmann, J.U. A gene expression map of Arabidopsis thaliana development. Nat. Genet. 2005, 37, 501–506. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.; Zentgraf, U. A HECT E3 ubiquitin ligase negatively regulates Arabidopsis leaf senescence through degradation of the transcription factor WRKY53. Plant J. 2010, 63, 179–188. [Google Scholar] [CrossRef]
- Patra, B.; Pattanaik, S.; Yuan, L. Ubiquitin protein ligase 3 mediates the proteasomal degradation of GLABROUS 3 and ENHANCER OF GLABROUS 3, regulators of trichome development and flavonoid biosynthesis in Arabidopsis. Plant J. 2013, 74, 435–447. [Google Scholar] [CrossRef]
- Bensussan, M.; Lefebvre, V.; Ducamp, A.; Trouverie, J.; Gineau, E.; Fortabat, M.-N.; Guillebaux, A.; Baldy, A.; Naquin, D.; Herbette, S.; et al. Suppression of Dwarf and irregular xylem Phenotypes Generates Low-Acetylated Biomass Lines in Arabidopsis. Plant Physiol. 2015, 168, 452–463. [Google Scholar] [CrossRef] [Green Version]
- Furniss, J.J.; Grey, H.; Wang, Z.; Nomoto, M.; Jackson, L.; Tada, Y.; Spoel, S.H. Proteasome-associated HECT-type ubiquitin ligase activity is required for plant immunity. PLoS Pathog. 2018, 14, e1007447. [Google Scholar] [CrossRef]
- Miller, C.; Wells, R.; McKenzie, N.; Trick, M.; Ball, J.; Fatihi, A.; Dubreucq, B.; Chardot, T.; Lepiniec, L.; Bevan, M.W. Variation in expression of the HECT E3 ligase UPL3 modulates LEC2 levels, seed size and crop yield in Brassica napus. Plant Cell 2019, 31, 2370–2385. [Google Scholar] [CrossRef]
- Ermolaeva, M.; Neri, F.; Ori, A.; Rudolph, K.L. Cellular and epigenetic drivers of stem cell ageing. Nat. Rev. Mol. Cell Boil. 2018, 19, 594–610. [Google Scholar] [CrossRef]
- Chrobok, D.; Law, S.R.; Brouwer, B.; Lindén, P.; Ziolkowska, A.; Liebsch, D.; Narsai, R.; Szal, B.; Moritz, T.; Rouhier, N.; et al. Dissecting the Metabolic Role of Mitochondria during Developmental Leaf Senescence1. Plant Physiol. 2016, 172, 2132–2153. [Google Scholar] [CrossRef] [PubMed]
- Keech, O. The conserved mobility of mitochondria during leaf senescence reflects differential regulation of the cytoskeletal components in Arabidopsis thaliana. Plant Signal. Behav. 2011, 6, 147–150. [Google Scholar] [CrossRef] [PubMed]
- Ruberti, C.; Barizza, E.; Bodner, M.; La Rocca, N.; De Michele, R.; Carimi, F.; Schiavo, F.L.; Zottini, M. Mitochondria Change Dynamics and Morphology during Grapevine Leaf Senescence. PLoS ONE 2014, 9, e102012. [Google Scholar] [CrossRef] [PubMed]
- Novikoff, A.B.; Beaufay, H.; De Duve, C. ELECTRON MICROSCOPY OF LYSOSOME-RICH FRACTIONS FROM RAT LIVER. J. Cell Boil. 1956, 2, 179–184. [Google Scholar] [CrossRef] [Green Version]
- De Duve, C.; Pressman, B.C.; Gianetto, R.; Wattiaux, R.; Appelmans, F. Tissue fractionation studies. 6. Intracellular distribution patterns of enzymes in rat-liver tissue. Biochem. J. 1955, 60, 604–617. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Denton, D.; Kumar, S. Autophagy-dependent cell death. Cell Death Differ. 2019, 26, 605–616. [Google Scholar] [CrossRef]
- Shpilka, T.; Haynes, C.M. The mitochondrial UPR: Mechanisms, physiological functions and implications in ageing. Nat. Rev. Mol. Cell Biol. 2018, 19, 109–120. [Google Scholar] [CrossRef]
- Leboucher, G.P.; Tsai, Y.C.; Yang, M.; Shaw, K.C.; Zhou, M.; Veenstra, T.D.; Glickman, M.H.; Weissman, A.M. Stress-induced phosphorylation and proteasomal degradation of mitofusin 2 facilitates mitochondrial fragmentation and apoptosis. Mol. Cell 2012, 47, 547–557. [Google Scholar] [CrossRef]
- Tanaka, A.; Cleland, M.M.; Xu, S.; Narendra, D.P.; Suen, D.-F.; Karbowski, M.; Youle, R.J. Proteasome and p97 mediate mitophagy and degradation of mitofusins induced by Parkin. J. Cell Boil. 2010, 191, 1367–1380. [Google Scholar] [CrossRef] [Green Version]
- Di Rita, A.; Peschiaroli, A.; D′Acunzo, P.; Strobbe, D.; Hu, Z.; Gruber, J.; Nygaard, M.; Lambrughi, M.; Melino, G.; Papaleo, E.; et al. HUWE1 E3 ligase promotes PINK1/PARKIN-independent mitophagy by regulating AMBRA1 activation via IKKalpha. Nat. Commun. 2018, 9, 3755. [Google Scholar] [CrossRef]
- Strappazzon, F.; Di Rita, A.; Peschiaroli, A.; Leoncini, P.P.; Locatelli, F.; Melino, G.; Cecconi, F. HUWE1 controls MCL1 stability to unleash AMBRA1-induced mitophagy. Cell Death Differ. 2019, 1–14. [Google Scholar]
- Liu, T.; Zhang, L.; Chen, G.; Shi, T. Identifying and Characterizing the Circular RNAs during the Lifespan of Arabidopsis Leaves. Front. Plant Sci. 2017, 8, 1278. [Google Scholar] [CrossRef] [PubMed]
- Brusslan, J.A.; Bonora, G.; Rus-Canterbury, A.M.; Tariq, F.; Jaroszewicz, A.; Pellegrini, M. A Genome-Wide Chronological Study of Gene Expression and Two Histone Modifications, H3K4me3 and H3K9ac, during Developmental Leaf Senescence. Plant Physiol. 2015, 168, 1246–1261. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qin, J.; Ma, X.; Yi, Z.; Tang, Z.; Meng, Y. A transcriptome-wide study on the microRNA- and the Argonaute 1-enriched small RNA-mediated regulatory networks involved in plant leaf senescence. Plant Biol. 2016, 18, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Woo, H.R.; Kim, H.J.; Lim, P.O.; Gil Nam, H. Leaf Senescence: Systems and Dynamics Aspects. Annu. Rev. Plant Boil. 2019, 70, 347–376. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, X.; Lu, L.; Mayer, K.S.; Scalf, M.; Qian, S.; Lomax, A.; Smith, L.M.; Zhong, X. POWERDRESS interacts with HISTONE DEACETYLASE 9 to promote aging in Arabidopsis. eLife 2016, 5, 6832. [Google Scholar] [CrossRef]
- Schmid, J.A.; Berti, M.; Walser, F.; Raso, M.C.; Schmid, F.; Krietsch, J.; Stoy, H.; Zwicky, K.; Ursich, S.; Freire, R.; et al. Histone Ubiquitination by the DNA Damage Response Is Required for Efficient DNA Replication in Unperturbed S Phase. Mol. Cell 2018, 71, 897–910. [Google Scholar] [CrossRef]
- Cole, A.J.; Clifton-Bligh, R.; Marsh, D.J. Histone H2B monoubiquitination: Roles to play in human malignancy. Endocr Relat. Cancer 2015, 22, T19–T33. [Google Scholar] [CrossRef]
- Bourbousse, C.; Ahmed, I.; Roudier, F.; Zabulon, G.; Blondet, E.; Balzergue, S.; Colot, V.; Bowler, C.; Barneche, F. Histone H2B Monoubiquitination Facilitates the Rapid Modulation of Gene Expression during Arabidopsis Photomorphogenesis. PLoS Genet. 2012, 8, 1002825. [Google Scholar] [CrossRef]
- Cao, J.; Yan, Q. Histone Ubiquitination and Deubiquitination in Transcription, DNA Damage Response, and Cancer. Front. Oncol. 2012, 2, 26. [Google Scholar] [CrossRef] [Green Version]
- Liu, Z.; Oughtred, R.; Wing, S.S. Characterization of E3Histone, a Novel Testis Ubiquitin Protein Ligase Which Ubiquitinates Histones. Mol. Cell. Boil. 2005, 25, 2819–2831. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Yang, H.; Wang, H. The Histone H2A Deubiquitinase USP16 Interacts with HERC2 and Fine-tunes Cellular Response to DNA Damage. J. Boil. Chem. 2014, 289, 32883–32894. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Danielsen, J.R.; Povlsen, L.K.; Villumsen, B.H.; Streicher, W.; Nilsson, J.; Wikström, M.; Bekker-Jensen, S.; Mailand, N. DNA damage–inducible SUMOylation of HERC2 promotes RNF8 binding via a novel SUMO-binding Zinc finger. J. Cell Boil. 2012, 197, 179–187. [Google Scholar] [CrossRef] [PubMed]
- Bekker-Jensen, S.; Rendtlew Danielsen, J.; Fugger, K.; Gromova, I.; Nerstedt, A.; Lukas, C.; Bartek, J.; Lukas, J.; Mailand, N. HERC2 coordinates ubiquitin-dependent assembly of DNA repair factors on damaged chromosomes. Nat. Cell Biol. 2010, 12, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Kan, S.; Huang, B.; Hao, Z.; Mak, T.W.; Zhong, Q. Mule determines the apoptotic response to HDAC inhibitors by targeted ubiquitination and destruction of HDAC2. Genes Dev. 2011, 25, 2610–2618. [Google Scholar] [CrossRef] [Green Version]
- Besche, H.C.; Sha, Z.; Kukushkin, N.V.; Peth, A.; Hock, E.-M.; Kim, W.; Gygi, S.; Gutierrez, J.A.; Liao, H.; Dick, L.; et al. Autoubiquitination of the 26S Proteasome on Rpn13 Regulates Breakdown of Ubiquitin Conjugates. EMBO J. 2014, 33, 1159–1176. [Google Scholar] [CrossRef]
- Chu, B.W.; Kovary, K.M.; Guillaume, J.; Chen, L.-C.; Teruel, M.N.; Wandless, T.J. The E3 Ubiquitin Ligase UBE3C Enhances Proteasome Processivity by Ubiquitinating Partially Proteolyzed Substrates. J. Boil. Chem. 2013, 288, 34575–34587. [Google Scholar] [CrossRef] [Green Version]
- Crosas, B.; Hanna, J.; Kirkpatrick, D.S.; Zhang, D.P.; Tone, Y.; Hathaway, N.A.; Buecker, C.; Leggett, D.S.; Schmidt, M.; King, R.W.; et al. Ubiquitin Chains Are Remodeled at the Proteasome by Opposing Ubiquitin Ligase and Deubiquitinating Activities. Cell 2006, 127, 1401–1413. [Google Scholar] [CrossRef] [Green Version]
- Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 2009, 78, 477–513. [Google Scholar] [CrossRef]



| Species | Subfamily | Members | Main Domain |
|---|---|---|---|
| In humans (28 members) | HERC family (6 members) | HERC1, HERC2, HERC3, HERC4, HERC5, HERC6 | RLDs, HECT domain |
| Nedd4 family (9 members) | Nedd4/Nedd4-1, NEDD4L/Nedd4-2, Smurf1, Smurf2, Itch/AIP4, WWP1/AIP5, WWP2, NEDL1/HECW1, NEDL2/HECW2 | C2, WW, HECT domain | |
| other HECTs family (13 members) | HACE1, HECTD1, HUWE1 a, UBE3A/E6-AP, UBE3B, UBE3C, UBR5/EDD1, G2E3, TRIP12, KIAA0317, HECTD3, HECTX/KIAA0614, HECTD2, | ANK, Arm-like, UBA, WWE, IQ, ZnF, PABC, PHD, RING, Filamin, DOC, HECT domain, etc. | |
| In Arabidopsis thaliana (7 members) | Subfamily I | UPL3, UPL4 | Arm, HECT domain |
| Subfamily II | UPL7 | IQ, HECT domain | |
| Subfamily III | UPL6 | IQ, HECT domain | |
| Subfamily V | UPL1, UPL2, UPL8 (lost in Arabidopsis thaliana) | Arm, UBA, UIM, HECT domain | |
| Subfamily VI | UPL5 | UBL, C-type lectin, LZ, HECT domain |
| Mutants | Targets | The Phenotype of Mutants | Reference |
|---|---|---|---|
| upl3 | a GL3/EGL3 (UPL3-N) | Trichrome development | [36] |
| upl3 | Unknown | Larger stem diameters | [37] |
| upl3 | LEC2 | Larger seed size | [39] |
| upl5 | WRKY53 | Premature | [35] |
| upl1, upl3, upl5 | Unknown | Plant immunity | [38] |
| upl3/upl4 | Unknown | Seed germination defect | Unpublished |
| upl3, upl3/upl4 | Unknown | Light response | Unpublished |
| upl2, upl3, upl4, upl6 | Unknown | Plant senescence | Unpublished |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lan, W.; Miao, Y. New Aspects of HECT-E3 Ligases in Cell Senescence and Cell Death of Plants. Plants 2019, 8, 483. https://doi.org/10.3390/plants8110483
Lan W, Miao Y. New Aspects of HECT-E3 Ligases in Cell Senescence and Cell Death of Plants. Plants. 2019; 8(11):483. https://doi.org/10.3390/plants8110483
Chicago/Turabian StyleLan, Wei, and Ying Miao. 2019. "New Aspects of HECT-E3 Ligases in Cell Senescence and Cell Death of Plants" Plants 8, no. 11: 483. https://doi.org/10.3390/plants8110483





