Plant Sulfate Transporters in the Low Phytic Acid Network: Some Educated Guesses
Abstract
1. Background
2. Sulfate Transporters: A Short Overview
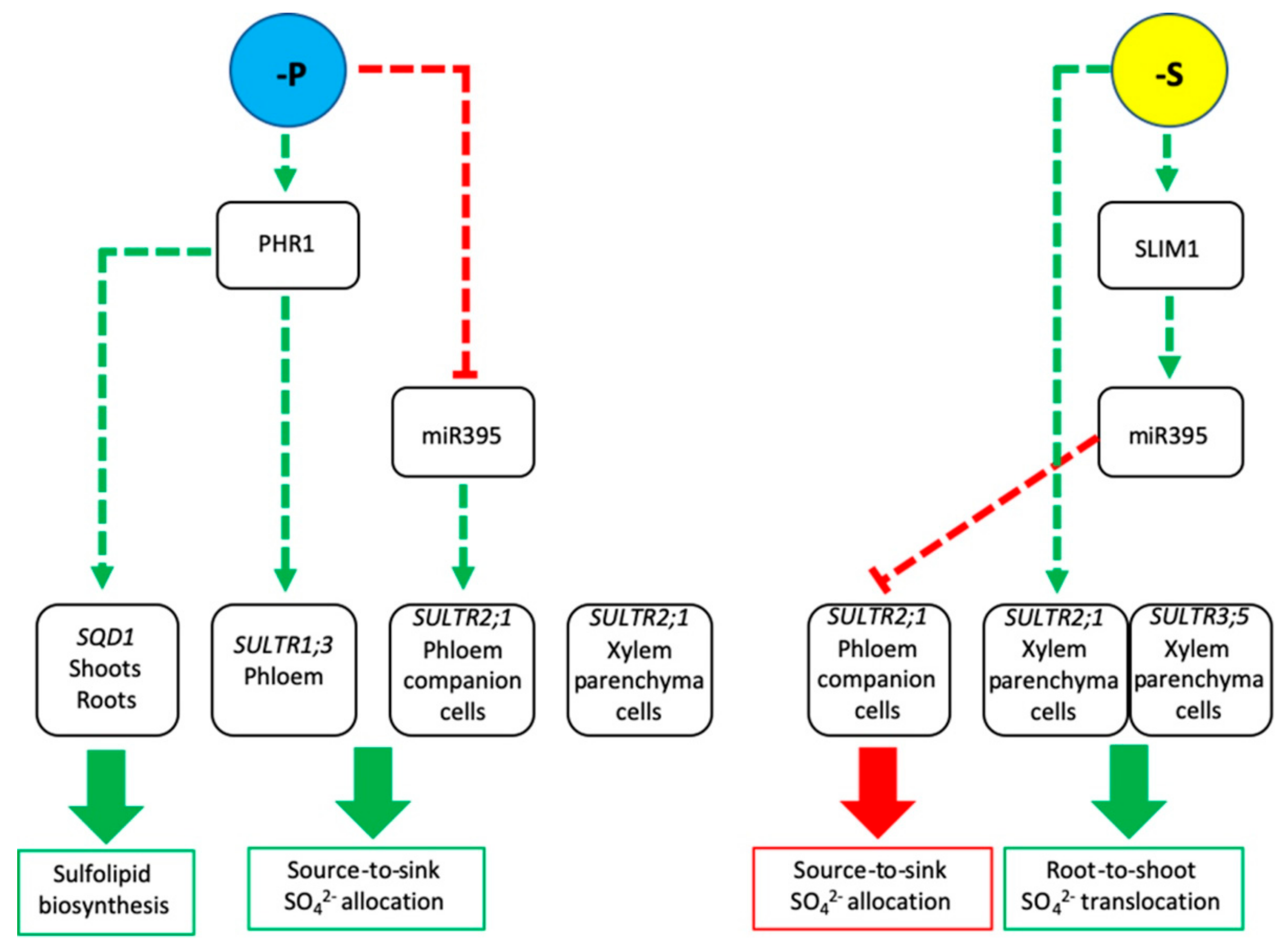
3. Sulfur and Phosphorous Interplay
4. SULTRs as Novel Elements in the lpa Network
5. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bohn, L.; Meyer, A.S.; Rasmussen, S.K. Phytate: Impact on environment and human nutrition. A challenge for molecular breeding. J. Zhejiang Univ. Sci. B 2008, 9, 165–191. [Google Scholar] [CrossRef] [PubMed]
- Raboy, V. Seeds for a better future: “Low phytate”, grains help to overcome malnutrition and reduce pollution. Trends Plant Sci. 2001, 6, 458–462. [Google Scholar] [CrossRef]
- Schlemmer, U.; Frølich, W.; Prieto, R.M.; Grases, F. Phytate in foods and significance for humans: Food sources, intake, processing, bioavailability, protective role and analysis. Mol. Nutr. Food Res. 2009, 53, S330–S375. [Google Scholar] [CrossRef]
- Raboy, V. Approaches and challenges to engineering seed phytate and total phosphorus. Plant Sci. 2009, 177, 281–296. [Google Scholar] [CrossRef]
- Sparvoli, F.; Cominelli, E. Seed biofortification and phytic acid reduction: A conflict of interest for the plant? Plants 2015, 4, 728–755. [Google Scholar] [CrossRef]
- Leustek, T.; Martin, M.N.; Bick, J.A.; Davies, J.P. Pathways and regulation of sulfur metabolism revealed through molecular and genetic studies. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2000, 51, 141–165. [Google Scholar] [CrossRef]
- Saito, K. Sulfur assimilatory metabolism. The long and smelling road. Plant Physiol. 2004, 136, 2443–2450. [Google Scholar] [CrossRef]
- Nocito, F.F.; Lancilli, C.; Giacomini, B.; Sacchi, G.A. Sulfur metabolism and cadmium stress in higher plants. Plant Stress 2007, 1, 142–156. [Google Scholar]
- Takahashi, H.; Kopriva, S.; Giordano, M.; Saito, K.; Hell, R. Sulfur assimilation in photosynthetic organisms: Molecular functions and regulations of transporters and assimilatory enzyme. Annu. Rev. Plant Biol. 2011, 62, 157–184. [Google Scholar] [CrossRef]
- Mugford, S.G.; Lee, B.R.; Koprivova, A.; Matthewman, C.; Kopriva, S. Control of sulfur partitioning between primary and secondary metabolism. Plant J. 2011, 65, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Koprivova, A.; Kopriva, S. Sulfation pathways in plants. Chem. Biol. Interact. 2016, 259, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Shibagaki, N.; Grossman, A.R. The role of the STAS domain in the function and biogenesis of a sulfate transporter as probed by random mutagenesis. J. Biol. Chem. 2006, 281, 22964–22973. [Google Scholar] [CrossRef] [PubMed]
- Shibagaki, N.; Grossman, A.R. Binding of cysteine synthase to the STAS domain of sulfate transporter and its regulatory consequences. J. Biol. Chem. 2010, 285, 25094–25102. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H. Sulfate transport systems in plants: Functional diversity and molecular mechanisms underlying regulatory coordination. J. Exp. Bot. 2019, 70, 4075–4087. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Asif, M.H.; Chakrabarty, D.; Tripathi, R.D.; Trivedi, P.K. Differential expression and alternative splicing of rice sulphate transporter family members regulate sulphur status during plant growth, development and stress conditions. Funct. Integr. Genom. 2011, 11, 259–273. [Google Scholar] [CrossRef]
- Takahashi, H.; Watanabe-Takahashi, A.; Smith, F.W.; Blake-Kalff, M.; Hawkesford, M.J.; Saito, K. The roles of three functional sulphate transporters involved in uptake and translocation of sulphate in Arabidopsis thaliana. Plant J. 2000, 23, 171–182. [Google Scholar] [CrossRef]
- Yoshimoto, N.; Takahashi, H.; Smith, F.W.; Yamaya, T.; Saito, K. Two distinct high-affinity sulfate transporters with different inducibilities mediate uptake of sulfate in Arabidopsis roots. Plant J. 2002, 29, 465–473. [Google Scholar] [CrossRef]
- Yoshimoto, N.; Inoue, E.; Watanabe-Takahashi, A.; Saito, K.; Takahashi, H. Posttranscriptional regulation of high-affinity sulfate transporters in Arabidopsis by sulfur nutrition. Plant Physiol. 2007, 145, 378–388. [Google Scholar] [CrossRef]
- Barberon, M.; Berthomieu, P.; Clairotte, M.; Shibagaki, N.; Davidian, J.C.; Gosti, F. Unequal functional redundancy between the two Arabidopsis thaliana high-affinity sulphate transporters SULTR1;1 and SULTR1;2. New Phytol. 2008, 180, 608–619. [Google Scholar] [CrossRef]
- El Kassis, E.; Cathala, N.; Rouached, H.; Fourcroy, P.; Berthomieu, P.; Terry, N.; Davidian, J.C. Characterization of a selenate-resistant Arabidopsis mutant. Root growth as a potential target for selenate toxicity. Plant Physiol. 2007, 143, 1231–1241. [Google Scholar] [CrossRef]
- Rouached, H.; Wirtz, M.; Alary, R.; Hell, R.; Arpat, A.B.; Davidian, J.C.; Fourcroy, P.; Berthomieu, P. Differential regulation of the expression of two high-affinity sulfate transporters, SULTR1.1 and SULTR1.2, in Arabidopsis. Plant Physiol. 2008, 147, 897–911. [Google Scholar] [CrossRef] [PubMed]
- Ferri, A.; Lancilli, C.; Maghrebi, M.; Lucchini, G.; Sacchi, G.A.; Nocito, F.F. The sulfate supply maximizing Arabidopsis shoot growth is higher under long- than short-term exposure to cadmium. Front. Plant Sci. 2017, 8, 854. [Google Scholar] [CrossRef] [PubMed]
- Maruyama-Nakashita, A.; Watanabe-Takahashi, A.; Inoue, E.; Yamaya, T.; Saito, K.; Takahashi, H. Sulfur-responsive elements in the 3’-nontranscribed intergenic region are essential for the induction of SULFATE TRANSPORTER 2;1 gene expression in Arabidopsis roots under sulfur deficiency. Plant Cell 2015, 27, 1279–1296. [Google Scholar] [CrossRef] [PubMed]
- Kawashima, C.G.; Yoshimoto, N.; Maruyama-Nakashita, A.; Tsuchiya, Y.N.; Saito, K.; Takahashi, H.; Dalmay, T. Sulphur starvation induces the expression of microRNA-395 and one of its target genes but in different cell types. Plant J. 2009, 57, 313–321. [Google Scholar] [CrossRef]
- Kawashima, C.G.; Matthewman, C.A.; Huang, S.; Lee, B.R.; Yoshimoto, N.; Koprivova, A.; Rubio-Somoza, I.; Todesco, M.; Rathjen, T.; Saito, K.; et al. Interplay of SLIM1 and miR395 in the regulation of sulfate assimilation in Arabidopsis. Plant J. 2011, 66, 863–876. [Google Scholar] [CrossRef]
- Maruyama-Nakashita, A.; Nakamura, Y.; Tohge, T.; Saito, K.; Takahashi, H. Arabidopsis SLIM1 is a central transcriptional regulator of plant sulfur response and metabolism. Plant Cell 2006, 18, 3235–3251. [Google Scholar] [CrossRef]
- Yoshimoto, N.; Inoue, E.; Saito, K.; Yamaya, T.; Takahashi, H. Phloem-localizing sulfate transporter, Sultr1;3, mediates re-distribution of sulfur from source to sink organs in Arabidopsis. Plant Physiol. 2003, 131, 1511–1517. [Google Scholar] [CrossRef]
- Kataoka, T.; Watanabe-Takahashi, A.; Hayashi, N.; Ohnishi, M.; Mimura, T.; Buchner, P.; Hawkesford, M.J.; Yamaya, T.; Takahashi, H. Vacuolar sulfate transporters are essential determinants controlling internal distribution of sulfate in Arabidopsis. Plant Cell 2004, 16, 2693–2704. [Google Scholar] [CrossRef]
- Cao, M.J.; Wang, Z.; Wirtz, M.; Hell, R.; Oliver, D.J.; Xiang, C.B. SULTR3;1 is a chloroplast-localized sulfate transporter in Arabidopsis thaliana. Plant J. 2013, 73, 607–616. [Google Scholar] [CrossRef]
- Chen, Z.; Zhao, P.X.; Miao, Z.Q.; Qi, G.F.; Wang, Z.; Yuan, Y.; Ahmad, N.; Cao, M.J.; Hell, R.; Wirtz, M.; et al. SULTR3s function in chloroplast sulfate uptake and affect ABA biosynthesis and the stress response. Plant Physiol. 2019, 180, 593–604. [Google Scholar] [CrossRef]
- Kataoka, T.; Hayashi, N.; Yamaya, T.; Takahashi, H. Root-to-shoot transport of sulfate in Arabidopsis. Evidence for the role of SULTR3;5 as a component of low-affinity sulfate transport system in the root vasculature. Plant Physiol. 2004, 136, 4198–4204. [Google Scholar] [CrossRef] [PubMed]
- Yamaji, N.; Takemoto, Y.; Miyaji, T.; Mitani-Ueno, N.; Yoshida, K.T.; Ma, J.F. Reducing phosphorus accumulation in rice grains with an impaired transporter in the node. Nature 2017, 541, 92–95. [Google Scholar] [CrossRef] [PubMed]
- Ding, G.; Lei, G.J.; Yamaji, N.; Yokosho, K.; Mitani-Ueno, N.; Huang, S.; Ma, J.F. Vascular cambium-localized AtSPDT mediates xylem-to-phloem transfer of phosphorus for its preferential distribution in Arabidopsis. Mol. Plant 2019. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Frank, T.; Tan, Y.; Zhou, C.; Mabnoune, M.; Arpat, A.B.; Cui, H.; Huang, J.; He, Z.; Poirier, Y.; et al. Disruption of OsSULTR3;3 reduces phytate and phosphorus concentrations and alters the metabolite profile in rice grains. New Phytol. 2016, 211, 926–939. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.L.; Xu, X.H.; Ren, X.L.; Fu, H.W.; Wu, D.X.; Shu, Q.Y. Generation and characterization of low phytic acid germplasm in rice (Oryza sativa L.). Theor. Appl. Genet. 2007, 114, 803–814. [Google Scholar] [CrossRef] [PubMed]
- Ye, H.; Zhang, X.Q.; Broughton, S.; Westcott, S.; Wu, D.; Lance, R.; Li, C. A nonsense mutation in a putative sulphate transporter gene results in low phytic acid in barley. Funct. Integr. Genom. 2011, 11, 103–110. [Google Scholar] [CrossRef]
- Wang, F.; Deng, M.; Xu, J.; Zhu, X.; Mao, C. Molecular mechanisms of phosphate transport and signaling in higher plants. Semin. Cell Dev. Biol. 2018, 74, 114–122. [Google Scholar] [CrossRef]
- Ham, B.K.; Chen, J.; Yan, Y.; Lucas, W.J. Insights into plant phosphate sensing and signaling. Curr. Opin. Biotechnol. 2018, 49, 1–9. [Google Scholar] [CrossRef]
- Veneklaas, E.J.; Lambers, H.; Bragg, J.; Finnegan, P.M.; Lovelock, S.E.; Plaxton, W.C.; Price, C.A.; Scheible, W.R.; Shane, M.W.; White, P.J.; et al. Opportunities for improving phosphorus-use efficiency in crop plants. New Phytol. 2012, 195, 306–320. [Google Scholar] [CrossRef]
- Rouached, H. Multilevel coordination of phosphate and sulfate homeostasis in plants. Plant Signal. Behav. 2011, 6, 952–955. [Google Scholar] [CrossRef]
- Härtel, H.; Essigmann, B.; Lokstein, H.; Hoffmann-Benning, S.; Peters-Kotting, M.; Benning, C. The phospholipid-deficient pho1 mutant of Arabidopsis thaliana is affected in the organization, but not in the light acclimation, of the thylakoid membrane. Biomembranes 1998, 1415, 205–218. [Google Scholar] [CrossRef][Green Version]
- Essigmann, B.; Guler, S.; Narang, R.A.; Linke, D.; Benning, C. Phosphate availability affects the thylakoid lipid composition and the expression of SQD1, a gene required for sulfolipid biosynthesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1998, 95, 1950–1955. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Xu, C.; Benning, C. Arabidopsis disrupted in SQD2 encoding sulfolipid synthase is impaired in phosphate-limited growth. Proc. Natl. Acad. Sci. USA 2002, 99, 5732–5737. [Google Scholar] [CrossRef] [PubMed]
- Sugimoto, K.; Sato, N.; Tsuzuki, M. Utilization of a chloroplast membrane sulfolipid as a major internal sulfur source for protein synthesis in the early phase of sulfur starvation in Chlamydomonas reinhardtii. FEBS Lett. 2007, 581, 4519–4522. [Google Scholar] [CrossRef] [PubMed]
- Tjellström, H.; Andersson, M.X.; Larsson, K.E.; Sandelius, A.S. Membrane phospholipids as a phosphate reserve: The dynamic nature of phospholipid-to-digalactosyl diacylglycerol exchange in higher plants. Plant Cell Environ. 2008, 31, 1388–1398. [Google Scholar] [CrossRef] [PubMed]
- Sugimoto, K.; Tsuzuki, M.; Sato, N. Regulation of synthesis and degradation of a sulfolipid under sulfur-starved conditions and its physiological significance in Chlamydomonas reinhardtii. New Phytol. 2010, 185, 676–686. [Google Scholar] [CrossRef]
- Martens, C.; Shekhar, M.; Borysik, A.J.; Lau, A.M.; Reading, E.; Tajkhorshid, E.; Booth, P.J.; Politis, A. Direct protein-lipid interactions shape the conformational landscape of secondary transporters. Nat. Commun. 2018, 9, 4151. [Google Scholar] [CrossRef]
- Rouached, H.; Secco, D.; Arpat, B.; Poirier, Y. The transcription factor PHR1 plays a key role in the regulation of sulfate shoot-to-root flux upon phosphate starvation in Arabidopsis. BMC Plant Biol. 2011, 11, 19. [Google Scholar] [CrossRef]
- Rubio, V.; Linhares, F.; Solano, R.; Martín, A.C.; Iglesias, J.; Leyva, A.; Paz-Ares, J. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 2001, 15, 2122–2133. [Google Scholar] [CrossRef]
- Bustos, R.; Castrillo, G.; Linhares, F.; Puga, M.I.; Rubio, V.; Pérez, J.; Solano, R.; Leyva, A.; Paz-Ares, J. A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet. 2010, 6, e1001102. [Google Scholar] [CrossRef]
- Franco-Zorrilla, J.M.; Gonzalez, E.; Bustos, R.; Linhares, F.; Leyva, A.; Paz-Ares, J. The transcriptional control of plant responses to phosphate limitation. J. Exp. Bot. 2004, 55, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Stefanovic, A.; Ribot, C.; Rouached, H.; Wang, Y.; Chong, J.; Belbahri, L.; Delessert, S.; Poirier, Y. Members of the PHO1 gene family show limited functional redundancy in phosphate transfer to the shoot, and are regulated by phosphate deficiency via distinct pathways. Plant J. 2007, 50, 982–994. [Google Scholar] [CrossRef] [PubMed]
- Khan, G.A.; Bouraine, S.; Wege, S.; Li, Y.; de Carbonnel, M.; Berthomieu, P.; Poirier, Y.; Rouached, H. Coordination between zinc and phosphate homeostasis involves the transcription factor PHR1, the phosphate exporter PHO1, and its homologue PHO1;H3 in Arabidopsis. J. Exp. Bot. 2014, 65, 871–884. [Google Scholar] [CrossRef] [PubMed]
- Bournier, M.; Tissot, N.; Mari, S.; Boucherez, J.; Lacombe, E.; Briat, J.F.; Gaymard, F. Arabidopsis ferritin 1 (AtFer1) gene regulation by the phosphate starvation response 1 (AtPHR1) transcription factor reveals a direct molecular link between iron and phosphate homeostasis. J. Biol. Chem. 2013, 288, 22670–22680. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, L.; Lin, S.; Shih, A.; Chen, J.; Lin, W.; Tseng, C.; Lin, W.Y.; Tseng, C.Y.; Li, H.Y.; Tzyy-Jen Chiou, T.J. Uncovering small RNA-mediated responses to phosphate-deficiency in Arabidopsis by deep sequencing. Plant Physiol. 2009, 151, 2120–2132. [Google Scholar] [CrossRef]
- Belgaroui, N.; Zaidi, I.; Farhat, A.; Chouayekh, H.; Bouain, N.; Chay, S.; Curie, C.; Mari, S.; Masmoudi, K.; Davidian, J.C.; et al. Over-expression of the bacterial phytase US417 in Arabidopsis reduces the concentration of phytic acid and reveals its involvement in the regulation of sulfate and phosphate homeostasis and signaling. Plant Cell Physiol. 2004, 55, 1912–1924. [Google Scholar] [CrossRef]
- Cominelli, E.; Pilu, R.; Sparvoli, F. Phytic acid, transporters and the generation of low phytic acid mutants in grain crops. Plants 2019. submitted for publication. [Google Scholar]
- Yamaji, N.; Ma, J.F. The node, a hub for mineral nutrient distribution in graminaceous plants. Trends Plant Sci. 2014, 19, 556–563. [Google Scholar] [CrossRef]
- Yamaji, N.; Ma, J.F. Node-controlled allocation of mineral elements in Poaceae. Curr. Opin. Plant Biol. 2017, 39, 18–24. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sacchi, G.A.; Nocito, F.F. Plant Sulfate Transporters in the Low Phytic Acid Network: Some Educated Guesses. Plants 2019, 8, 616. https://doi.org/10.3390/plants8120616
Sacchi GA, Nocito FF. Plant Sulfate Transporters in the Low Phytic Acid Network: Some Educated Guesses. Plants. 2019; 8(12):616. https://doi.org/10.3390/plants8120616
Chicago/Turabian StyleSacchi, Gian Attilio, and Fabio Francesco Nocito. 2019. "Plant Sulfate Transporters in the Low Phytic Acid Network: Some Educated Guesses" Plants 8, no. 12: 616. https://doi.org/10.3390/plants8120616
APA StyleSacchi, G. A., & Nocito, F. F. (2019). Plant Sulfate Transporters in the Low Phytic Acid Network: Some Educated Guesses. Plants, 8(12), 616. https://doi.org/10.3390/plants8120616





