Perspectives on microRNAs and Phased Small Interfering RNAs in Maize (Zea mays L.): Functions and Big Impact on Agronomic Traits Enhancement
Abstract
:1. Introduction
2. MiRNA and phasiRNA Biogenesis in Maize
2.1. Core Components of sRNA Biogenesis in Plants
2.2. MiRNA-Mediated Gene Silencing in Maize
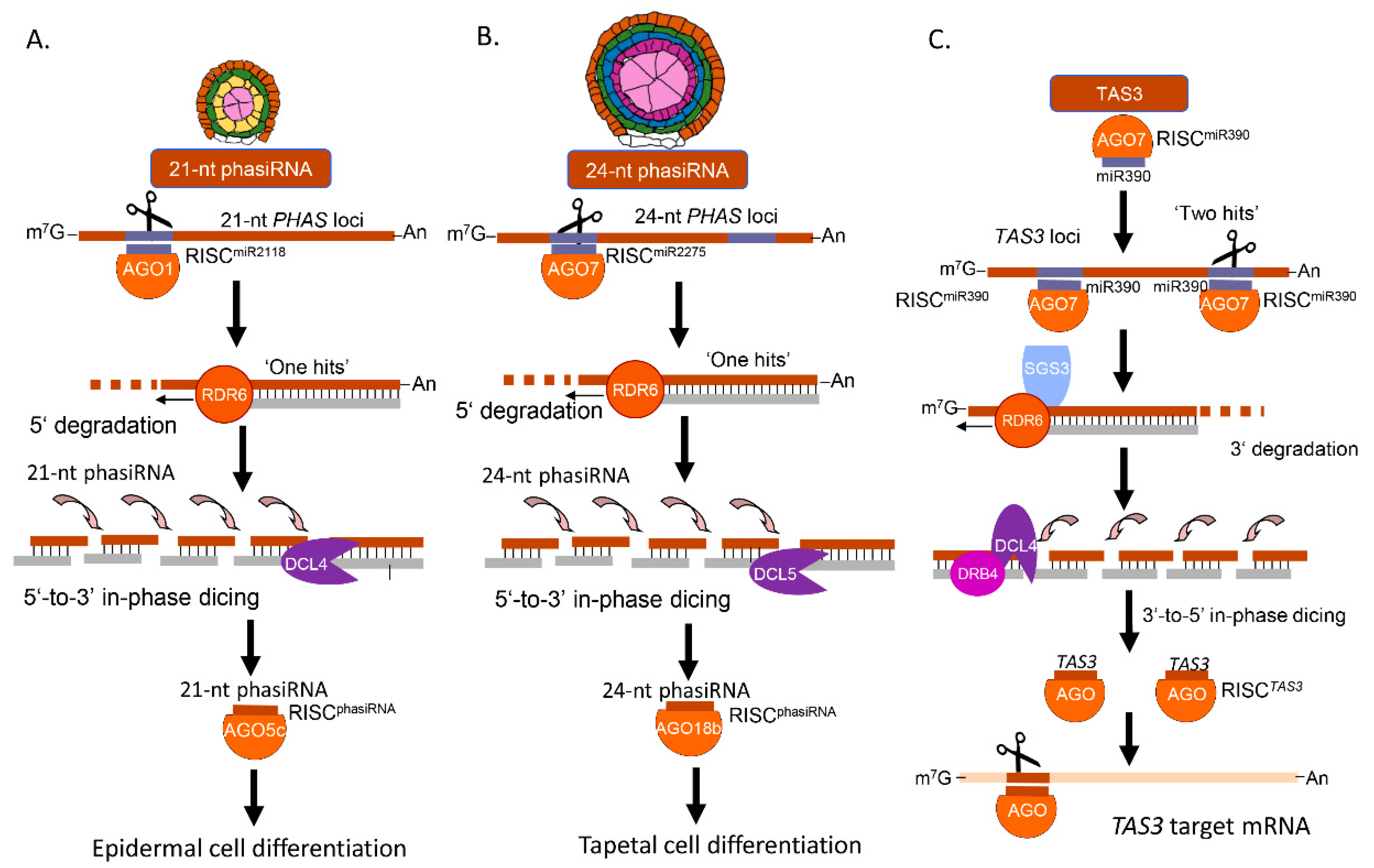
2.3. Origin and Biogenesis of phasiRNAs in Maize
2.4. Functional Redundancy and Divergence of the Key Components in Maize sRNA Biogenesis Pathways
2.4.1. DCLs
2.4.2. AGOs
2.4.3. RDRs
3. Functions of miRNAs and phasiRNAs in Maize
3.1. The Interaction of miR156 and miR172 Fine Tunes Plant Developmental Timing
3.2. Plant Architecture Modulated by miR156 and miR319
3.3. Roles of miR172, miR156 and miR159 in Sex Determination
3.4. Leaf Patterns Are Shaped by miR166 and miR390-TAS3
3.5. PhasiRNAs and Maize Male Fertility
3.6. Other miRNA Functions in Maize
4. Exploiting the Roles of Maize Small RNAs in Important Agronomic Traits Improvement
5. Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chen, X. Small RNAs and their roles in plant development. Annu. Rev. Cell Dev. Biol. 2009, 25, 21–44. [Google Scholar] [CrossRef] [PubMed]
- Kim, V.N.; Han, J.; Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 2009, 10, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Fei, Q.; Xia, R.; Meyers, B.C. Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks. Plant Cell 2013, 25, 2400–2415. [Google Scholar] [CrossRef] [PubMed]
- Axtell, M.J. Classification and comparison of small RNAs from plants. Annu. Rev. Plant Biol. 2013, 64, 137–159. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ma, Z.; Castillo-González, C.; Sun, D.; Li, Y.; Yu, B.; Zhao, B.; Li, P.; Zhang, X. SWI2/SNF2 ATPase CHR2 remodels pri-miRNAs via Serrate to impede miRNA production. Nature 2018, 557, 516–521. [Google Scholar] [CrossRef]
- Rogers, K.; Chen, X. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell 2013, 25, 2383–2399. [Google Scholar] [CrossRef] [PubMed]
- Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 2009, 136, 669–687. [Google Scholar] [CrossRef]
- Rhoades, M.W.; Reinhart, B.J.; Lim, L.P.; Burge, C.B.; Bartel, B.; Bartel, D.P. Prediction of plant microRNA targets. Cell 2002, 110, 513–520. [Google Scholar] [CrossRef]
- Cuperus, J.T.; Fahlgren, N.; Carrington, J.C. Evolution and functional diversification of MIRNA Genes. Plant Cell 2011, 23, 431–442. [Google Scholar] [CrossRef]
- Jones-Rhoades, M.W.; Bartel, D.P.; Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 2006, 57, 19–53. [Google Scholar] [CrossRef]
- Yu, Y.; Zhou, Y.; Zhang, Y.; Chen, Y. Grass phasiRNAs and male fertility. Sci. China Life Sci. 2018, 61, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Zhai, J.; Zhang, H.; Arikit, S.; Huang, K.; Nan, G.L.; Walbot, V.; Meyers, B.C. Spatiotemporally dynamic, cell-type-dependent premeiotic and meiotic phasiRNAs in maize anthers. Proc. Natl. Acad. Sci. USA 2015, 112, 3146–3151. [Google Scholar] [CrossRef]
- Juarez, M.T.; Twigg, R.W.; Timmermans, M.C. Specification of adaxial cell fate during maize leaf development. Development 2004, 131, 4533–4544. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, H.; Wang, H. The miR156/SPL module, a regulatory hub and versatile toolbox, gears up crops for enhanced agronomic traits. Mol. Plant 2015. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Chu, C. MicroRNAs in crop improvement: Fine-tuners for complex traits. Nat. Plants 2017, 3, 17077. [Google Scholar] [CrossRef] [PubMed]
- Gore, M.A.; Chia, J.M.; Elshire, R.J.; Sun, Q.; Ersoz, E.S.; Hurwitz, B.L.; Peiffer, J.A.; McMullen, M.D.; Grills, G.S.; Ross-Ibarra, J.; et al. A first-generation haplotype map of maize. Science 2009, 326, 1115–1117. [Google Scholar] [CrossRef] [PubMed]
- Brutnell, T.P. Transposon tagging in maize. Funct. Integr. Genom. 2002, 2, 4–12. [Google Scholar] [CrossRef]
- Arteaga-Vazquez, M.A.; Chandler, V.L. Paramutation in maize: RNA mediated trans-generational gene silencing. Curr. Opin. Genet. Dev. 2010, 20, 156–163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, W.; Xiang, X.; Zhai, L.; Zhang, D.; Cao, Z.; Liu, L.; Zhang, Z. AGO18b negatively regulates determinacy of spikelet meristems on the tassel central spike in maize. J. Integr. Plant Biol. 2018, 60, 65–78. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Liu, X.; Yang, J.; Liu, W.; Du, Q.; Wang, H.; Fu, C.; Li, W.X. MicroRNA528 affects lodging resistance of maize by regulating lignin biosynthesis under nitrogen-luxury conditions. Mol. Plant 2018, 11, 806–814. [Google Scholar] [CrossRef]
- Field, S.; Thompson, B. Analysis of the Maize dicer-like1 Mutant, fuzzy tassel, Implicates MicroRNAs in Anther Maturation and Dehiscence. PLoS ONE 2016, 11, e0146534. [Google Scholar] [CrossRef] [PubMed]
- Thompson, B.E.; Basham, C.; Hammond, R.; Ding, Q.; Kakrana, A.; Lee, T.F.; Simon, S.A.; Meeley, R.; Meyers, B.C.; Hake, S. The dicer-like1 homolog fuzzy tassel is required for the regulation of meristem determinacy in the inflorescence and vegetative growth in maize. Plant Cell 2014, 26, 4702–4717. [Google Scholar] [CrossRef] [PubMed]
- Petsch, K.; Manzotti, P.S.; Tam, O.H.; Meeley, R.; Hammell, M.; Consonni, G.; Timmermans, M.C. Novel DICER-LIKE1 siRNAs bypass the requirement for DICER-LIKE4 in maize development. Plant Cell 2015, 27, 2163–2177. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Yang, H.; Zou, C.; Li, W.X.; Xu, Y.; Xie, C. Identification and functional characterization of the AGO1 ortholog in maize. J. Integr. Plant Biol. 2016, 58, 749–758. [Google Scholar] [CrossRef] [PubMed]
- Chuck, G.; Cigan, A.M.; Saeteurn, K.; Hake, S. The heterochronic maize mutant Corngrass1 results from overexpression of a tandem microRNA. Nat. Genet. 2007, 39, 544–549. [Google Scholar] [CrossRef] [PubMed]
- Juarez, M.T.; Kui, J.S.; Thomas, J.; Heller, B.A.; Timmermans, M.C. MicroRNA-mediated repression of rolled leaf1 specifies maize leaf polarity. Nature 2004, 428, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Lauter, N.; Kampani, A.; Carlson, S.; Goebel, M.; Moose, S.P. MicroRNA172 down-regulates glossy15 to promote vegetative phase change in maize. Proc. Natl. Acad. Sci. USA 2005, 102, 9412–9417. [Google Scholar] [CrossRef] [PubMed]
- Luan, M.; Xu, M.; Lu, Y.; Zhang, L.; Fan, Y.; Wang, L. Expression of zma-miR169 miRNAs and their target ZmNF-YA genes in response to abiotic stress in maize leaves. Gene 2015, 555, 178–185. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Wang, X.; Huang, C.; Xu, G.; Liang, Y.; Chen, Q.; Wang, C.; Li, D.; Tian, J.; Wu, L.; et al. Glossy15 plays an important role in the divergence of the vegetative transition between maize and its progenitor, teosinte. Mol. Plant 2017, 10, 1579–1583. [Google Scholar] [CrossRef] [PubMed]
- Chuck, G.S.; Brown, P.J.; Meeley, R.; Hake, S. Maize SBP-box transcription factors unbranched2 and unbranched3 affect yield traits by regulating the rate of lateral primordia initiation. Proc. Natl. Acad. Sci. USA 2014, 111, 18775–18780. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Guo, G.; Guo, W.; Guo, G.; Tong, D.; Ni, Z.; Sun, Q.; Yao, Y. miRNA164-directed cleavage of ZmNAC1 confers lateral root development in maize (Zea mays L.). BMC Plant Biol. 2012, 12, 220. [Google Scholar] [CrossRef] [PubMed]
- Qian, Y.; Cheng, Y.; Cheng, X.; Jiang, H.; Zhu, S.; Cheng, B. Identification and characterization of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families in maize. Plant Cell Rep. 2011, 30, 1347–1363. [Google Scholar] [CrossRef] [PubMed]
- Chuck, G.; Whipple, C.; Jackson, D.; Hake, S. The maize SBP-box transcription factor encoded by tasselsheath4 regulates bract development and the establishment of meristem boundaries. Development 2010, 137, 1243–1250. [Google Scholar] [CrossRef] [PubMed]
- Hultquist, J.F.; Dorweiler, J.E. Feminized tassels of maize mop1 and ts1 mutants exhibit altered levels of miR156 and specific SBP-box genes. Planta 2008, 229, 99–113. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, M.; Arora, R.; Lama, T.; Nijhawan, A.; Khurana, J.P.; Tyagi, A.K.; Kapoor, S. Genome-wide identification, organization and phylogenetic analysis of Dicer-like, Argonaute and RNA-dependent RNA Polymerase gene families and their expression analysis during reproductive development and stress in rice. BMC Genom. 2008, 9, 451. [Google Scholar] [CrossRef] [PubMed]
- Matzke, M.A.; Birchler, J.A. RNAi-mediated pathways in the nucleus. Nat. Rev. Genet. 2005, 6, 24–35. [Google Scholar] [CrossRef]
- He, J.; Dong, Z.; Jia, Z.; Wang, J.; Wang, G. Isolation, expression and functional analysis of a putative RNA-dependent RNA polymerase gene from maize (Zea mays L.). Mol. Biol. Rep. 2010, 37, 865–874. [Google Scholar] [CrossRef]
- Douglas, R.N.; Wiley, D.; Sarkar, A.; Springer, N.; Timmermans, M.C.; Scanlon, M.J. Ragged seedling2 Encodes an ARGONAUTE7-like protein required for mediolateral expansion, but not dorsiventrality, of maize leaves. Plant Cell 2010, 22, 1441–1451. [Google Scholar] [CrossRef]
- Dorweiler, J.E.; Carey, C.C.; Kubo, K.M.; Hollick, J.B.; Kermicle, J.L.; Chandler, V.L. Mediator of paramutation1 is required for establishment and maintenance of paramutation at multiple maize loci. Plant Cell 2000, 12, 2101–2118. [Google Scholar] [CrossRef]
- Ramachandran, V.; Chen, X. Small RNA metabolism in Arabidopsis. Trends Plant Sci. 2008, 13, 368–374. [Google Scholar] [CrossRef]
- Yu, B.; Yang, Z.; Li, J.; Minakhina, S.; Yang, M.; Padgett, R.W.; Steward, R.; Chen, X. Methylation as a crucial step in plant microRNA biogenesis. Science 2005, 307, 932–935. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Weinstein, E.G.; Rhoades, M.W.; Bartel, B.; Bartel, D.P. MicroRNAs in plants. Genes Dev. 2002, 16, 1616–1626. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Llave, C.; Xie, Z.; Kasschau, K.D.; Carrington, J.C. Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 2002, 297, 2053–2056. [Google Scholar] [CrossRef]
- Zhang, Z.; Hu, F.; Sung, M.W.; Shu, C.; Castillo-Gonzalez, C.; Koiwa, H.; Tang, G.; Dickman, M.; Li, P.; Zhang, X. RISC-interacting clearing 3′-5′exoribonucleases (RICEs) degrade uridylated cleavage fragments to maintain functional RISC in Arabidopsis thaliana. eLife 2017, 6, e24466. [Google Scholar] [CrossRef] [PubMed]
- Huntzinger, E.; Izaurralde, E. Gene silencing by microRNAs: Contributions of translational repression and mRNA decay. Nat. Rev. Genet. 2011, 12, 99–110. [Google Scholar] [CrossRef]
- Pontes, O.; Costa-Nunes, P.; Vithayathil, P.; Pikaard, C.S. RNA polymerase V functions in Arabidopsis interphase heterochromatin organization independently of the 24-nt siRNA-directed DNA methylation pathway. Mol. Plant 2009, 2, 700–710. [Google Scholar] [CrossRef] [PubMed]
- Teotia, S.; Singh, D.; Tang, G. DNA Methylation in plants by microRNAs. In Plant Epigenetics; Rajewsky, N., Jurga, S., Barciszewski, J., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 247–262. [Google Scholar]
- Axtell, M.J. Lost in translation? microRNAs at the rough ER. Trends Plant Sci. 2017, 22, 273–274. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Ji, L.; Le, B.H.; Zhai, J.; Chen, J.; Luscher, E.; Gao, L.; Liu, C.; Cao, X.; Mo, B.; et al. ARGONAUTE10 promotes the degradation of miR165/6 through the SDN1 and SDN2 exonucleases in Arabidopsis. PLoS Biol. 2017, 15, e2001272. [Google Scholar] [CrossRef] [PubMed]
- Borges, F.; Martienssen, R.A. The expanding world of small RNAs in plants. Nat. Rev. Mol. Cell Biol. 2015, 16, 727–741. [Google Scholar] [CrossRef] [Green Version]
- Arikit, S.; Zhai, J.; Meyers, B.C. Biogenesis and function of rice small RNAs from non-coding RNA precursors. Curr. Opin. Plant Biol. 2013, 16, 170–179. [Google Scholar] [CrossRef] [PubMed]
- Axtell, M.J.; Jan, C.; Rajagopalan, R.; Bartel, D.P. A two-hit trigger for siRNA biogenesis in plants. Cell 2006, 127, 565–577. [Google Scholar] [CrossRef] [PubMed]
- Allen, E.; Howell, M.D. miRNAs in the biogenesis of trans-acting siRNAs in higher plants. Semin. Cell Dev. Biol. 2010, 21, 798–804. [Google Scholar] [CrossRef] [PubMed]
- De Felippes, F.F.; Marchais, A.; Sarazin, A.; Oberlin, S.; Voinnet, O. A single miR390 targeting event is sufficient for triggering TAS3-tasiRNA biogenesis in Arabidopsis. Nucleic Acids Res. 2017, 45, 5539–5554. [Google Scholar] [CrossRef]
- Dotto, M.C.; Petsch, K.A.; Aukerman, M.J.; Beatty, M.; Hammell, M.; Timmermans, M.C. Genome-wide analysis of leafbladeless1-regulated and phased small RNAs underscores the importance of the TAS3 ta-siRNA pathway to maize development. PLoS Genet. 2014, 10, e1004826. [Google Scholar] [CrossRef] [PubMed]
- Park, W.; Li, J.; Song, R.; Messing, J.; Chen, X. CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr. Biol. 2002, 12, 1484–1495. [Google Scholar] [CrossRef]
- Parent, J.S.; Bouteiller, N.; Elmayan, T.; Vaucheret, H. Respective contributions of Arabidopsis DCL2 and DCL4 to RNA silencing. Plant J. Cell Mol. Biol. 2015, 81, 223–232. [Google Scholar] [CrossRef]
- Xie, Z.; Johansen, L.K.; Gustafson, A.M.; Kasschau, K.D.; Lellis, A.D.; Zilberman, D.; Jacobsen, S.E.; Carrington, J.C. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2004, 2, e104. [Google Scholar] [CrossRef]
- Xie, Z.; Allen, E.; Wilken, A.; Carrington, J.C. DICER-LIKE 4 functions in trans-acting small interfering RNA biogenesis and vegetative phase change in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2005, 102, 12984–12989. [Google Scholar] [CrossRef]
- Margis, R.; Fusaro, A.F.; Smith, N.A.; Curtin, S.J.; Watson, J.M.; Finnegan, E.J.; Waterhouse, P.M. The evolution and diversification of Dicers in plants. FEBS Lett. 2006, 580, 2442–2450. [Google Scholar] [CrossRef]
- Vaucheret, H.; Vazquez, F.; Crete, P.; Bartel, D.P. The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes Dev. 2004, 18, 1187–1197. [Google Scholar] [CrossRef]
- Borges, F.; Pereira, P.A.; Slotkin, R.K.; Martienssen, R.A.; Becker, J.D. MicroRNA activity in the Arabidopsis male germline. J. Exp. Bot. 2011, 62, 1611–1620. [Google Scholar] [CrossRef] [PubMed]
- Harvey, J.J.; Lewsey, M.G.; Patel, K.; Westwood, J.; Heimstadt, S.; Carr, J.P.; Baulcombe, D.C. An antiviral defense role of AGO2 in plants. PLoS ONE 2011, 6, e14639. [Google Scholar] [CrossRef]
- Zilberman, D.; Cao, X.; Johansen, L.K.; Xie, Z.; Carrington, J.C.; Jacobsen, S.E. Role of Arabidopsis ARGONAUTE4 in RNA-directed DNA methylation triggered by inverted repeats. Curr. Biol. 2004, 14, 1214–1220. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Yang, Z.; Wang, Y.; Zheng, L.; Ye, R.; Ji, Y.; Zhao, S.; Ji, S.; Liu, R.; Xu, L.; et al. Viral-inducible Argonaute18 confers broad-spectrum virus resistance in rice by sequestering a host microRNA. eLife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Zhai, L.; Sun, W.; Zhang, K.; Jia, H.; Liu, L.; Liu, Z.; Teng, F.; Zhang, Z. Identification and characterization of Argonaute gene family and meiosis-enriched Argonaute during sporogenesis in maize. J. Integr. Plant Biol. 2014, 56, 1042–1052. [Google Scholar] [CrossRef] [PubMed]
- Marin, E.; Jouannet, V.; Herz, A.; Lokerse, A.S.; Weijers, D.; Vaucheret, H.; Nussaume, L.; Crespi, M.D.; Maizel, A. miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 2010, 22, 1104–1117. [Google Scholar] [CrossRef] [PubMed]
- Montgomery, T.A.; Howell, M.D.; Cuperus, J.T.; Li, D.; Hansen, J.E.; Alexander, A.L.; Chapman, E.J.; Fahlgren, N.; Allen, E.; Carrington, J.C. Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 2008, 133, 128–141. [Google Scholar] [CrossRef] [PubMed]
- Xia, R.; Meyers, B.C.; Liu, Z.; Beers, E.P.; Ye, S.; Liu, Z. MicroRNA superfamilies descended from miR390 and their roles in secondary small interfering RNA Biogenesis in Eudicots. Plant Cell 2013, 25, 1555–1572. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Higgins, D.G. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 2018, 27, 135–145. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life v2: Online annotation and display of phylogenetic trees made easy. Nucleic Acids Res. 2011, 39, W475–W478. [Google Scholar] [CrossRef]
- Zaratiegui, M.; Irvine, D.V.; Martienssen, R.A. Noncoding RNAs and gene silencing. Cell 2007, 128, 763–776. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.H.; Coruh, C.; Axtell, M.J. miR156 and miR390 regulate tasiRNA accumulation and developmental timing in Physcomitrella patens. Plant Cell 2012, 24, 4837–4849. [Google Scholar] [CrossRef]
- Aukerman, M.J.; Sakai, H. Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 2003, 15, 2730–2741. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.H.; Seo, Y.H.; Seo, P.J.; Reyes, J.L.; Yun, J.; Chua, N.H.; Park, C.M. The GIGANTEA-regulated microRNA172 mediates photoperiodic flowering independent of CONSTANS in Arabidopsis. Plant Cell 2007, 19, 2736–2748. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Poethig, R.S. Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development 2006, 133, 3539–3547. [Google Scholar] [CrossRef] [PubMed]
- Chuck, G.S.; Tobias, C.; Sun, L.; Kraemer, F.; Li, C.; Dibble, D.; Arora, R.; Bragg, J.N.; Vogel, J.P.; Singh, S.; et al. Overexpression of the maize Corngrass1 microRNA prevents flowering, improves digestibility, and increases starch content of switchgrass. Proc. Natl. Acad. Sci. USA 2011, 108, 17550–17555. [Google Scholar] [CrossRef] [PubMed]
- Doebley, J.; Stec, A.; Hubbard, L. The evolution of apical dominance in maize. Nature 1997, 386, 485–488. [Google Scholar] [CrossRef] [PubMed]
- Kidner, C.A.; Martienssen, R.A. Spatially restricted microRNA directs leaf polarity through ARGONAUTE1. Nature 2004, 428, 81–84. [Google Scholar] [CrossRef] [PubMed]
- Palatnik, J.F.; Allen, E.; Wu, X.; Schommer, C.; Schwab, R.; Carrington, J.C.; Weigel, D. Control of leaf morphogenesis by microRNAs. Nature 2003, 425, 257–263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schommer, C.; Palatnik, J.F.; Aggarwal, P.; Chetelat, A.; Cubas, P.; Farmer, E.E.; Nath, U.; Weigel, D. Control of jasmonate biosynthesis and senescence by miR319 targets. PLoS Biol. 2008, 6, e230. [Google Scholar] [CrossRef]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Yu, H.; Xiong, G.; Wang, J.; Jiao, Y.; Liu, G.; Jing, Y.; Meng, X.; Hu, X.; Qian, Q.; et al. Genome-wide binding analysis of the transcription activator ideal plant architecture1 reveals a complex network regulating rice plant architecture. Plant Cell 2013, 25, 3743–3759. [Google Scholar] [CrossRef] [PubMed]
- Moreno, M.A.; Harper, L.C.; Krueger, R.W.; Dellaporta, S.L.; Freeling, M. liguleless1 encodes a nuclear-localized protein required for induction of ligules and auricles during maize leaf organogenesis. Genes Dev. 1997, 11, 616–628. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Du, Y.; Shen, X.; Li, M.; Sun, W.; Huang, J.; Liu, Z.; Tao, Y.; Zheng, Y.; Yan, J.; et al. KRN4 controls quantitative variation in maize kernel row number. PLoS Genet. 2015, 11, e1005670. [Google Scholar] [CrossRef] [PubMed]
- Chuck, G.; Meeley, R.; Irish, E.; Sakai, H.; Hake, S. The maize tasselseed4 microRNA controls sex determination and meristem cell fate by targeting Tasselseed6/indeterminate spikelet1. Nat. Genet. 2007, 39, 1517–1521. [Google Scholar] [CrossRef] [PubMed]
- Banks, J.A. MicroRNA, sex determination and floral meristem determinacy in maize. Genome Biol. 2008, 9, 204. [Google Scholar] [CrossRef] [PubMed]
- Peng, T.; Qiao, M.; Liu, H.; Teotia, S.; Zhang, Z.; Zhao, Y.; Wang, B.; Zhao, D.; Shi, L.; Zhang, C.; et al. A resource for inactivation of microRNAs using Short Tandem Target Mimic technology in model and crop plants. Mol. Plant 2018, 11, 1400–1417. [Google Scholar] [CrossRef] [PubMed]
- Donaire, L.; Barajas, D.; Martinez-Garcia, B.; Martinez-Priego, L.; Pagan, I.; Llave, C. Structural and genetic requirements for the biogenesis of tobacco rattle virus-derived small interfering RNAs. J. Virol. 2008, 82, 5167–5177. [Google Scholar] [CrossRef] [PubMed]
- Tsuji, H.; Aya, K.; Ueguchi-Tanaka, M.; Shimada, Y.; Nakazono, M.; Watanabe, R.; Nishizawa, N.K.; Gomi, K.; Shimada, A.; Kitano, H.; et al. GAMYB controls different sets of genes and is differentially regulated by microRNA in aleurone cells and anthers. Plant J. Cell Mol. Biol. 2006, 47, 427–444. [Google Scholar] [CrossRef] [PubMed]
- Archak, S.; Nagaraju, J. Computational prediction of rice (Oryza sativa) miRNA targets. Genom. Proteom. Bioinform. 2007, 5, 196–206. [Google Scholar] [CrossRef]
- Nogueira, F.T.; Madi, S.; Chitwood, D.H.; Juarez, M.T.; Timmermans, M.C. Two small regulatory RNAs establish opposing fates of a developmental axis. Genes Dev. 2007, 21, 750–755. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McConnell, J.R.; Emery, J.; Eshed, Y.; Bao, N.; Bowman, J.; Barton, M.K. Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 2001, 411, 709–713. [Google Scholar] [CrossRef] [PubMed]
- Emery, J.F.; Floyd, S.K.; Alvarez, J.; Eshed, Y.; Hawker, N.P.; Izhaki, A.; Baum, S.F.; Bowman, J.L. Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr. Biol. 2003, 13, 1768–1774. [Google Scholar] [CrossRef] [PubMed]
- Nogueira, F.T.; Chitwood, D.H.; Madi, S.; Ohtsu, K.; Schnable, P.S.; Scanlon, M.J.; Timmermans, M.C. Regulation of small RNA accumulation in the maize shoot apex. PLoS Genet. 2009, 5, e1000320. [Google Scholar] [CrossRef] [PubMed]
- Shkibbe, D.S.; Schnable, P.S. Male sterility in maize. Maydica 2005, 50, 367–376. [Google Scholar]
- Wang, H.; Nussbaum-Wagler, T.; Li, B.; Zhao, Q.; Vigouroux, Y.; Faller, M.; Bomblies, K.; Lukens, L.; Doebley, J.F. The origin of the naked grains of maize. Nature 2005, 436, 714–719. [Google Scholar] [CrossRef]
- Wang, H.; Studer, A.J.; Zhao, Q.; Meeley, R.; Doebley, J.F. Evidence that the origin of naked kernels during maize domestication was caused by a single amino acid substitution in tga1. Genetics 2015, 200, 965–974. [Google Scholar] [CrossRef]
- Wu, L.; Zhang, D.; Xue, M.; Qian, J.; He, Y.; Wang, S. Overexpression of the maize GRF10, an endogenous truncated growth-regulating factor protein, leads to reduction in leaf size and plant height. J. Integr. Plant Biol. 2014, 56, 1053–1063. [Google Scholar] [CrossRef]
- Yan, J.; Zhao, C.; Zhou, J.; Yang, Y.; Wang, P.; Zhu, X.; Tang, G.; Bressan, R.A.; Zhu, J.K. The miR165/166 mediated regulatory module plays critical roles in ABA homeostasis and response in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006416. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, H.; Srivastava, A.K.; Pan, Y.; Bai, J.; Fang, J.; Shi, H.; Zhu, J.K. Knockdown of rice microRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol. 2018, 176, 2082–2094. [Google Scholar] [CrossRef]
- Hochholdinger, F.; Yu, P.; Marcon, C. Genetic control of root system development in maize. Trends Plant Sci. 2018, 23, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Du, Q.; Wang, K.; Zou, C.; Xu, C.; Li, W.X. The PILNCR1-miR399 regulatory module is important for low phosphate tolerance in maize. Plant Physiol. 2018, 177, 1743–1753. [Google Scholar] [CrossRef]
- Duvick, D.N. The contribution of breeding to yield advances in maize (Zea mays L.). Adv. Agron. 2005, 86, 83–145. [Google Scholar] [CrossRef]
- Buckler, E.S.; Holland, J.B.; Bradbury, P.J.; Acharya, C.B.; Brown, P.J.; Browne, C.; Ersoz, E.; Flint-Garcia, S.; Garcia, A.; Glaubitz, J.C.; et al. The genetic architecture of maize flowering time. Science 2009, 325, 714–718. [Google Scholar] [CrossRef]
- Lackey, E.; Ng, D.W.; Chen, Z.J. RNAi-mediated down-regulation of DCL1 and AGO1 induces developmental changes in resynthesized Arabidopsis allotetraploids. New Phytol. 2010, 186, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.; Candela, H.; Hake, S. The Liguleless narrow mutation affects proximal-distal signaling and leaf growth. Development 2013, 140, 405–412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luan, M.; Xu, M.; Lu, Y.; Zhang, Q.; Zhang, L.; Zhang, C.; Fan, Y.; Lang, Z.; Wang, L. Family-wide survey of miR169s and NF-YAs and their expression profiles response to abiotic stress in maize roots. PLoS ONE 2014, 9, e91369. [Google Scholar] [CrossRef]
- Portwood, J.L., 2nd; Woodhouse, M.R.; Cannon, E.K.; Gardiner, J.M.; Harper, L.C.; Schaeffer, M.L.; Walsh, J.R.; Sen, T.Z.; Cho, K.T.; Schott, D.A.; et al. MaizeGDB 2018: The maize multi-genome genetics and genomics database. Nucleic Acids Res. 2019, 47, D1146–D1154. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Gu, Y.; Jia, X.; Kang, W.; Pan, S.; Tang, X.; Chen, X.; Tang, G. Effective small RNA destruction by the expression of a short tandem target mimic in Arabidopsis. Plant Cell 2012, 24, 415–427. [Google Scholar] [CrossRef]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; Garcia, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef]
- Jeong, D.H.; Park, S.; Zhai, J.; Gurazada, S.G.; De Paoli, E.; Meyers, B.C.; Green, P.J. Massive analysis of rice small RNAs: Mechanistic implications of regulated microRNAs and variants for differential target RNA cleavage. Plant Cell 2011, 23, 4185–4207. [Google Scholar] [CrossRef] [PubMed]




| Gene | Accession Number | Chromosomal Location (5′-3′) | Type |
|---|---|---|---|
| 1. ZmDCLs | |||
| ZmDCL1 | GRMZM2G040762_P01 | Chr. 1: 4,600,841–4,608,248 | DCL1 |
| ZmDCL2 | GRMZM2G301405_P01 | Chr. 5: 19,916,753–19,927,967 | DCL2 |
| ZmDCL3a | GRMZM5G814985_P01 | Chr. 3: 164,415,209–164,418,189 | DCL3 |
| ZmDCL5/3b | GRMZM2G413853_P01 | Chr. 1: 229,801,762–229,819,069 | DCL3 |
| ZmDCL4 | GRMZM2G160473_P01 | Chr. 10: 129,990,456–129,992,917 | DCL4 |
| 2. ZmAGOs | |||
| ZmAGO1a | GRMZM2G441583_P01 | Chr. 6: 43,253,105–43,261,555 | AGO1 |
| ZmAGO1b | AC209206.3_FGP011 | Chr. 10: 137,506,877–137,513,415 | AGO1 |
| ZmAGO1c | GRMZM2G039455_P01 | Chr. 2: 17,563,301–17,573,156 | AGO1 |
| ZmAGO1d | GRMZM2G361518_P01 | Chr. 5: 64,791,077–64,796,881 | AGO1 |
| ZmAGO2a | GRMZM2G007791_P01 | Chr. 2: 9,973,816–9,981,340 | ZIPPY |
| ZmAGO2b | GRMZM2G354867_P01 | Chr. 1:142,397,812–142,403,450 | ZIPPY |
| ZmAGO4 | GRMZM2G589579_P01 | Chr. 8: 2,511,663–2,519,008 | AGO4 |
| ZmAGO5a | GRMZM2G461936_P02 | Chr. 5: 13,611,800–13,618,698 | MEL1 |
| ZmAGO5b | GRMZM2G059033_P01 | Chr. 2: 233,385,077–233,392,000 | MEL1 |
| ZmAGO5c | GRMZM2G347402_P01 | Chr. 7: 72,044,775–72,053,779 | MEL1 |
| ZmAGO5d | GRMZM2G123063_P01 | Chr. 5:4,000,995–4,009,425 | MEL1 |
| ZmAGO7 | GRMZM2G354867_P01 | Chr. 10: 141,823,070–141,828,449 | ZIPPY |
| ZmAGO9 | GRMZM2G141818_P03 | Chr. 6: 168,642,369–168,650,358 | AGO4 |
| ZmAGO10a | AC189879.3_FG003 | Chr. 9: 87,408,375–87,414,276 | AGO1 |
| ZmAGO10b | GRMZM2G079080_P02 | Chr. 6: 103,286,236–103,293,200 | AGO1 |
| ZmAGO18a | GRMZM2G105250_P01 | Chr. 2: 199,510,528–199,516,085 | OsAGO18 |
| ZmAGO18b | GRMZM2G457370_P01 | Chr. 1: 250,132,189–250,137,737 | OsAGO18 |
| ZmAGO18c | GRMZM2G457370_P02 | Chr. 1: 250,132,189–250,137,737 | OsAGO18 |
| 3. ZmRDRs | |||
| ZmRDR1 | GRMZM2G481730_P01 | Chr. 5: 205,385,818–205,389,710 | RDR1 |
| ZmMOP1 | GRMZM2G042443_P01 | Chr. 2: 41,131,324–41,136,928 | RDR2 |
| ZmRDR6a | GRMZM2G357825_P01 | Chr. 9: 109,055,576–109,093,885 | RDR6 |
| ZmRDR6b | GRMZM2G145201_P01 | Chr. 3: 102,532,883–102,536,036 | RDR6 |
| ZmRDR6c | GRMZM2G347931_P01 | Chr. 9: 106,302,354–106,306,175 | RDR6 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Z.; Teotia, S.; Tang, J.; Tang, G. Perspectives on microRNAs and Phased Small Interfering RNAs in Maize (Zea mays L.): Functions and Big Impact on Agronomic Traits Enhancement. Plants 2019, 8, 170. https://doi.org/10.3390/plants8060170
Zhang Z, Teotia S, Tang J, Tang G. Perspectives on microRNAs and Phased Small Interfering RNAs in Maize (Zea mays L.): Functions and Big Impact on Agronomic Traits Enhancement. Plants. 2019; 8(6):170. https://doi.org/10.3390/plants8060170
Chicago/Turabian StyleZhang, Zhanhui, Sachin Teotia, Jihua Tang, and Guiliang Tang. 2019. "Perspectives on microRNAs and Phased Small Interfering RNAs in Maize (Zea mays L.): Functions and Big Impact on Agronomic Traits Enhancement" Plants 8, no. 6: 170. https://doi.org/10.3390/plants8060170




