Genetic Characterization of Apulian Olive Germplasm as Potential Source in New Breeding Programs
Abstract
:1. Introduction
2. Results
2.1. Allelic Variation of SSR Markers
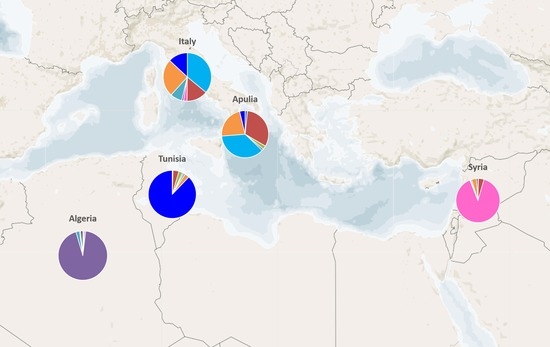
2.2. Genetic Characterization of Olive Germplasm
2.3. Synonymies Discovery
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. SSR Molecular Analysis
4.3. Molecular Marker Diversity and Population Structure
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Belaj, A.; del Carmen Dominguez-García, M.; Atienza, S.G.; Urdíroz, N.M.; De la Rosa, R.; Satovic, Z.; Martín, A.; Kilian, A.; Trujillo, I.; Valpuesta, V.; et al. Developing a core collection of olive (Olea europaea L.) based on molecular markers (DArTs, SSRs, SNPs) and agronomic traits. Tree Genet. Genomes 2012, 8, 365–378. [Google Scholar] [CrossRef]
- Sardaro, R.; Bozzo, F.; Petrontino, A.; Fucilli, V. Community preferences in support of a conservation programme for olive landraces in the Mediterranean area. Acta Hortic. 2018, 1199, 183–188. [Google Scholar] [CrossRef]
- Famiani, F.; Farinelli, D.; Gardi, T.; Rosati, A. The cost of flowering in olive (Olea europaea L.). Sci. Hortic. 2019, 252, 268–273. [Google Scholar] [CrossRef]
- Boskou, D.; Camposeo, S.; Clodoveo, M.L. Table Olives as Sources of Bioactive Compounds. In Olives and Olive Oil Bioactive Constituents; Boskou, D., Ed.; AOCS Press: Urbana, IL, USA, 2015; pp. 217–260. ISBN 978-1-630670-41-2. [Google Scholar]
- Clodoveo, M.L.; Camposeo, S.; Amirante, R.; Dugo, G.; Cicero, N.; Boskou, D. Research and Innovative Approaches to Obtain Virgin Olive Oils with a Higher Level of Bioactive Constituents. In Olives and Olive Oil Bioactive Constituents; Boskou, D., Ed.; AOCS Press: Urbana, IL, USA, 2015; pp. 179–216. ISBN 978-1-630670-41-2. [Google Scholar]
- Sardaro, R.; Fucilli, V.; Acciani, C. Measuring the value of rural landscape in support of preservation policies. Scienze Regionali 2015, 14, 125–138. [Google Scholar] [CrossRef]
- ISTAT 2018. Available online: http://dati.istat.it (accessed on 20 May 2019).
- Besnard, G.; Khadari, B.; Navascués, M.; Fernández-Mazuecos, M.; El Bakkali, A.; Arrigo, N.; Baali-Cherif, D.; Brunini-Bronzini de Caraffa, V.; Santoni, S.; Vargas, P.; et al. The complex history of the olive tree: From late quaternary diversification of mediterranean lineages to primary domestication in the northern Levant. Proc. R. Soc. B Biol. Sci. 2013, 280. [Google Scholar] [CrossRef] [PubMed]
- Vossen, P. Olive oil: History, production, and characteristics of the world’s classic oils. HortScience 2007, 42, 1093–1100. [Google Scholar] [CrossRef]
- Diez, C.M.; Trujillo, I.; Martinez-Uriroz, N.; Barranco, D.; Rallo, L. Olive domestication and diversification in the mediterranean basin. New Phytol. 2015, 206, 436–447. [Google Scholar] [CrossRef] [PubMed]
- Besnard, G.; Rubio de Casas, R. Single vs multiple independent olive domestications: The jury is (still) out. New Phytol. 2016, 209, 466–470. [Google Scholar] [CrossRef] [PubMed]
- Muzzalupo, I.; Vendramin, G.G.; Chiappetta, A. Genetic Biodiversity of Italian Olives (Olea europaea) Germplasm Analyzed by SSR Markers. Sci. World J. 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- D’Agostino, N.D.; Taranto, F.; Camposeo, S.; Mangini, G.; Fanelli, V.; Gadaleta, S.; Miazzi, M.M.; Pavan, S.; di Rienzo, V.; Sabetta, W.; et al. GBS-derived SNP catalogue unveiled wide genetic variability and geographical relationships of Italian olive cultivars. Sci. Rep. 2018, 26. [Google Scholar] [CrossRef]
- Di Rienzo, V.; Miazzi, M.M.; Fanelli, V.; Sabetta, W.; Montemurro, C. The preservation and characterization of Apulian olive germplasm biodiversity. Acta Hortic. 2018, 1199, 1–6. [Google Scholar] [CrossRef]
- Binetti, G.; Del Coco, L.; Ragone, R.; Zelasco, S.; Perri, E.; Montemurro, C.; Valentini, R.; Naso, D.; Fanizzi, F.P.; Schena, F.P. Cultivar classification of Apulian olive oils: Use of artificial neural networks for comparing NMR, NIR and merceological data. Food Chem. 2017, 219, 131–138. [Google Scholar] [CrossRef] [PubMed]
- Sabetta, W.; Miazzi, M.M.; di Rienzo, V.; Fanelli, V.; Pasqualone, A.; Montemurro, C. Development and application of protocols to certify the authenticity and the traceability of Apulian typical products in olive sector. Rivista Italiana Delle Sostanze Grasse 2017, 94, 37–43. [Google Scholar]
- Pasqualone, A.; Montemurro, C.; di Rienzo, V.; Summo, C.; Paradiso, V.M.; Caponio, F. Evolution and perspectives of cultivar identification and traceability from tree to oil and table olives by means of DNA markers. J. Sci. Food Agric. 2016, 96, 3642–3657. [Google Scholar] [CrossRef] [PubMed]
- di Rienzo, V.; Sion, S.; Taranto, F.; D’Agostino, N.; Montemurro, C.; Fanelli, V.; Sabetta, W.; Boucheffa, S.; Tamendjari, A.; Pasqualone, A.; et al. Genetic flow among olive populations within the Mediterranean basin. Peer J. 2018, 6, e5260. [Google Scholar] [CrossRef] [Green Version]
- Tanasijevic, L.; Todorovic, M.; Pereira, L.S.; Pizzigalli, C.; Lionello, P. Impacts of climate change on olive crop evapotranspiration and irrigation requirements in the Mediterranean region. Agric. Water Manag. 2014, 144, 54–68. [Google Scholar] [CrossRef]
- Ponti, L.; Gutierrez, A.P.; Ruti, P.M.; Dell’Aquila, A. Fine-scale ecological and economic assessment of climate change on olive in the Mediterranean Basin reveals winners and losers. Proc. Natl. Acad. Sci. USA 2014, 111, 5598–5603. [Google Scholar] [CrossRef] [Green Version]
- Martelli, G.P. The current status of the quick decline syndrome of olive in southern Italy. Phytoparasitica 2016, 44, 1–10. [Google Scholar] [CrossRef]
- Cornara, D.; Cavalieri, V.; Dongiovanni, C.; Altamura, G.; Palmisano, F.; Bosco, D.; Porcelli, F.; Almeida, R.P.P.; Saponari, M. Transmission of Xylella fastidiosa by naturally infected Philaenus spumarius (Hemiptera, Aphrophoridae) to different host plants. J. Appl. Entomol. 2017, 141, 80–87. [Google Scholar] [CrossRef]
- Giampetruzzi, A.; Morelli, M.; Saponari, M.; Loconsole, G.; Chiumenti, M.; Boscia, D.; Savino, V.N.; Martelli, G.P.; Saldarelli, P. Transcriptome profiling of two olive cultivars in response to infection by the CoDiRO strain of Xylella fastidiosa subsp. pauca. BMC Genom. 2016, 17, 475. [Google Scholar] [CrossRef]
- Saponari, M.; Boscia, D.; Altamura, G.; Loconsole, G.; Zicca, S.; D’Attoma, G.; Morelli, M.; Palmisano, F.; Saponari, A.; Tavano, D.; et al. Isolation and pathogenicity of Xylella fastidiosa associated to the olive quick decline syndrome in southern Italy. Sci. Rep. 2017, 7, 17723. [Google Scholar] [CrossRef] [PubMed]
- Bau, A.; Delbianco, A.; Stancanelli, G.; Tramontini, S. Susceptibility of Olea europaea L. varieties to Xylella fastidiosa subsp. pauca ST53: Systematic literature search up to 24 March 2017. EFSA J. 2017, 15, e04772. [Google Scholar] [CrossRef]
- Taranto, F.; Francese, G.; Di Dato, F.; D’Alessandro, A.; Greco, B.; Onofaro Sanajà, V.; Pentangelo, A.; Mennella, G.; Tripodi, P. Leaf Metabolic, Genetic, and Morphophysiological Profiles of Cultivated and Wild Rocket Salad (Eruca and Diplotaxis Spp.). J. Agric. Food Chem. 2016, 64, 5824–5836. [Google Scholar] [CrossRef] [PubMed]
- De Giovanni, C.; Pavan, S.; Taranto, F.; Di Rienzo, V.; Miazzi, M.M.; Marcotrigiano, A.R.; Mangini, G.; Ricciardi, L.; Lotti, C. Genetic variation of a global germplasm collection of chickpea (Cicer arietinum L.) including Italian accessions at risk of genetic erosion. Physiol. Mol. Biol. Plants 2017, 23, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Boucheffa, S.; Miazzi, M.M.; di Rienzo, V.; Mangini, G.; Fanelli, V.; Tamendjari, A.; Pignone, D.; Montemurro, C. The coexistence of oleaster and traditional varieties affects genetic diversity and population structure in Algerian olive (Olea europaea) germplasm. Genet. Resour. Crop Evol. 2017, 64, 379–390. [Google Scholar] [CrossRef]
- Mariotti, R.; Cultrera, N.G.M.; Mousavi, S.; Baglivo, F.; Rossi, M.; Albertini, E.; Alagna, F.; Perrotta, G.; Baldoni, L. Development, evaluation, and validation of new EST-SSR markers in olive (Olea europaea L.). Tree Genet. Genomes 2016, 12, 120. [Google Scholar] [CrossRef]
- Veloso, M.M.; Simões-Costa, M.C.; Carneiro, L.C.; Guimarães, J.B.; Mateus, C.; Fevereiro, P.; Candido, P.R. Olive Tree (Olea europaea L.) diversity in traditional small farms of Ficalho, Portugal. Diversity 2018, 10, 5. [Google Scholar] [CrossRef]
- Besnard, G.; Baradat, P.; Bervillé, A. Genetic relationships in the olive (Olea europaea L.) reflect multilocal selection of cultivars. Theor. Appl. Genet. 2001, 102, 251–258. [Google Scholar] [CrossRef]
- Besnard, G.; Baradat, P.; Breton, C.; Khadari, B.; Berville, A. Olive domestication from structure of oleasters and cultivars using nuclear RAPDs and mitochondrial RFLPs. Genet. Sel. Evol. 2001, 33, 251–268. [Google Scholar]
- Rao, R.; La Mura, M.; Corrado, G.; Ambrosino, O.; Foroni, I.; Perri, E.; Pugliano, G. Molecular diversity and genetic relationships of southern Italian olive cultivars as depicted by AFLP and morphological traits. J. Hortic. Sci. Biotechnol. 2009, 84, 261–266. [Google Scholar] [CrossRef]
- Montemurro, C.; Simeone, R.; Pasqualone, A.; Ferrara, E.; Blanco, A. Genetic relationships and cultivar identification among 112 olive accessions using AFLP and SSR markers. J. Hortic. Sci. Biotechnol. 2005, 80, 105–110. [Google Scholar] [CrossRef]
- Taranto, F.; D’Agostino, N.; Greco, B.; Cardi, T.; Tripodi, P. Genome-wide SNP discovery and population structure analysis in pepper (Capsicum annuum) using genotyping by sequencing. BMC Genom. 2016, 17, 943. [Google Scholar] [CrossRef]
- Pavan, S.; Marcotrigiano, A.R.; Ciani, E.; Mazzeo, R.; Zonno, V.; Ruggieri, V.; Lotti, C.; Ricciardi, L. Genotyping-by-sequencing of a melon (Cucumis melo L.) germplasm collection from a secondary center of diversity highlights patterns of genetic variation and genomic features of different gene pools. BMC Genom. 2017, 18, 59. [Google Scholar] [CrossRef] [PubMed]
- Taranto, F.; D’Agostino, N.; Fanelli, V.; di Rienzo, V.; Miazzi, M.M.; Pavan, S.; Zelasco, S.; Perri, E.; Montemurro, C. SNP diversity in an olive germplasm collection. Acta Hortic. 2018, 1199, 27–32. [Google Scholar] [CrossRef]
- Boucheffa, S.; Tamendjari, A.; Sanchez-Gimeno, A.C.; Rovellini, P.; Venturini, S.; di Rienzo, V.; Miazzi, M.M.; Montemurro, C. Diversity Assessment of Algerian Wild and Cultivated Olives (Olea europaea L.) by Molecular, Morphological, and Chemical Traits. Eur. J. Lipid Sci. Technol. 2019, 121. [Google Scholar] [CrossRef]
- Baldoni, L.; Cultrera, N.G.; Mariotti, R.; Ricciolini, C.; Arcioni, S.; Vendramin, G.G. A consensus list of microsatellite markers for olive genotyping. Mol. Breed. 2009, 24, 213–231. [Google Scholar] [CrossRef]
- Cipriani, G.; Marrazzo, M.T.; Marconi, R.; Cimato, A.; Testolin, R. Microsatellite markers isolated in olive (Olea europaea L.) are suitable for individual fingerprinting and reveal polymorphism within ancient cultivars. Theor. Appl. Genet. 2002, 104, 223–228. [Google Scholar] [CrossRef]
- Pasqualone, A.; Di Rienzo, V.; Nasti, R.; Blanco, A.; Gomes, T.; Montemurro, C. Traceability of PDO Italian table olives by means of microsatellite molecular markers. J. Agric. Food Chem. 2013, 61, 3068–3073. [Google Scholar] [CrossRef]
- Montemurro, C.; Miazzi, M.M.; Pasqualone, A.; Fanelli, V.; Sabetta, W.; Di Rienzo, V. Traceability of PDO olive oil “terra di Bari” using high resolution melting. J. Chem. 2015, 2015, 496986. [Google Scholar] [CrossRef]
- Pasqualone, A.; Rienzo, V.D.; Miazzi, M.M.; Fanelli, V.; Caponio, F.; Montemurro, C. High resolution melting analysis of DNA microsatellites in olive pastes and virgin olive oils obtained by talc addition. Eur. J. Lipid Sci. Technol. 2015, 117, 2044–2048. [Google Scholar] [CrossRef]
- Rugini, E.; Baldoni, L.; Muleo, R.; Sebastiani, L. The Olive Tree Genome; Springer: Berlin, Germany, 2016; Volume 978, pp. 3–319. [Google Scholar] [CrossRef]
- Dominguez-Garcia, M.C.; Laib, M.; de la Rosa, R.; Belaji, A. Characterisation and identification of olive cultivars from North-eastern Algeria using molecular markers. J. Hortic. Sci. Biotechnol. 2012, 87, 95–100. [Google Scholar] [CrossRef]
- Rekik, I.; Salimonti, A.; Kamoun, N.G.; Muzzalupo, I.; Lepais, O.; Gerber, S.; Perri, E.; Rebai, A. Characterization and identification of tunisian olive tree varieties by microsatellite markers. HortScience 2008, 43, 1371–1376. [Google Scholar] [CrossRef]
- Díez, C.M.; Moral, J.; Barranco, D.; Rallo, L. Genetic diversity and conservation of olive genetic resources. In Genetic Diversity and Erosion in Plants: Case Histories; Ahuja, M.R., Mohan Jain, S., Eds.; Sustainable development and biodiversity series; Springer: Basel, Switzerland, 2016; Volume 8, pp. 337–356. [Google Scholar] [CrossRef]
- Montemurro, C.; Dambruoso, G.; Bottalico, G.; Sabetta, W. Self-incompatibility assessment of some Italian olive genotypes (Olea europaea L.) and cross-derived seedling selection by SSR markers on seed endosperms. Front. Plant Sci. 2019, 451, 10. [Google Scholar] [CrossRef] [PubMed]
- Bracci, T.; Sebastiani, L.; Busconi, M.; Fogher, C.; Belaj, A.; Trujillo, I. SSR markers reveal the uniqueness of olive cultivars from the Italian region of Liguria. Sci Hortic. 2009, 122, 209–215. [Google Scholar] [CrossRef]
- D’Imperio, M.; Viscosi, V.; Scarano, M.T.; D’Andrea, M.; Zullo, B.A.; Pilla, F. Integration between molecular and morphological markers for the exploitation of olive germoplasm (Olea europaea). Sci. Hortic. (Amst.) 2011, 130, 229–240. [Google Scholar] [CrossRef]
- Marra, F.P.; Caruso, T.; Costa, F.; Di Vaio, C.; Mafrica, R.; Marchese, A. Genetic relationships, structure and parentage simulation among the olive tree (Olea europaea L. subsp. europaea) cultivated in Southern Italy revealed by SSR markers. Tree Genet. Genomes 2013, 9, 961–973. [Google Scholar] [CrossRef]
- Hauville, A. La répartition des variétés d’olivers en Algérie et ses conséquences pratiques. Bulletin de la Société desAgriculteurs d’ Álgérie 1953, 580, 1–8. [Google Scholar]
- Abdessemed, S.; Muzzalupo, I.; Benbouza, H. Assessment of genetic diversity among Algerian olive (Olea europaea L.) cultivars using SSR marker. Sci. Hortic. (Amst.) 2015, 192, 10–20. [Google Scholar] [CrossRef]
- Tubeileh, A.; Abdeen, M.; Al-Ibrahem, A. Morphological and productive aspects offour syrian olive cultivars. Acta Hortic. 2008, 791, 415–418. [Google Scholar] [CrossRef]
- Abdelhamid, S.; Grati-Kammoun, N.; Marra, F.; Caruso, T. Genetic similarity among Tunisian cultivated olive estimated through SSR markers. Sci. Agric. 2013, 70, 33–38. [Google Scholar] [CrossRef] [Green Version]
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 2010, 11, 94. [Google Scholar] [CrossRef] [PubMed]
- Taranto, F.; Nicolia, A.; Pavan, S.; De Vita, P.; D’Agostino, N. Biotechnological and digital revolution for climate-smart plant breeding. Agronomy 2018, 8, 277. [Google Scholar] [CrossRef]
- Wright, S. The genetical structure of populations. Ann. Eugen. 1949, 15, 323–354. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenALEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Botstein, D.; White, R.L.; Skolnick, M.; Davis, R.W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 1980, 32, 314–331. [Google Scholar] [PubMed]
- Kalinowski, S.T.; Taper, M.L.; Marshall, T.C. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 2007, 16, 1099–1106. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.; Ritland, K. Estimation of pairwise relatedness with molecular markers. Genetics 1999, 152, 1753–1766. [Google Scholar] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; von Holdt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence Limits on Phylogenies: An Approach Using the Bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]






| Locus | Na | Ne | Ho | He | PIC | F |
|---|---|---|---|---|---|---|
| DCA03 | 15 | 7.70 | 0.88 | 0.87 | 0.86 | −0.02 |
| DCA05 | 12 | 6.46 | 0.78 | 0.85 | 0.83 | 0.08 |
| EMOL | 17 | 3.49 | 0.32 | 0.71 | 0.92 | 0.56 |
| DCA18 | 32 | 10.90 | 0.75 | 0.91 | 0.72 | 0.17 |
| DCA09 | 23 | 13.63 | 0.84 | 0.93 | 0.86 | 0.1 |
| DCA15 | 20 | 3.64 | 0.42 | 0.73 | 0.9 | 0.43 |
| GAPU101 | 22 | 8.44 | 0.83 | 0.88 | 0.87 | 0.06 |
| DCA17 | 30 | 7.44 | 0.57 | 0.87 | 0.87 | 0.34 |
| EMO90 | 18 | 8.09 | 0.78 | 0.88 | 0.69 | 0.11 |
| Total | 189 | 69.78 | ||||
| Mean | 21 | 7.75 | 0.68 | 0.85 | 0.83 | 0.2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sion, S.; Taranto, F.; Montemurro, C.; Mangini, G.; Camposeo, S.; Falco, V.; Gallo, A.; Mita, G.; Saddoud Debbabi, O.; Ben Amar, F.; et al. Genetic Characterization of Apulian Olive Germplasm as Potential Source in New Breeding Programs. Plants 2019, 8, 268. https://doi.org/10.3390/plants8080268
Sion S, Taranto F, Montemurro C, Mangini G, Camposeo S, Falco V, Gallo A, Mita G, Saddoud Debbabi O, Ben Amar F, et al. Genetic Characterization of Apulian Olive Germplasm as Potential Source in New Breeding Programs. Plants. 2019; 8(8):268. https://doi.org/10.3390/plants8080268
Chicago/Turabian StyleSion, S., F. Taranto, C. Montemurro, G. Mangini, S. Camposeo, V. Falco, A. Gallo, G. Mita, O. Saddoud Debbabi, F. Ben Amar, and et al. 2019. "Genetic Characterization of Apulian Olive Germplasm as Potential Source in New Breeding Programs" Plants 8, no. 8: 268. https://doi.org/10.3390/plants8080268