1. Introduction
The pressure to increase and sustain food production has been felt for a long time. Tools have thus been developed to guarantee greater accuracy in selection. The currently used methods of selection have been enhanced by the achievements of molecular biology and statistical models, enabling identification of both the markers of individual traits resulting from the action of individual genes and those conditioned by many QTLs that explain the phenotypic traits to various extents [
1].
The DArT marker can be used in genomic selection (GS) [
2]. GS allows for plant selection based on the total pool of DNA markers for the selected statistical model. It reduces the need for phenotyping and shortens the culture cycle. Meuwissen first described this method; he examined the accuracy of genomic selection carried out using the DArT technique and compared this with phenotypic selection and with selection supported by molecular markers (marker assisted selection, MAS). Genomic selection proved to be 28% more accurate than traditional marker-assisted selection, though slightly less accurate than phenotypic selection. The results of his study demonstrate that GS can be used to increase the profitability of breeding [
3]. The method has been successfully used in barley [
4] and oats [
5], and also works well in improving the efficiency of breeding perennial species, such as eucalyptus (Eucalyptus L’Her) [
6].
Modern methods for identifying single nucleotide polymorphisms (SNPs) make use of next generation sequencing (NGS) methods. These refer to sequencing techniques developed in the twenty-first century that provide higher performance and throughput than the Sanger [
7] sequencing technique commonly used before. The most common NGS techniques are pyrosequencing 454 [
8], the Solex technique (Ilumina), the SOLiD platform (Applied Biosystems), the Polonator system (Dover/Harvard), and the HeliScope single molecule sequencer (Helicos). These technologies provide inexpensive whole-genome sequence readings through methods such as chromatin immunoprecipitation, mutation mapping, detection of polymorphisms, and detection of noncoding RNA sequences [
9]. Modern sequencing methods enable the identification of a large number of markers and also allow more accurate examination of many loci.
Modern genotyping technologies can also shed new light on the genetic basis of heterosis. The use of heterosis to increase and stabilize yield has become one of the major drivers of increased agricultural production over the last few decades. Despite the huge significance of heterosis and the growing tendency to use hybrid vigor even in inbred crops like bread wheat, the molecular and genetic mechanisms underlying this phenomenon have still not been fully explained [
10].
Song and Messing [
11] isolated a specific region of the genome of two crossed inbred corn lines, which were subsequently sequenced and mapped. They found that the size of this area and the presence of genes from a given gene family in it were significantly different. Genes that were present in one line were absent in the other, although phenotypic symptoms of their expression were visible in the other line. This is evidence that genes from the same gene family that produce similar phenotypic effects were located in different parts of the genome in each of the tested lines. According to Song and Mesing, heterosis can therefore be a consequence of differences in the structure of the genome, especially in the distribution and presence of certain genes from a given gene family in crossed inbred lines. Predicting the magnitude of the heterosis effect in hybrids based on molecular marker analysis has been widely discussed. According to the literature, there is regression of either hybrid performance or heterosis with increasing molecular genetic distance and estimation of correlations between these variables [
11,
12,
13,
14] or estimation of marker effects and marker associations with hybrid performance, heterosis, or specific combining ability [
12,
15].
The aim of this study was to identify single nucleotide polymorphism (SNP) and SilicoDArT markers associated with yield traits and to predict the heterosis effect for yield traits in maize (Zea mays L.). This topic was selected because the decreasing cost of next-generation sequencing means that these methods are beginning to be used in applied research to identify feature markers or even to select on the whole-genome level. This publication is one of a number to recently have suggested the possibility of using the latest molecular techniques (such as SNP and SilicoDArT) to select parental materials for heterosis crosses.
3. Discussion
Maize is a major crop species characterized by very high yield efficiency and versatility in utilizing the whole plant. Modern breeding programs focus on hybrid cultivars with the greatest heterosis effect, thanks to which it is possible to obtain much higher yields through appropriate selection of parental components.
Heterosis or hybrid vigor is a phenotypic result of gene interaction due to the effect of heterozygotes of hybrids in the F1 generation. According to the dominance hypothesis cited by Ruebenbauer, having many heterozygous genes causes an increase in hybrid performance due to the dominant alleles [
16]. The more homozygous alleles are complementary in the parental forms, the greater the effect of heterosis in the hybrid forms. Hence, the greatest theoretical heterosis can be expected when there is a large allele diversity of individual genes in the parent plants. Such diversity occurs when the crossed genotypes are less related and the genetic distance is greater. Progeny of genotypes with a large genetic distance should thus show a significant heterosis effect.
We used the SilicoDArT and SNP markers to assess genetic diversity. The DArTseq NGS analysis of the tested maize lines allowed us to identify 49,911 polymorphisms (33,452 SilicoDArT and 16,459 SNP). In total, 8192 of these markers (including 8189 SilicoDArT and three SNPs) were selected for GWAM. Of all the analyzed markers, 76 were selected as being significantly associated with at least six traits observed in 2013 and 2014 at both Łagiewniki and Smolice.
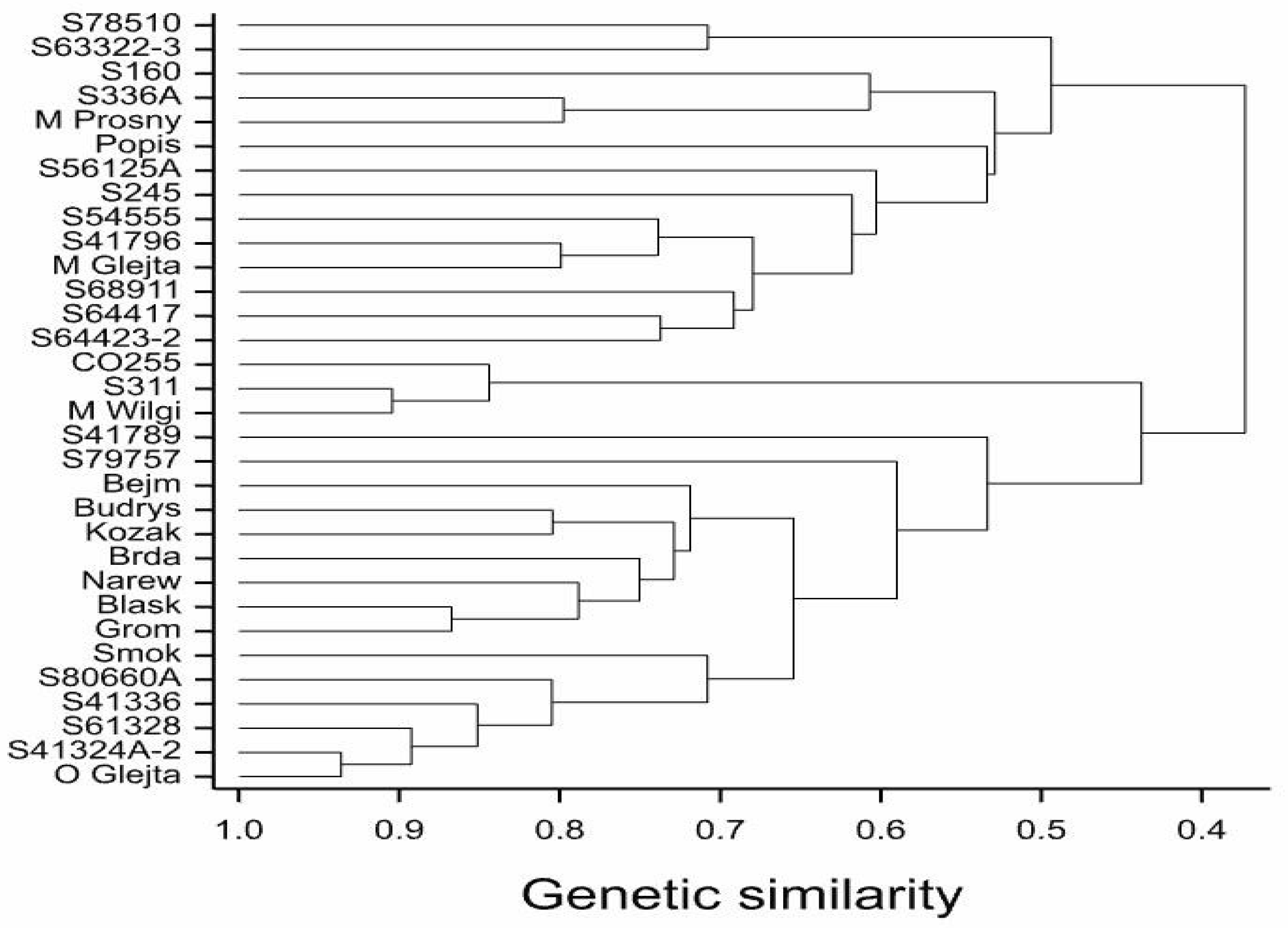
In the present study, the genetic distance between parental components, as determined by the SNP and SilicoDArT markers, reflected their degree of relationship and was significantly correlated with the heterosis effect observed in the majority of the yield structure features, as well as the yield itself. Genotypes grouped according to specific patterns are shown on the dendrogram, with the first group including all inbred lines, except for the S41324A-2, S160, and O Glejt hybrid lines. The second group consists of hybrid forms that have the same paternal components. The third group consists of hybrids (M Glejt, M Prosny, Budrys, and Popis) whose parental components were not related to each other, or which were only related to a small percentage.
As the results indicate, the parental components for heterosis crosses can be selected on the basis of genetic distance between the parental components, as determined using SNP and SilicoDArT markers, supported with information on the origin of the parental forms.
In this study, Narew, Popis, Kozak, M Glejt, and Grom were the hybrids that showed the highest significant heterosis effect for the majority of the yield structure traits, at both sites in both years. Importantly, the parental components of these hybrids (except for the Grom hybrid) were either not related to each other or else showed only a low degree of relationship due to origin (Narew: 4% relationship between parents; Popis: 0%; Kozak: 0%; and M Glejt: 13%). The similarity between parental components determined on the basis of SNP and SilicoDArT marker analysis did not exceed 33% (Narew: 18% similarity between parents; Popis: 26%; Kozak: 26%; and M Glejt: 33%). Many researchers attribute the dependence of the heterosis effect on the genetic distance of parental forms, taking into account their degree of relationship [
17,
18,
19,
20,
21]. In our own research, the genetic distance between parental components, as estimated with SNP and SilicoDArT markers, reflected their relationship and translated into the magnitude of the heterosis effect. We observed that the lower the similarity and the degree of relationship between parental components, the greater the effect of heterosis was in the hybrid forms.
In recent years, methods have been sought to allow initial selection of lines intended for heterosis crossing. The dependence of the heterosis effect on genetic distance, as determined using molecular markers, has been analyzed by many researchers in various species, including [
22], pepper [
23], cocoa [
24], barley or sunflower [
25]. Factors associated with the hybrid heterosis effect resulting from the crossing of inbred maize lines were discovered in 1992 [
26]. Research conducted using molecular RFLP markers on 148 inbred lines of maize supported the use of these markers when the breeding material was clustered into heterotic groups [
27]. The AFLP system was used to select parental components for maize heterosis crosses [
28]. Five primer pairs generated 56 polymorphic bands, allowing the degree of similarity to be determined, which was then correlated with the effect of heterosis. Shehata et al. [
29] demonstrated the usefulness of the SSR system in assessing the genetic distance of eight inbred maize lines. Berilli et al. [
30] studied the genetic distance between two maize populations (CYMMYT and Piranao), which was estimated using molecular ISSR markers. Thirteen primers generated as many as 140 products, of which 84.4% were polymorphic. The genotypes tested were divided into two main groups, which contained mainly individuals from a single population.
DArT technology also works as an efficient diagnostic tool for analyzing genetic diversity [
31]. DArT markers have been successfully used to study the genetic diversity and structure of Chinese common wheat (
Triticulum aestivum L). A total of 111 cultivars and breeding lines from northern China were examined, with the results providing information that allowed further selection of parental forms and the establishment of heterozygous materials for the needs of the Chinese wheat breeding program [
32]. The DArT method has found broad application in relationship analysis, such as in oats (
Avena sp.), where 134 cultivars were examined and groups corresponding to winter and spring forms were identified [
33]. However, research into 232 forms of the pigeon pea (
Cajanus cajan) showed a low degree of material differentiation. Of 696 DArT markers, only 64 turned out to be polymorphic, with the wild forms being the most diverse [
34].
Genome profiling in large hybrid populations currently offers unprecedented resolution for the dissection of loci and genes involved in heterotic expression. Huang et al. [
35] recently published a study in which an extensive population of 1495 elite hybrid rice varieties, along with their inbred parental lines, were subject to detailed genome-wide sequence analysis in order to investigate genomic effects on hybrid vigor for 38 agronomic traits. The resequenced genomes of all parental lines harbored around 1.3 million polymorphic SNP markers, which were subsequently used to study population genetic parameters and perform GWAS at an unprecedented resolution. This approach revealed heterozygous chromosome regions that contributed to trait expression in the F1 hybrids. Elucidation of the corresponding genomic effects on phenotypic traits demonstrated that the pyramiding of multiple loci facilitated the accumulation of many rare superior alleles with positive effects. In other words, dominance complementation contributes most to the heterosis effect in the hybrid rice production. A combination of forward and background selection using high-throughput genome screening tools [
36,
37] can thus significantly increase the breeding gain potential through the efficient exploitation of hybrid vigor.
The idea of genomic hybrid breeding, in which a genome-based prediction strategy using genomic sequence data is used to estimate the performance of the F1 progeny in hybrid breeding, was introduced in rice by Xu et al. [
38]. These authors used over 250,000 SNP markers generated by resequencing 210 parental inbred lines from a training set of 278 randomly selected hybrids; this study demonstrated the power of marker-directed estimation of F1 hybrid yields in rice. The top one hundred predicted hybrids, from a total of 21,945 possible combinations between the parental accessions, were estimated to exceed the overall average yield by 16%. This means there was a significant improvement in the average selection gains, compared to conventional breeding and accelerated hybrid rice production.
4. Materials and Methods
4.1. Plant Material
The plant material used for the research consisted of 19 inbred maize lines derived from a range of starting materials, and 13 hybrids resulting from their crossing. Maize lines and hybrids came from Hodowla Roślin Smolice (
Table 3).
4.2. Phenotyping
A two-year field experiment (2012, 2013) with inbred lines and hybrids was established on 10 m2 plots in a randomized block design in three replicates at two breeding stations owned by Plant Breeding Smolice, part of the Plant Breeding Acclimatization Institute Group, at Smolice (51°42′20.813′’N, 17°9′57.405′’E) and Łagiewniki (50°47′27′’N, 16°50′40′’E), Poland. One cob each was selected from ten plants of each replicate to perform biometric measurements on. Biometric measurements were carried out in the first half of November each year and included cob length (LC), cob diameter (DC), core length (LCO), core diameter (DCO), number of rows of grain (NRG), number of grains in a row (NGR), mass of grain from the cob (MGC), weight of one thousand grains (WTG), and Yield.
4.3. Genotyping and SilicoDArT and SNP Data Processing
The genotypic data for association mapping were derived from polymorphisms identified in DArT and candidate gene sequences.
4.4. DArT Sequences
Thirty-two genotypes were genotyped. The total genomic DNA extraction from the young leaves of the analyzed forms was performed using the GenElute Plant Mini Kit (Sigma-Aldrich, Darmstadt, Germany). DNA purity and concentration were determined spectrophotometrically (Thermo Scientific, Waltham, MA, USA), and the quality was determined electrophoretically in a 1% agarose gel. The concentration of all DNA samples was adjusted to 100 ng µl
–1. DArTseq analysis was performed at Diversity Arrays Technology, Australia. The GBS procedure involves several stages, which include preparation of DNA samples, digestion of genomic DNA with restriction enzymes, ligation of adapters, and independent creation of individual libraries and their final assembly. Next, the products are amplified, and the results are sequenced and analyzed. The detailed methodology is as follows: DNA samples were processed in digestion/ligation reactions principally as per Kilian et al. [
39], but replacing a single PstI-compatible adapter with two different adapters corresponding to two different restriction enzyme (RE) overhangs, and transferring the assay onto the sequencing platform, as described by Sansaloni et al. [
40]. The PstI-compatible adapter was designed to include the Illumina flowcell attachment sequence, sequencing primer sequence, and a “staggered” barcode region of varying length, similar to the sequence reported by Elshire et al. [
41]. The reverse adapter contained the flowcell attachment region and the NspI-compatible overhang sequence.
Only “mixed fragments” (PstI-NspI) were effectively amplified in 30 PCR cycles under the following reaction conditions: denaturation for 1 min at 94 °C, followed by 30 cycles of 20 sec at 94 °C, 30 sec at 58 °C, 45 sec at 72 °C, and final elongation for 7 min at 72 °C. After PCR, equimolar amounts of the amplification products from each sample of a 96-well microtiter plate were bulked and applied to c-Bot Illumina bridge PCR, before sequencing on an Illumina Hiseq2500. Single read sequencing was run for 77 cycles.
The sequences generated from each lane were processed using proprietary DArT analytical pipelines. In the primary pipeline, the fastq files were first processed to filter away poor quality sequences, applying more stringent selection criteria to the barcode region than to the rest of the sequence. In this way, assigning sequences to the specific samples carried in the “barcode split” step is very reliable. Approximately 2,500,000 (± 7%) sequences per barcode/sample were used in marker calling. Finally, identical sequences were collapsed into “fastqcall files”. These files were used in the secondary pipeline for DArT PL’s proprietary SNP and SilicoDArT calling algorithms (presence/absence of restriction fragments in representation; DArTsoft14). Only DArT sequences meeting the following criteria were selected for the association analysis: one SilicoDArT and SNP within a given sequence (69 nt), minor allele frequency (MAF) > 0.25, and < 10% missing observation fractions.
4.5. Statistical Analysis and Association Mapping
The Henderson method [
42] was used to construct a relationship matrix using the full pedigree information. Firstly, the normality of trait distribution was tested using the Shapiro–Wilk normality test [
43]. Relationships between the traits were estimated using correlation coefficients on the basis of means of genotypes for each location and year independently. The results were also examined using multivariate methods. The canonical variate analysis was applied in order to present a multitrait assessment of the similarity of the tested genotypes in a lower number of dimensions with the least possible loss of information [
44]. This allows the genotype variation to be illustrated in a graphic form in terms of all observed traits. The Mahalanobis distance was suggested as a measure of “polytrait” genotype similarity [
45], whose significance was verified by means of the critical D
α value referred to as the least significant distance [
46]. The Mahalanobis distances were calculated for species. The coefficients of genetic similarity (S) of the investigated lines were calculated using the Nei and Li [
47] formulas. The lines were grouped hierarchically using the unweighted pair group method of arithmetic means (UPGMA) based on the calculated coefficients. The relationship between lines was presented in the form of a dendrogram. Association mapping was performed using a method based on a mixed linear model with the population structure estimated by eigenanalysis (principal component analysis applied to all markers) and modeled by random effects [
48,
49]. All analyses were conducted in Genstat 18.2. The significance of associations between traits and SilicoDArT and SNP markers was assessed on the basis of
P-values corrected for multiple testing using the Benjamini–Hochberg method [
50].
4.6. Prediction of the Heterosis Effect
Heterosis effects for hybrids for each trait were estimated and tested by comparing a particular hybrid with the trait mean of both parents. Analysis was carried out using the GenStat 18 statistical package.