Abstract
Kalmegh (Andrographis paniculata (Burm. F.) Nees) is one of the most important medicinal plants and has been widely explored as traditional medicine. To exploit its natural genetic diversity and initiations of molecular breeding to develop novel cultivars or varieties, developments of genomic resources are essential. Four microsatellite-enriched genomic libraries—(CT)14, (GT)12, (AG)15 and (AAC)8—were constructed using the genomic DNA of A. paniculata. Initially, 183 recombinant colonies were screened for the presence of CT, GT, AG, and AAC microsatellite repeats, out of which 47 clones found positive for the desired simple sequence repeats (SSRs). It was found that few colonies had more than one desirable SSR. Thus, a sum of 67 SSRs were designed and synthesized for their validation among 42 A. paniculata accessions. Out of the 67 SSRs used for genotyping, only 41 were found to be polymorphic. The developed set of g-SSR markers showed substantial genetic variability among the selected A. paniculata accessions, with an average polymorphic information content (PIC) value of 0.32. Neighbor-joining tree analysis, population structure analysis, analysis of molecular variance (AMOVA), and principal coordinate analysis (PCoA) illustrated the considerable genetic diversity among them. The novel g-SSR markers developed in the present study could be important genomic resources for future applications in A. paniculata.
1. Introduction
The Kalmegh is an important medicinal crop species that is botanically known as Andrographis paniculate (Burm. F.) Nees and belongs to the family Acanthaceae [1]. This is an annual herb, about a meter in height, and bitter in taste, like Neem [2]; thus, Kalmegh is often called the king of bitter plants [3]. The plant is diploid (2n = 2x = 50) and found in both cultivated and wild forms in India [4]. A. paniculata is widely distributed in Asian countries like India, Sri Lanka, and China [5]. Andrographolide is one of the major bitter-tasting secondary metabolites derived from Kalmegh, a major bioactive substance responsible for the therapeutic interest [6]. This crop species is widely used as a traditional medicine in different parts of the world due to its versatile biological properties like immune-stimulatory [7], hepatoprotection [8], antibacterial [9], antimalarial [10], antithrombotic [11], antitumor [12], and anti-inflammatory [13]. Thus, this crop species has been used to treat various human diseases such as diabetes, hepatitis, leprosy, HIV, bronchitis, hypertension, cancer, and kidney disorders [14]. Currently, Kalmegh has been declared by the Indian National Medicinal Plants Board to be one of the most prioritized plant species among medicinal crop species for the exploitation of its potential use in human disease control and therapeutics [15].
Harnessing the genetic variability and development of the superior Kalmegh varieties with enhanced medicinal value is one of the prime objectives among researchers. Hence, genetic characterization of germplasm is one of the essential and primary steps in crop breeding programs. An in-depth characterization of germplasm allows effective selection of diverse parents in varietal improvement, besides helping efficient germplasm management in any crop species. Since distinguishing the genotypes based on morphological traits is time-consuming [16], the use of molecular markers can overcome its limitations. In the past, molecular markers like RAPD (randomly amplified polymorphic DNA) [3,4,17], AFLP (amplified fragment length polymorphism) [17], ISSR (inter simple sequence repeats), SCoT (start codon targeted polymorphism), and CBDP (CAAT box-derived polymorphism) [4] have been used for the genetic characterization of A. paniculate accessions. However, these dominant marker systems have low reproducibility and low consistency [18], which is a significant impediment for their further utilization in A. paniculate genomics and molecular breeding. Moreover, these dominant marker systems, namely, microsatellite or simple sequence repeat (SSR) markers, are one of the most preferred marker systems in studies of plant breeding and genetics due to the abundance in genomes, codominant natures, high reproducibility, multiallelic traits, and high transferability across the species [19]. Only the plastid genome sequence of A. paniculata has been carried out, and the whole genome of this crop is still not sequenced. It is, therefore, essential to develop microsatellite markers in A. paniculata.
Several methods have been employed to develop SSR markers in different plant species, and the microsatellite enrichment method is considered one of the most robust, reproducible, and cost-effective techniques [20]. This technique has been exploited for the development of SSR markers in several plant species including medicinal crops viz., Paeonia lactiflora [21], Centella asiatica [22], Tinospora cordifolia [23], and Bauhinia strychnifolia [24]. Therefore, the development of novel SSR markers was undertaken in A. paniculata using the microsatellite enrichment technique. The efficacy and informativeness of the developed SSRs were validated through genetic diversity studies among the A. paniculata accessions collected from different Indian states and maintained at ICAR-NBPGR, New Delhi, in the National Gene Bank.
2. Materials and Methods
2.1. Plant Materials
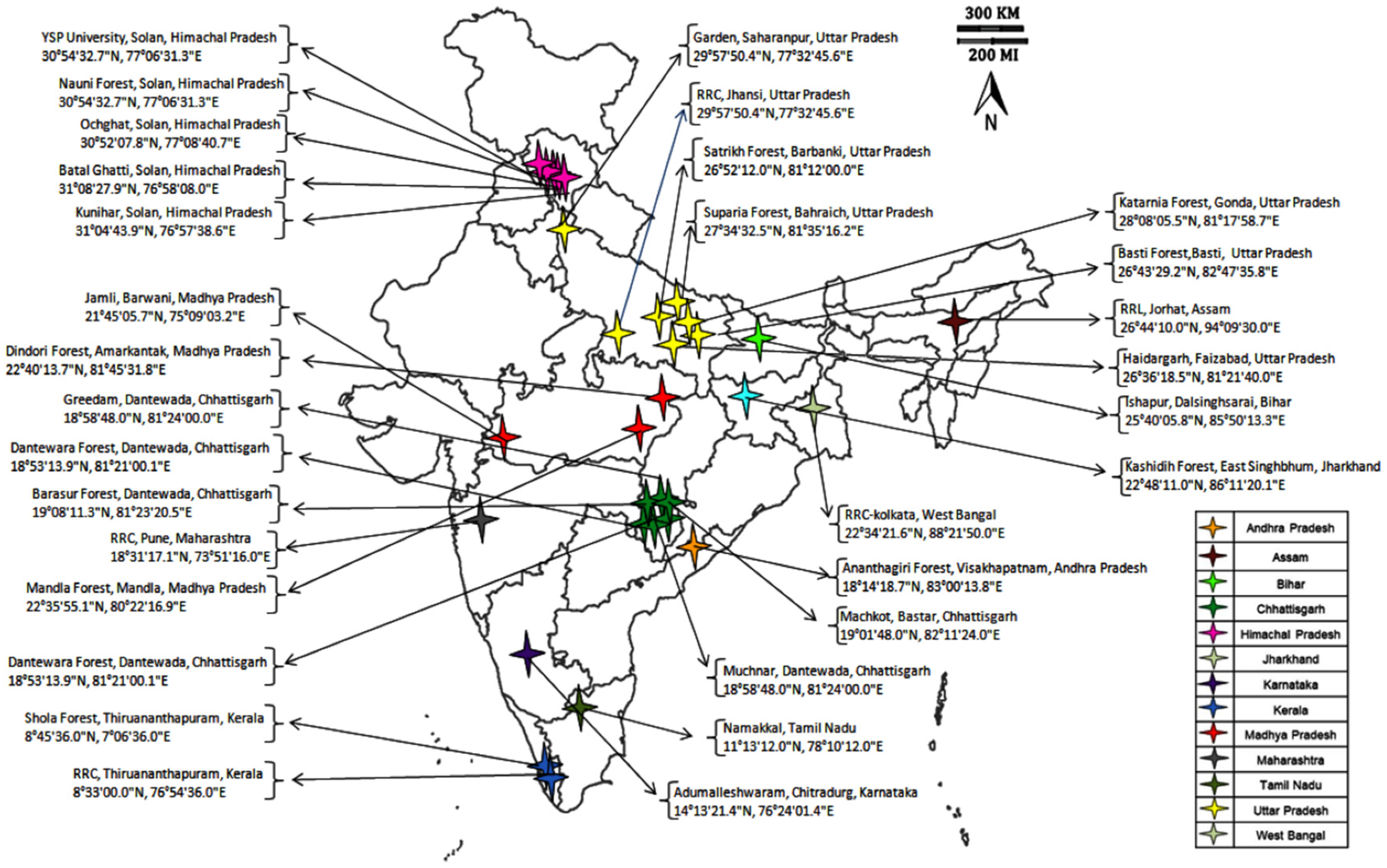
Seeds of 42 A. paniculata accessions were collected from the National Gene Bank ICAR-NBPGR, New Delhi, which were earlier collected from different geographical regions of Indian states (Figure 1). The collected seeds were sown in the experimental field at Issapur Research Farm (situated at 28.24 N latitude and 76.50 E longitude and an elevation of 190.7 m above sea level). The details on the A. paniculata accessions and their collection places are given in Table 1.
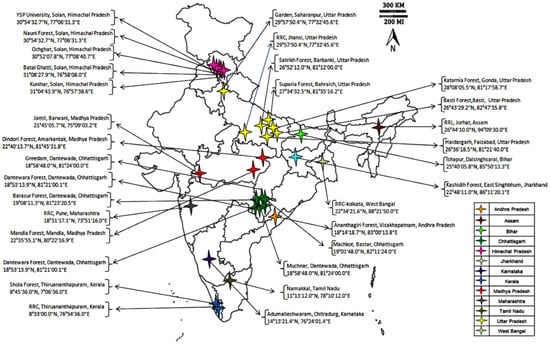
Figure 1.
Andrographis paniculata accessions collected from different geographical regions of India.

Table 1.
The details of Andrographis paniculata accessions studied.
2.2. Plant Genomic DNA (gDNA) Isolation
The young and healthy leaves of each accession were collected at 45 days after sowing, snap-frozen, and stored at −80 °C for further use. The genomic DNA was isolated, following the CTAB method described by Doyle and Doyle [25], with minor alterations. Since A. paniculata is rich in phenolic compounds, 3% polyvinylpyrrolidone (PVP) was also used to reduce the phenolics and facilitate quality gDNA extraction. To eliminate RNA contamination, 2.5 U of RNaseA enzyme (Himedia, West Chester, PA, USA) was used. The DNA quality was evaluated on 0.8% agarose gel, and its concentration was determined using a NanoDrop instrument (Thermo Fisher Scientific, Waltham, MA, USA).
2.3. Construction of the Microsatellite-Enriched Library
The microsatellite-enriched genomic libraries in A. paniculata were developed using the altered biotin-capture method, as suggested by Fischer and Bachmann [26]. The gDNA of A. paniculata accession IC111291 (1000 ng) was digested using the restriction enzyme Sau3A1 (New England Bio Labs, Knowl Piece Wilbury Way Hitchin UK) by incubation at 37 °C for 2 h and, later, inactivating it at 65 °C for 20 min. The restriction digestion of gDNA and the adapter-ligation of DNA were done as suggested by Bloor et al. [27].
SSR-containing DNA fragments were acquired by hybridization reaction of an adaptor-attached DNA fragment with prewashed (1X washing buffer and 2X washing buffer, respectively) streptavidin-coated magnetic beads and 3′-biotinylated oligonucleotide probes ((CT)14, (GT)12, (AG)15 and (AAC)8) at 60 °C for 30 min, with frequent shaking at 5 min intervals in 6X SSC buffer(Saline Sodium Citrate buffer). After the hybridization reaction, the magnetic beads were separated using the magnetic stand and the incubation of hybridization products in 2X SSC and 1XSSC, respectively, followed by final boiling at 95 °C for 15 min in TE buffer (Tris-EDTA). The concentration of enriched DNA was enhanced by performing a PCR reaction [27]. Thereafter, the PCR-amplified products were ligated into pCR 2.1 Cloning Vector (Thermo Fisher Scientific, Waltham, MA, USA) overnight at 16 °C and transformed into E. coli (Escherichia coli) DH5α competent cells. As per the X-gal/IPTG (5-Bromo-4-chloro-3-indolyl β-D-galactopyranoside (X-gal)/ Isopropyl-β-D-thiogalactoside (IPTG))selection method, a total of 183 white colonies were screened, from which 119 positive clones were selected while performing colony PCR (using M13 universal primer). The plasmid DNA from selected positive clones was extracted using a plasmid isolation kit (Zymo Research, Irvine, California, USA). The plasmids were subsequently sequenced, along with M13 primers, using Sanger’s dideoxy sequencing approach (Macrogen Inc., Seoul, Korea).
2.4. SSR Finding and Primer Designing
After the trimming of vector sequences, the sequencing results of positive clones were searched for desirable microsatellite repeats using an online SSR finder tool (http://www.csufresno.edu/ssrfinder/). The microsatellite primer pairs were designed based on the sequences flanking the SSR motifs using an online tool, Primer 3.0 input version 0.4.0 (http://bioinfo.ut.ee/primer3-0.4.0/). Finally, primer pairs were designed in the range of 18–25 nucleotides having an amplicon size ranging from 100 to 500 base pairs.
2.5. Polymerase Chain Reaction
A set of 67 developed SSR primers were selected for genetic diversity analysis, and these primers were amplified on 42 A. paniculata accessions, out of which 41 were found reproducible and polymorphic. The gDNA of selected 42 accessions was isolated, and its final working concentrations were kept at 10 ng/µL. The PCR reaction was performed in the total volume of 25 µL containing 7 µL gDNA (70 ng) as a template, 2.5 µL of 10X DreamTaq buffer, 3 µL of 2.5 mM MgCl2, 2.5 µL of 2.5 mM dNTPs, 0.8 µL of each primer (10 nmol), and 0.4 µL of DreamTaq DNA polymerase enzyme (Thermo Scientific, Waltham, MA, USA), with 8.8 µL Milli-Q water added to make the final volume. PCR amplification was performed in a thermocycler (Gstorm, Essex, England) using following the PCR cycle: initial denaturation at 94 °C for 4 min, followed by 36 cycles of denaturation at 94 °C for the 30 s, annealing temperature (standardize by gradient PCR) for 45 s, extension at 72 °C for 2 min, and a final extension at 72 °C for 10 min. The PCR products were checked on 4% metaphor agarose gel (Lonza, Rockland, ME, USA) for 4 h at a constant supply of 120 V, and gel images were captured using a gel documentation system (Alpha Imager®, Bengaluru, Karnataka, India).
2.6. Data Scoring and Statistical Analyses
The amplified PCR products of each primer pair among the A. paniculata accessions were scored using PyElph 1.4 [28]. The genetic diversity statistics viz. the dominant allele frequency, gene diversity, heterozygosity, and polymorphic information content (PIC) were calculated using Power Marker 3.5 [29]. An unrooted neighbor-joining (N-J) tree was generated, and the genetic distances between the A. paniculata accessions were also estimated using Power Marker 3.5 software [29]. Model-based population structure analysis was performed using STRUCTURE software version 2.3.4 [30]; the software was run multiple times by setting k (the number of populations) from 2 to 10, the length of burn-in period and number Markov Chain Monte Carlo (MCMC) replications were set at 100,000 for each run for all 42 genotypes to evaluate the number of populations [31]. An online tool, Structure Harvester (http://taylor0.biology.ucla.edu), was used to calculate the most probable genetic population groups of the studied A. paniculate accessions. Principal coordinate analysis (PCoA), analysis of molecular variance (AMOVA), and the Mantel test were done using the program GenAlEx 6.5 [32].
3. Results and Discussion
3.1. Development of SSR Markers from Enriched Genomic Libraries
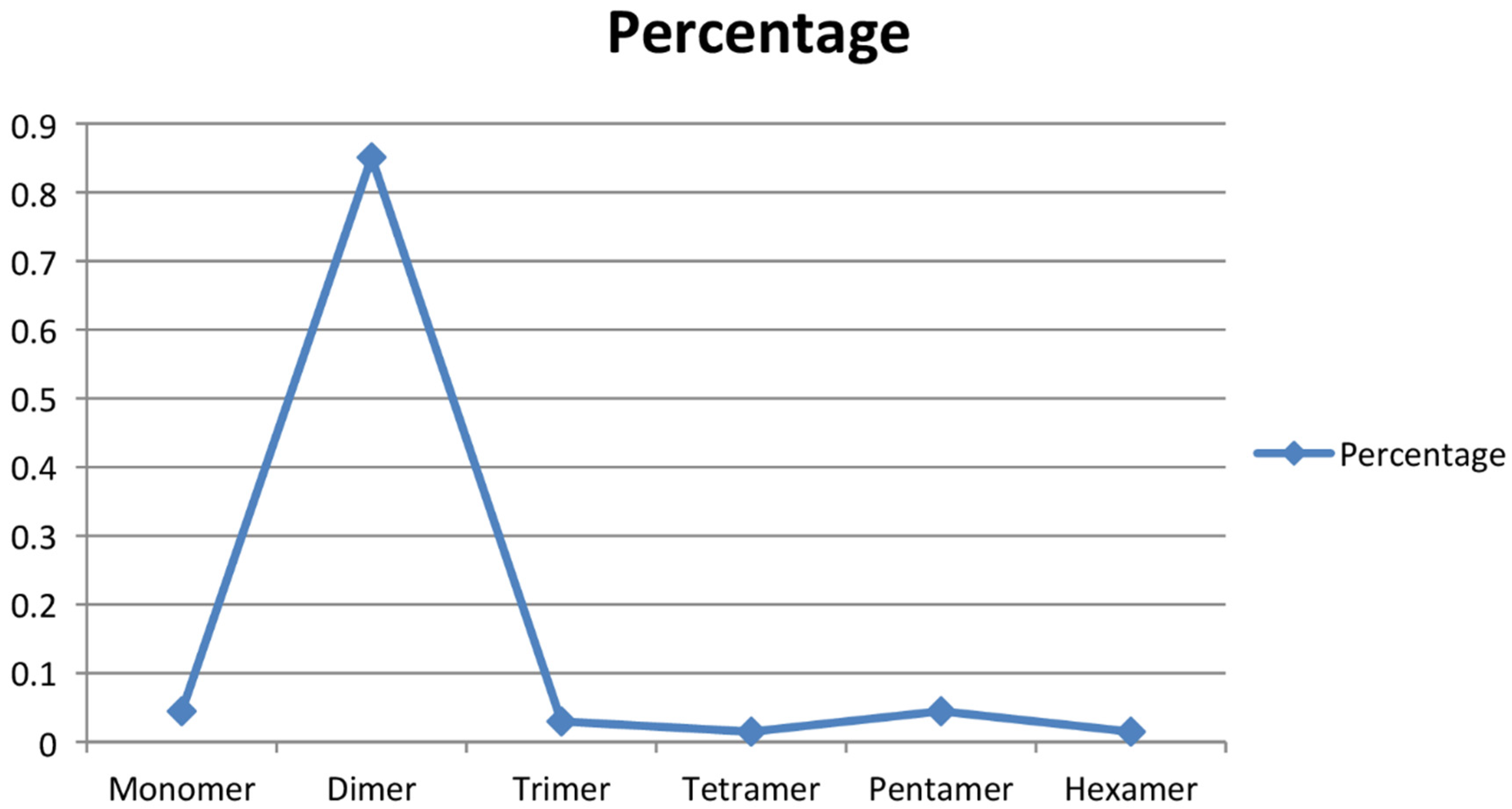
To obtain the microsatellite-enriched libraries, the genomic DNA of A. paniculata (IC 111291) was digested with restriction enzyme Sau3A1, which was further enriched with four types of 3′ biotinylated oligonucleotide probes ((CT)14, (GT)12, (AG)15 and (AAC)8). Altogether, 183 recombinant colonies were screened for the presence of CT, GT, AG, and AAC repeats, of which 119 were confirmed as positive clones (65%) through colony PCR and submitted for Sanger sequencing. The SSR finder tool was used to identify the perfect SSR markers, and 47 positive clones (39%) with perfect microsatellite repeats were identified. It was noticed that a few positive clones had more than one microsatellite repeat, and thus, a total of 67 primer pairs were developed (Table 2). The developed and synthesized microsatellite markers had motif-length groups, varying from monomer to hexamer, and their occurrence percentage varied from 1.49 to 85.07 (Figure 2). The tetramer and hexamer motif-length groups had a 1.49% occurrence; trimer had 2.98%, monomer and pentamer had 4.47%, while the dimer motif-length group had a maximum occurrence of 85.07%. Earlier, Wee et al. [33] sequenced 192 clones, and 102 colonies were obtained with desirable SSRs. Furthermore, Kaliswamy et al. [34] also reported that di- and trinucleotide repeats had more occurrences in the Acanthaceae family, which is similar to our findings. In addition to that, Lagercrantz et al. [35] and La Rota et al. [36] noticed that GA/CT microsatellite motifs are more abundant than the CA/GT motif in the plant species, which is similar to the present investigation. Marker-assisted breeding essentially requires a robust and informative marker system in the crop of interest [37]. Microsatellite markers are one of the choicest marker systems in molecular breeding of crop species due to its versatile applications in crop genetics and breeding, including cultivar identification [38], genetic diversity assessment [39], genetic mapping [40], gene tagging [41], gene flow [42], and molecular evolution studies [43] on plant species. In A. paniculate, the availability of microsatellite markers is lacking, which is a major limitation for its marker-assisted breeding. The screening of microsatellite-enriched libraries and the sequencing of microsatellite-positive clones are effective methods for the development of SSR markers [44].

Table 2.
Details of 67 novel genomic simple sequence repeat (SSR) loci developed through microsatellite-enriched genomic libraries.
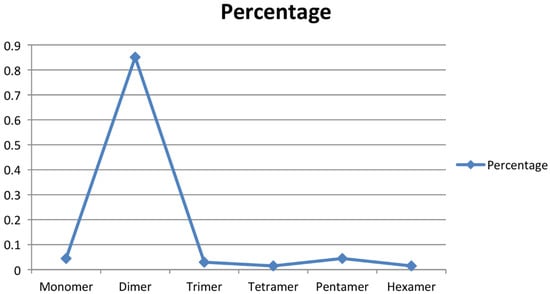
Figure 2.
SSR motif-length groups and their percentage occurrence.
3.2. Validation of g-SSR Loci and Genetic Diversity Statistics
A genetic diversity study among 42 A. paniculata accessions was performed using the 67 genomic SSR loci developed from four microsatellite-enriched libraries. The developed g-SSRs were screened for their amplification among the A. paniculata accessions, out of which 41 SSRs were found to be polymorphic (Table 3). These SSRs had substantial variations in allele number, which ranged from 2 to 8 with an average of 3.95 alleles per locus, and allele sizes, which ranged from 100 to 870 bp (Figures S4–S6). Similarly, Geng et al. [45] also recorded a range in the number of alleles, from 2 to 8, in Acanthus ilicifolius, which is congruent with our results. The PIC value varied from 0.09 for primer Ando4-36-2 to 0.38 for primer Ando5-29, with an average of 0.32. The observed heterozygosity was calculated as 0.00 for several markers, and the highest value was 0.21 for the marker Ando5-12-1, with a mean value of 0.02. Gene diversity (expected heterozygosity) ranged from 0.10 (Ando4-36-2) to 0.50 (Ando5-29, Ando5-26-2, Ando5-14-2, Ando4-27-2, Ando4-26, and Ando2-31-2), with a mean value of 0.40. Similarly, Geng et al. [45] calculated the observed and expected heterozygosity in Acanthus ilicifolius using SSR markers, which ranged from 0.200 to 0.875 and 0.227 to 0.798, respectively. Furthermore, Suárez-Montes et al. [46] also calculated the observed and expected heterozygosity values among the Aphelandra aurantiaca genotypes, ranging from 0.22 to 0.96 and 0.20 to 0.87, respectively, which is higher than the present study. The present investigation also deciphered large differences between the observed and expected heterozygosity, which indicates that the selected population of A. paniculata deviates from Hardy Weinberg’s equilibrium, which might be due to inbreeding, population bottleneck, or random genetic drift [17,45].

Table 3.
Details of allele number, major allele frequency, gene diversity, heterozygosity, and PIC values of developed g-SSRs.
3.3. Cluster Analysis
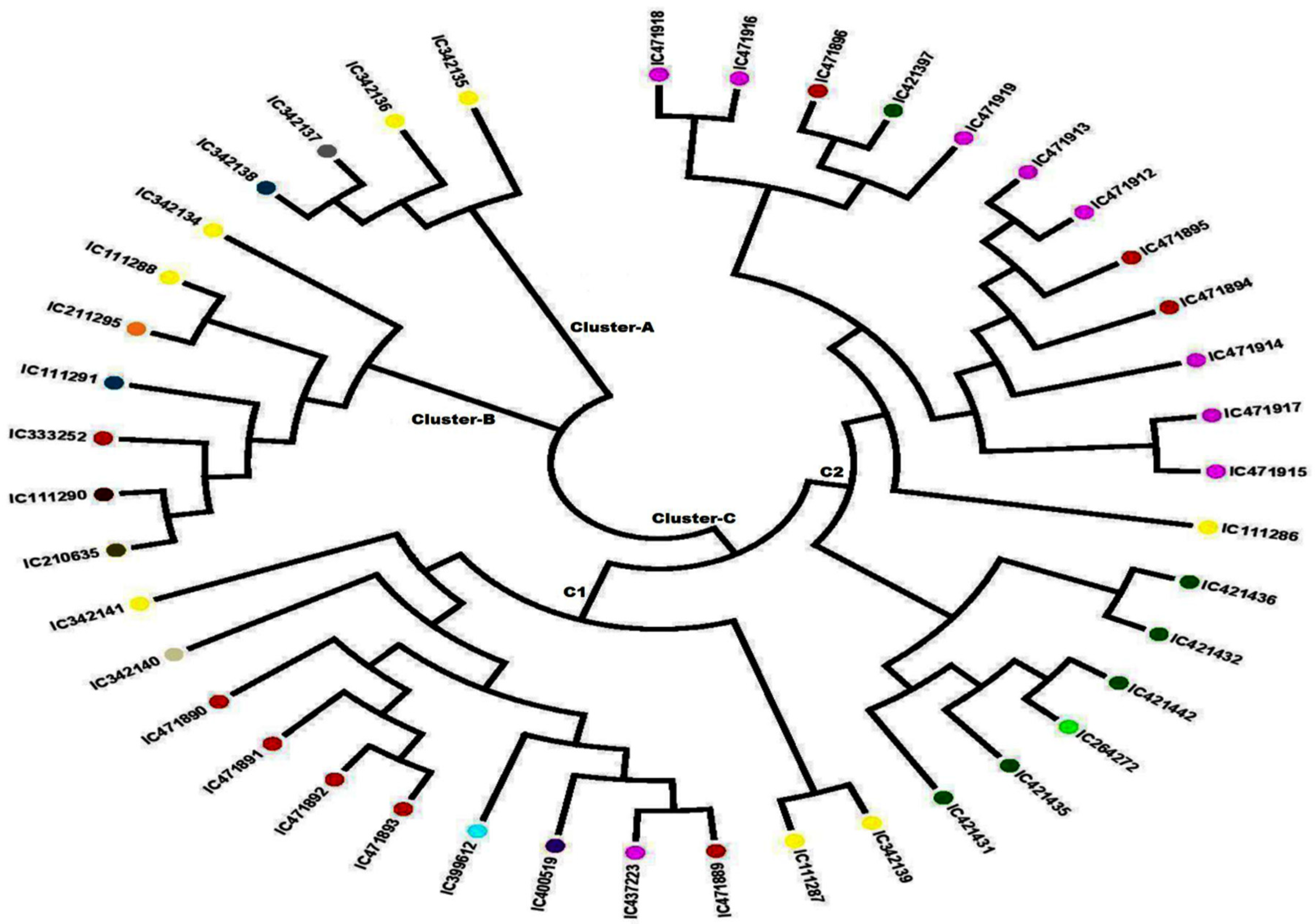
The unrooted N-J tree was constructed based on 41 developed SSR loci, which clustered all the 42 A. paniculata accessions into three major clusters (Figure 3). Earlier, Wijarat et al. [17] clustered 58 A. paniculata accessions into two major clusters using SSR markers that are lower than our findings. The genetic distance between the A. paniculata accessions ranged from 0.010–0.810, with an average of 0.400. The minimum genetic distance (0.010) was estimated between accessions IC 111291 & IC 211295 and IC 412436 & IC 421432, while the maximum (0.810) was between IC 471917 & IC 471891. Cluster A contained four accessions of A. paniculata, out of which two samples were from Uttar Pradesh, one each from Kerala, and one from Maharashtra. Cluster B constituted seven individuals of A. paniculata, out of which two were from Uttar Pradesh and one sample each from Andhra Pradesh, Kerala, Madhya Pradesh, Assam, and Tamil Nadu. Cluster C was further divided into two subclusters; subcluster C-1 contained 12 individuals, while C-2 had 19 individuals. Subcluster C-1 showed a tight grouping of four samples from Madhya Pradesh (IC 471890, IC 471891, IC 471892, and IC 471893) and two samples from Uttar Pradesh (IC 111287 and IC 342139). In subcluster C-2, there was a close grouping of two genotypes collected from Chhattisgarh (IC 421436 and IC 421432), and three tight groups from Himachal Pradesh were observed (Tight Group 1—IC 471,915 and IC 471917, Tight Group 2—IC 471912 and IC 471913, Tight Group 3—IC 471916 and IC 471918). Furthermore, A. paniculate accessions viz. IC 471890, IC 471891, IC 471892, and IC 471893 were in Cluster C-1 while IC 421431, IC 421435, IC 264272, IC 421442, IC 421432 and IC 421436 were in Cluster C2, grouped according to their natural habitat, which is possibly due to less human intervention in their natural habitats. Thus, a few of the A. paniculate accessions were tightly grouped according to their habitat and agro-geographical regions, while most of the accessions did not group according to their habitat and agro-geographical regions, which might be due to gene flow in the form of either gamete or genotype.
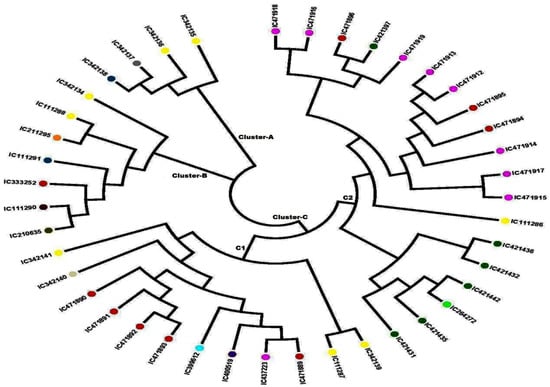
Figure 3.
Neighbor-joining (N-J) phylogenetic tree showing the grouping of 42 accessions of Andrographis paniculata based on the data of 41 SSR markers.
3.4. Population Structure
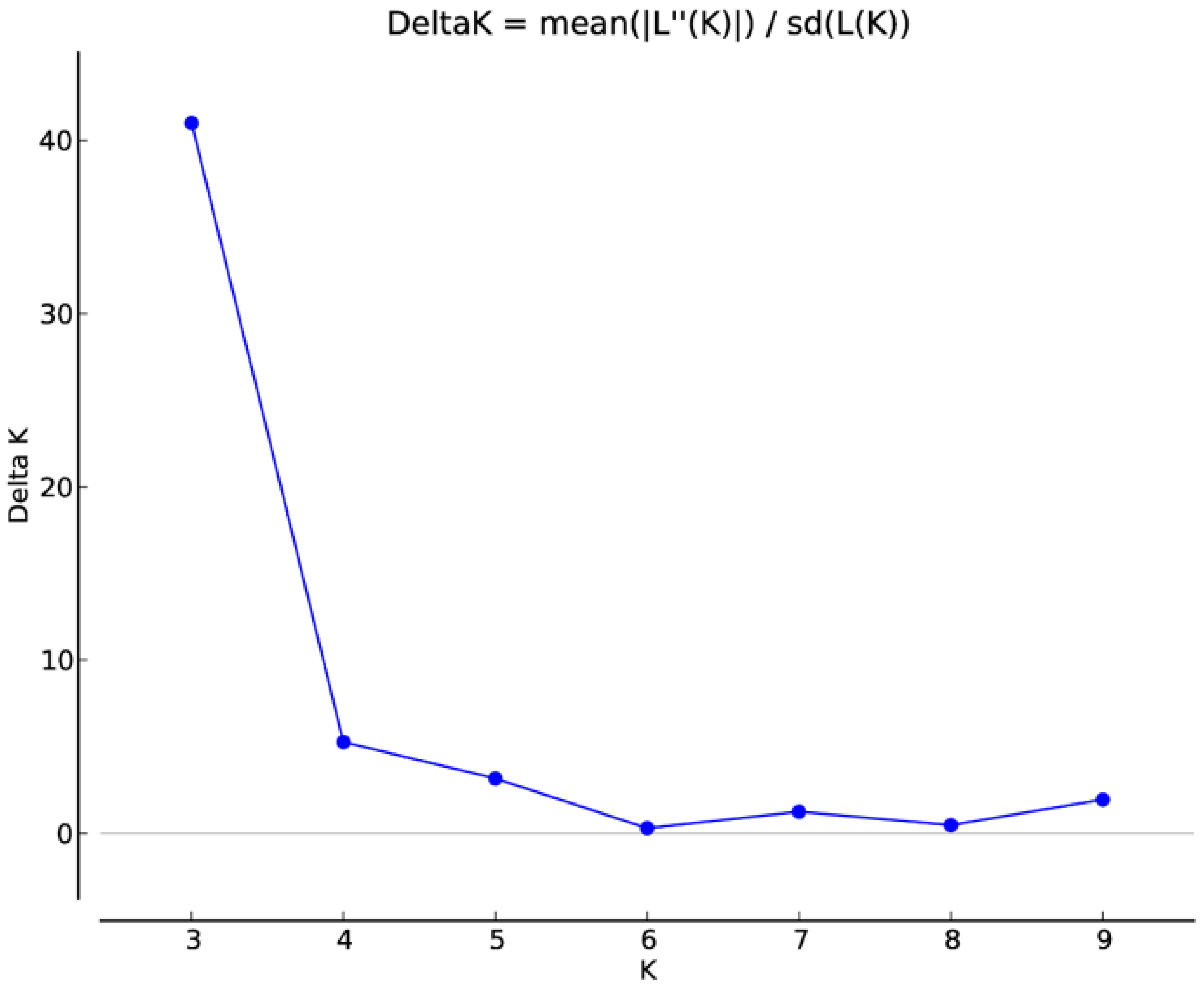
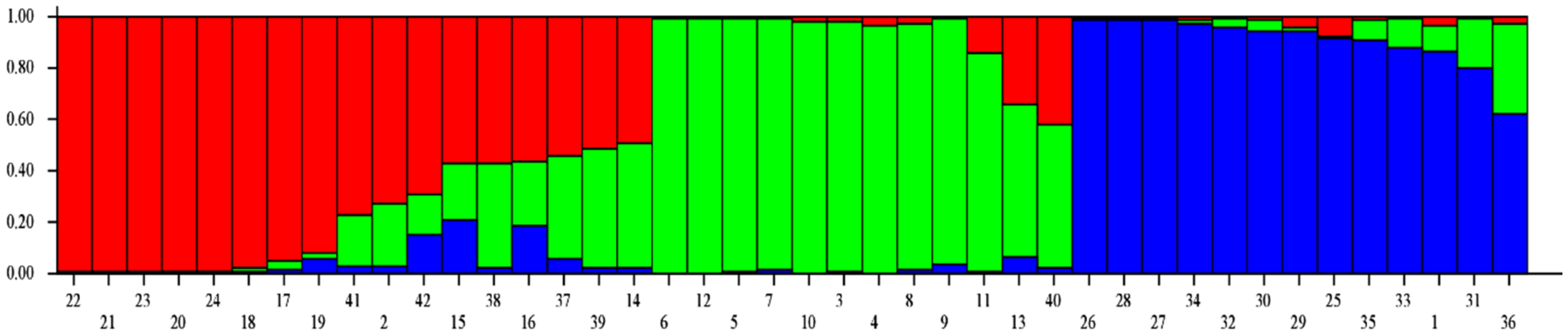
Model-based population structure analysis was utilized to rebuild the genetic relationship among 42 A. paniculata accessions using 41 developed SSR markers. Structure Harvester identified three genetic populations in the present set of A. paniculate accessions (Figure 4, Figure 5, Figures S1 and S2). The individuals with a probability score of more than 0.80 are considered genetically pure accession, while a score of <0.80, as an admixture accession. Population I showed eight pure accessions (IC471891, IC 471890, IC 471892, IC 471889, IC 471893, IC 400519, IC 399612, IC 437223) and nine admixed accessions (IC 421442, IC 111287, IC 264272, IC 342140, IC 421432, IC 342141, IC 421431, IC 421435, IC 342139). Population II showed ten pure accessions (IC 210635, IC 342137, IC 111291, IC 211295, IC 342135, IC 111288, IC 111290, IC 333252, IC 342134, IC 342136) and two admixed accessions (IC 342138, IC 421436). Population III showed twelve pure accessions (IC 471895, IC 471913, IC 471912, IC 471919, IC 471917, IC 471915, IC 471914, IC 471894, IC 471896, IC 471918, IC 111286, IC 471916) and one admixed accession (IC 421397). The genetic population differentiation of plant species is the consequence of various processes such as mating strategies, selection, mutations, and gene flow [47]. The genetic population differentiation among the A. paniculata accessions might be due to their mating behavior since the crop is self-pollinated and up to 4% cross-pollination occurs through insects [4,17]. The mean Fst value of Population I, Population II, and Population III were 0.4847, 0.5563, and 0.5090, respectively, and the mean alpha value was 0.1075 (Table S2). The allele-frequency divergence between Population I and Population II, Populations II and III, and Populations I and III were 0.2288, 0.1639, and 0.2277, respectively (Table S3). The population structure study indicated the genetic differentiation of A. paniculata accessions, which amply suggested that the developed gSSR markers were suitable for population structure studies.
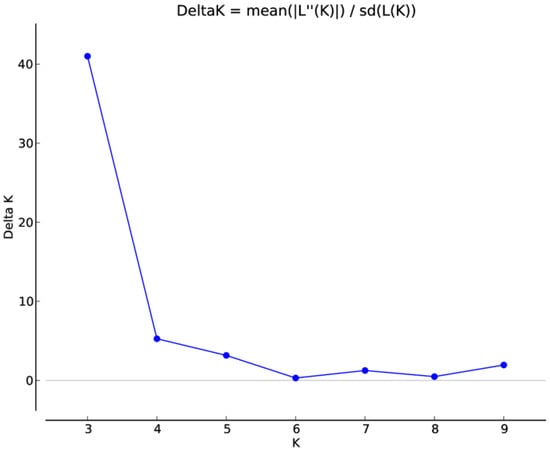
Figure 4.
Estimation of the number of populations using LnP(D)-derived Delta k from 2 to 10 using SSR data.
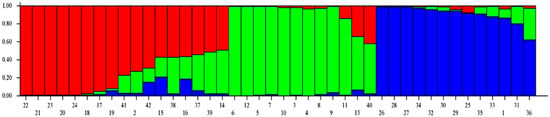
Figure 5.
Model-based population structure analysis of 42 accessions of Andrographis paniculata.
3.5. AMOVA, PCoA, and Mantel Test
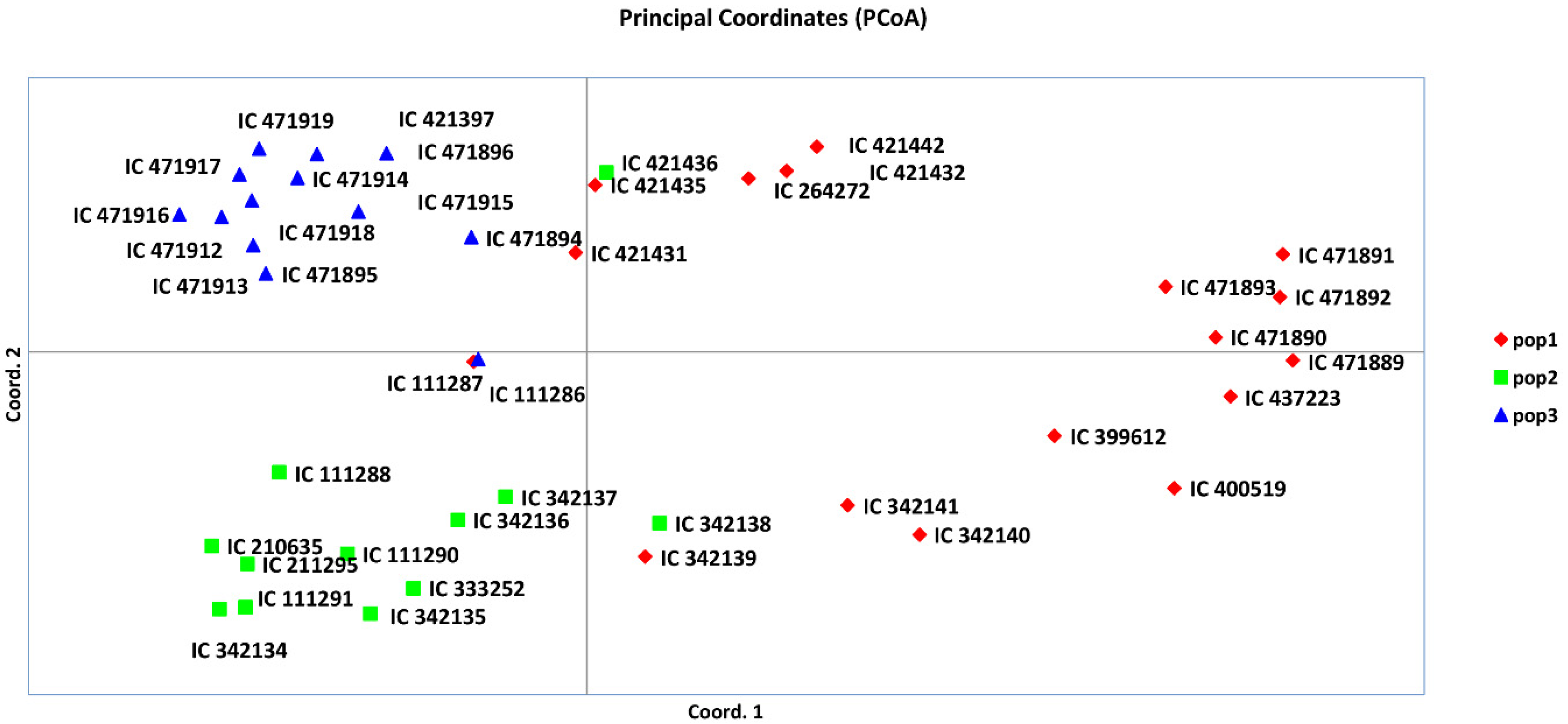
An analysis of molecular variance (AMOVA) was undertaken using the three genetic population groups of A. paniculate, as deciphered by model-based population structure analysis. The AMOVA illustrated 22% variance among the populations, with 77% variance among the individuals and 1% variance within the individuals of the populations (Table 4 and Figure S3). The first three principal coordinate analyses (PCoA) explained 34.88% cumulative variance, whereas the first, second and third axes explained 14.16%, 11.81%, and 8.91% of genetic variation, respectively (Table S4). Furthermore, the grouping of the A. paniculata accessions is depicted in three colors on the coordinates, supplementing the results of the model-based population structure analysis (Figure 6). AMOVA and PCoA explained the substantial genetic diversity among the A. paniculate accessions. Furthermore, the Mantel test was performed to obtain the correlation between genetic distance and geographical distance of A. paniculata accessions. Overall, a correlation coefficient with a low value (Rxy = 0.046) was observed (Table S5, Figure S7), indicating very little correlation between the genetic and geographical distances of A. paniculata accessions [48]. This might be due to the gene flow of A. paniculata accessions, in the form of either genotype or gamete.

Table 4.
Summary of analysis of molecular variance of 41 genomic SSRs among 42 Kalmegh accessions.
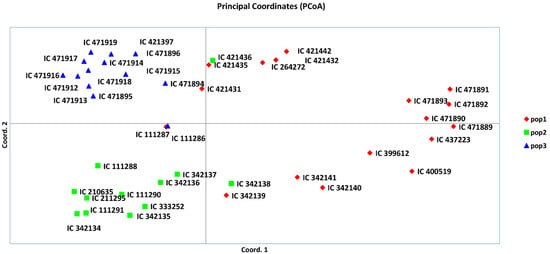
Figure 6.
Principal coordinate analysis (PCoA) based on model-based population structure.
4. Conclusions
Based on this study, it can be concluded that the novel set of g-SSR primer pairs developed in the present study were found to be efficient for molecular characterization of A. paniculate accessions. Thus, it can be added as new genomic resources for A. paniculata and further utilized in germplasm management and basic population genetics and plant-breeding studies.
Supplementary Materials
The following are available online at https://www.mdpi.com/2223-7747/9/12/1734/s1. Figure S1: Analysis of molecular variance (AMOVA) of 42 accessions of Andrographis paniculata based on SSR marker data. Figure S2: Plot of mean likelihood L (K) and variance per K value from STRUCTURE on SSR dataset. Figure S3: Table output of the Evanno method results; yellow highlight is performed dynamically on the Structure Harvester online tool and shows the highest value in the Delta K column. Figure S4: Gel image of primer Ando4-34-1 among the 42 accessions of A. paniculate; M = 100 bp marker; polymorphic bands are indicated with a yellow arrow. Figure S5: Gel image of primer Ando2-40-2 among the 42 accessions of A. paniculata. Figure S6: Gel image of primer Ando4-26 among the 42 accessions of A. paniculata. Figure S7: Relationship between genetic and geographic distances for 42 A. paniculata accessions. Table S1: Nanodrop DNA quantification result of 42 A. paniculata accessions. Table S2: Mean value of Fst1, Fst2, Fst3, and alpha concluded from a model-based approach. Table S3: Allele-frequency divergence among populations of A. paniculata accessions. Table S4: Percentage of variation explained by the first three axes among the A. paniculata accessions. Table S5: Mantel results for geographic distance vs. genetic distance.
Author Contributions
R.S. conceived and designed the experiments; R.K., C.K., D.R.C., and R.P. performed the experiments; R.K., I.S., and R.S. analyzed the data; A.K. (Ashok Kumar) contributed reagents/materials/analysis tools; R.K., A.K. (Abha Kumari), and R.S. contributed to the writing of the manuscript. All authors have read and agreed to the final version of the manuscript.
Funding
The research work was fully supported by the ICAR-National Bureau of Plant Genetic Resources, New Delhi.
Acknowledgments
We are grateful to the Indian Council of Agricultural Research (ICAR), New Delhi, for financial support, and the University Grant Commission (UGC) for providing fellowship to the senior author. We are also thankful to the Director, ICAR-NBPGR, New Delhi, and the In-charge Head, Division of Genomic Resources, NBPGR, New Delhi, for providing research facilities.
Conflicts of Interest
There is no conflict of interest for this manuscript.
References
- Chandrasekaran, C.; Thiyagarajan, P.; Sundarajan, K.; Goudar, K.S.; Deepak, M.; Murali, B.; Joseph, J.A.; Agarwal, A. Evaluation of the genotoxic potential and acute oral toxicity of standardized extract of Andrographis paniculata (KalmCold™). Food Chem. Toxicol. 2009, 47, 1892–1902. [Google Scholar] [CrossRef] [PubMed]
- Niranjan, A.; Tewari, S.K.; Lehri, A. Biological activities of Kalmegh (Andrographis paniculata Nees). Indian J. Nat. Prod. Resour. 2010, 1, 125–135. [Google Scholar]
- Sharma, S.N.; Sinha, R.K.; Sharma, D.K.; Zenu, J. Assessment of intra-specific variability at morphological, molecular and biochemical level of Andrographis paniculata (Kalmegh). Curr. Sci. 2009, 10, 402–408. [Google Scholar]
- Tiwari, G.; Singh, R.; Singh, N.; Choudhury, D.R.; Paliwal, R.; Kumar, A.; Gupta, V. Study of arbitrarily amplified (RAPD and ISSR) and gene targeted (SCoT and CBDP) markers for genetic diversity and population structure in Kalmegh [Andrographis paniculata (Burm. f.) Nees]. Ind. Crop. Prod. 2016, 86, 1–11. [Google Scholar] [CrossRef]
- Lattoo, S.K.; Dhar, R.S.; Khan, S.; Bamotra, S.; Bhan, M.K.; Dhar, A.K.; Gupta, K.K. Comparative analysis of genetic diversity using molecular and morphometric markers in Andrographis paniculata (Burm. f.) Nees. Genet. Resour. Crop. Evol. 2007, 55, 33–43. [Google Scholar] [CrossRef]
- Jayakumar, T.; Hsieh, C.-Y.; Lee, J.-J.; Sheu, J.-R. Experimental and Clinical Pharmacology of Andrographis paniculate and Its Major Bioactive Phytoconstituent Andrographolide. Evidence-Based Complement. Altern. Med. 2013, 2013, 1–16. [Google Scholar] [CrossRef]
- Puri, A.; Saxena, R.; Saxena, K.C.; Srivastava, V.; Tandon, J.S. Immunostimulant Agents from Andrographis paniculata. J. Nat. Prod. 1993, 56, 995–999. [Google Scholar] [CrossRef]
- Kapil, A.; Koul, I.; Banerjee, S.; Gupta, B. Antihepatotoxic effects of major diterpenoid constituents of Andrographis paniculata. Biochem. Pharmacol. 1993, 46, 182–185. [Google Scholar] [CrossRef]
- Singha, P.K.; Roy, S.; Dey, S. Antimicrobial activity of Andrographis paniculata. Fitoterapia 2003, 74, 692–694. [Google Scholar] [CrossRef]
- Dua, V.; Ojha, V.; Roy, R.; Joshi, B.; Valecha, N.; Devi, C.U.; Bhatnagar, M.; Sharma, V.; Subbarao, S. Anti-malarial activity of some xanthones isolated from the roots of Andrographis paniculata. J. Ethnopharmacol. 2004, 95, 247–251. [Google Scholar] [CrossRef]
- Thisoda, P.; Rangkadilok, N.; Pholphana, N.; Worasuttayangkurn, L.; Ruchirawat, S.; Satayavivad, J. Inhibitory effect of Andrographis paniculata extract and its active diterpenoids on platelet aggregation. Eur. J. Pharmacol. 2006, 553, 39–45. [Google Scholar] [CrossRef]
- Zhou, J.; Zhang, S.; Choon-Nam, O.; Shen, H.-M. Critical role of pro-apoptotic Bcl-2 family members in andrographolide-induced apoptosis in human cancer cells. Biochem. Pharmacol. 2006, 72, 132–144. [Google Scholar] [CrossRef]
- Liu, J.; Wang, Z.; Ge, B.-X. Andrograpanin, isolated from Andrographis paniculata, exhibits anti-inflammatory property in lipopolysaccharide-induced macrophage cells through down-regulating the p38 MAPKs signaling pathways. Int. Immunopharmacol. 2008, 8, 951–958. [Google Scholar] [CrossRef]
- Valdiani, A.; Kadir, M.A.; Tan, S.G.; Talei, D.; Abdullah, M.P.; Nikzad, S. Nain-e Havandi Andrographis paniculata present yesterday, absent today: A plenary review on underutilized herb of Iran’s pharmaceutical plants. Mol. Biol. Rep. 2011, 39, 5409–5424. [Google Scholar] [CrossRef]
- Verma, H.; Negi, M.S.; Mahapatra, B.S.; Shukla, A.; Paul, J. Evaluation of an emerging medicinal crop Kalmegh [Andrographis paniculata (Burm. F.) Wall. Ex. Nees] for commercial cultivation and pharmaceutical & industrial uses: A review. J. Pharmacogn. Phytochem. 2019, 8, 835–848. [Google Scholar]
- Rukhsar; Patel, M.P.; Parmar, D.J.; Kalola, A.D.; Kumar, S. Morphological and molecular diversity patterns in castor germplasm accessions. Ind. Crop. Prod. 2017, 97, 316–323. [Google Scholar] [CrossRef]
- Wijarat, P.; Keeratinijakal, V.; Toojinda, T.; Vnavichit, A.; Tragoonrung, S. Genetic evaluation of Andrographis paniculata (Burm. f.) Nees revealed by SSR, AFLP and RAPD markers. J. Med. Plant Res. 2012, 6, 2777–2788. [Google Scholar] [CrossRef]
- Bharti, R.; Kumar, S.; Parekh, M.J. Development of genomic simple sequence repeat (gSSR) markers in cumin and their application in diversity analyses and cross-transferability. Ind. Crop. Prod. 2018, 111, 158–164. [Google Scholar] [CrossRef]
- Kalia, R.K.; Rai, M.K.; Kalia, S.; Singh, R.; Dhawan, A.K. Microsatellite markers: An overview of the recent progress in plants. Euphytica 2011, 177, 309–334. [Google Scholar] [CrossRef]
- Billotte, N.; Risterucci, A.M.; Barcelos, E.; Noyer, J.L.; Amblard, P.; Baurens, F.-C. Development, characterisation, and across-taxa utility of oil palm (Elaeis guineensis Jacq.) microsatellite markers. Genome 2001, 44, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y. Development of simple sequence repeat (SSR) markers to study diversity in the herbaceous peony (Paeonia lactiflora). J. Med. Plants Res. 2011, 5. [Google Scholar] [CrossRef]
- Rakotondralambo, S.O.R.; Lussert, A.; Rivallan, R.; Danthu, P.; Noyer, J.-L.; Baurens, F.-C.; He, X.; Wang, Y.; Li, F.; Weng, Q.; et al. Microsatellite markers isolated from the wild medicinal plant Centella asiatica (Apiaceae) from an enriched genomic library. Am. J. Bot. 2012, 99, e176–e178. [Google Scholar] [CrossRef]
- Paliwal, R.; Kumar, R.; Choudhury, D.R.; Singh, A.K.; Kumar, S.; Kumar, A.; Bhatt, K.; Singh, R.; Mahato, A.K.; Singh, N.; et al. Development of genomic simple sequence repeats (g-SSR) markers in Tinospora cordifolia and their application in diversity analyses. Plant. Gene 2016, 5, 118–125. [Google Scholar] [CrossRef]
- Sraphet, S.; Chantravisut, P.; Srisawad, N.; Suksee, N.; Smith, D.R.; Triwitayakorn, K. Development and application of SSR markers derived from Bauhinia Strychnifolia a semi-endemic plant in Thailand. J. Herbs Spices Med. Plants 2018, 24, 386–393. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 13–15. [Google Scholar]
- Fischer, D.; Bachmann, K. Microsatellite Enrichment in Organisms with Large Genomes (Allium cepa L.). Biotechniques 1998, 24, 796–802. [Google Scholar] [CrossRef]
- Bloor, P.A.; Barker, F.S.; Watts, P.C.; Noyes, H.A.; Kemp, S.J. Microsatellite Libraries by Enrichment 2001. Available online: http://www.genomics.liv.ac.uk/animal/research/microsatellite.pdf (accessed on 4 February 2010).
- Pavel, A.B.; Vasile, C.I. PyElph—A software tool for gel images analysis and phylogenetics. BMC Bioinform. 2012, 13, 9. [Google Scholar] [CrossRef]
- Liu, K.; Muse, S.V. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. genalex 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Wee, A.K.S.; Takayama, K.; Kajita, T.; Webb, E.L. Microsatellite loci for Avicennia alba (acanthaceae), Sonneratia alba (lythraceae) and Rhizophora mucronata (rhizophoraceae). J. Trop. For. Sci. 2013, 25, 131–136. [Google Scholar]
- Selvaraj, S.; Kaliswamy, P.; Vellingiri, S.; Nathan, B. Microsatellite analysis in the genome of Acanthaceae: An in silico approach. Pharmacogn. Mag. 2015, 11, 152–156. [Google Scholar] [CrossRef]
- Lagercrantz, U.; Ellegren, H.; Andersson, L. The abundance of various polymorphic microsatellite motifs differs between plants and vertebrates. Nucleic Acids Res. 1993, 21, 1111–1115. [Google Scholar] [CrossRef]
- La Rota, M.; Kantety, R.V.; Yu, J.-K.; Sorrells, M.E. Nonrandom distribution and frequencies of genomic and EST-derived microsatellite markers in rice, wheat, and barley. BMC Genom. 2005, 6. [Google Scholar] [CrossRef]
- Saha, M.C.; Cooper, J.D.; Mian, M.A.R.; Chekhovskiy, K.; May, G.D. Tall fescue genomic SSR markers: Development and transferability across multiple grass species. Theor. Appl. Genet. 2006, 113, 1449–1458. [Google Scholar] [CrossRef]
- Rongwen, J.; Akkaya, M.S.; Bhagwat, A.A.; Lavi, U.; Cregan, P.B. The use of microsatellite DNA markers for soybean genotype identification. Theor. Appl. Genet. 1995, 90, 43–48. [Google Scholar] [CrossRef]
- Goldstein, D.B.; Linares, A.R.; Cavalli-Sforza, L.L.; Feldman, M.W. An Evaluation of Genetic Distances for Use with Microsatellite Loci. Genetics 1995, 139, 463–471. [Google Scholar]
- Chen, H.; Song, Y.; Li, L.; Khan, M.A.; Li, X.-G.; Korban, S.S.; Wu, J.; Zhang, S.-L. Construction of a High-Density Simple Sequence Repeat Consensus Genetic Map for Pear (Pyrus spp.). Plant. Mol. Biol. Rep. 2014, 33, 316–325. [Google Scholar] [CrossRef]
- Yu, Y.G. RFLP and Microsatellite Mapping of a Gene for Soybean Mosaic Virus Resistance. Phytopathology 1994, 84. [Google Scholar] [CrossRef]
- Moe, A.M.; Weiblen, G.D. Development and characterization of microsatellite loci in dioecious figs (Ficus, Moraceae). Am. J. Bot. 2011, 98, e25–e27. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-L.; Yang, J.; Boykin, L.M.; Zhao, Q.-Y.; Wang, Y.-J.; Liu, S.-S.; Wang, X.-W. Developing conversed microsatellite markers and their implications in evolutionary analysis of the Bemisia tabaci complex. Sci. Rep. 2014, 4, 6351. [Google Scholar] [CrossRef] [PubMed]
- Rafalski, J.A.; Vogel, J.M.; Morgante, M.; Powell, W.; Andre, C.; Tingey, S.V. Generating and Using DNA Markers in Plants. In Nonmammalian Genomic Analysis; Academic Press, 1996; pp. 75–134. [Google Scholar]
- Geng, Q.F.; Lian, C.L.; Tao, J.M.; Hogetsu, T. Permanent Genetic Resources: Development of microsatellite markers for two nonviviparous mangrove species, Acanthus ilicifolius and Lumnitzera racemosa. Mol. Ecol. Resour. 2008, 8, 377–380. [Google Scholar] [CrossRef]
- Suárez-Montes, P.; Tapia-López, R.; Núaez-Farfán, J. Genetic variation at microsatellite loci in the tropical herb Aphelandra aurantiaca (Acanthaceae)1. Appl. Plant. Sci. 2015, 3, 1500058. [Google Scholar] [CrossRef]
- Schaal, B.A.; Hayworth, D.A.; Olsen, K.M.; Rauscher, J.T.; Smith, W.A. Phylogeographic studies in plants: Problems and prospects. Mol. Ecol. 1998, 7, 465–474. [Google Scholar] [CrossRef]
- Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 1967, 27, 209–220. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).