The bRPS6-Family Protein RFC3 Prevents Interference by the Splicing Factor CFM3b during Plastid rRNA Biogenesis in Arabidopsis thaliana
Abstract
:1. Introduction
2. Results
2.1. Identification of Sprt Mutants
2.2. Most rfc3 Mutant Phenotypes were Recovered by the sprt2-1 Mutation
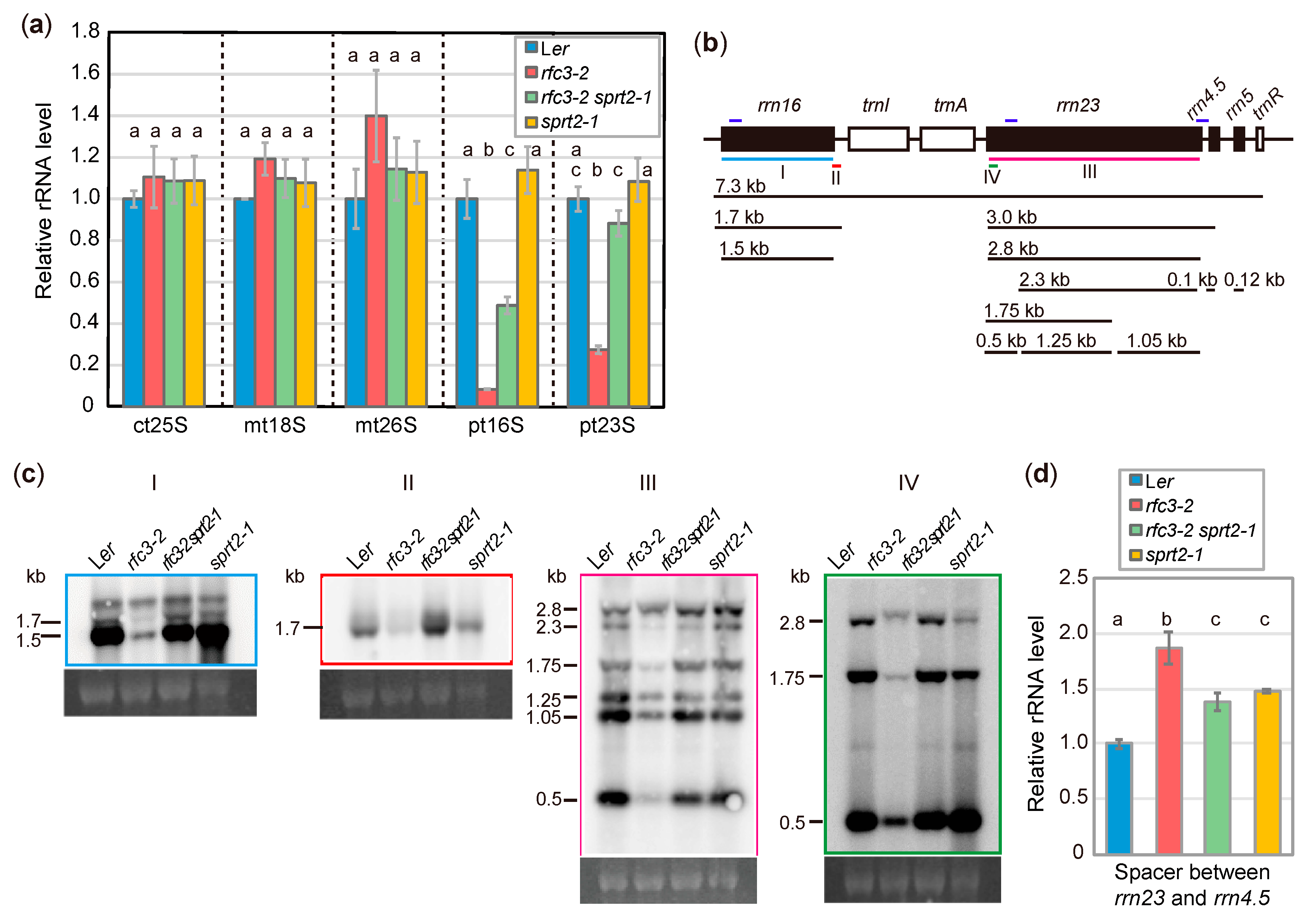
2.3. rfc3-2 sprt2-1 Mutants Exhibit Restoration of Plastid rRNA Accumulation
2.4. Effect of the sprt2-1 Mutation on Plastid Ribosome Biogenesis
2.5. Identification of the sprt2-1 Mutation Site
2.6. Expression Pattern of CFM3b and Subcellular Localization of its Product
2.7. RFC3 Interacts with CFM3b in Plastids
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Isolation of Suppressor Mutants of rfc3-2
4.3. Genetic Mapping of the sprt2-1 Mutation Site
4.4. Complementation Test of sprt2
4.5. Microscopy
4.6. Transient Assays
4.7. Northern Blot Analysis of Plastid rRNAs
4.8. RT-qPCR
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Andersson, S.G.E.; Karlberg, O.; Canbäck, B.; Kurland, C.G. On the origin of mitochondria: A genomics perspective. Phil. Trans. R. Soc. Lond. B 2003, 358, 165–179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raven, J.A.; Allen, J.F. Genomics and chloroplast evolution: What did cyanobacteria do for plants? Genome Biol. 2003, 4, 209. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harris, E.H.; Boynton, J.E.; Gillham, N.W. Chloroplast ribosomes and protein synthesis. Microbiol. Rev. 1994, 58, 700–754. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, K.; von Knoblauch, K.; Subramanian, A.R. The plastid ribosomal proteins. Identification of all the proteins in the 30 S subunit of an organelle ribosome (chloroplast). J. Biol. Chem. 2000, 275, 28455–28465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamaguchi, K.; Subramanian, A.R. The plastid ribosomal proteins. Identification of all the proteins in the 50 S subunit of an organelle ribosome (chloroplast). J. Biol. Chem. 2000, 275, 28466–28482. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamaguchi, K.; Subramanian, A.R. Proteomic identification of all plastid-specific ribosomal proteins in higher plant chloroplast 30S ribosomal subunit. Eur. J. Biochem. 2003, 270, 190–205. [Google Scholar] [CrossRef]
- Tiller, N.; Bock, R. The translational apparatus of plastids and its role in plant development. Mol. Plant 2014, 7, 1105–1120. [Google Scholar] [CrossRef] [Green Version]
- Wakasugi, T.; Tsudzuki, T.; Sugiura, M. The genomics of land plant chloroplasts: Gene content and alteration of genomic information by RNA editing. Photosynth. Res. 2001, 70, 107–118. [Google Scholar] [CrossRef]
- Stern, D.B.; Goldschmidt-Clermont, M.; Hanson, M.R. Chloroplast RNA metabolism. Annu. Rev. Plant Biol. 2010, 61, 125–155. [Google Scholar] [CrossRef]
- Germain, A.; Hotto, A.M.; Barkan, A.; Stern, D.B. RNA processing and decay in plastids. WIREs RNA 2013, 4, 295–316. [Google Scholar] [CrossRef]
- Sun, L.; Xu, Y.; Bai, S.; Bai, X.; Zhu, H.; Dong, H.; Wang, W.; Zhu, X.; Hao, F.; Song, C.P. Transcriptome-wide analysis of pseudouridylation of mRNA and non-coding RNAs in Arabidopsis. J. Exp. Bot. 2019, 70, 5089–5100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.; Tang, K.; Zhang, D.; Wan, Y.; Wen, Y.; Lu, Q.; Wang, L. High-throughput m6A-seq reveals RNA m6A methylation patterns in the chloroplast and mitochondria transcriptomes of Arabidopsis thaliana. PLoS ONE 2017, 12, e0185612. [Google Scholar] [CrossRef] [PubMed]
- Burgess, A.L.; David, R.; Searle, I.R. Conservation of tRNA and rRNA 5-methylcytosine in the kingdom Plantae. BMC Plant Biol. 2015, 15, 199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nishimura, K.; Ashida, H.; Ogawa, T.; Yokota, A. A DEAD box protein is required for formation of a hidden break in Arabidopsis chloroplast 23S rRNA. Plant J. 2010, 63, 766–777. [Google Scholar] [CrossRef] [PubMed]
- Bonen, L.; Vogel, J. The ins and outs of group II introns. Trends Genet. 2001, 17, 322–331. [Google Scholar] [CrossRef]
- Ostersetzer, O.; Cooke, A.M.; Watkins, K.P.; Barkan, A. CRS1, a chloroplast group II intron splicing factor, promotes intron folding through specific interactions with two intron domains. Plant Cell 2005, 17, 241–255. [Google Scholar] [CrossRef] [Green Version]
- De Longevialle, A.F.; Small, I.D.; Lurin, C. Nuclearly encoded splicing factors implicated in RNA splicing in higher plant organelles. Mol. Plant 2010, 3, 691–705. [Google Scholar] [CrossRef] [Green Version]
- Barkan, A.; Klipcan, L.; Ostersetzer, O.; Kawamura, T.; Asakura, Y.; Watkins, K.P. The CRM domain: An RNA binding module derived from an ancient ribosome-associated protein. RNA 2007, 13, 55–64. [Google Scholar] [CrossRef] [Green Version]
- Gagarinova, A.; Stewart, G.; Samanfar, B.; Phanse, S.; White, C.A.; Aoki, H.; Deineko, V.; Beloglazova, N.; Yakunin, A.F.; Golshani, A.; et al. Systematic genetic screens reveal the dynamic global functional organization of the bacterial translation machinery. Cell Rep. 2016, 17, 904–916. [Google Scholar] [CrossRef] [Green Version]
- Ostheimer, G.J.; Williams-Carrier, R.; Belcher, S.; Osborne, E.; Gierke, J.; Barkan, A. Group II intron splicing factors derived by diversification of an ancient RNA-binding domain. EMBO J. 2003, 22, 3919–3929. [Google Scholar] [CrossRef] [Green Version]
- Jenkins, B.D.; Barkan, A. Recruitment of a peptidyl-tRNA hydrolase as a facilitator of group II intron splicing in chloroplasts. EMBO J. 2001, 20, 872–879. [Google Scholar] [CrossRef] [PubMed]
- Till, B.; Schmitz-Linneweber, C.; Williams-Carrier, R.; Barkan, A. CRS1 is a novel group II intron splicing factor that was derived from a domain of ancient origin. RNA 2001, 7, 1227–1238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asakura, Y.; Bayraktar, O.A.; Barkan, A. Two CRM protein subfamilies cooperate in the splicing of group IIB introns in chloroplasts. RNA 2008, 14, 2319–2332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asakura, Y.; Barkan, A. A CRM domain protein functions dually in group I and group II intron splicing in land plant chloroplasts. Plant Cell 2007, 19, 3864–3875. [Google Scholar] [CrossRef] [Green Version]
- Zmudjak, M.; des Francs-Small, C.C.; Keren, I.; Shaya, F.; Belausov, E.; Small, I.; Ostersetzer-Biran, O. mCSF1, a nucleus-encoded CRM protein required for the processing of many mitochondrial introns, is involved in the biogenesis of respiratory complexes I and IV in Arabidopsis. New Phytol. 2013, 199, 379–394. [Google Scholar] [CrossRef]
- Lee, K.; Park, S.J.; Park, Y.I.; Kang, H. CFM9, a mitochondrial CRM protein, is crucial for mitochondrial intron splicing, mitochondria function and Arabidopsis growth and stress responses. Plant Cell Physiol. 2019, 60, 2538–2549. [Google Scholar] [CrossRef]
- Lee, K.; Lee, H.J.; Kim, D.H.; Jeon, Y.; Pai, H.S.; Kang, H. A nuclear-encoded chloroplast protein harboring a single CRM domain plays an important role in the Arabidopsis growth and stress response. BMC Plant Biol. 2014, 14, 98. [Google Scholar] [CrossRef] [Green Version]
- Sakamoto, W.; Miyagishima, S.; Jarvis, P. Chloroplast biogenesis: Control of plastid development, protein import, division and inheritance. Arab. Book 2008, 6, e0110. [Google Scholar] [CrossRef] [Green Version]
- Waters, M.T.; Langdale, J.A. The making of a chloroplast. EMBO J. 2009, 28, 2861–2873. [Google Scholar] [CrossRef]
- Kobayashi, K.; Baba, S.; Obayashi, T.; Sato, M.; Toyooka, K.; Keränen, M.; Aro, E.M.; Fukaki, H.; Ohta, H.; Sugimoto, K.; et al. Regulation of root greening by light and auxin/cytokinin signaling in Arabidopsis. Plant Cell 2012, 24, 1081–1095. [Google Scholar] [CrossRef] [Green Version]
- Pyke, K.A.; Page, A.M. Plastid ontogeny during petal development in Arabidopsis. Plant Physiol. 1998, 116, 797–803. [Google Scholar] [CrossRef] [Green Version]
- Bramham, L.; Pyke, K. Changing plastid dynamics within early root and shoot apical meristem-derived tissue of A. thaliana. Biosci. Horizons 2017, 10, hz002. [Google Scholar] [CrossRef] [Green Version]
- Horiguchi, G.; Kodama, H.; Iba, K. Characterization of Arabidopsis mutants that are associated with altered C18 unsaturated fatty acid metabolism. Plant Sci. 2001, 161, 1117–1123. [Google Scholar] [CrossRef]
- Horiguchi, G.; Kodama, H.; Iba, K. Mutations in a gene for plastid ribosomal protein S6-like protein reveal a novel developmental process required for the correct organization of lateral root meristem in Arabidopsis. Plant J. 2003, 33, 521–529. [Google Scholar] [CrossRef] [PubMed]
- Nakata, M.T.; Sato, M.; Wakazaki, M.; Sato, N.; Kojima, K.; Sekine, A.; Nakamura, S.; Shikanai, T.; Toyooka, K.; Tsukaya, H.; et al. Plastid translation is essential for lateral root stem cell patterning in Arabidopsis thaliana. Biol. Open 2018, 7, bio028175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Malamy, J.E.; Benfey, P.N. Organization and cell differentiation in lateral roots of Arabidopsis thaliana. Development 1997, 124, 33–44. [Google Scholar] [PubMed]
- Okushima, Y.; Fukaki, H.; Onoda, M.; Theologis, A.; Tasaka, M. ARF7 and ARF19 Regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 2007, 19, 118–130. [Google Scholar] [CrossRef] [Green Version]
- De Rybel, B.; Vassileva, V.; Parizot, B.; Demeulenaere, M.; Grunewald, W.; Audenaert, D.; Van Campenhout, J.; Overvoorde, P.; Jansen, L.; Vanneste, S.; et al. A novel Aux/IAA28 signaling cascade activates GATA23-dependent specification of lateral root founder cell identity. Curr. Biol. 2010, 20, 1697–1706. [Google Scholar] [CrossRef]
- Goh, T.; Joi, S.; Mimura, T.; Fukaki, H. The establishment of asymmetry in Arabidopsis lateral root founder cells is regulated by LBD16/ASL18 and related LBD/ASL proteins. Development 2012, 139, 883–893. [Google Scholar] [CrossRef] [Green Version]
- Du, Y.; Scheres, B. PLETHORA transcription factors orchestrate de novo organ patterning during Arabidopsis lateral root outgrowth. Proc. Natl. Acad. Sci. USA 2017, 114, 11709–11714. [Google Scholar] [CrossRef] [Green Version]
- Goh, T.; Toyokura, K.; Wells, D.M.; Swarup, K.; Yamamoto, M.; Mimura, T.; Weijers, D.; Fukaki, H.; Laplaze, L.; Bennett, M.J.; et al. Quiescent center initiation in the Arabidopsis lateral root primordia is dependent on the SCARECROW transcription factor. Development 2016, 143, 3363–3371. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimotohno, A.; Heidstra, R.; Blilou, I.; Scheres, B. Root stem cell niche organizer specification by molecular convergence of PLETHORA and SCARECROW transcription factor modules. Genes Dev. 2018, 32, 1085–1100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Araya, T.; Miyamoto, M.; Wibowo, J.; Suzuki, A.; Kojima, S.; Tsuchiya, Y.N.; Sawa, S.; Fukuda, H.; von Wirén, N.; Takahashi, H. CLE-CLAVATA1 peptide-receptor signaling module regulates the expansion of plant root systems in a nitrogen-dependent manner. Proc. Natl. Acad. Sci. USA 2014, 111, 2029–2034. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hofhuis, H.; Laskowski, M.; Du, Y.; Prasad, K.; Grigg, S.; Pinon, V.; Scheres, B. Phyllotaxis and rhizotaxis in Arabidopsis are modified by three PLETHORA transcription factors. Curr. Biol. 2013, 23, 956–962. [Google Scholar] [CrossRef] [Green Version]
- Romani, I.; Tadini, L.; Rossi, F.; Masiero, S.; Pribil, M.; Jahns, P.; Kater, M.; Leister, D.; Pesaresi, P. Versatile roles of Arabidopsis plastid ribosomal proteins in plant growth and development. Plant J. 2012, 72, 922–934. [Google Scholar] [CrossRef]
- Ahmed, T.; Shi, J.; Bhushan, S. Unique localization of the plastid-specific ribosomal proteins in the chloroplast ribosome small subunit provides mechanistic insights into the chloroplastic translation. Nucleic Acids Res. 2017, 45, 8581–8595. [Google Scholar] [CrossRef]
- Wang, W.J.; Zheng, K.L.; Gong, X.D.; Xu, J.L.; Huang, J.R.; Lin, D.Z.; Dong, Y.J. The rice TCD11 encoding plastid ribosomal protein S6 is essential for chloroplast development at low temperature. Plant Sci. 2017, 259, 1–11. [Google Scholar] [CrossRef]
- Espinar-Marchena, F.J.; Babiano, R.; de la Cruz, J. Placeholder factors in ribosome biogenesis: Please, pave my way. Microb. Cell 2017, 4, 144–168. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Yuan, H.; Yang, Y.; Fish, T.; Lyi, S.M.; Thannhauser, T.W.; Zhang, L.; Li, L. Plastid ribosomal protein S5 is involved in photosynthesis, plant development, and cold stress tolerance in Arabidopsis. J. Exp. Bot. 2016, 67, 2731–2744. [Google Scholar] [CrossRef] [Green Version]
- Tiller, N.; Weingartner, M.; Thiele, W.; Maximova, E.; Schöttler, M.A.; Bock, R. The plastid-specific ribosomal proteins of Arabidopsis thaliana can be divided into non-essential proteins and genuine ribosomal proteins. Plant J. 2012, 69, 302–316. [Google Scholar] [CrossRef]
- Aryamanesh, N.; Ruwe, H.; Vincis Pereira Sanglard, L.; Eshraghi, L.; Bussell, J.D.; Howell, K.A.; Small, I.; des Francs-Small, C.C. The pentatricopeptide repeat protein EMB2654 is essential for trans-splicing of a chloroplast small ribosomal subunit transcript. Plant Physiol. 2017, 173, 1164–1176. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Zheng, M.; Wang, R.; Wang, R.; An, L.; Rodermel, S.R.; Yu, F. Genetic interactions reveal that specific defects of chloroplast translation are associated with the suppression of var2-mediated leaf variegation. J. Integr. Plant Biol. 2013, 55, 979–993. [Google Scholar] [PubMed]
- Kleinknecht, L.; Wang, F.; Stübe, R.; Philippar, K.; Nickelsen, J.; Bohne, A.V. RAP, the sole octotricopeptide repeat protein in Arabidopsis, is required for chloroplast 16S rRNA maturation. Plant Cell 2014, 26, 777–787. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fristedt, R.; Scharff, L.B.; Clarke, C.A.; Wang, Q.; Lin, C.; Merchant, S.S.; Bock, R. RBF1, a plant homolog of the bacterial ribosome-binding factor RbfA, acts in processing of the chloroplast 16S ribosomal RNA. Plant Physiol. 2014, 164, 201–215. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Yu, F.; Rodermel, S. An Arabidopsis pentatricopeptide repeat protein, SUPPRESSOR OF VARIEGATION7, is required for FtsH-mediated chloroplast biogenesis. Plant Physiol. 2010, 154, 1588–1601. [Google Scholar] [CrossRef] [Green Version]
- Tadini, L.; Pesaresi, P.; Kleine, T.; Rossi, F.; Guljamow, A.; Sommer, F.; Mühlhaus, T.; Schroda, M.; Masiero, S.; Pribil, M.; et al. GUN1 controls accumulation of the plastid ribosomal protein S1 at the protein level and interacts with proteins involved in plastid protein homeostasis. Plant Physiol. 2016, 170, 1817–1830. [Google Scholar] [CrossRef] [Green Version]
- Bobik, K.; Fernandez, J.C.; Hardin, S.R.; Ernest, B.; Ganusova, E.E.; Staton, M.E.; Burch-Smith, T.M. The essential chloroplast ribosomal protein uL15c interacts with the chloroplast RNA helicase ISE2 and affects intercellular trafficking through plasmodesmata. New Phytol. 2019, 221, 850–865. [Google Scholar] [CrossRef] [Green Version]
- Ehrnthaler, M.; Scharff, L.B.; Fleischmann, T.T.; Hasse, C.; Ruf, S.; Bock, R. Synthetic lethality in the tobacco plastid ribosome and its rescue at elevated growth temperatures. Plant Cell 2014, 26, 765–776. [Google Scholar] [CrossRef] [Green Version]
- Meurer, J.; Schmid, L.M.; Stoppel, R.; Leister, D.; Brachmann, A.; Manavski, N. PALE CRESS binds to plastid RNAs and facilitates the biogenesis of the 50S ribosomal subunit. Plant J. 2017, 92, 400–413. [Google Scholar] [CrossRef]
- Wu, W.; Liu, S.; Ruwe, H.; Zhang, D.; Melonek, J.; Zhu, Y.; Hu, X.; Gusewski, S.; Yin, P.; Small, I.D.; et al. SOT1, a pentatricopeptide repeat protein with a small MutS-related domain, is required for correct processing of plastid 23S–4.5S rRNA precursors in Arabidopsis thaliana. Plant J. 2016, 85, 607–621. [Google Scholar] [CrossRef]
- Köhler, D.; Helm, S.; Agne, B.; Baginsky, S. Importance of translocon subunit Tic56 for rRNA processing and chloroplast ribosome assembly. Plant Physiol. 2016, 172, 2429–2444. [Google Scholar] [CrossRef] [Green Version]
- Yagi, Y.; Ishizaki, Y.; Nakahira, Y.; Tozawa, Y.; Shiina, T. Eukaryotic-type plastid nucleoid protein pTAC3 is essential for transcription by the bacterial-type plastid RNA polymerase. Proc. Natl. Acad. Sci. USA 2012, 109, 7541–7546. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Rodermel, S.R.; Yu, F. A var2 leaf variegation suppressor locus, SUPPRESSOR OF VARIEGATION3, encodes a putative chloroplast translation elongation factor that is important for chloroplast development in the cold. BMC Plant Biol. 2010, 10, 287. [Google Scholar] [CrossRef] [Green Version]
- Romani, I.; Manavski, N.; Morosetti, A.; Tadini, L.; Maier, S.; Kühn, K.; Ruwe, H.; Schmitz-Linneweber, C.; Wanner, G.; Leister, D.; et al. A Member of the Arabidopsis mitochondrial transcription termination factor family is required for maturation of chloroplast transfer RNAIle(GAU). Plant Physiol. 2015, 169, 627–646. [Google Scholar] [CrossRef] [Green Version]
- Huang, W.; Zhu, Y.; Wu, W.; Li, X.; Zhang, D.; Yin, P.; Huang, J. The pentatricopeptide repeat protein SOT5/EMB2279 is Required for plastid rpl2 and trnK intron splicing. Plant Physiol. 2018, 177, 684–697. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, T.; Zhang, J.; Rong, L.; Feng, Y.; Wang, Q.; Song, Q.; Zhang, L.; Ouyang, M. ECD1 functions as an RNA-editing trans-factor of rps14-149 in plastids and is required for early chloroplast development in seedlings. J. Exp. Bot. 2018, 69, 3037–3051. [Google Scholar] [CrossRef] [PubMed]
- Bollenbach, T.J.; Lange, H.; Gutierrez, R.; Erhardt, M.; Stern, D.B.; Gagliardi, D. RNR1, a 3’-5′ exoribonuclease belonging to the RNR superfamily, catalyzes 3′ maturation of chloroplast ribosomal RNAs in Arabidopsis thaliana. Nucleic. Acids. Res. 2005, 33, 2751–2763. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, J.; Zhou, W.; Liu, G.; Yang, C.; Sun, Y.; Wu, W.; Cao, S.; Wang, C.; Hai, G.; Wang, Z.; et al. The conserved endoribonuclease YbeY is required for chloroplast ribosomal RNA processing in Arabidopsis. Plant Physiol. 2015, 168, 205–221. [Google Scholar] [CrossRef] [Green Version]
- Parker, M.D.; Collins, J.C.; Korona, B.; Ghalei, H.; Karbstein, K. A kinase-dependent checkpoint prevents escape of immature ribosomes into the translating pool. PLoS. Biol. 2019, 17, e3000329. [Google Scholar] [CrossRef] [Green Version]
- Pratte, D.; Singh, U.; Murat, G.; Kressler, D. Mak5 and Ebp2 act together on early pre-60S particles and their reduced functionality bypasses the requirement for the essential pre-60S factor Nsa1. PLoS ONE 2013, 8, e82741. [Google Scholar] [CrossRef] [Green Version]
- Konikkat, S.; Woolford, J.L. Principles of 60S ribosomal subunit assembly emerging from recent studies in yeast. Biochem. J. 2017, 474, 195–214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pestov, D.G.; Strezoska, Z.; Lau, L.F. Evidence of p53-dependent cross-talk between ribosome biogenesis and the cell cycle: Effects of nucleolar protein Bop1 on G1/S transition. Mol. Cell. Biol. 2001, 21, 4246–4255. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deisenroth, C.; Franklin, D.A.; Zhang, Y. The evolution of the ribosomal protein–MDM2–p53 pathway. Cold Spring Harb. Perspect. Med. 2016, 6, a026138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohbayashi, I.; Lin, C.Y.; Shinohara, N.; Matsumura, Y.; Machida, Y.; Horiguchi, G.; Tsukaya, H.; Sugiyama, M. Evidence for a role of ANAC082 as a ribosomal stress response mediator leading to growth defects and developmental alterations in Arabidopsis. Plant Cell 2017, 29, 2644–2660. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maekawa, S.; Ishida, T.; Yanagisawa, S. Reduced expression of APUM24, encoding a novel rRNA processing factor, induces sugar-dependent nucleolar stress and altered sugar responses in Arabidopsis thaliana. Plant Cell 2018, 30, 209–227. [Google Scholar] [CrossRef] [Green Version]
- Kleine, T.; Leister, D. Retrograde signaling: Organelles go networking. Biochim. Biophys. Acta 2016, 1857, 1313–1325. [Google Scholar] [CrossRef]
- Koussevitzky, S.; Nott, A.; Mockler, T.C.; Hong, F.; Sachetto-Martins, G.; Surpin, M.; Lim, J.; Mittler, R.; Chory, J. Signals from chloroplasts converge to regulate nuclear gene expression. Science 2007, 316, 715–719. [Google Scholar] [CrossRef]
- Tameshige, T.; Fujita, H.; Watanabe, K.; Toyokura, K.; Kondo, M.; Tatematsu, K.; Matsumoto, N.; Tsugeki, R.; Kawaguchi, M.; Nishimura, M.; et al. Pattern dynamics in adaxial-abaxial specific gene expression are modulated by a plastid retrograde signal during Arabidopsis thaliana leaf development. PLoS Genet. 2013, 9, e1003655. [Google Scholar] [CrossRef] [Green Version]
- Blilou, I.; Xu, J.; Wildwater, M.; Willemsen, V.; Paponov, I.; Friml, J.; Heidstra, R.; Aida, M.; Palme, K.; Scheres, B. The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 2005, 433, 39–44. [Google Scholar] [CrossRef]
- Galinha, C.; Hofhuis, H.; Luijten, M.; Willemsen, V.; Blilou, I.; Heidstra, R.; Scheres, B. PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 2007, 449, 1053–1057. [Google Scholar] [CrossRef] [Green Version]
- Sabatini, S.; Beis, D.; Wolkenfelt, H.; Murfett, J.; Guilfoyle, T.; Malamy, J.; Benfey, P.; Leyser, O.; Bechtold, N.; Weisbeek, P.; et al. An auxin-dependent distal organizer of pattern and polarity in the Arabidopsis root. Cell 1999, 99, 463–472. [Google Scholar] [CrossRef] [Green Version]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Edwards, K.; Johnstone, C.; Thompson, C. A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res. 1991, 19, 1349. [Google Scholar] [CrossRef] [PubMed]
- Horiguchi, G.; Gonzalez, N.; Beemster, G.T.S.; Inzé, D.; Tsukaya, H. Impact of segmental chromosomal duplications on leaf size in the grandifolia-D mutants of Arabidopsis thaliana. Plant J. 2009, 60, 122–133. [Google Scholar] [CrossRef] [PubMed]
- Kojima, K.; Tamura, J.; Chiba, H.; Fukada, K.; Tsukaya, H.; Horiguchi, G. Two nucleolar proteins, GDP1 and OLI2, function as ribosome biogenesis factors and are preferentially involved in promotion of leaf cell proliferation without strongly affecting leaf adaxial–abaxial patterning in Arabidopsis thaliana. Front. Plant Sci. 2018, 8, 2240. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [Green Version]
- Truernit, E.; Bauby, H.; Dubreucq, B.; Grandjean, O.; Runions, J.; Barthélémy, J.; Palauqui, J.C. High-resolution whole-mount imaging of three-dimensional tissue organization and gene expression enables the study of phloem development and structure in Arabidopsis. Plant Cell 2008, 20, 1494–1503. [Google Scholar] [CrossRef] [Green Version]
- Hasegawa, J.; Sakamoto, Y.; Nakagami, S.; Aida, M.; Sawa, S.; Matsunaga, S. Three-dimensional imaging of plant organs using a simple and rapid transparency technique. Plant Cell Physiol. 2016, 57, 462–472. [Google Scholar] [CrossRef] [Green Version]
- Kapila, J.; De Rycke, R.; Van Montagu, M.; Angenon, G. An Agrobacterium-mediated transient gene expression system for intact leaves. Plant Sci. 1997, 122, 101–108. [Google Scholar] [CrossRef]
- D’Aoust, M.A.; Lavoie, P.O.; Belles-Isles, J.; Bechtold, N.; Martel, M.; Vézina, L.P. Transient expression of antibodies in plants using syringe agroinfiltration. Methods Mol. Biol. 2009, 483, 41–50. [Google Scholar]
- Tanaka, Y.; Kimura, T.; Hikino, K.; Goto, S.; Nishimura, M.; Mano, S.; Nakagawa, T. Gateway vectors for plant genetic engineering: Overview of plant vectors, application for bimolecular fluorescence complementation (BiFC) and multigene construction. In Genetic Engineering—Basics, New Applications and Responsibilities; Barrera-Saldaña, H.A., Ed.; InTech: Rijeka, Croatia, 2012. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]








© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nagashima, Y.; Ohshiro, K.; Iwase, A.; Nakata, M.T.; Maekawa, S.; Horiguchi, G. The bRPS6-Family Protein RFC3 Prevents Interference by the Splicing Factor CFM3b during Plastid rRNA Biogenesis in Arabidopsis thaliana. Plants 2020, 9, 328. https://doi.org/10.3390/plants9030328
Nagashima Y, Ohshiro K, Iwase A, Nakata MT, Maekawa S, Horiguchi G. The bRPS6-Family Protein RFC3 Prevents Interference by the Splicing Factor CFM3b during Plastid rRNA Biogenesis in Arabidopsis thaliana. Plants. 2020; 9(3):328. https://doi.org/10.3390/plants9030328
Chicago/Turabian StyleNagashima, Yumi, Katsutomo Ohshiro, Akiyasu Iwase, Miyuki T. Nakata, Shugo Maekawa, and Gorou Horiguchi. 2020. "The bRPS6-Family Protein RFC3 Prevents Interference by the Splicing Factor CFM3b during Plastid rRNA Biogenesis in Arabidopsis thaliana" Plants 9, no. 3: 328. https://doi.org/10.3390/plants9030328




