Diversity and Metabolic Potential of a PAH-Degrading Bacterial Consortium in Technogenically Contaminated Haplic Chernozem, Southern Russia
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Sites
2.2. Methods
2.2.1. Sample Collection and Processing
2.2.2. Soil Analysis
PAHs Determination in the Soil Samples
DNA Extraction and Sequencing
Metagenome Quality Control, Assembly, and Annotation
Extraction of PAH Catabolic Genes from Shotgun Data
Statistical Analysis
3. Results
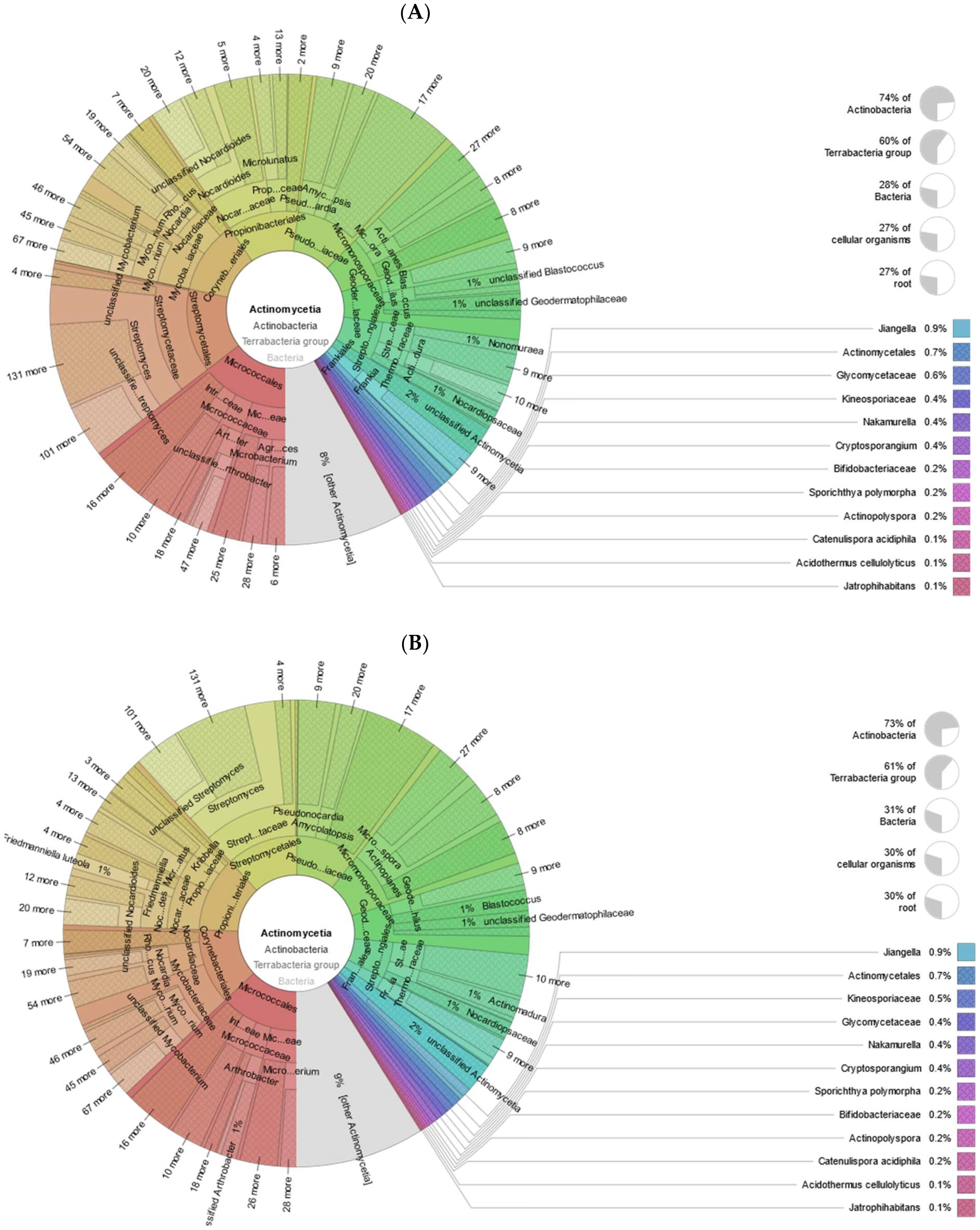
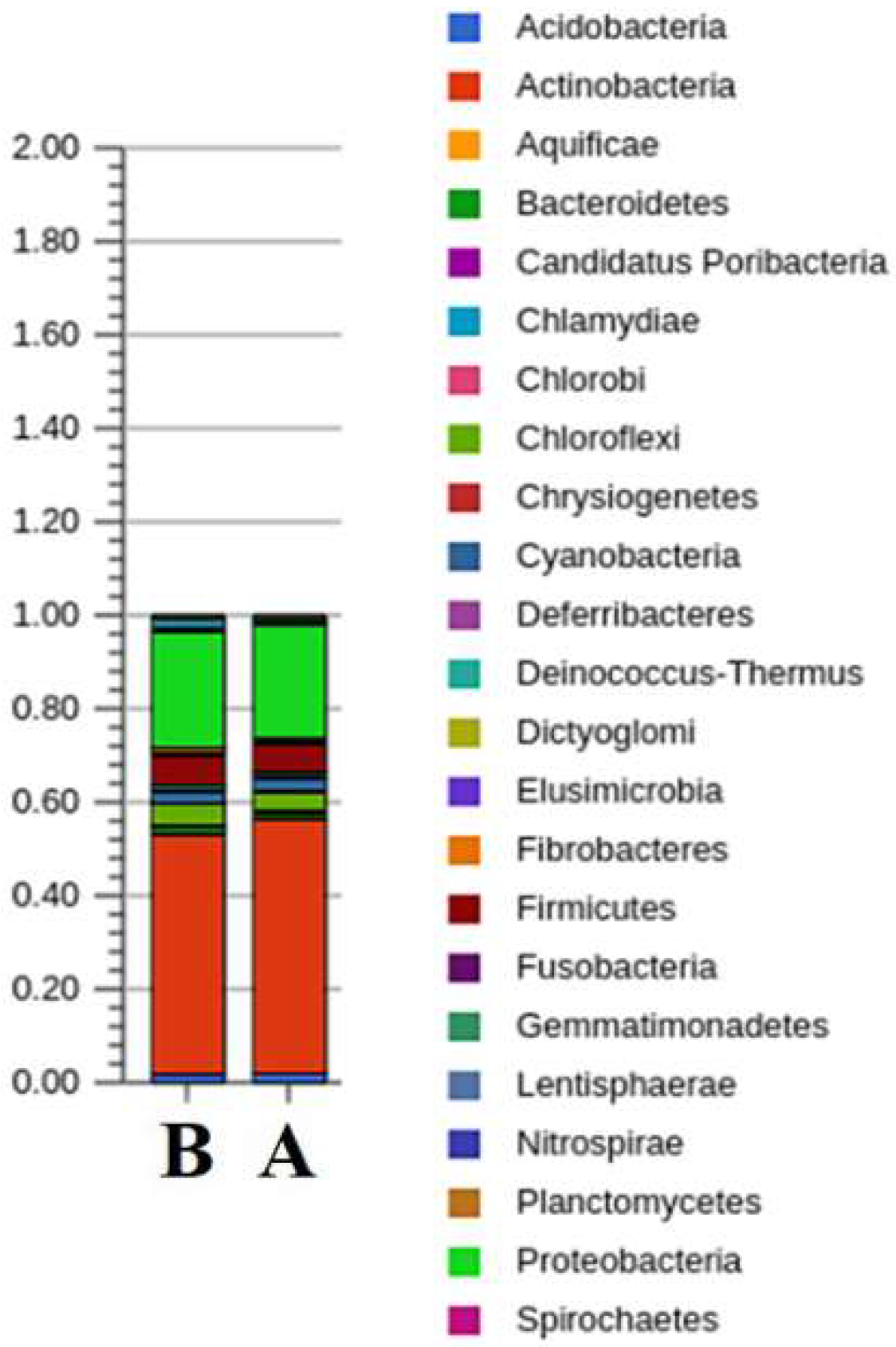
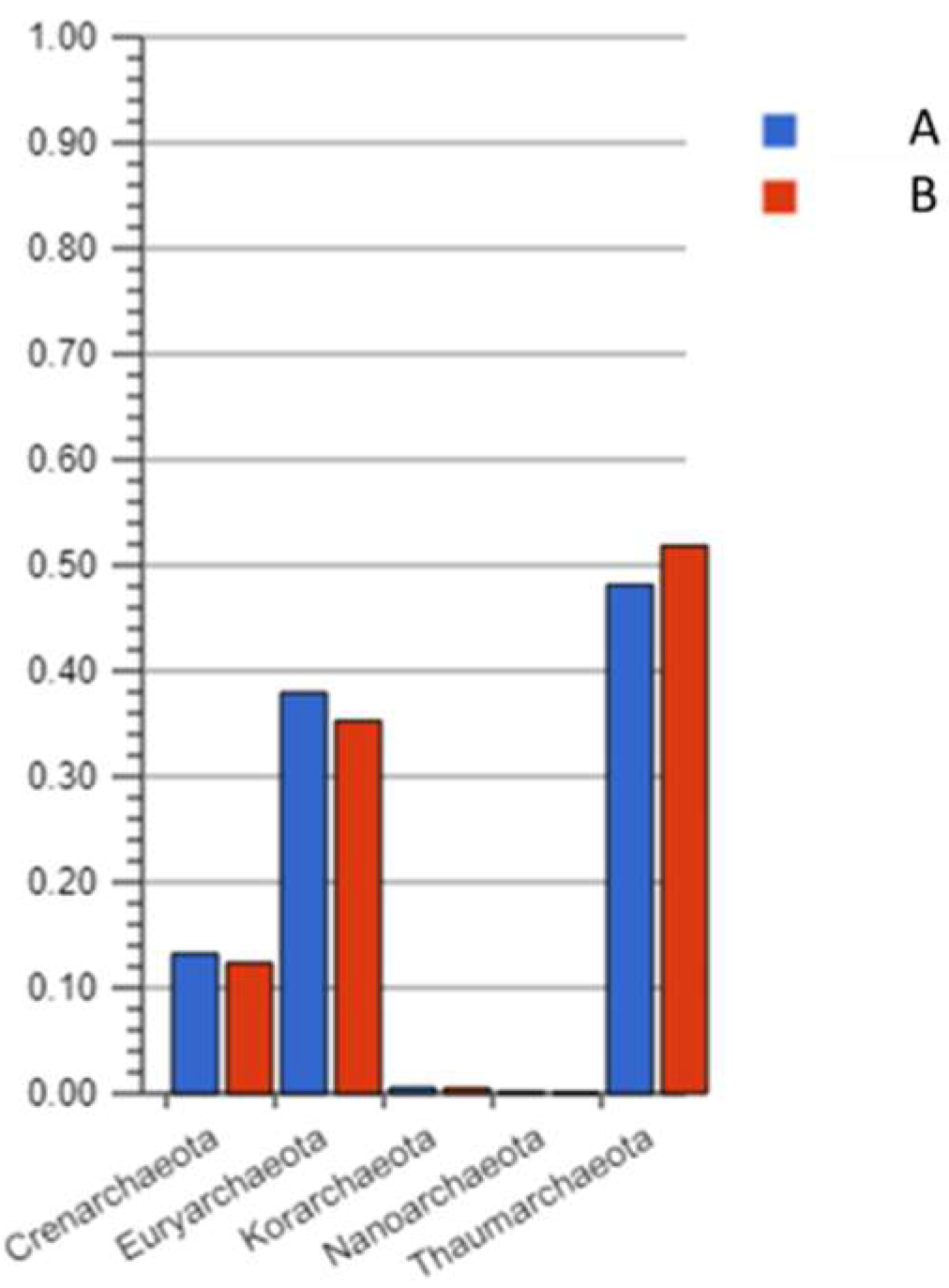
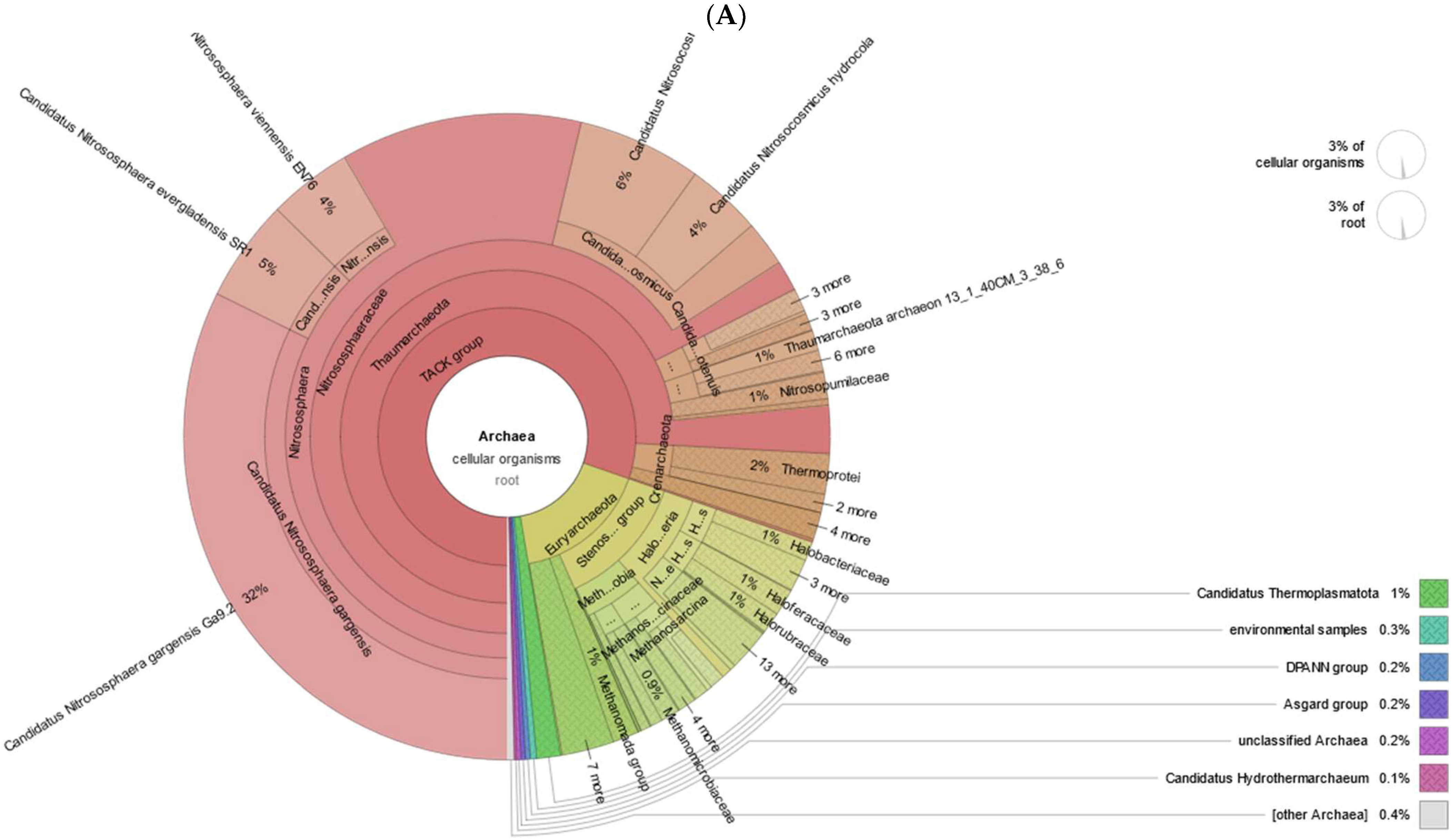
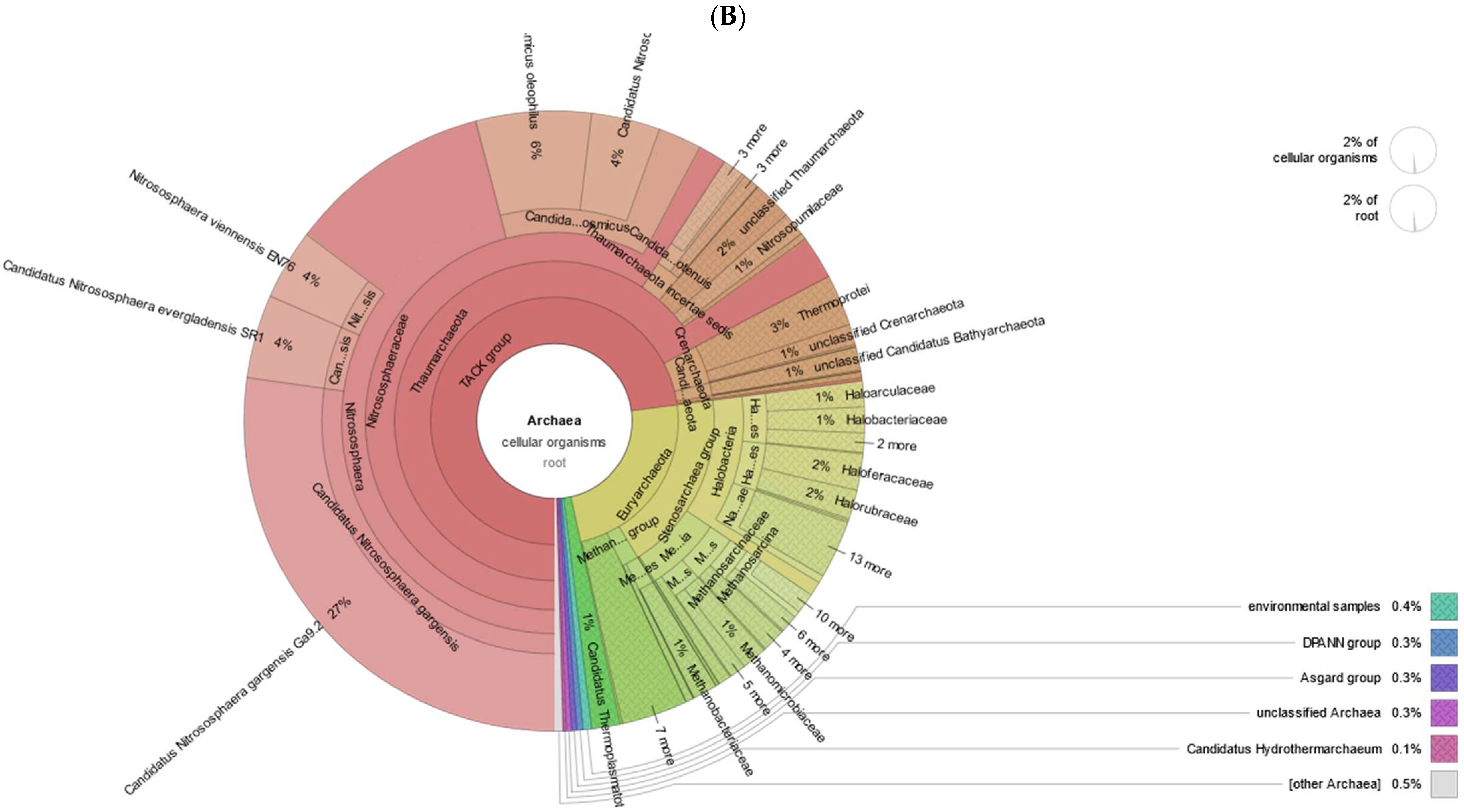
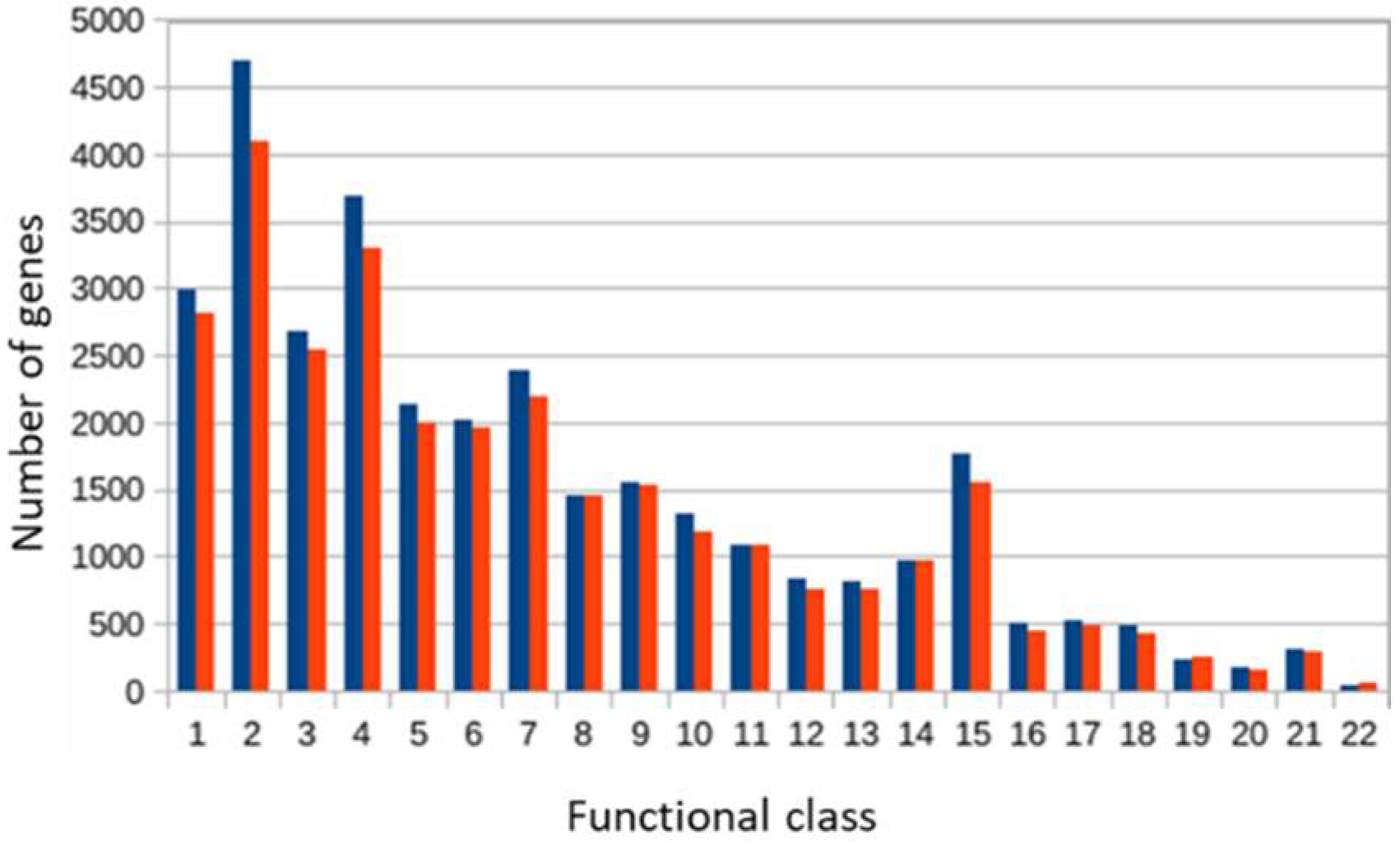
3.1. Taxonomic and Functional Data Classification
3.2. Reconstruction of Metagenome-Assembled Genomes
3.3. Identification of Aromatic Hydrocarbon-Degrading Coding DNA Sequences
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Grimmer, G. PAH—Their contribution to the carcinogenicity of various emissions. Toxicol. Environ. Chem. 1985, 10, 171–181. [Google Scholar] [CrossRef]
- Hodson, P.V. The toxicity to fish embryos of PAH in crude and refined oils. Arch. Environ. Contam. Toxicol. 2017, 73, 12–18. [Google Scholar] [CrossRef]
- Chang, Y.; Siddens, L.K.; Heine, L.K.; Sampson, D.A.; Yu, Z.; Fischer, K.A.; Löhr, C.V.; Tilton, S.C. Comparative mechanisms of PAH toxicity by benzo[a]pyrene and dibenzo [def, p] chrysene in primary human bronchial epithelial cells cultured at air-liquid interface. Toxicol. Appl. Pharmacol. 2019, 379, 114644. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Luqueño, F.; Valenzuela-Encinas, C.; Marsch, R.; Martínez-Suárez, C.; Vázquez-Núñez, E.; Dendooven, L. Microbial communities to mitigate contamination of PAHs in soil—Possibilities and challenges: A review. Environ. Sci. Pollut. Res. 2011, 18, 12–30. [Google Scholar] [CrossRef]
- Sushkova, S.; Minkina, T.; Dudnikova, T.; Barbashev, A.; Mazarji, M.; Chernikova, N.; Lobzenko, I.; Deryabkina, I.; Kizilkaya, R. Influence of carbon-containing and mineral sorbents on the toxicity of soil contaminated with benzo[a]pyrene during phytotesting. Environ. Geochem. Health 2021, 44, 179–193. [Google Scholar] [CrossRef] [PubMed]
- Minkina, T.; Vasilyeva, G.; Popileshko, Y.; Bauer, T.; Sushkova, S.; Fedorenko, A.; Antonenko, E.; Pinskii, D.; Mazarji, M.; Ferreira, C.S.S. Sorption of benzo [a] pyrene by Chernozem and carbonaceous sorbents: Comparison of kinetics and interaction mechanisms. Environ. Geochem. Health 2021, 44, 133–148. [Google Scholar] [CrossRef] [PubMed]
- DeBruyn, J.M.; Mead, T.J.; Sayler, G.S. Horizontal transfer of PAH catabolism genes in Mycobacterium: Evidence from comparative genomics and isolated pyrene-degrading bacteria. Environ. Sci. Technol. 2012, 46, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Minkina, T.; Fedorenko, A.; Hassan, T.; Fedorenko, G.; Chaplygin, V.; Chernikova, N.; Nevidomskaya, D.; Sushkova, S. Morphological Studies of Tyhpa Australis under Stress Environmental Factor; IOP Publishing: Bristol, UK, 2021; p. 012210. [Google Scholar]
- Haritash, A.K. A comprehensive review of metabolic and genomic aspects of PAH-degradation. Arch. Microbiol. 2020, 202, 2033–2058. [Google Scholar]
- Tongpim, S.; Pickard, M.A. Growth of Rhodococcus S1 on anthracene. Can. J. Microbiol. 1996, 42, 289–294. [Google Scholar] [CrossRef]
- Liu, T.-T.; Xu, Y.; Liu, H.; Luo, S.; Yin, Y.-J.; Liu, S.-J.; Zhou, N.-Y. Functional characterization of a gene cluster involved in gentisate catabolism in Rhodococcus sp. strain NCIMB 12038. Appl. Microbiol. Biotechnol. 2011, 90, 671–678. [Google Scholar] [CrossRef] [PubMed]
- Kulakova, A.N.; Reid, K.A.; Larkin, M.J.; Allen, C.C.; Kulakov, L.A. Isolation of Rhodococcus rhodochrous NCIMB13064 derivatives with new biodegradative abilities. FEMS Microbiol. Lett. 1996, 145, 227–231. [Google Scholar] [CrossRef] [PubMed]
- Kulakov, L.A.; Chen, S.; Allen, C.C.; Larkin, M.J. Web-type evolution of Rhodococcus gene clusters associated with utilization of naphthalene. Appl. Environ. Microbiol. 2005, 71, 1754–1764. [Google Scholar] [CrossRef] [PubMed]
- Kulakov, L.A.; Allen, C.C.; Lipscomb, D.A.; Larkin, M.J. Cloning and characterization of a novel cis-naphthalene dihydrodiol dehydrogenase gene (narB) from Rhodococcus sp. NCIMB12038. FEMS Microbiol. Lett. 2000, 182, 327–331. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-L.; Shen, F.-T.; Tan, C.-C.; Huang, C.-C.; Chen, B.-Y.; Arun, A.; Young, C.-C. Characterization of Gordonia sp. strain CC-NAPH129-6 capable of naphthalene degradation. Microbiol. Res. 2012, 167, 395–404. [Google Scholar] [CrossRef]
- Fedorenko, A.G.; Chernikova, N.; Minkina, T.; Sushkova, S.; Dudnikova, T.; Antonenko, E.; Fedorenko, G.; Bauer, T.; Mandzhieva, S.; Barbashev, A. Effects of benzo[a]pyrene toxicity on morphology and ultrastructure of Hordeum sativum. Environ. Geochem. Health 2021, 43, 1551–1562. [Google Scholar] [CrossRef]
- Gorovtsov, A.; Demin, K.; Sushkova, S.; Minkina, T.; Grigoryeva, T.; Dudnikova, T.; Barbashev, A.; Semenkov, I.; Romanova, V.; Laikov, A. The effect of combined pollution by PAHs and heavy metals on the topsoil microbial communities of Spolic Technosols of the lake Atamanskoe, Southern Russia. Environ. Geochem. Health 2021, 44, 1299–1315. [Google Scholar] [CrossRef]
- Minkina, T.; Konstantinova, E.; Bauer, T.; Mandzhieva, S.; Sushkova, S.; Chaplygin, V.; Burachevskaya, M.; Nazarenko, O.; Kizilkaya, R.; Gülser, C. Environmental and human health risk assessment of potentially toxic elements in soils around the largest coal-fired power station in Southern Russia. Environ. Geochem. Health 2021, 43, 2285–2300. [Google Scholar] [CrossRef]
- Mishra, S.; Lin, Z.; Pang, S.; Zhang, W.; Bhatt, P.; Chen, S. Recent advanced technologies for the characterization of xenobiotic-degrading microorganisms and microbial communities. Front. Bioeng. Biotechnol. 2021, 9, 31. [Google Scholar] [CrossRef]
- Sushkova, S.N.; Yakovleva, E.V.; Minkina, T.M.; Gabov, D.N.; Antonenko, E.M.; Dudnikova, T.S.; Rajput, V.D. Accumulation of benzo[a]pyrene in plants of different species and organogenic horizon of soils of steppe phytocenosis under technogenic pollution. [НАКОПЛЕНИЕ БЕНЗ[А]ПИРЕНА В РАСТЕНИЯХ РАЗНЫХ ВИДОВ И ОРГАНОГЕННОМ ГОРИЗОНТЕ ПОЧВ СТЕПНЫХ ФИТОЦЕНОЗОВ ПРИ ТЕХНОГЕННОМ ЗАГРЯЗНЕНИИ]. Bull. Tomsk Polytech. Univ. Geo Assets Eng. 2021, 331, 200–214. [Google Scholar] [CrossRef]
- Quince, C.; Walker, A.W.; Simpson, J.T.; Loman, N.J.; Segata, N. Shotgun metagenomics, from sampling to analysis. Nat. Biotechnol. 2017, 35, 833–844. [Google Scholar] [CrossRef] [PubMed]
- Rajput, V.D.; Gorovtsov, A.V.; Fedorenko, G.M.; Minkina, T.M.; Fedorenko, A.G.; Lysenko, V.S.; Sushkova, S.S.; Mandzhieva, S.S.; Elinson, M.A. The influence of application of biochar and metal-tolerant bacteria in polluted soil on morpho-physiological and anatomical parameters of spring barley. Environ. Geochem. Health 2021, 43, 1477–1489. [Google Scholar] [CrossRef] [PubMed]
- ISO 10381-1; Soil Quality—Sampling. Part 1: Guidance on the Design of Sampling Programmes. International Organization for Standardization: Geneva, Switzerland, 2002.
- Directive document 52.10.556-95. Methodical Instructions. In Definition of Polluting Substances in Sediments and Suspension; Roshydromet: Moscow, Russia, 2002. (In Russian) [Google Scholar]
- Sushkova, S.; Minkina, T.; Turina, I.; Mandzhieva, S.; Bauer, T.; Kizilkaya, R.; Zamulina, I. Monitoring of benzo[a]pyrene content in soils under the effect of long-term technogenic poluttion. J. Geochem. Explor. 2017, 174, 100–106. [Google Scholar] [CrossRef]
- Sushkova, S.; Minkina, T.; Tarigholizadeh, S.; Antonenko, E.; Konstantinova, E.; Gülser, C.; Dudnikova, T.; Barbashev, A.; Kizilkaya, R. PAHs accumulation in soil-plant system of Phragmites australis Cav. in soil under long-term chemical contamination. Eurasian J. Soil Sci. 2020, 9, 242–253. [Google Scholar] [CrossRef]
- Procedure of Measurements Benz(a)pyrene Content in Soils, Sediments, and Sludges by Highly Effective Liquid Chromatography Method; Certificate 27-08; Rosstandart: Moscow, Russia, 2008; p. 27. (In Russian)
- ISO 13859-2014; Soil Quality. Determination of Polycyclic Aromatic Hydrocarbons (PAH) by Gas Chromatography (GC) and High Performance Liquid Chromatography (HPLC). ISO: Geneva, Switzerland, 2014.
- Rotmistrovsky, K.; Agarwala, R. BMTagger: Best Match Tagger for Removing Human Reads from Metagenomics Datasets. 2011. Available online: ftp://ftp.ncbi.nlm.nih.gov/pub/agarwala/bmtagger/ (accessed on 8 June 2021).
- Sourceforge.net. Available online: https://sourceforge.net/projects/bbmap/ (accessed on 20 June 2021).
- Bioinformatics.babraham.ac.uk. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 14 May 2021).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.; Korobeynikov, A.; Lapidus, A.; Prjibelsky, A.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling genomes and mini-metagenomes from highly chimeric reads. In Annual International Conference on Research in Computational Molecular Biology; Springer: Berlin, Heidelberg, 2013; pp. 158–170. [Google Scholar] [CrossRef]
- Hyatt, D.; Chen, G.L.; LoCascio, P.F.; Land, M.L.; Larimer, F.W.; Hauser, L.J. Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010, 11, 1–11. [Google Scholar] [CrossRef]
- Buchfink, B.; Huson, D.H.; Xie, C. MetaScope-Fast and accurate identification of microbes in metagenomic sequencing data. arXiv 2015, arXiv:1511.08753. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Menzel, P.; Ng, K.L.; Krogh, A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat. Commun. 2016, 7, 1–9. [Google Scholar] [CrossRef]
- Ondov, B.D.; Bergman, N.H.; Phillippy, A.M. Interactive metagenomic visualization in a Web browser. BMC Bioinform. 2011, 12, 1–10. [Google Scholar] [CrossRef]
- Zhang, P.; Chen, Y. Polycyclic aromatic hydrocarbons contamination in surface soil of China: A review. Sci. Total Environ. 2017, 605, 1011–1020. [Google Scholar] [CrossRef]
- Sosa, D.; Hilber, I.; Faure, R.; Bartolomé, N.; Fonseca, O.; Keller, A.; Schwab, P.; Escobar, A.; Bucheli, T.D. Polycyclic aromatic hydrocarbons and polychlorinated biphenyls in soils of Mayabeque, Cuba. Environ. Sci. Pollut. Res. 2017, 24, 12860–12870. [Google Scholar] [CrossRef] [PubMed]
- Gabov, D.; Yakovleva, E.; Vasilevich, R. Vertical distribution of PAHs during the evolution of permafrost peatlands of the European arctic zone. Appl. Geochem. 2020, 123, 104790. [Google Scholar] [CrossRef]
- Pikovskii, Y.I.; Smirnova, M.; Gennadiev, A.; Zavgorodnyaya, Y.A.; Zhidkin, A.; Kovach, R.; Koshovskii, T. Parameters of the native hydrocarbon status of soils in different bioclimatic zones. Eurasian Soil Sci. 2019, 52, 1333–1346. [Google Scholar] [CrossRef]
- GN 2.1.7.2041-06; Maximum permissible concentration (MPC) of chemicals in soils. Rospotrebnadzor: Moscow, Russia, 2006; p. 15. (In Russian)
- Soukarieh, B.; El Hawari, K.; El Husseini, M.; Budzinski, H.; Jaber, F. Impact of Lebanese practices in industry, agriculture and urbanization on soil toxicity. Eval. Polycycl. Aromat. Hydrocarb. (PAHs) Levels Soil. Chemosphere 2018, 210, 85–92. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, Q.; Xie, S. Molecular characterization of phenanthrene-degrading methanogenic communities in leachate-contaminated aquifer sediment. Int. J. Environ. Sci. Technol. 2012, 9, 705–712. [Google Scholar] [CrossRef]
- Krzmarzick, M.J.; Taylor, D.K.; Fu, X.; McCutchan, A.L. Diversity and niche of archaea in bioremediation. Archaea 2018, 2018, 3194108. [Google Scholar] [CrossRef]
- Rochman, F.F.; Sheremet, A.; Tamas, I.; Saidi-Mehrabad, A.; Kim, J.-J.; Dong, X.; Sensen, C.W.; Gieg, L.M.; Dunfield, P.F. Benzene and naphthalene degrading bacterial communities in an oil sands tailings pond. Front. Microbiol. 2017, 8, 1845. [Google Scholar] [CrossRef]
- Benedek, T.; Szentgyörgyi, F.; Szabó, I.; Farkas, M.; Duran, R.; Kriszt, B.; Táncsics, A. Aerobic and oxygen-limited naphthalene-amended enrichments induced the dominance of Pseudomonas spp. from a groundwater bacterial biofilm. Appl. Microbiol. Biotechnol. 2020, 104, 6023–6043. [Google Scholar] [CrossRef]
- Lee, Y.; Lee, Y.; Jeon, C.O. Biodegradation of naphthalene, BTEX, and aliphatic hydrocarbons by Paraburkholderia aromaticivorans BN5 isolated from petroleum-contaminated soil. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef]
- Weissenfels, W.D.; Beyer, M.; Klein, J. Degradation of phenanthrene, fluorene and fluoranthene by pure bacterial cultures. Appl. Microbiol. Biotechnol. 1990, 32, 479–484. [Google Scholar] [CrossRef] [PubMed]
- Willumsen, P.A.; Arvin, E. Kinetics of degradation of surfactant-solubilized fluoranthene by a Sphingomonas paucimobilis. Environ. Sci. Technol. 1999, 33, 2571–2578. [Google Scholar] [CrossRef]
- Reddy, P.V.; Karegoudar, T.; Monisha, T.; Mukram, I.; Nayak, A.S. Biodegradation of fluoranthene by Paenibacillus sp. strain PRNK-6: A pathway for complete mineralization. Arch. Microbiol. 2018, 200, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Y.; Luan, T.; Lin, L.; Liu, H.; Tam, N.F. Production of metabolites in the biodegradation of phenanthrene, fluoranthene and pyrene by the mixed culture of Mycobacterium sp. and Sphingomonas sp. Bioresour. Technol. 2011, 102, 2965–2972. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.-x.; Hu, X.; Cao, Y.; Pang, W.-j.; Huang, J.-y.; Guo, P.; Huang, L. Biodegradation of phenanthrene and heavy metal removal by acid-tolerant Burkholderia fungorum FM-2. Front. Microbiol. 2019, 10, 408. [Google Scholar] [CrossRef]
- Bankole, P.O.; Semple, K.T.; Jeon, B.-H.; Govindwar, S.P. Biodegradation of fluorene by the newly isolated marine-derived fungus, Mucor irregularis strain bpo1 using response surface methodology. Ecotoxicol. Environ. Saf. 2021, 208, 111619. [Google Scholar] [CrossRef]
- Pinyakong, O.; Habe, H.; Omori, T. The unique aromatic catabolic genes in sphingomonads degrading polycyclic aromatic hydrocarbons (PAHs). J. Gen. Appl. Microbiol. 2003, 49, 1–19. [Google Scholar] [CrossRef]
- Wu, P.; Wang, Y.-S. Fluorene degradation by Rhodococcus sp. A2-3 isolated from hydrocarbon contaminated sediment of the Pearl River estuary, China. Ecotoxicology 2021, 30, 929–935. [Google Scholar] [CrossRef]
- Wang, L.; Ji, G.; Huang, S. Contribution of the Kodama and 4S pathways to the dibenzothiophene biodegradation in different coastal wetlands under different C/N ratios. Res. J. Environ. Sci. 2019, 76, 217–226. [Google Scholar] [CrossRef]
- Jové, P.; Olivella, M.À.; Camarero, S.; Caixach, J.; Planas, C.; Cano, L.; De Las Heras, F.X. Fungal biodegradation of anthracene-polluted cork: A comparative study. J. Environ. Sci. Health A 2016, 51, 70–77. [Google Scholar] [CrossRef]
- Bhatt, K.K.; Lily, M.K.; Joshi, G.; Dangwal, K. Benzo[a]pyrene degradation pathway in Bacillus subtilis BMT4i (MTCC 9447). Turk. J. Biochem. 2018, 43, 693–701. [Google Scholar] [CrossRef]
- Johnsen, A.R.; Wick, L.Y.; Harms, H. Principles of microbial PAH-degradation in soil. Environ. Pollut. 2005, 133, 71–84. [Google Scholar] [CrossRef]
- Cochrane, R.V.; Vederas, J.C. Highly selective but multifunctional oxygenases in secondary metabolism. Acc. Chem. Res. 2014, 47, 3148–3161. [Google Scholar] [CrossRef] [PubMed]
- Musumeci, M.A.; Loviso, C.L.; Lozada, M.; Ferreira, F.V.; Dionisi, H.M. Substrate specificities of aromatic ring-hydroxylating oxygenases of an uncultured gammaproteobacterium from chronically-polluted subantarctic sediments. Int. Biodeterior. Biodegrad. 2019, 137, 127–136. [Google Scholar] [CrossRef]
- AbuBakr, S.; Macmil, S.L.; Nanny, M.A.; Duncan, K.E. Enzymatic transformation of humic substances by NDO. Soil Biol. Biochem. 2008, 40, 2055–2062. [Google Scholar] [CrossRef]
- Wu, Y.; Teng, Y.; Li, Z.; Liao, X.; Luo, Y. Potential role of polycyclic aromatic hydrocarbons (PAHs) oxidation by fungal laccase in the remediation of an aged contaminated soil. Soil Biol. Biochem. 2008, 40, 789–796. [Google Scholar] [CrossRef]
- Kumar, A.; Schimmelmann, A.; Sauer, P.E.; Brassell, S.C. Distribution and sources of polycyclic aromatic hydrocarbons (PAHs) in laminated Santa Barbara Basin sediments. Org. Geochem. 2017, 113, 303–314. [Google Scholar] [CrossRef]
- Holman, A.I.; Grice, K. δ13C of aromatic compounds in sediments, oils and atmospheric emissions: A review. Org. Geochem. 2018, 123, 27–37. [Google Scholar] [CrossRef]
- Gao, P.; Li, H.; Wilson, C.P.; Townsend, T.G.; Xiang, P.; Liu, Y.; Ma, L.Q. Source identification of PAHs in soils based on stable carbon isotopic signatures. Crit. Rev. Environ. Sci. Technol. 2018, 48, 923–948. [Google Scholar] [CrossRef]
- Bakhtiari, A.R.; Zakaria, M.P.; Yaziz, M.I.; Lajis, M.N.H.; Bi, X.; Abd Rahim, M.C. Vertical distribution and source identification of polycyclic aromatic hydrocarbons in anoxic sediment cores of Chini Lake, Malaysia: Perylene as indicator of land plant-derived hydrocarbons. Appl. Geochem. 2009, 24, 1777–1787. [Google Scholar] [CrossRef]
- Wakeham, S.G.; Canuel, E.A. Biogenic polycyclic aromatic hydrocarbons in sediments of the San Joaquin River in California (USA), and current paradigms on their formation. Environ. Sci. Pollut. Res. 2016, 23, 10426–10442. [Google Scholar] [CrossRef] [PubMed]
- Bixiong, Y.; Zhihuan, Z.; Ting, M. Pollution sources identification of polycyclic aromatic hydrocarbons of soils in Tianjin area, China. Chemosphere 2006, 64, 525–534. [Google Scholar] [CrossRef] [PubMed]
- Lima, E.A.; Neves, P.A.; Patchineelam, S.R.; da Silva, J.F.B.; Takiyama, L.R.; Martins, C.C.; Lourenço, R.A.; Taniguchi, S.; Elias, V.O.; Bícego, M.C. Anthropogenic and natural inputs of polycyclic aromatic hydrocarbons in the sediment of three coastal systems of the Brazilian Amazon. Environ. Sci. Pollut. Res. 2021, 28, 19485–19496. [Google Scholar] [CrossRef] [PubMed]
- Cabrerizo, A.; Tejedo, P.; Dachs, J.; Benayas, J. Anthropogenic and biogenic hydrocarbons in soils and vegetation from the South Shetland Islands (Antarctica). Sci. Total Environ. 2016, 569, 1500–1509. [Google Scholar] [CrossRef] [PubMed]
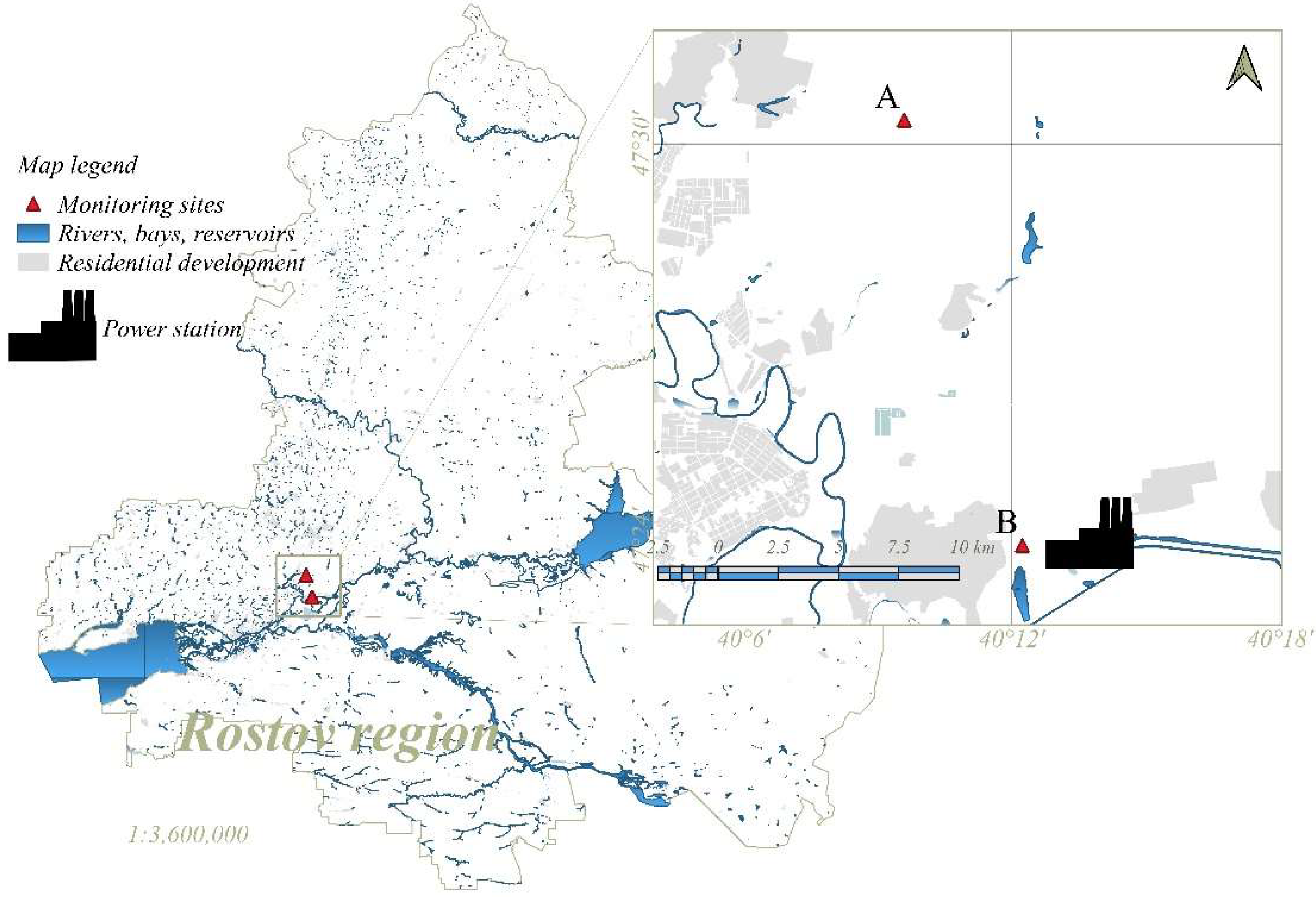
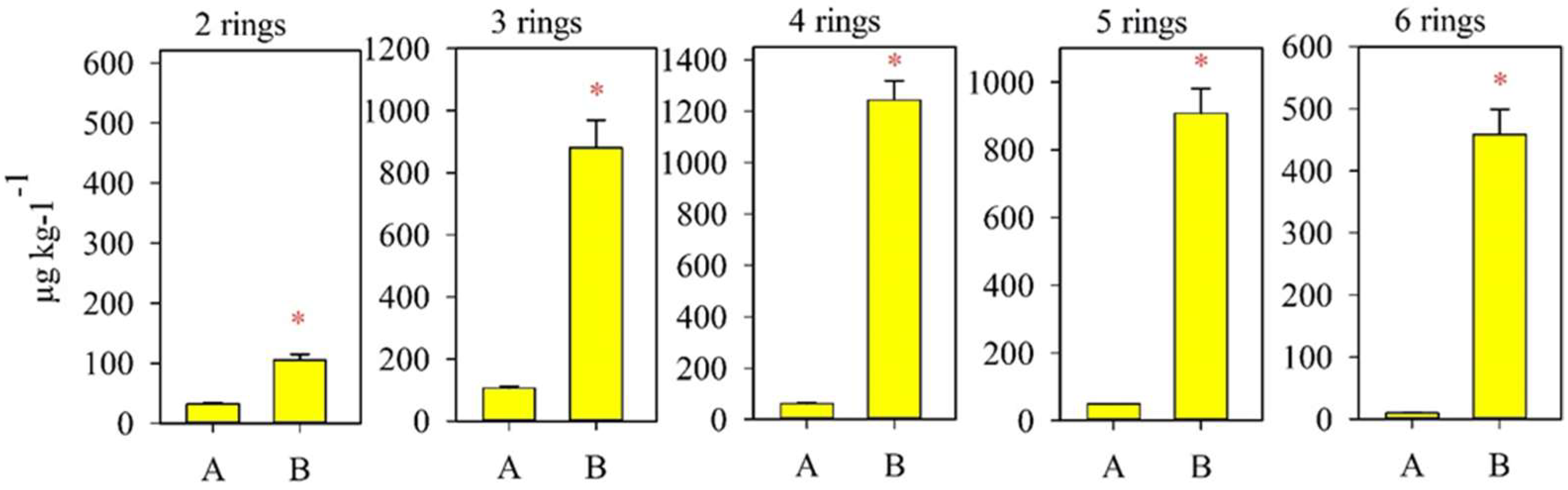
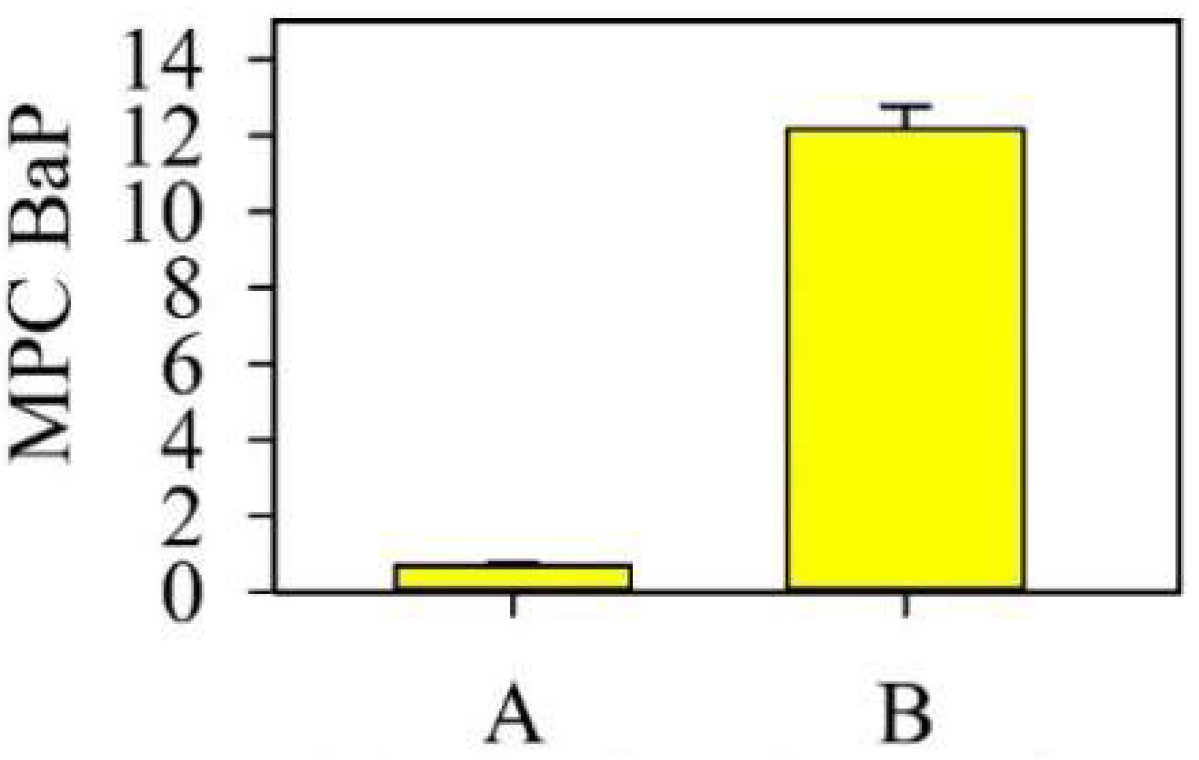
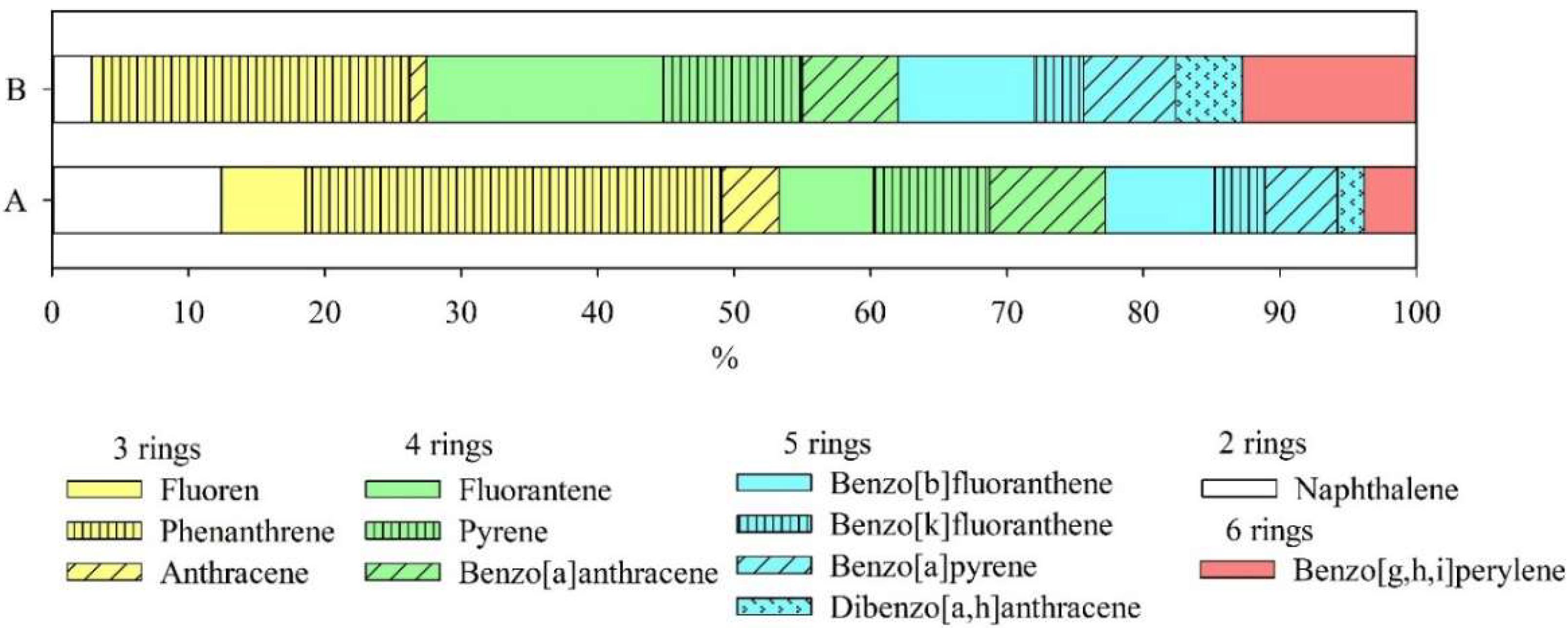
| Monitoring Sites | Soil Type | pH | Physical Clay Content (Particle ˂ 0.01 mm) | Silt (Particle ˂ 0.001 mm) | Corg,% |
|---|---|---|---|---|---|
| A | Haplic Chernozem | 7.57 ± 0.12 | 48.1 ± 1.1 | 28.6 ± 0.5 | 3.7 ± 0.2 |
| B | 7.62 ± 0.10 | 58.4 ± 1.0 | 31.2 ± 0.2 | 2.6 ± 0.2 |
| Soil Samples | Readings before QC | Readings after QC |
|---|---|---|
| A | 79,329,568 | 71,292,041 |
| B | 92,191,436 | 83,604,114 |
| Characteristics | A | B |
|---|---|---|
| The number of contigs in the assembly (>500 bp) | 49,461 | 56,636 |
| First contig length, bp | 76,132 | 26,183 |
| Affiliation of the first contig according to RefSeq database | Candidatus Nitrososphaera gargensis Ga9.2 | Bradyrhizobium sp. CCBAU 53421 |
| Sample | Alpha-Diversity |
|---|---|
| A | 378 |
| B | 372 |
| Samples | Genes Predicted | Genes Functionally Annotated | Genes Functionally Annotated, % |
|---|---|---|---|
| A | 90,889 | 30,457 | 33.5 |
| B | 90,520 | 32,865 | 36.3 |
| Gene Types | Gene Function Code | A | B |
|---|---|---|---|
| nahY (chemotaxis protein) | K03406 | 6 | 3 |
| nagA (naphthalene dioxygenase) | K01443 | 12 | 5 |
| nagG_1 (salicylate hydroxylase) | K00499 | 4 | 4 |
| Rieske 2Fe-2S domain-containing protein | K05710 | 12 | 8 |
| homogentisate 1,2-dioxygenase | K00451 | 5 | 8 |
| FAD dependent oxidoreductase | K00111 | 16 | 15 |
| Number of genes with assigned KO number | 30,457 | 32,865 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Delegan, Y.; Sushkova, S.; Minkina, T.; Filonov, A.; Kocharovskaya, Y.; Demin, K.; Gorovtsov, A.; Rajput, V.D.; Zamulina, I.; Grigoryeva, T.; et al. Diversity and Metabolic Potential of a PAH-Degrading Bacterial Consortium in Technogenically Contaminated Haplic Chernozem, Southern Russia. Processes 2022, 10, 2555. https://doi.org/10.3390/pr10122555
Delegan Y, Sushkova S, Minkina T, Filonov A, Kocharovskaya Y, Demin K, Gorovtsov A, Rajput VD, Zamulina I, Grigoryeva T, et al. Diversity and Metabolic Potential of a PAH-Degrading Bacterial Consortium in Technogenically Contaminated Haplic Chernozem, Southern Russia. Processes. 2022; 10(12):2555. https://doi.org/10.3390/pr10122555
Chicago/Turabian StyleDelegan, Yanina, Svetlana Sushkova, Tatiana Minkina, Andrey Filonov, Yulia Kocharovskaya, Konstantin Demin, Andrey Gorovtsov, Vishnu D. Rajput, Inna Zamulina, Tatiana Grigoryeva, and et al. 2022. "Diversity and Metabolic Potential of a PAH-Degrading Bacterial Consortium in Technogenically Contaminated Haplic Chernozem, Southern Russia" Processes 10, no. 12: 2555. https://doi.org/10.3390/pr10122555
APA StyleDelegan, Y., Sushkova, S., Minkina, T., Filonov, A., Kocharovskaya, Y., Demin, K., Gorovtsov, A., Rajput, V. D., Zamulina, I., Grigoryeva, T., Dudnikova, T., Barbashev, A., & Maksimov, A. (2022). Diversity and Metabolic Potential of a PAH-Degrading Bacterial Consortium in Technogenically Contaminated Haplic Chernozem, Southern Russia. Processes, 10(12), 2555. https://doi.org/10.3390/pr10122555










