Abstract
Leaf litter is an essential functional aspect of forest ecosystems, acting as a source of organic matter, a protective layer in forest soils, and a nurturing habitat for micro- and macro-organisms. Through their successional occurrence, litter-inhabiting microfungi play a key role in litter decomposition and nutrient recycling. Despite their importance in terrestrial ecosystems and their abundance and diversity, information on the taxonomy, diversity, and host preference of these decomposer taxa is scarce. This study aims to clarify the taxonomy and phylogeny of four saprobic fungal taxa inhabiting Dipterocarpus alatus leaf litter. Leaf litter samples were collected from Doi Inthanon National Park in Chiang Mai, northern Thailand. Fungal isolates were characterized based on morphology and molecular phylogeny of the nuclear ribosomal DNA (ITS, LSU) and protein-coding genes (tub2, tef1-α, rpb2). One novel saprobic species, Ciliochorella dipterocarpi, and two new host records, Pestalotiopsis dracontomelon and Robillarda australiana, are introduced. The newly described taxa are compared with similar species, and comprehensive descriptions, micrographs, and phylogenetic trees are provided.
1. Introduction
Leaf litter is abundantly found in forest ecosystems with a rich source of organic matter [1,2], where fungi act as key players in decomposing leaf litter by the production of extracellular enzymes [3,4,5,6]. Fungi are cosmopolitan organisms that thrive in diverse ecosystems and environments [7,8,9,10,11]. They are involved in fundamental ecological processes as pathogens, mutualists, and decomposers [12,13]. Saprobic fungi are involved in carbon cycling and nutrient recycling [7,8,9,10,11]; mediating the functioning and balance in forest ecosystems [14]. Despite their role and abundance, fungal diversity and its associations with host species are poorly studied; thus, exploring tropical ecosystems for unidentified fungal species has gained attention [15,16,17].
Sordariomycetes is the second-largest class in Ascomycota [18,19,20], with members with a cosmopolitan distribution [21,22]. They act as pathogens, endophytes [23,24], and saprobes [25]. Additionally, some taxa have been reported as opportunistic pathogens [26] and fungicolous [27]. In this group, appendage-bearing coelomycetes containing fusoid multi-septate conidia are circumscribed as pestalotioid fungi [23,28,29] and classified as Sporocadaceae [30]. They are characterized by their asexual morphology: acervular conidiomata, conidiogenesis, and conidia [29]. However, due to the absence of strong morphological markers for some genera, multilocus phylogenetic analyses are essential for their taxonomic placements [29,31,32,33].
Nevertheless, within appendage-bearing fungi, Pestalotiopsis is comprised mostly of asexual morphs, rich with cryptic species [34,35], with the morphological species delimitation taxonomically insignificant [36]. To solve this issue, in the most recent review of resolving the genus taxonomy, Maharachchikumbura et al. [23] used multilocus phylogeny based on internal transcribed spacer (ITS) region, partial translation elongation factor (tef1-α), and β-tubulin (tub2) genes coupled with morphology and set the species boundaries within the genus. On the other hand, Ciliochorella (Xylariomycetidae incertae sedis) and Robillarda (Sporocadaceae) have distinctive morphological characteristics while lacking complete molecular information for most of the species.
Assessing the tropical forest mycobiome to recover novel fungal taxa and document their host associations, life modes, abundance through seasonal variations, and ecological preference is critical in elucidating fungal biodiversity. In contrast, taxonomy to assess biodiversity is also important [37]. The present study aimed to investigate the taxonomy and phylogeny of three saprobic appendage-bearing Sordariomycetes from the leaf litter of Dipterocarpus alatus collected from Doi Inthanon National Park, Thailand. One novel species, Ciliochorella dipterocarpi, and two new host records, Pestalotiopsis dracontomelon and Robillarda australiana, were identified and described using morpho-molecular analysis.
2. Materials and Methods
2.1. Sample Collection and Fungal Isolation
Dipterocarpus alatus leaf litter was collected from Doi Inthanon National Park, Chiang Mai, Thailand. The samples were stored in separate paper bags, brought back to the laboratory, and pressed in between newspaper sheets. After two days, leaf samples were placed in a sterile moisture chamber to favor the emergence of saprobic fungal taxa. The specimens were examined using a dissecting microscope (Motic SMZ-168, Wetzlar, Germany) and a light microscope (Nikon ECLIPSE 80i, Tokyo, Japan). Single-spore isolation was carried out in potato dextrose agar (PDA) plates as described by Senanayake et al. [38]. The obtained cultures were incubated at 25 °C for 2–3 weeks with frequent observation to detect and record the sporulation and colony characteristics.
2.2. Morphological Observation
Freehand sectioning through the conidiomata was performed to observe the morphological characteristics (e.g., conidiomata shape, conidiomatal wall cell structure, conidiophores, conidiogenous cells, and conidia). Colony characteristics were examined and recorded after 7 days of incubation at 25 °C. Fungal fruiting bodies were mounted on water and examined with a light microscope (Nikon Eclipse E600), and digital images were captured with a Nikon DS-U2 and Cannon 750D camera (Tokyo, Japan). All measurements were made using the Tarosoft (R) Image Framework software v.0.9.0.7, and Adobe Photoshop CS6 v. 12.0 (Adobe Systems, San Jose, CA, USA) was used for making photo plates.
2.3. Material Deposition
All the herbarium materials were deposited in the Mae Fah Luang University Herbarium (MFLU), Chiang Rai, Thailand, and living cultures were deposited in the Mae Fah Luang University Culture Collection (MFLUCC). Faces of Fungi and Index Fungorum numbers were obtained as in Jayasiri et al. [39] and Index Fungorum [40]. The obtained sequences were deposited in the GenBank database. The illustrations and descriptions were submitted to the GMS MICROFUNGI database [41]. The novel species was introduced following the guidelines by Maharachchikumbura et al. [42].
2.4. DNA Extraction, PCR Amplification, and Sequencing
Genomic DNA was extracted from the margins of 2-week-old cultures grown on PDA plates at 25 °C using a Forensic DNA Kit (OMEGA, Biotek, Norcross, GA, USA), following the manufacturer’s instructions. Polymerase chain reactions (PCR) were conducted in an Applied Biosystems C1000 Touch TM Thermal Cycler with the PCR conditions specified in Table 1. The PCR reaction mixture was composed of 25 μL of Taq DNA polymerase, 2 μL of genomic DNA, 2 μL (20 μM) of each primer, and 19 μL of double-distilled H2O. PCR products were visualized via agarose gel electrophoresis with 1% agarose gel under the Biorad gel documentation system (Hercules, CA, USA). Sequencing of PCR products was carried out bi-directionally by Biogenomed, Seoul, Korea. All the sequences generated in this study were deposited in GenBank.

Table 1.
Primers and PCR conditions used in this study.
2.5. Phylogenetic Analyses
The obtained sequence chromatograms were checked with Chromas 2.6.6 (Technelysium Pty Ltd., South Brisbane, Australia), and the low-quality regions were trimmed. The sequences were subjected to BLASTn searches against the NCBI nucleotide non-redundant databases, with the option “sequences from type material” selected. Separate data sets were used to perform phylogenetic analyses for Pestalotiopsis, Ciliochorella, and Robillarda. Reference sequences were obtained from recent literature [50,51] and downloaded from GenBank (www.ncbi.nlm.nih.gov/genbank/ (accessed on 18 November 2022)) (Table 2, Table 3 and Table 4). Taxon sampling for Pestalotiopsis was performed by selecting the type strains and a duplicate strain if available from all the recorded species, excluding unverified sequences, to obtain better topology for the phylogenetic tree. The single locus of each data set was aligned by MUSCLE [52] implemented in MEGA (v. 7.0.26), applying the default settings. The aligned sequences were automatically trimmed using trimAl 1.2 [53] under the default -gappyout option for all loci and concatenated using BioEdit v. 7.0.5.2 [54]. Phylogenetic analyses were based on maximum likelihood (ML), maximum parsimony (MP), and Bayesian inference (BI) posterior probability (PP). ML, MP, and BI were performed for Pestalotiopsis, while Ciliochorella, and Robillarda were resolved using model-based methods (ML, BI) following the previous literature [23,29,50]. ML and BI were performed in the CIPRES Science Gateway platform [55]. ML was executed by RAxML-HPC2 on XSEDE v. 8.2.8 [56,57], with 1000 bootstrap replicates under the GTRGAMMA nucleotide substitution model.

Table 2.
Newly generated sequences in this study are indicated in bold. * indicates type strains.

Table 3.
Newly generated sequences in this study are indicated in bold. * indicates type strains.

Table 4.
Newly generated sequences in this study are indicated in bold. * indicate type strains.
Bayesian analysis was performed using MrBayes v. 3.2.7a on XSEDE [58] Markov Chain Monte Carlo (MCMC) sampling to calculate the posterior probabilities [59,60], under the nucleotide evolutionary models calculated by jModelTest v.2.1.6 (Table 5). Then, 2 parallel independent runs with 6 MCMCs were run for 15,000,000 (Pestalotiopsis) and 3,000,000 generations (Ciliochorella and Robillarda), with trees sampled every 1000th generation. Twenty-five percent of the trees representing the burn-in phase were discarded, and the remaining trees were used to calculate the PP in the majority-rule consensus tree.

Table 5.
The nucleotide evolutionary models calculated for Bayesian inference.
Maximum-parsimony trees were generated by PAUP v4.0b10 [59], using the heuristic search option with 1000 random sequence additions, with Maxtrees set to 1000. Branches of zero length were collapsed, and all maximum parsimony trees were saved. Descriptive tree statistics for parsimony-tree length (TL), consistency index (CI), retention index (RI), relative consistency index (RC), and homoplasy index (HI) were calculated following the Kishino–Hasegawa test (KHT) criteria [61].
The phylogenetic trees were visualized and exported using FigTree v.1.4.0 [62], and the phylograms were edited and annotated in Microsoft PowerPoint (2013) and Adobe Photoshop CS6.
3. Results
3.1. Phylogenetic Analyses
The combined alignment (ITS, tef1-α, and tub2) of Pestalotiopsis comprised 146 taxa, including 5 outgroup taxa. The best-scoring ML tree (Figure 1) had an ML optimization likelihood value of −16,987.384789. There were 1,025 distinct alignment patterns with 26.01% undetermined characters or gaps. The estimated base frequencies were as follows: A = 0.238425, C = 0.294335, G = 0.217098, and T = 0.250142; substitution rates, AC = 1.142015, AG = 3.088078, AT = 1.117450, CG = 0.907663, CT = 4.297777, and GT = 1.000000; gamma distribution shape parameter α = 0.286165; and tree length = 1.911747. The MP analysis for the combined dataset had 576 parsimony-informative, 1389 constant, and 259 parsimony-uninformative characters and yielded the single most parsimonious tree with the parameters TL = 2471, CI = 0.492, RI = 0.791, RC = 0.389, and HI = 0.508. The Bayesian analysis calculated the average standard deviation of split frequencies at the end of 15,000,000 MCMC generations with a stop value of 0.009. The ML, MP, and BI trees were similar in topology. The resulting phylogenetic tree from the concatenated alignment resolved our Pestalotiopsis isolate into a novel host record, P. dracontomelon (MFLUCC 22-0119), with strong statistical support (99% ML, 99% MP, 1.00 PP) (Figure 1).


Figure 1.
Maximum-likelihood consensus tree inferred by RAxML analysis of combined ITS, tub2, and tef1-a sequence data for Pestalotiopsis. Bootstrap support values for ML ≥ 60%, MP ≥ 60%, and PP ≥ 0.90 are given at the nodes (ML/MP/PP). The tree is rooted in Neopestalotiopsis magna (MFLUCC 12-0652) and N. saprophytica (MFLUCC 12-0282). Type and ex-type strains are shown in bold, and newly identified strain is in red.
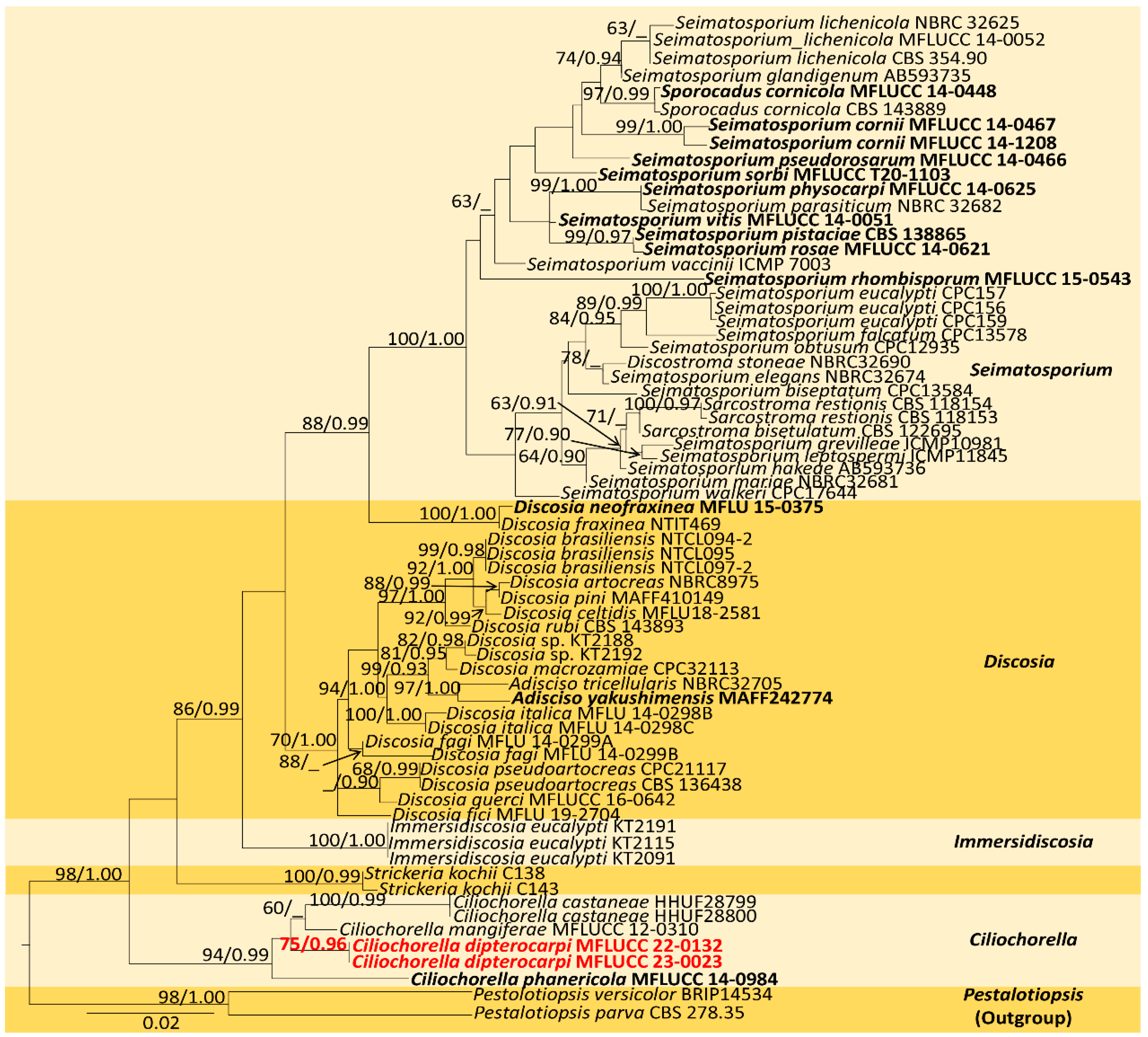
The combined sequence alignment (LSU, ITS, and tub2) of Ciliochorella comprised 71 taxa, including Pestalotiopsis versicolor (BRIP 14534) and P. parva (CBS 278.35) as the outgroup taxa. The matrix had 599 distinct alignment patterns with 34.06% undetermined characters or gaps. The ML analysis of the combined dataset yielded a best-scoring tree with a final ML optimization likelihood value of −8593.236182. The estimated base frequencies were as follows: A = 0.252388, C = 0.224809, G = 0.259858, and T = 0.262945; substitution rates, AC = 1.255639, AG = 2.934468, AT = 1.259383, CG = 1.113458, CT = 5.503206, and GT = 1.000000; gamma distribution shape parameter α = 0.120262; and tree length = 0.748074. In the Bayesian analysis, the average standard deviation of split frequencies at the end of 3,000,000 MCMC generations was calculated with a stop value of 0.009935. The ML and BI trees were similar in topology. The resulting phylogenetic tree from the concatenated alignment resolved C. dipterocarpi (MFLUCC 22-0132) isolate into a well-supported distinct lineage with strong statistical support (94% ML, 0.95 PP) (Figure 2).
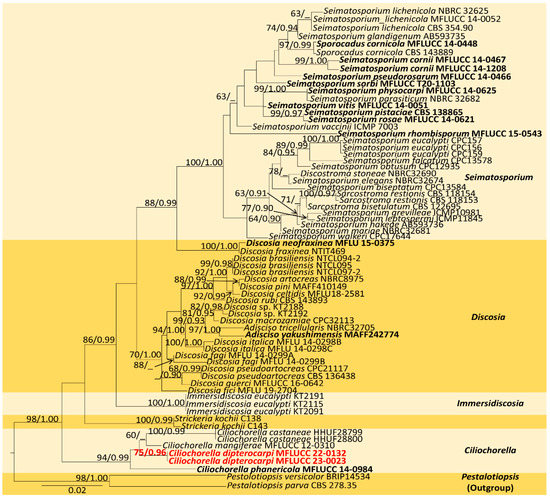
Figure 2.
Maximum-likelihood consensus tree inferred by RAxML analysis of combined LSU, ITS, and tub2 sequence data. Bootstrap support values for ML ≥ 60% and PP ≥ 0.90 are given at the nodes (BS/PP). The tree is rooted with Pestalotiopsis versicolor (BRIP14534) and P. parva (CBS 278.35). The type species are shown in bold, and the newly identified strain is in red.
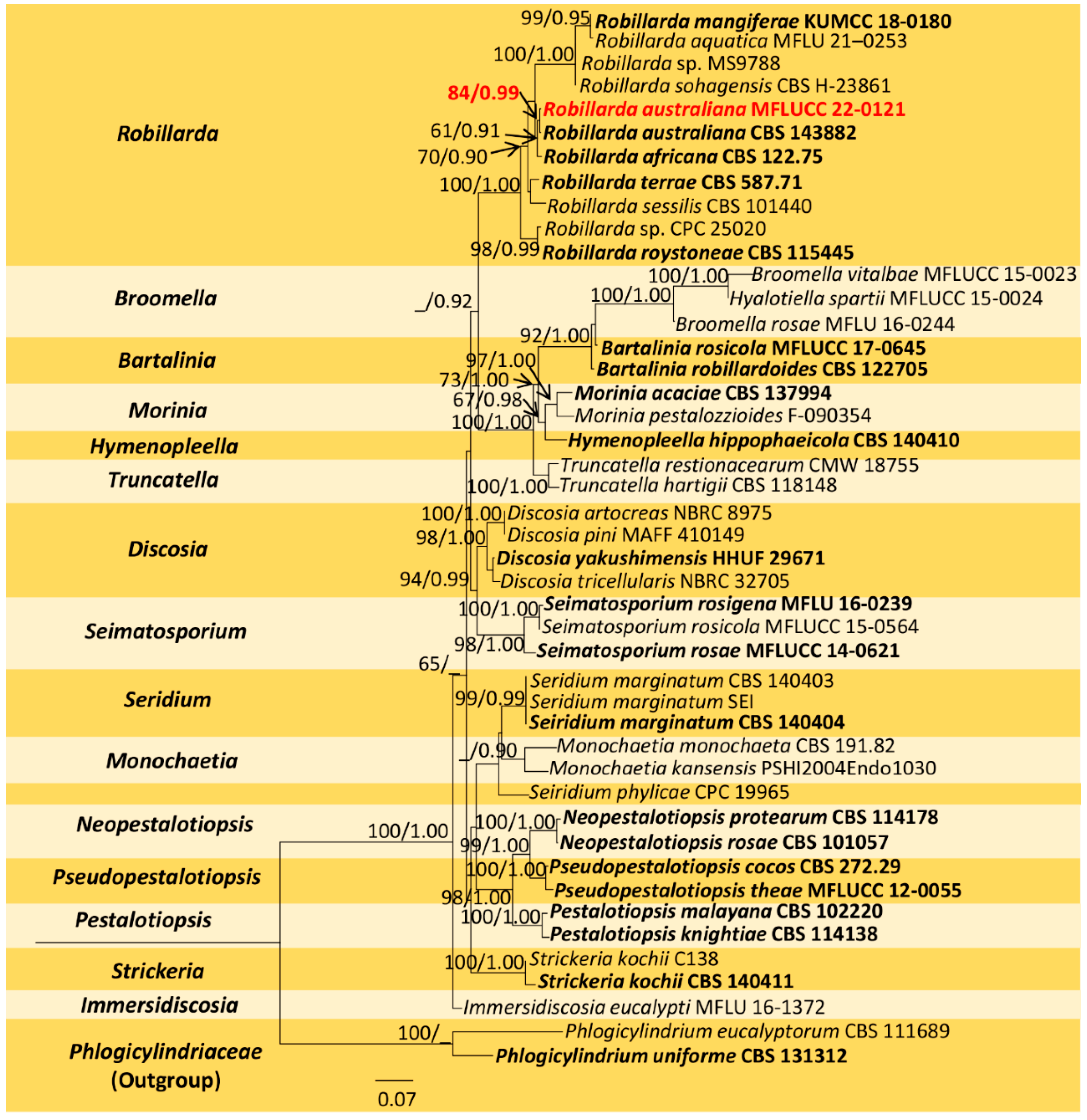
The combined sequence alignment (ITS, LSU, rpb2, and tef1-α) of Robillarda comprised 45 taxa, including Phlogicylindrium eucalyptorum (CBS 111689) and P. uniforme (CBS 131312) as the outgroup taxa. The matrix had 1482 distinct alignment patterns with 44.64% undetermined characters or gaps. The ML analysis of the combined dataset yielded a best-scoring tree with a final ML optimization likelihood value of −26214.966097. The estimated base frequencies were as follows: A = 0.250239, C = 0.240486, G = 0.252222, and T = 0.257053; substitution rates, AC = 1.409959, AG = 3.149767, AT = 1.321223, CG = 1.040419, CT = 5.807309, and GT = 1.000000; gamma distribution shape parameter α = 0.244419; and tree length = 3.825788. In the Bayesian analysis, the average standard deviation of split frequencies at the end of 3,000,000 MCMC generations was calculated with a stop value of 0.009951. The ML and BI trees were similar in topology. The resulting phylogenetic tree from the concatenated alignment grouped the isolate MFLUCC 22-0121 with the ex-type strain R. australiana (CBS 143882) and formed a well-supported clade with strong statistical support (84% ML, 0.99 PP) (Figure 3).
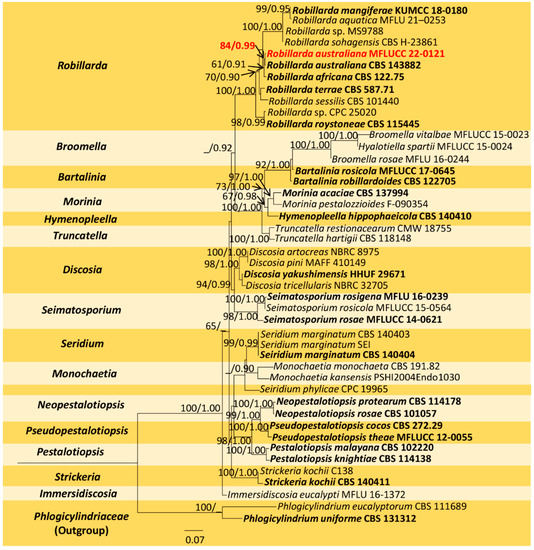
Figure 3.
Maximum-likelihood consensus tree inferred by RAxML analysis of combined ITS, LSU, rpb2, and tef1-a sequence data. Bootstrap support values for ML ≥ 60% and PP ≥ 0.90 are given at the nodes (BS/PP). The tree is rooted with Phlogicylindrium eucalyptorum (CBS 111689) and P. uniforme (CBS 131312). Type species are shown in bold, and the newly generated strain is in red.
3.2. Taxonomy
3.2.1. Pestalotiopsis dracontomelon Maharachch. and K.D. Hyde
Index Fungorum number: IF550943; Faces of Fungi number: FoF00457 (Figure 4)
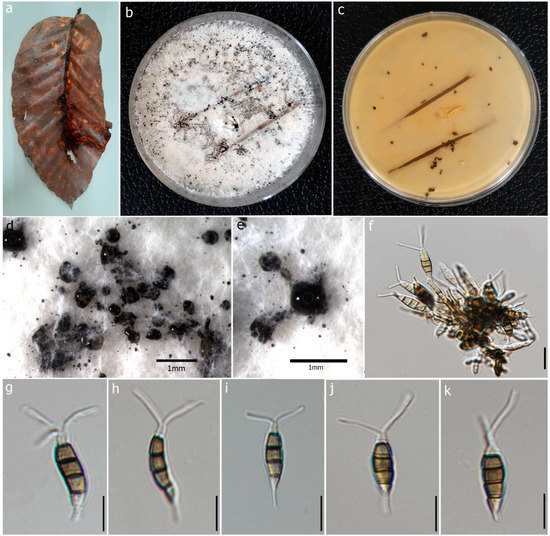

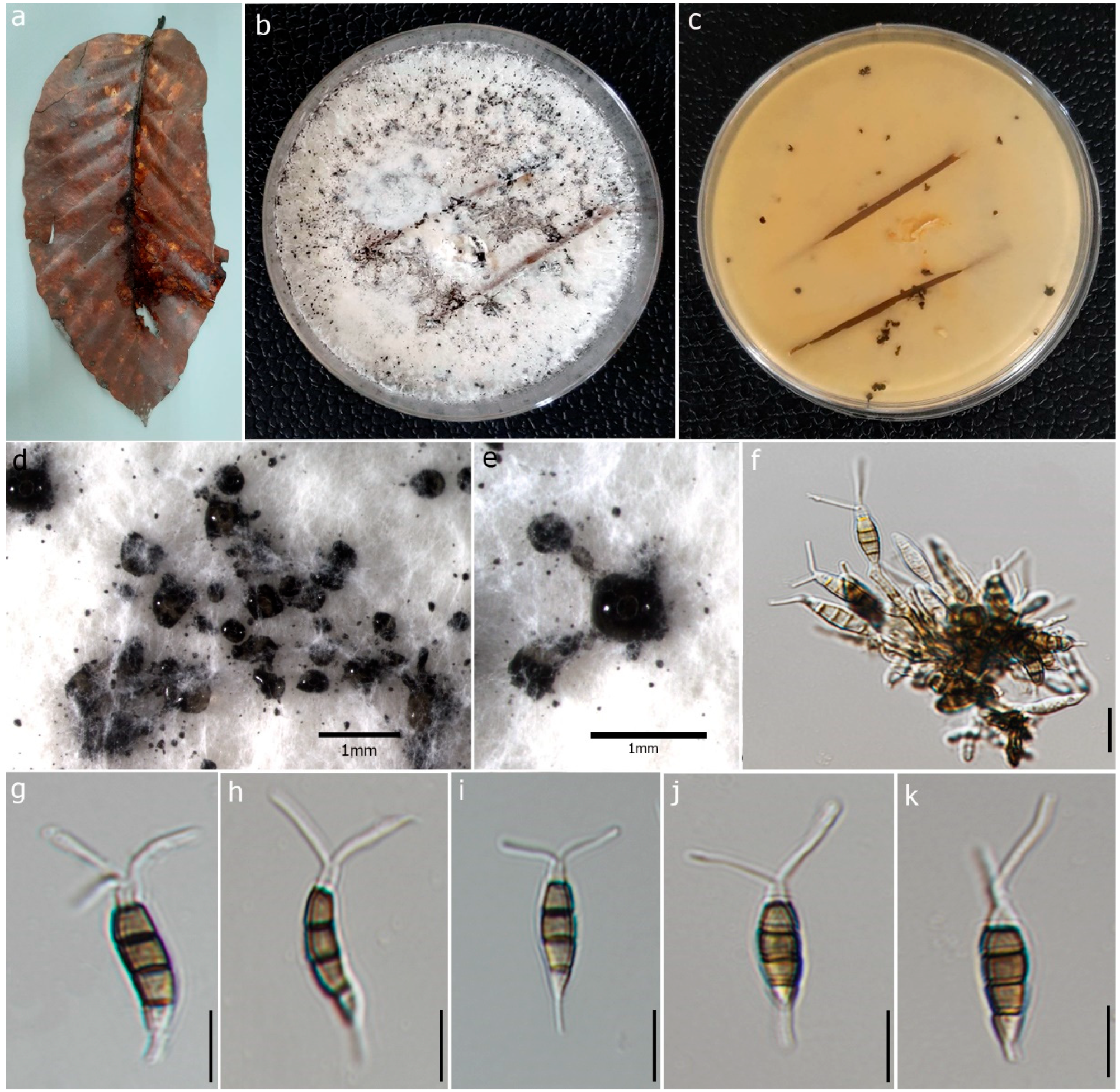
Figure 4.
Pestalotiopsis dracontomelon (MFLU 22-0195, new host record). (a) Dead leaf of Dipterocarpus alatus. (b,c) Upper and reverse view of the colonies on PDA. (d,e) Conidiomata on PDA. (f) Conidiogenous cells with developing conidia. (g–k) Conidia. Scale bars: (d,e) = 1 mm, (f) = 5 μm, (g–k) = 10 μm.
Description: Saprobic on dead leaves of D. alatus. Sexual morph: Undetermined. Asexual morph: Conidiomata 60–75 × 40–60 μm (x = 68 × 50 μm, n = 20), pycnidial, black, globose, scattered on PDA culture media. Conidiophores subcylindrical, branched base with 2–3 septa. Conidiogenous cells 4–10 × 1.7–2.3 μm (x = 6.05 × 1.98 μm, n = 10) with percurrent proliferation of 1–2 times, integrated, cylindrical. Conidia 14–22 × 4–6 μm (x = 18 × 5 μm, n = 50), straight to slightly curved, fusoid, 4-septate; basal cell 2–5 μm long, conic, truncate base, hyaline, thin-walled; three median cells 10–13 μm (x = 12 μm) long, doliform or subcylindrical, olivaceous, concolorous and verrucose wall; second cell from the base 3–5 μm (x = 3 μm); third cell 2–4 μm (x = 3 μm); fourth cell 3–5 μm (x = 4 μm); apical cell 2–3 μm long, subcylindrical, hyaline, thin-walled, rugose; 2 appendages 10–19 μm long (x = 14 μm), tubular, filiform, unbranched, flexuous, appendages arise from apical crest; single basal appendage 2–6 μm long (x = 4 μm), unbranched, tubular, centric.
Culture characteristics: Colonies on PDA reaching 4.5 cm diam. after 7 days at 25 °C, aerial, whitish, smooth edge, entire margin, with gregarious black spore masses. Colonies creamy white from above and reverse without producing pigmentation on media.
Material examined: Thailand, Chiang Mai Province, Doi Inthanon National Park, (18.5356° N, 98.5221° E), on dead leaves of D. alatus (Dipterocarpaceae), 25 October 2021, Milan C. Samarakoon, D6a (MFLU 22-0195); living culture, MFLUCC 22-0119.
Known host: Pathogenic on leaves of Dracontomelon mangifera (Anacardiaceae) [63], leaf spots of Podocarpus sp. (Podocarpaceae) [27], saprobic on D. alatus leaf litter (this study).
Known distribution: Thailand.
Notes:Pestalotiopsis dracontomelon was introduced by Liu et al. [63], as a pathogenic species of Dracontomelon mangiferum in Thailand. Our isolate (MFLUCC 22-0119) shares the morphological characteristics of the type of P. dracontomelon (MFLUCC 10-0149), having septate conidiophores, conidiogenous cells with percurrent proliferation, 4-septate conidia, 2–3 tubular apical appendages, and a single centric basal appendage [63] with slight variations in the sizes of conidiogenous cells and conidia (x = 18 × 5 μm vs. 20 × 6.5 μm). According to the multilocus phylogeny (ITS, tub2, and tef1-α), our isolate clustered with the type P. dracontomelon with solid statistical support (99% ML, 99% MP, 1.00 PP). Thus, we introduce it as a new host record from D. alatus in Thailand.
3.2.2. Ciliochorella dipterocarpi Samaradiwakara, Lumyong and K.D. Hyde, sp. nov.
Index Fungorum number: IF900181; Faces of Fungi number: FoF13617 (Figure 5).
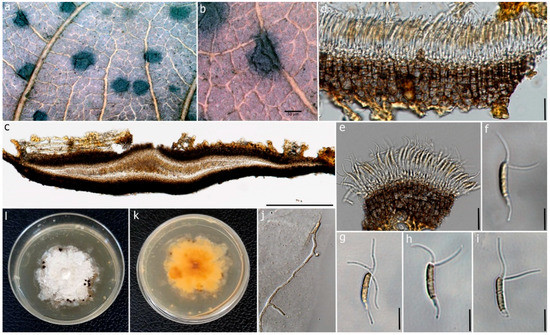
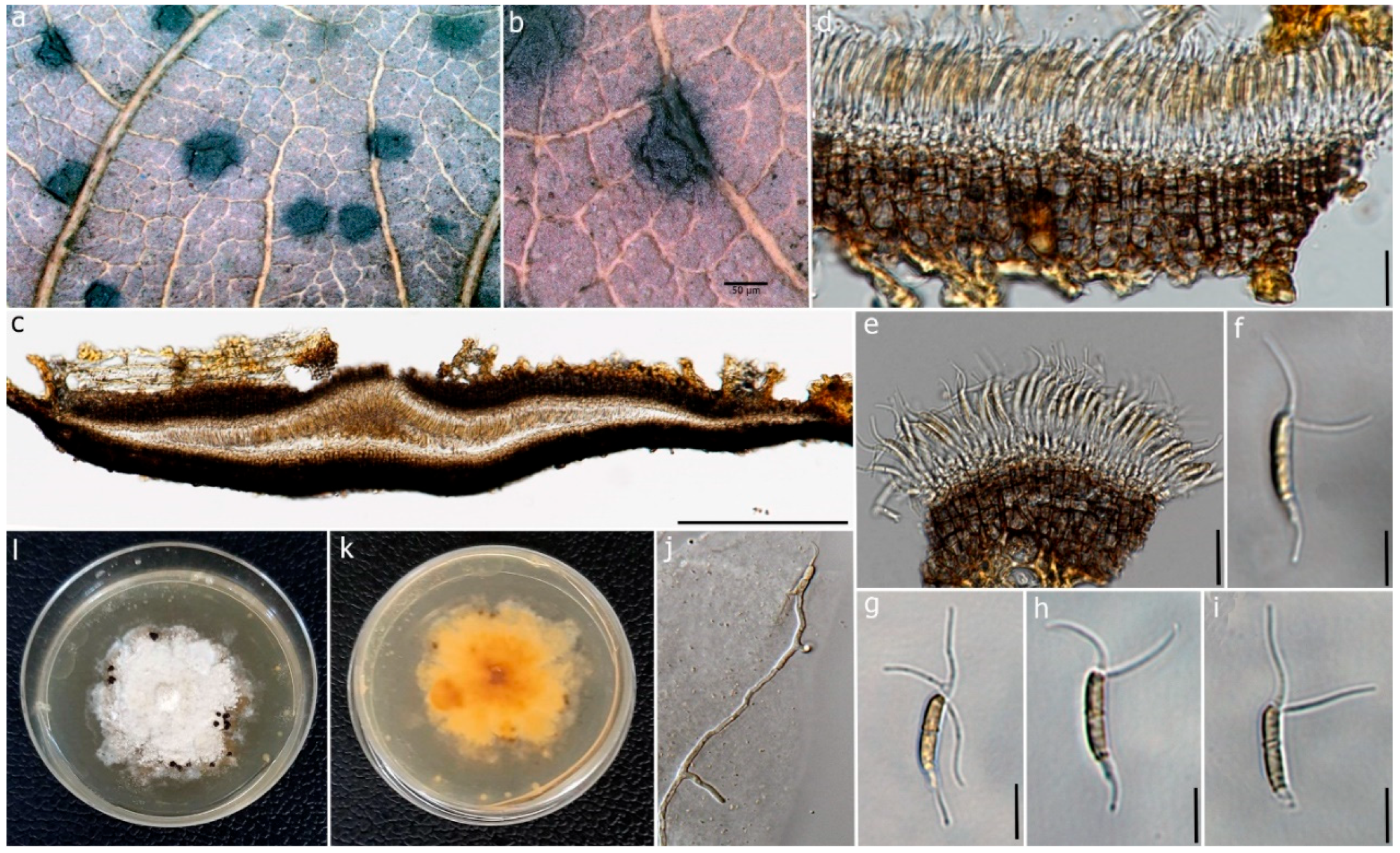
Figure 5.
Ciliochorella dipterocarpi (MFLUCC 22-0132, holotype). (a) Conidiomata on the leaf. (b) Close-up of conidioma. (c) Vertical section of the conidioma. (d) Peridium. (e) Conidiogenous cells with developing conidia. (f–i) Conidia. (j) Germinating conidium. (k,l) Reverse and front view of the colonies on PDA. Scale bars: (b) = 50 μm, (c) = 50 μm, (d) = 15 μm, (e) = 10 μm, (f–i) = 15 μm, (j) = 20 μm.
Etymology. The epithet “dipterocarpi” refers to the host genus.
Holotype. MFLU 22-0197.
Description: Saprobic on dead leaves of D. alatus. Sexual morph: Undetermined. Asexual morph: Coelomycetous. Conidiomata 120–150 μm high, 650–800 μm diam., acervulus, superficial or semi-immersed, mostly solitary, unilocular, horizontally fusiform. Peridium 20–35 μm wide, 6–8 layered, cells of textura angularis, composed of thin-walled pale-brown inner layer, thick-walled, dark brown to black outer layer. Conidiophores reduced to conidiogenous cells. Conidiogenous cells 0.6–1.3 μm (x = 0.95 μm, n = 30), hyaline to pale brown, smooth. Conidia 9–18 × 1–3 μm (x = 14 × 2 µm, n = 50), fusiform to naviculate, allantoid to sub-cylindrical, slightly curved, slightly guttulate, 1-septate, septum thicker and slightly darker, hyaline to pale brown, thick-walled, narrow towards the base, bearing appendages on both apical and basal ends; wide middle cell, basal end narrowly obconic, with truncate base terminated into a filiform appendage; apical end forked, hyaline at apex and sub-hyaline below with 2–3 apical appendages. Appendages tubular, filiform, flexuous; appendage on apical cell 12–15 μm (x = 13 μm) long, 2–3 tubular, unequal; basal appendage 8–17 µm (x = 10 µm) long, single.
Culture characteristics: Colonies on PDA, reaching 3.5 cm diam. after 7 days at 25 °C. Mycelium dense, flat, circular, or round, with smooth, erose, entire margin. Colonies white to cream from front, pale yellow from reverse.
Material examined: Thailand, Chiang Mai Province, Doi Inthanon National Park (18.5356° N, 98.5221° E), on dead leaves of D. alatus (Dipterocarpaceae), 25 October 2021, Milan C. Samarakoon, NP009 (MFLU 22-0197, Holotype), ex-type living culture, MFLUCC 22-0132.
Notes: Ciliochorella belongs to Xylariomycetidae incertae sedis in Sordariomycetes, and it produces appendage-bearing conidia [29]. The multilocus phylogeny indicates that our isolate (MFLUCC 22-0132) has a precise placement and clusters basal to C. mangiferae (MFLUCC 12-310), C. castaneae (HHUF28800), and C. castaneae (HHUF28799), with 81% ML and 0.95 PP statistical support (Figure 2). In the BLAST search in GenBank, the closest match for the ITS region of C. dipterocarpi (MFLUCC 22-0132) was C. phanericola (MFLUCC 12-310) with 99.45% similarity across 95% query coverage, LSU with a 99.77% similarity to C. phanericola (MFLUCC 12-310) across 97% query coverage, and tub2 with a 98.97% similarity to Ciliochorella sp. (MFLUCC 12-310) across 90% query coverage. The major difference in base pairs of the ITS region between C. dipterocarpi (MFLUCC 22-0132) and C. castaneae was 1.18% (10/841), and between C. dipterocarpi and C. mangiferae, it was 1.71% (7/408). The base pair difference of the tub2 between C. dipterocarpi and C. phanericola was 11.93% (37/310). The values agree with the new species concept outlined by Maharachchikumbura et al. [42].
Morphologically, C. dipterocarpi can be distinguished from other Ciliochorella species based on having conidiogenous cells bearing conidia with mostly 2-septate, slightly guttulate, and shorter apical appendages, 12–15 μm (x = 13 μm) long. In comparison, C. castaneae has conidia that are 0–2-septate with branched conidiophores [64], whereas C. mangiferae and C. phanericola have conidia in the range of 9–18 ×1–3 μm [50,65]. Ciliochorella dipterocarpi conidial sizes include 9–18 × 1–3 μm, with 2-3 apical appendages, and 12–15 μm long, with single basal appendage 8–17 µm long. In this study, we isolated C. dipterocarpi from the leaf litter of D. alatus, and considering the variations in morphological characteristics and phylogenetic placement, C. dipterocarpi (MFLUCC 22-0132) is introduced as a novel species.
3.2.3. Robillarda australiana F. Liu, L. Cai & Crous
Index Fungorum number: IF828387; Faces of Fungi Number: FoF13618 (Figure 6)
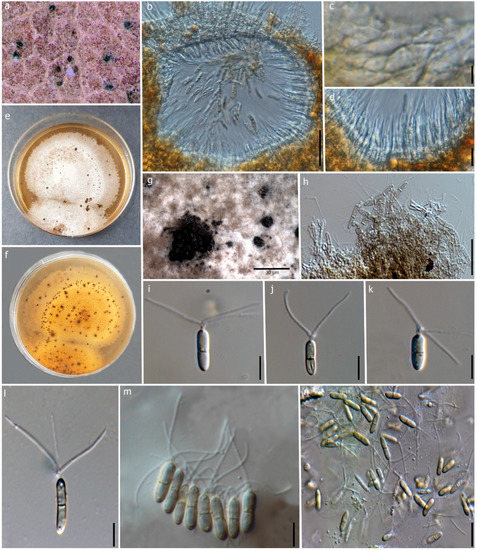
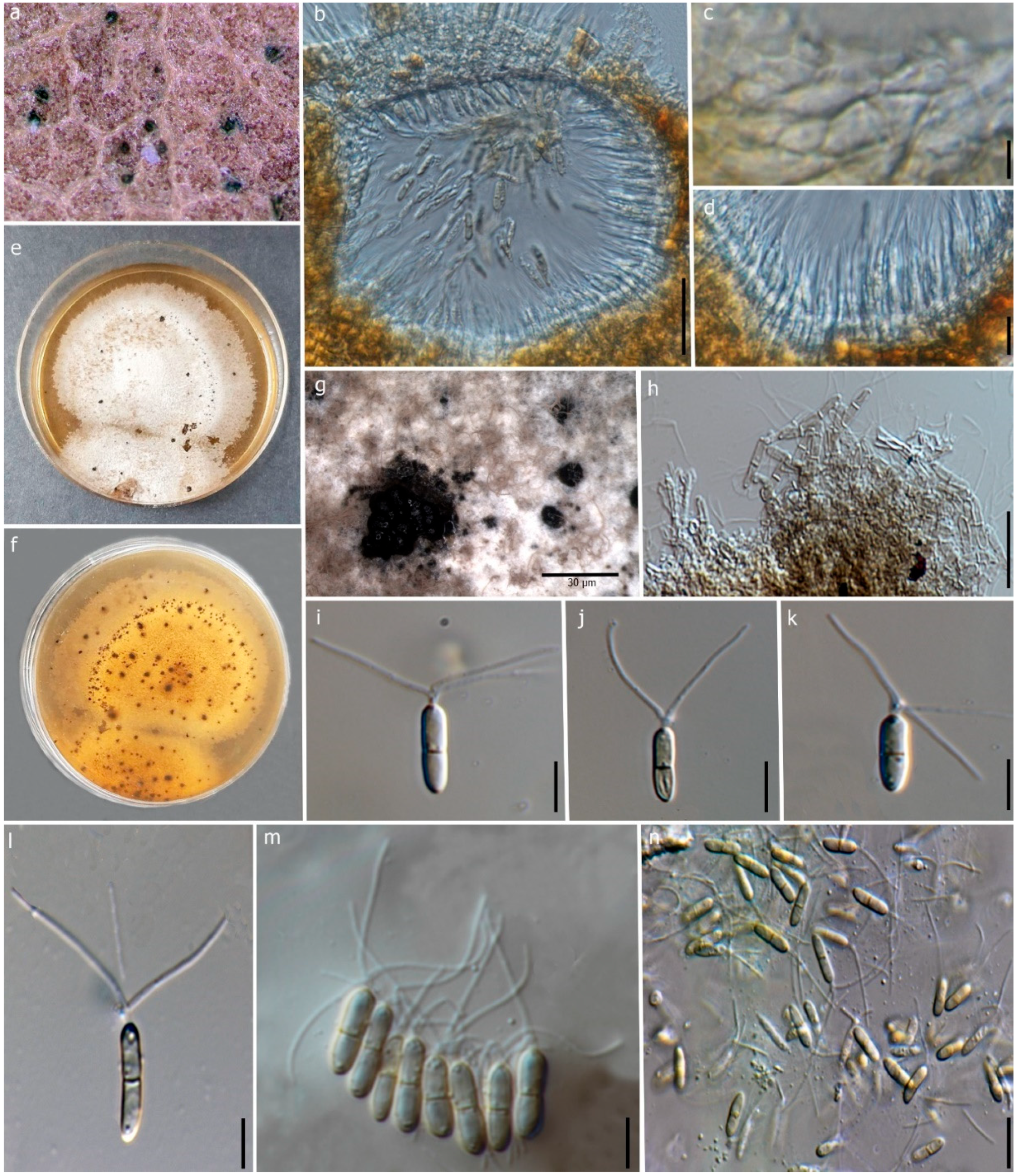
Figure 6.
Robillarda australiana (MFLU 22-0198, new host record). (a) Conidiomata on the leaf. (b) Vertical section of the conidioma. (c) Conidiomata wall at the base. (d) Conidiogenous cells and developing conidia. (e,f) Front and reverse view of the colonies on PDA. (g) Conidiomata on PDA. (h) Conidiophores and conidiogenous cells. (i–n) Conidia. Scale bars: (b) = 50 μm, (c) = 15 μm, (d) = 5 μm, (g) = 30 μm, (h–n) = 10 μm.
Description: Saprobic on dead leaves of D. alatus. Sexual morph: Undetermined. Asexual morph: Coelomycetous. Conidiomata 120–150 × 100–150 μm (x = 137 × 119 μm, n = 10), mostly erumpent, semi-immersed, pycnidial, mostly solitary, irregular shape, thick-walled, black. Conidiomatal wall 10–15 μm wide, thick-walled at upper wall, 1-2 layers of dark brown outer cells, inner cells hyaline, arranged in textura angularis. Conidiophores reduced to conidiogenous cells. Conidiogenous cells 4–10 × 2–4 μm (x = 7 × 3 μm, n = 10), guttulate, smooth, thin-walled, hyaline, discrete, irregular or ampulliform, lageniform, 2–3 distinct, small protuberances at the apex. Conidia 7–10 × 2–3 μm (x = 8 × 2.5 μm, n = 25), guttulate, cylindrical, 1-septate, slightly constricted at the median septum, smooth, straight, hyaline to pale brown, symmetrical, cylindrical apical cell modified into three appendages. Apical appendages unbranched, 3 divergent branches, attenuated at the apex, 9–17 μm (x = 14 μm, n = 15) long; basal appendages absent.
Culture characteristics: Conidia germinated on PDA within 24 h. Colonies on PDA, reaching 3 cm after 14 days at 25 °C. Mycelia superficial, flat, entire margin, surface rough, form black pycnidia after one month of incubation. Colonies white to light brown from front, pale yellow from reverse without producing pigmentation.
Material examined: Thailand, Chiang Mai Province, Doi Inthanon National Park, (18.5356° N, 98.5221° E), on dead leaves of D. alatus, 25 October 2021, Milan C. Samarakoon, NPSC 04 (MFLU 22-0198) living culture, MFLUCC 22-0121.
Known hosts: D. alatus (this study).
Known distribution: Australia, Thailand.
Notes: Phylogeny shows that our isolate (MFLUCC 22-0121) clusters with the type R. australiana (CBS 143882) with 84% ML and 0.99 BYPP (Figure 3). Both isolates share similar morphology, having conidia without constriction at the median septum with larger apical cells (R. australiana MFLUCC 22-0121, 8 × 2.5 μm vs. CBS 143882 11 × 2 μm) [29]. Therefore, we introduce our collection as a new host record of R. australiana from dead leaves of D. alatus in Thailand.
4. Discussion
Appendages are informative morphological characteristics in delineating fungal species [28]. Additionally, they support the adherence of spores to their substrate and dispersal, characterizing important ecological functions [28]. In this study, we used a polyphasic approach for the species boundaries of novel collections of leaf-litter-inhabiting fungi [42,66,67,68]. With this approach, out of three appendage-bearing Sordariomycetes collected from Doi Inthanon National Park, northern Thailand, one was placed as Ciliochorella dipterocarpi sp. nov, and two (P. dracontomelon and R. australiana) were placed as new records on D. alatus.
Ciliochorella is characterized by its distinct morphological characteristics [64,65] and molecular phylogeny based on the ITS, LSU, and tub2 sequence data [50]. Nevertheless, many species and isolates lack the sequence data for one or more loci, which may cause biases in the tree topology and phylogenetic placement [69,70,71,72] [e.g., 71,72,73,74]. Still, our newly introduced taxon C. dipterocarpi is clustered sister to C. mangiferae (MFLUCC 12-0310) (Figure 2), which has all the loci with available sequence data (Table 3). Like C. dipterocarpi, Ciliochorella species have been mainly reported as saprobes on various plant litter. In the present study, using a taxonomic approach (Figure 2), we expand the diversity of Ciliochorella with the introduction of C. dipterocarpi, isolated from the dead leaves of D. alatus.
Regarding the new litter host association of P. dracontomelon (MFLUCC 22-0119) and D. alatus, Pestalotiopsis species have been reported as endophytes producing important bioactive secondary metabolites, pathogens on several economically important crops, and saprobes on different plant litter [36,73,74,75,76]. Since the last detailed morphology and molecular phylogeny update [23,24], several species have been introduced, and around 100 Pestalotiopsis species are currently accepted [29,30,77,78,79]. However, the taxonomic placement is doubtful, mainly due to the lack of strong morphological characteristics [36] and the poor resolution of the employed molecular markers (ITS, tub2, and tef1-α), which presumably identify species rather than populations within the genus [80]. In our analysis, as taxon sampling is fundamental for the taxonomic placement and delimitation [81,82], to obtain better phylogenetic resolution (Figure 1), we excluded distantly related species and those with inconsistent sequence data and included only representative type strains (Table 2). Thus, a taxonomic revision of this genus is urgently needed, especially using the polyphasic approach with additional genomic regions [24,42,80,83,84].
Unlike Pestalotiopsis, Robillarda is characterized by its distinct morphological features [85,86], coupled whenever possible with the multilocus phylogeny of the ITS, rpb2, tub2, and tef1-α barcodes [29]. Nevertheless, out of its 41 species listed in the Index Fungorum database [40], only 8 have molecular data available [29,67,85,87,88]. Robillarda species are found as saprobic on decaying leaves in terrestrial and aquatic habitats, dust particles, and soil [85]. Here, we contribute with the expansion of the host and geographical record of R. australiana, which was found as a saprobe on decaying leaves of D. alatus (Figure 6) at Doi Inthanon National Park, Thailand. This species was only reported in Australia on unidentified plant litter [29].
Many fungal species have been discovered in Thailand, using polyphasic approaches [89]. As evident from the present study, much work remains to be done in this regard. This includes examining unstudied areas, such as protected national parks and hosts, to address whether fungi are host-specific and at which level [13]. Tennakoon et al. [90] contributed significantly to this matter. This is important in predicting the number of species [19] and in assessing the diversity in a given area or ecosystem [91], which have the same significance [92]. Currently, only approximately 10% of the 2.2–3.8 million species estimated by Hawksworth and Lucking [93] have been described [13,16,94], highlighting the potential for unraveling novel fungal taxa in largely untapped ecosystems. In this regard, this study contributes to the above topics, and the number of teleomorph species of Ascomycota estimated by Senanayake et al. [95] was between 1.37 and 2.56 million. Finally, the findings of the present study provide additional insights into fungal diversity in pristine tropical environments, as predicted by Hyde et al. [89]. However, given the different lifestyles of the species, elucidating their evolution, host and lifestyle shifting, and environmental adaptations is needed for a better understanding of their roles [13].
5. Conclusions
Ecological preference and evolutionary relationships can support the emergence of novel species in contrasting life modes [4,13]. This hypothesis expands the scope of saprobic fungal habitats and their host preferences [4,13]. This scenario is supported by identifying saprobic lifestyles of different genera from the present study. Moreover, we also identified fungal species associated with novel hosts and geographical locations (P. dracontomelon and R. australiana), along with the novel species C. dipterocarpi.
Author Contributions
Conceptualization, N.P.S. and D.S.T.; Formal analysis, N.P.S., D.S.T. and J.V.S.A.; Funding acquisition, K.W.T.C., J.K. and S.L.; Investigation, A.R.G.d.F. and C.S.B.; Methodology, A.R.G.d.F., D.S.T. and J.V.S.A.; Resources, J.K. and S.L.; Software, A.R.G.d.F.; Supervision, A.R.G.d.F. and S.L.; Validation, C.S.B.; Writing—original draft, N.P.S.; Writing—review and editing, N.P.S., A.R.G.d.F., D.S.T., C.S.B., J.V.S.A., K.W.T.C. and S.L. All authors have read and agreed to the published version of the manuscript.
Funding
The authors would like to thank the National Research Council of Thailand for the project entitled “Comparison of diversity and biogeographical distribution of Ascomycetous fungi from two protected areas in Turkey and Thailand” (NRCT: Project No. P-19-52624) and for providing permission to conduct research in the Doi Inthanon National Park, Thailand (No. 0402-2803).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All sequences generated in this study were submitted to GenBank (https://www.ncbi.nlm.nih.gov (accessed on 4 April 2023)).
Acknowledgments
Nethmini P. Samaradiwakara is grateful for the Chiang Mai University Presidential Scholarship 2020. Chitrabhanu S. Bhunjun would like to express thanks for the National Research Council of Thailand (NRCT) grant “Total fungal diversity in a given forest area with implications towards species numbers, chemical diversity and biotechnology” (grant no. N42A650547). The authors would like to thank Chiang Mai University for their partial support to conduct the study. Milan C. Samarakoon and Achala J. Gajanayake are also thanked for their valuable support. The authors thank Shaun Pennycook for nomenclatural clarification of the new species. All the staff and colleagues at the Centre of Excellence in Fungal Research, Mae Fah Luang University, Thailand are acknowledged.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Luo, Y.; Berg, B.; Laskowski, R. Litter Decomposition: A Guide to Carbon and Nutrient Turnover; Luo, Y., Laskowski, R., Eds.; Elsevier Science: Amsterdam, The Netherlands, 2006; ISBN 012373617X, 9780123736178. [Google Scholar]
- Matsukura, K.; Hirose, D.; Kagami, M.; Osono, T.; Yamaoka, Y. Geographical distributions of rhytismataceous fungi on Camellia japonica leaf litter in Japan. Fungal Ecol. 2017, 26, 37–44. [Google Scholar] [CrossRef]
- Promputtha, I.; Hyde, K.D.; McKenzie, E.H.C.; Peberdy, J.F.; Lumyong, S. Can leaf degrading enzymes provide evidence that endophytic fungi becoming saprobes? Fungal Divers. 2010, 41, 89–99. [Google Scholar] [CrossRef]
- Zhang, Y.; Jeewon, R.; Fournier, J.; Hyde, K.D. Multi-gene phylogeny and morphotaxonomy of Amniculicola Lignicola: A novel freshwater fungus from France and its relationships to the Pleosporales. Mycol. Res. 2008, 112, 1186–1194. [Google Scholar] [CrossRef] [PubMed]
- Tennakoon, D.; Gentekaki, E.; Jeewon, R.; Kuo, C.; Promputtha, I.; Hyde, K. Life in leaf litter: Fungal community succession during decomposition. Mycosphere 2021, 12, 406–429. [Google Scholar] [CrossRef]
- Mapook, A.; Hyde, K.D.; Hassan, K.; Kemkuignou, B.M.; Čmoková, A.; Surup, F.; Kuhnert, E.; Paomephan, P.; Cheng, T.; de Hoog, S.; et al. Ten decadal advances in fungal biology leading towards human well-being. Fungal Divers. 2022, 116, 547–614. [Google Scholar] [CrossRef] [PubMed]
- Wainwright, M.; Wickramasinghe, N.; Narlikar, J.; Rajaratnam, P. Microorganisms cultured from stratospheric air samples obtained at 41 Km. FEMS Microbiol. Lett. 2003, 218, 161–165. [Google Scholar] [CrossRef] [PubMed]
- Bucher, V.V.C.; Pointing, S.B.; Hyde, K.D.; Reddy, C.A. Production of wood decay enzymes, loss of mass, and lignin solubilization in wood by diverse tropical freshwater fungi. Microb. Ecol. 2004, 48, 331–337. [Google Scholar] [CrossRef]
- Pointing, S.B.; Pelling, A.L.; Smith, G.J.D.; Hyde, K.D.; Reddy, C.A. Screening of Basidiomycetes and Xylariaceous fungi for lignin peroxidase and laccase gene-specific sequences. Mycol. Res. 2005, 109, 115–124. [Google Scholar] [CrossRef]
- Jobard, M.; Rasconi, S.; Sime-Ngando, T. Diversity and functions of microscopic fungi: A missing component in pelagic food webs. Aquat. Sci. 2010, 72, 255–268. [Google Scholar] [CrossRef]
- Nagahama, T.; Takahashi, E.; Nagano, Y.; Abdel-Wahab, M.A.; Miyazaki, M. Molecular evidence that deep-branching fungi are major fungal components in deep-sea methane cold-seep sediments. Environ. Microbiol. 2011, 13, 2359–2370. [Google Scholar] [CrossRef]
- Schmit, J.P.; Mueller, G.M. An estimate of the lower limit of global fungal diversity. Biodivers. Conserv. 2007, 16, 99–111. [Google Scholar] [CrossRef]
- Hyde, K.D.; Jeewon, R.; Chen, Y.-J.; Bhunjun, C.S.; Calabon, M.S.; Jiang, H.-B.; Lin, C.-G.; Norphanphoun, C.; Sysouphanthong, P.; Pem, D.; et al. The numbers of fungi: Is the descriptive curve flattening? Fungal Divers. 2020, 103, 219–271. [Google Scholar] [CrossRef]
- Mohamed, D.J.; Martiny, J.B. Patterns of fungal diversity and composition along a salinity gradient. ISME J. 2011, 5, 379–388. [Google Scholar] [CrossRef]
- Phukhamsakda, C.; McKenzie, E.H.C.; Phillips, A.J.L.; Jones, E.B.G.; Bhat, D.J.; Stadler, M.; Bhunjun, C.S.; Wanasinghe, D.N.; Thongbai, B.; Camporesi, E.; et al. Microfungi associated with Clematis (Ranunculaceae) with an integrated approach to delimiting species boundaries. Fungal Divers. 2020, 102, 1–203. [Google Scholar] [CrossRef]
- Phukhamsakda, C.; Nilsson, R.H.; Bhunjun, C.S.; de Farias, A.R.G.; Sun, Y.-R.; Wijesinghe, S.N.; Raza, M.; Bao, D.-F.; Lu, L.; Tibpromma, S.; et al. The numbers of fungi: Contributions from traditional taxonomic studies and challenges of metabarcoding. Fungal Divers. 2022, 114, 327–386. [Google Scholar] [CrossRef]
- de Silva, N.I.; Thambugala, K.M.; Tennakoon, D.S.; Karunarathna, S.C.; Kumla, J.; Suwannarach, N.; Lumyong, S. Taxonomy and phylogenetic appraisal of dothideomycetous fungi associated with Magnolia, Lilium Longiflorum and Hedychium Coronarium. J. Fungi 2022, 8, 1094. [Google Scholar] [CrossRef]
- Zhang, N.; Castlebury, L.A.; Miller, A.N.; Huhndorf, S.M.; Schoch, C.L.; Seifert, K.A.; Rossman, A.Y.; Rogers, J.D.; Kohlmeyer, J.; Volkmann-Kohlmeyer, B.; et al. An overview of the systematics of the Sordariomycetes based on a four-gene phylogeny. Mycologia 2006, 98, 1076–1087. [Google Scholar] [CrossRef]
- Hyde, K.; Norphanphoun, C.; Maharachchikumbura, S.S.N.; Bhat, D.J.; Jones, E.B.G.; Bundhun, D.; Chen, Y.J.; Bao, D.F.; Boonmee, S.; Calabon, M.S.; et al. Refined families of Sordariomycetes. Mycosphere 2020, 11, 305–1059. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Jones, E.B.G.; McKenzie, E.H.C.; Bhat, J.D.; Dayarathne, M.C.; Huang, S.-K.; Norphanphoun, C.; Senanayake, I.C.; Perera, R.H.; et al. Families of Sordariomycetes. Fungal Divers. 2016, 79, 1–317. [Google Scholar] [CrossRef]
- Pratibha, J.; Prabhugaonkar, A.; Hyde, K.D.; Bhat, D.J. Phylogenetic placement of Bahusandhika, Cancellidium and Pseudoepicoccum (Asexual Ascomycota). Phytotaxa 2014, 176, 68. [Google Scholar] [CrossRef]
- Jones, E.B.G.; Suetrong, S.; Sakayaroj, J.; Bahkali, A.H.; Abdel-Wahab, M.A.; Boekhout, T.; Pang, K.-L. Classification of marine Ascomycota, Basidiomycota, Blastocladiomycota and Chytridiomycota. Fungal Divers. 2015, 73, 1–72. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Groenewald, J.Z.; Xu, J.; Crous, P.W. Pestalotiopsis Revisited. Stud. Mycol. 2014, 79, 121–186. [Google Scholar] [CrossRef] [PubMed]
- Norphanphoun, C.; Jayawardena, R.S.; Chen, Y.; Wen, T.C.; Meepol, W.; Hyde, K.D. Morphological and phylogenetic characterization of novel pestalotioid species associated with mangroves in Thailand. Mycosphere 2019, 10, 531–578. [Google Scholar] [CrossRef]
- Zhang, N.; Wang, Z. 3 Pezizomycotina: Sordariomycetes and Leotiomycetes. In Systematics and Evolution; Springer: Berlin/Heidelberg, Germany, 2015; pp. 57–88. [Google Scholar]
- Gostinčar, C.; Zajc, J.; Lenassi, M.; Plemenitaš, A.; de Hoog, S.; Al-Hatmi, A.M.S.; Gunde-Cimerman, N. Fungi between extremotolerance and opportunistic pathogenicity on humans. Fungal Divers. 2018, 93, 195–213. [Google Scholar] [CrossRef]
- Sun, J.-Z.; Liu, X.-Z.; McKenzie, E.H.C.; Jeewon, R.; Liu, J.-K.; Zhang, X.-L.; Zhao, Q.; Hyde, K.D. Fungicolous fungi: Terminology, diversity, distribution, evolution, and species checklist. Fungal Divers. 2019, 95, 337–430. [Google Scholar] [CrossRef]
- Nag Raj, T.R. Coelomycetous Anamorphs with Appendage-Bearing Conidia; Mycologue Publications: Sidney, BC, Canada, 1993; ISBN 0969223722. [Google Scholar]
- Liu, F.; Bonthond, G.; Groenewald, J.Z.; Cai, L.; Crous, P.W. Sporocadaceae, A family of coelomycetous fungi with appendage-bearing conidia. Stud. Mycol. 2019, 92, 287–415. [Google Scholar] [CrossRef]
- Wijayawardene, N.N.; Dai, D.Q.; Zhang, G.Q.; Gao, Y.; Wijayawardene, N.N.; Hyde, K.D.; Pem, D.; Thiyagaraja, V.; Hyde, K.D.; Dong, W.; et al. Outline of fungi and fungus-like taxa—2021. Mycosphere 2022, 13, 53–453. [Google Scholar] [CrossRef]
- Barber, P.A.; Crous, P.W.; Groenewald, J.Z.; Pascoe, I.G.; Keane, P. Reassessing Vermisporium (Amphisphaeriaceae), A genus of foliar pathogens of Eucalypts. Pers. Mol. Phylogeny Evol. Fungi 2011, 27, 90–118. [Google Scholar] [CrossRef]
- Tanaka, K.; Endo, M.; Hirayama, K.; Okane, I.; Hosoya, T.; Sato, T. Phylogeny of Discosia and Seimatosporium, and introduction of Adisciso and Immersidiscosia genera nova. Pers. Mol. Phylogeny Evol. Fungi 2011, 26, 85–98. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Gardiennet, A.; Voglmayr, H. Resolution of morphology-based taxonomic delusions: Acrocordiella, Basiseptospora, Blogiascospora, Clypeosphaeria, Hymenopleella, Lepteutypa, Pseudapiospora, Requienella, Seiridium and Strickeria. Pers. Mol. Phylogeny Evol. Fungi 2016, 37, 82–105. [Google Scholar] [CrossRef]
- Barr, M.E. Pestalosphaeria, A new genus in the Amphisphaeriaceae. Mycologia 1975, 67, 187–194. [Google Scholar] [CrossRef]
- Barr, M.E. Prodromus to nonlichenized, pyrenomycetous members of class Hymenoascomycetes. Mycotaxon 1990, 39, 43–184. [Google Scholar]
- Hu, H.; Jeewon, R.; Zhou, D.; Zhou, T.; Hyde, K.D. Phylogenetic diversity of endophytic Pestalotiopsis species in Pinus Armandii and Ribes spp.: Evidence from rdna and β-tubulin gene phylogenies. Fungal Divers. 2007, 24, 1–22. [Google Scholar]
- Lücking, R.; Aime, M.C.; Robbertse, B.; Miller, A.N.; Aoki, T.; Ariyawansa, H.A.; Cardinali, G.; Crous, P.W.; Druzhinina, I.S.; Geiser, D.M.; et al. Fungal taxonomy and sequence-based nomenclature. Nat. Microbiol. 2021, 6, 540–548. [Google Scholar] [CrossRef]
- Senanayake, I.; Rathnayaka, A.; Sandamali, D.; Calabon, M.; Gentekaki, E. Morphological approaches in studying fungi: Collection, examination, isolation, sporulation and preservation. Mycosphere 2020, 11, 2678–2754. [Google Scholar] [CrossRef]
- Jayasiri, S.C.; Hyde, K.D.; Ariyawansa, H.A.; Bhat, J.; Buyck, B.; Cai, L.; Dai, Y.-C.; Abd-Elsalam, K.A.; Ertz, D.; Hidayat, I.; et al. The faces of fungi database: Fungal names linked with morphology, phylogeny and human impacts. Fungal Divers. 2015, 74, 3–18. [Google Scholar] [CrossRef]
- Index Fungorum. Available online: https://www.indexfungorum.org/names/Names.asp (accessed on 8 March 2023).
- Chaiwan, N.; Gomdola, D.; Wang, S.; Monkai, J.; Tibpromma, S.; Doilom, M.; Wanasinghe, D.N.; Mortimer, P.E.; Lumyong, S.; Hyde, K.D. An online database providing updated information of microfungi in the greater mekong subregion. Mycosphere 2021, 12, 1513–1526. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Chen, Y.; Ariyawansa, H.A.; Hyde, K.D.; Haelewaters, D.; Perera, R.H.; Samarakoon, M.C.; Wanasinghe, D.N.; Bustamante, D.E.; Liu, J.-K.; et al. Integrative approaches for species delimitation in Ascomycota. Fungal Divers. 2021, 109, 155–179. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification, and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols; Elsevier: Amsterdam, The Netherlands, 1990; pp. 315–322. [Google Scholar]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [CrossRef]
- Rehner, S.A.; Samuels, G.J. Taxonomy and phylogeny of Gliocladium analysed from nuclear large subunit ribosomal DNA sequences. Mycol. Res. 1994, 98, 625–634. [Google Scholar] [CrossRef]
- Rehner, S.A.; Buckley, E.A. Beauveria phylogeny inferred from nuclear ITS and EF1-sequences: Evidence for cryptic diversification and links to cordyceps teleomorphs. Mycologia 2005, 97, 84–98. [Google Scholar] [CrossRef] [PubMed]
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among Ascomycetes: Evidence from an RNA polymerse ii subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef] [PubMed]
- Sung, G.-H.; Sung, J.-M.; Hywel-Jones, N.L.; Spatafora, J.W. A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): Identification of localized incongruence using a combinational bootstrap approach. Mol. Phylogenet. Evol. 2007, 44, 1204–1223. [Google Scholar] [CrossRef]
- Hyde, K.D.; Hongsanan, S.; Jeewon, R.; Bhat, D.J.; McKenzie, E.H.C.; Jones, E.B.G.; Phookamsak, R.; Ariyawansa, H.A.; Boonmee, S.; Zhao, Q.; et al. Fungal diversity notes 367–490: Taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers. 2016, 80, 1–270. [Google Scholar] [CrossRef]
- Li, W.L.; Liu, Z.P.; Zhang, T.; Dissanayake, A.J.; Luo, Z.L.; Su, H.Y.; Liu, J.K. Additions to Distoseptispora (Distoseptisporaceae) associated with submerged decaying wood in China. Phytotaxa 2021, 520, 75–86. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Capella-Gutierrez, S.; Silla-Martinez, J.M.; Gabaldon, T. TrimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES science gateway for inference of large phylogenetic trees. In Proceedings of the 2010 Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; IEEE: Amsterdam, The Netherlands, 2010; pp. 1–8. [Google Scholar]
- Stamatakis, A.; Hoover, P.; Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst. Biol. 2008, 57, 758–771. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML Version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Rannala, B.; Yang, Z. Probability distribution of molecular evolutionary trees: A new method of phylogenetic inference. J. Mol. Evol. 1996, 43, 304–311. [Google Scholar] [CrossRef]
- Swofford, D.L. Phylogenetic analysis using parsimony. J. Mol. Evol. 2002, 42, 294–307. [Google Scholar] [CrossRef]
- Kishino, H.; Hasegawa, M. Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in Hominoidea. J. Mol. Evol. 1989, 29, 170–179. [Google Scholar] [CrossRef]
- Rambaut, A. FigTree version 1.4.0. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 12 December 2022).
- Liu, J.K.; Hyde, K.D.; Jones, E.B.G.; Ariyawansa, H.A.; Bhat, D.J.; Boonmee, S.; Maharachchikumbura, S.S.N.; McKenzie, E.H.C.; Phookamsak, R.; Phukhamsakda, C.; et al. Fungal diversity notes 1–110: Taxonomic and phylogenetic contributions to fungal species. Fungal Divers. 2015, 72, 1–197. [Google Scholar] [CrossRef]
- Allegrucci, N.; Elíades, L.; Cabello, M.; Arambarri, A. New species Koorchaloma and Ciliochorella from xeric forests in Argentina. Mycotaxon 2011, 115, 175–181. [Google Scholar] [CrossRef]
- Tangthirasunun, N.; Silar, P.; Bhat, D.J.; Maharachchikumbura, S.S.N.; Wijayawardene, N.N.; Bahkali, A.H.; Hyde, K.D. Morphology and phylogeny of two appendaged genera of coelomycetes: Ciliochorella and Discosia. Sydowia 2015, 67, 217–226. [Google Scholar] [CrossRef]
- Thilini Chethana, K.W.; Jayawardena, R.S.; Chen, Y.J.; Konta, S.; Tibpromma, S.; Abeywickrama, P.D.; Gomdola, D.; Balasuriya, A.; Xu, J.; Lumyong, S.; et al. Diversity and function of appressoria. Pathogens 2021, 10, 746. [Google Scholar] [CrossRef]
- Manawasinghe, I.S.; Calabon, M.S.; Jones, E.B.G.; Zhang, Y.X.; Liao, C.F.; Xiong, Y.R.; Chaiwan, N.; Kularathnage, N.D.; Liu, N.G.; Tang, S.M.; et al. Mycosphere notes 345–386. Mycosphere 2022, 13, 454–557. [Google Scholar] [CrossRef]
- Jayawardena, R.S.; Hyde, K.D.; de Farias, A.R.G.; Bhunjun, C.S.; Ferdinandez, H.S.; Manamgoda, D.S.; Udayanga, D.; Herath, I.S.; Thambugala, K.M.; Manawasinghe, I.S.; et al. What is a species in fungal plant pathogens? Fungal Divers. 2021, 109, 239–266. [Google Scholar] [CrossRef]
- Simmons, M.P. A confounding effect of missing data on character conflict in maximum likelihood and bayesian MCMC phylogenetic analyses. Mol. Phylogenet. Evol. 2014, 80, 267–280. [Google Scholar] [CrossRef] [PubMed]
- Hovmöller, R.; Lacey Knowles, L.; Kubatko, L.S. Effects of missing data on species tree estimation under the coalescent. Mol. Phylogenet. Evol. 2013, 69, 1057–1062. [Google Scholar] [CrossRef] [PubMed]
- Streicher, J.W.; Schulte, J.A.; Wiens, J.J. How should genes and taxa be sampled for phylogenomic analyses with missing data? An empirical study in Iguanian lizards. Syst. Biol. 2016, 65, 128–145. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.T.; Mauck, W.M.; Benz, B.W.; Andersen, M.J. Uneven missing data skew phylogenomic relationships within the lories and lorikeets. Genome Biol. Evol. 2020, 12, 1131–1147. [Google Scholar] [CrossRef]
- Debbab, A.; Aly, A.H.; Proksch, P. Endophytes and associated marine derived fungi—Ecological and chemical perspectives. Fungal Divers. 2012, 57, 45–83. [Google Scholar] [CrossRef]
- Ko, T.W.; Stephenson, S.L.; Bahkali, A.H.; Hyde, K.D. From morphology to molecular biology: Can we use sequence data to identify fungal endophytes? Fungal Divers. 2011, 50, 113–120. [Google Scholar] [CrossRef]
- Xu, J.; Ebada, S.S.; Proksch, P. Pestalotiopsis a highly creative genus: Chemistry and bioactivity of secondary metabolites. Fungal Divers. 2010, 44, 15–31. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Chukeatirote, E.; Guo, L.D.; Crous, P.W.; Mckenzie, E.H.C.; Hyde, K.D. Pestalotiopsis species associated with Camellia Sinensis (Tea). Mycotaxon 2013, 123, 47–61. [Google Scholar] [CrossRef]
- Rashmi, M.A. Worldwide list of endophytic fungi with notes on ecology and diversity. Mycosphere 2019, 10, 798–1079. [Google Scholar] [CrossRef]
- Shu, J.; Yu, Z.; Sun, W.; Zhao, J.; Li, Q.; Tang, L.; Guo, T.; Huang, S.; Mo, J.; Hsiang, T.; et al. Identification and characterization of pestalotioid fungi causing leaf spots on Mango in southern China. Plant Dis. 2020, 104, 1207–1213. [Google Scholar] [CrossRef]
- Monteiro, P.; Gonçalves, M.F.M.; Pinto, G.; Silva, B.; Martín-García, J.; Diez, J.J.; Alves, A. Three novel species of fungi associated with pine species showing needle blight-like disease symptoms. Eur. J. Plant Pathol. 2022, 162, 183–202. [Google Scholar] [CrossRef]
- Liu, F.; Hou, L.; Raza, M.; Cai, L. Pestalotiopsis and allied genera from Camellia, with description of 11 new species from China. Sci. Rep. 2017, 7, 866. [Google Scholar] [CrossRef]
- Watanabe, K.; Motohashi, K.; Ono, Y. Description of Pestalotiopsis pallidotheae: A new species from Japan. Mycoscience 2010, 51, 182–188. [Google Scholar] [CrossRef]
- Bhunjun, C.S.; Phukhamsakda, C.; Jayawardena, R.S.; Jeewon, R.; Promputtha, I.; Hyde, K.D. Investigating species boundaries in Colletotrichum. Fungal Divers. 2021, 107, 107–127. [Google Scholar] [CrossRef]
- Bhunjun, C.S.; Dong, Y.; Jayawardena, R.S.; Jeewon, R.; Phukhamsakda, C.; Bundhun, D.; Hyde, K.D.; Sheng, J.A. Polyphasic approach to delineate species in Bipolaris. Fungal Divers. 2020, 102, 225–256. [Google Scholar] [CrossRef]
- Bhunjun, C.S.; Phillips, A.J.L.; Jayawardena, R.S.; Promputtha, I.; Hyde, K.D. Importance of molecular data to identify fungal plant pathogens and guidelines for pathogenicity testing based on Koch’s postulates. Pathogens 2021, 10, 1096. [Google Scholar] [CrossRef]
- Crous, P.W.; Carris, L.M.; Giraldo, A.; Groenewald, J.Z.; Hawksworth, D.L.; Hemández-Restrepo, M.; Jaklitsch, W.M.; Lebrun, M.H.; Schumacher, R.K.; Stielow, J.B.; et al. The genera of fungi—Fixing the application of the type species of generic names—G2: Allantophomopsis, Latorua, Macrodiplodiopsis, Macrohilum, Milospium, Protostegia, Pyricularia, Robillarda, Rotula, Septoriella, Torula, and Wojnowicia. IMA Fungus 2015, 6, 163–198. [Google Scholar] [CrossRef]
- Wijayawardene, N.N.; Hyde, K.D.; Wanasinghe, D.N.; Papizadeh, M.; Goonasekara, I.D.; Camporesi, E.; Bhat, D.J.; McKenzie, E.H.C.; Phillips, A.J.L.; Diederich, P.; et al. Taxonomy and phylogeny of dematiaceous coelomycetes. Fungal Divers. 2016, 77, 1–316. [Google Scholar] [CrossRef]
- Phookamsak, R.; Hyde, K.D.; Jeewon, R.; Bhat, D.J.; Jones, E.B.G.; Maharachchikumbura, S.S.N.; Raspé, O.; Karunarathna, S.C.; Wanasinghe, D.N.; Hongsanan, S.; et al. Fungal diversity notes 929–1035: Taxonomic and phylogenetic contributions on genera and species of fungi. Fungal Divers. 2019, 95, 1–273. [Google Scholar] [CrossRef]
- Haelewaters, D.; Dima, B.; Abdel-Hafiz, A.; Abdel-Wahab, M.; Abul-Ezz, S.; Acar, I.; Aguirre-Acosta, E.; Aime, M.; Aldemir, S.; Ali, M.; et al. Fungal systematics and evolution 6. Sydowia 2020, 72, 231–356. [Google Scholar]
- Hyde, K.D.; Norphanphoun, C.; Chen, J.; Dissanayake, A.J.; Doilom, M.; Hongsanan, S.; Jayawardena, R.S.; Jeewon, R.; Perera, R.H.; Thongbai, B.; et al. Thailand’s Amazing diversity: Up to 96% of fungi in northern Thailand may be novel. Fungal Divers. 2018, 93, 215–239. [Google Scholar] [CrossRef]
- Tennakoon, D.S.; Kuo, C.H.; Maharachchikumbura, S.S.N.; Thambugala, K.M.; Gentekaki, E.; Phillips, A.J.L.; Bhat, D.J.; Wanasinghe, D.N.; de Silva, N.I.; Promputtha, I.; et al. Taxonomic and phylogenetic contributions to Celtis Formosana, Ficus Ampelas, F. Septica, Macaranga Tanarius and Morus Australis leaf litter inhabiting microfungi. Fungal Divers. 2021, 108, 1–215. [Google Scholar] [CrossRef]
- Bermudez, G.; Lindemann, M.P. “What matters is species richness” High school students’ understanding of the components of biodiversity. Res. Sci. Educ. 2020, 50, 2159–2187. [Google Scholar] [CrossRef]
- Shtilerman, E.; Thompson, C.J.; Stone, L.; Bode, M.; Burgman, M. A novel method for estimating the number of species within a region. Proc. R. Soc. B Biol. Sci. 2014, 281, 2013–3009. [Google Scholar] [CrossRef]
- Hawksworth, D.L.; Lücking, R. Fungal diversity revisited: 2.2 to 3.8 million species. Microbiol. Spectr. 2017, 5, 4. [Google Scholar] [CrossRef]
- Bhunjun, C.S.; Niskanen, T.; Suwannarach, N.; Wannathes, N.; Chen, Y.-J.; McKenzie, E.H.C.; Maharachchikumbura, S.S.N.; Buyck, B.; Zhao, C.L.; Fan, Y.G.; et al. The numbers of fungi: Are the most speciose genera truly diverse? Fungal Divers. 2022, 114, 387–462. [Google Scholar] [CrossRef]
- Senanayake, I.C.; Pem, D.; Rathnayaka, A.R.; Wijesinghe, S.N.; Tibpromma, S.; Wanasinghe, D.N.; Phookamsak, R.; Kularathnage, N.D.; Gomdola, D.; Harishchandra, D.; et al. Predicting global numbers of teleomorphic Ascomycetes. Fungal Divers. 2022, 114, 237–278. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).