Abstract
This study aimed to evaluate the potential of Large Language Models (LLMs) in healthcare diagnostics, specifically their ability to analyze symptom-based prompts and provide accurate diagnoses. The study focused on models including GPT-4, GPT-4o, Gemini, o1 Preview, and GPT-3.5, assessing their performance in identifying illnesses based solely on provided symptoms. Symptom-based prompts were curated from reputable medical sources to ensure validity and relevance. Each model was tested under controlled conditions to evaluate their diagnostic accuracy, precision, recall, and decision-making capabilities. Specific scenarios were designed to explore their performance in both general and high-stakes diagnostic tasks. Among the models, GPT-4 achieved the highest diagnostic accuracy, demonstrating strong alignment with medical reasoning. Gemini excelled in high-stakes scenarios requiring precise decision-making. GPT-4o and o1 Preview showed balanced performance, effectively handling real-time diagnostic tasks with a focus on both precision and recall. GPT-3.5, though less advanced, proved dependable for general diagnostic tasks. This study highlights the strengths and limitations of LLMs in healthcare diagnostics. While models such as GPT-4 and Gemini exhibit promise, challenges such as privacy compliance, ethical considerations, and the mitigation of inherent biases must be addressed. The findings suggest pathways for responsibly integrating LLMs into diagnostic processes to enhance healthcare outcomes.
1. Introduction
The advancement of Large Language Models (LLMs) has transformed natural language processing, unlocking new possibilities for healthcare and clinical applications [1,2,3,4,5,6,7]. In the healthcare sector, LLMs are emerging as potential tools for enhancing diagnostic accuracy and streamlining patient interactions by interpreting and generating human-like text [8,9]. Given the high demand for healthcare services and the need for efficient patient management, these models could automate parts of the diagnostic process, making healthcare services more accessible while reducing provider workloads [10]. LLMs have already demonstrated utility in data-intensive fields like radiology and pathology, where rapid interpretation of diagnostic data aids in early disease detection and treatment planning [11].
Beyond diagnostics, LLMs contribute to patient engagement by offering personalized health consultations and symptom assessments, potentially improving trust and satisfaction. By analyzing diverse data sources, including patient histories, imaging, and sensor data from wearable devices, these models enable more informed decision-making for healthcare providers [12]. Applications extend to mental health care, where trends in speech and behavior can be analyzed, chronic disease management supported through continuous data monitoring, and vision care improved through early detection of conditions like diabetic retinopathy [13].
However, the deployment of LLMs in healthcare raises significant challenges, including concerns about patient privacy, model transparency, and ethical implications of automated decision-making. Meeting data privacy standards, such as those outlined by HIPAA, is critical [14]. Additionally, the inherent biases in LLMs must be addressed to mitigate potential negative consequences for patient care and diagnostic accuracy [15]. Addressing these technical, ethical, and regulatory concerns is vital for ensuring the safe implementation of LLMs in clinical environments.
This study systematically evaluates the diagnostic capabilities of five advanced Large Language Models (LLMs)—GPT-4, GPT-4o, Gemini, o1 Preview, and GPT-3.5—using a structured framework designed to ensure consistency and fairness in comparing their performance. The results demonstrate GPT-4’s exceptional diagnostic accuracy, driven by its extensive training on diverse datasets, making it effective in handling complex medical scenarios. Gemini stands out in high-stakes situations, prioritizing precision and minimizing false positives, while GPT-4o and o1 Preview exhibit a balanced performance, combining real-time applicability with reliable accuracy. GPT-3.5, while less advanced, proves useful for general diagnostic tasks and resource-limited settings. The study also highlights critical challenges faced by these models, including difficulties in interpreting ambiguous or nuanced symptoms, the need for robust privacy safeguards, and the risk of perpetuating biases inherent in their training data. These findings advance the understanding of LLMs’ diagnostic potential and establish a foundation for their responsible integration into clinical workflows.
2. Related Work
2.1. Advancements in LLM Capabilities for Healthcare Applications
The integration of Large Language Models (LLMs) into healthcare is transforming medical diagnostics and patient care [16]. These models enhance the precision and speed of diagnostic processes, especially in fields like radiology and pathology, which benefit from the detailed analysis and early detection these models enable [11]. LLMs also play a role in improving patient interactions by offering personalized consultations and symptom assessments, which foster trust and patient satisfaction. Beyond diagnostics, these models analyze diverse data sources—patient histories, imaging, and sensor data from wearable devices—to assist healthcare providers in informed decision-making [12]. For example, Meskó et al. [17] highlight how models like GPT-4 can process large datasets and deliver human-like responses, creating a multidimensional approach to healthcare by integrating textual, visual, and sensor-based data streams. This integration enables more comprehensive health assessments and ultimately improves patient outcomes. Other studies [13,18] have demonstrated how LLMs extend into mental health care by analyzing behavioral data to predict outcomes, a valuable application that supports mental health professionals in creating personalized treatment plans.
2.2. Challenges in Integrating LLMs into Healthcare
The integration of generative AI and LLMs in healthcare brings diverse and complex challenges [19,20,21,22,23,24,25,26]. Yu et al. [27] underscore the necessity for robust data privacy measures, precise model fine-tuning, and thorough implementation strategies to ensure AI deployment aligns with healthcare needs without compromising security or efficiency. They emphasize the role of collaborative co-design—engaging both clinicians and patients in AI development—to ensure tools are tailored to medical requirements while safeguarding data security and patient privacy [14]. Singh et al. [28] address psychological challenges, such as the cognitive biases in LLMs, which may lead to overconfidence or underestimation in diagnostic outputs. These biases could lead to errors in clinical decisions, underscoring the need for mechanisms that assess and adjust AI confidence levels. Ullah et al. [29] further discuss technical challenges in diagnostic medicine, particularly issues of contextual understanding and interpretability in fields like digital pathology. The inherent “black-box” nature of LLMs, combined with biases in training datasets, complicates their clinical acceptance and reliability [30]. However, the practical deployment of such systems faces significant challenges, including the reliability of AI applications in medical settings, the imperative for extensive clinical trials, and ongoing concerns about the confidentiality and security of patient data [31].
2.3. LLMs in Clinical Trials and Patient Monitoring
LLMs have shown potential in optimizing clinical trials and patient monitoring by enhancing data processing and predictive capabilities. Yuan et al. [32] demonstrate how LLMs can improve clinical trial efficiency through patient-trial matching by utilizing Electronic Health Records (EHRs). Their privacy-aware data augmentation strategy resulted in a 7.32% improvement in matching accuracy, enhancing both the speed and precision of patient selection for trials, thereby expediting research and treatment discovery. Jin et al. [33] and Kim et al. [34] discuss the Health-LLM system, which uses complex algorithms to analyze data from wearable sensors in real time. This system provides personalized health predictions that adapt to changes in patient conditions, especially for chronic disease management. These studies illustrate the capacity of LLMs to support continuous health monitoring and contribute to preventive care by encouraging adherence to prescribed health regimens. Xu et al. [35] explore Mental-LLM applications, which assess language patterns in real time to predict mental health trends, adding valuable insights into patient monitoring for mental health conditions. These challenges necessitate continuous advancements in AI technologies to improve their accuracy, reliability, and applicability in diverse clinical environments [36,37].
2.4. Applications and Limitations of LLMs in Diagnostics
The use of LLMs in diagnostics highlights both their potential and limitations. Meskó et al. [17] and Kusa et al. [12] provide foundational insights into LLMs’ capabilities for synthesizing multimodal data, which can enhance diagnostic accuracy and improve patient outcomes. However, Kusa et al. also point out challenges with LLM sensitivity to user input variations, which can lead to discrepancies in diagnostic results. These variations often stem from differing symptom descriptions and entrenched patient beliefs, underscoring the need for systems that can adjust to such differences to avoid diagnostic errors. Additionally, research by Abbasian et al. [23] examined how conversational health agents powered by LLMs enhanced patient experience in diagnostics. The reliance of LLMs on the quality of input data remains a significant barrier, as inconsistent input can affect model performance and reliability [27]. The insights from these studies are critical for understanding LLMs’ role in diagnostics and the steps needed to manage sensitivity and variability in user inputs.
2.5. Ethical and Technical Considerations for LLM Deployment
Deploying LLMs in healthcare requires rigorous attention to ethical and technical standards. Montagna et al. [11] emphasize the challenges of implementing LLM-based chatbots for chronic disease management, particularly within decentralized systems where data privacy and security are critical. Their proposed architecture for chronic disease management supports diverse medical conditions while adhering to strict privacy regulations, but its deployment faces challenges in reliability and the need for extensive clinical trials to validate performance. Similarly, Kim et al. [34] illustrate the technical difficulties of processing multimodal data in real time, especially in specialized healthcare contexts. Addressing these issues requires advancements in both AI technology and regulatory frameworks. Humphrey et al. [14] and Yuan et al. [38] argue for ethical AI practices and rigorous regulatory compliance to protect patient privacy and data integrity. Meeting these standards will be essential to maximize LLM utility in healthcare while ensuring responsible, safe, and secure integration.
3. Method
3.1. Research Strategy
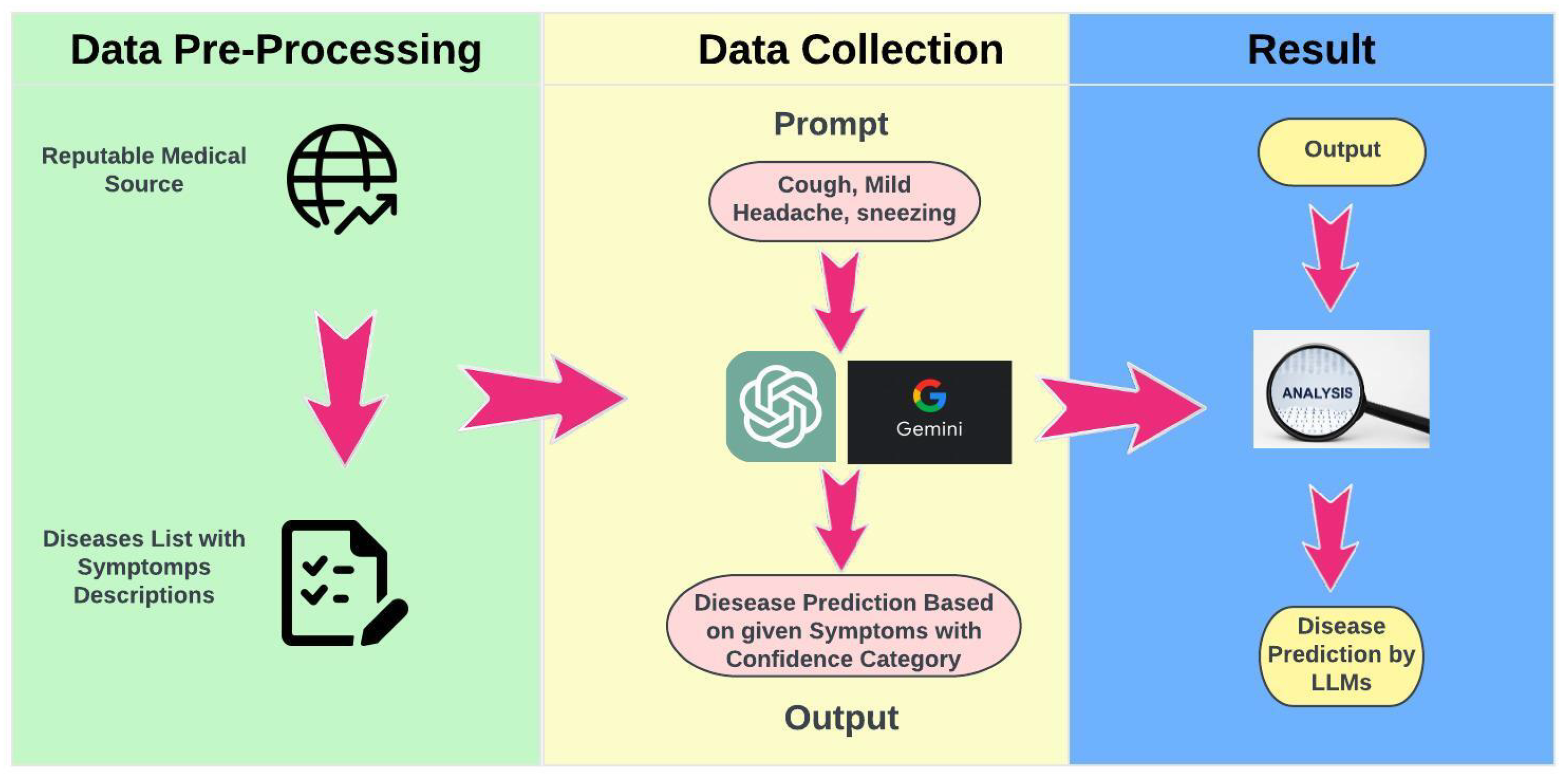
This study utilized a structured framework to evaluate the diagnostic capabilities of Large Language Models (LLMs) using symptom-based prompts derived from a carefully curated dataset (Figure 1). Each LLM was presented with the same sequence of prompts to ensure that observed differences in responses were attributable solely to the models’ unique interpretive capabilities. This standardized procedure facilitated consistent comparisons across models, maintaining uniformity in prompt presentation and response recording, while mitigating variability introduced by differing input formats or evaluation protocols. The curated dataset was constructed from reputable medical sources, including publicly available clinical guidelines and verified mappings of diseases to symptoms. Each prompt was designed to replicate real-world diagnostic scenarios by incorporating commonly reported symptom combinations, such as “cough, mild headache, sneezing”. The inclusion of diverse symptom profiles ensured that the models were evaluated on a wide spectrum of diagnostic complexities.
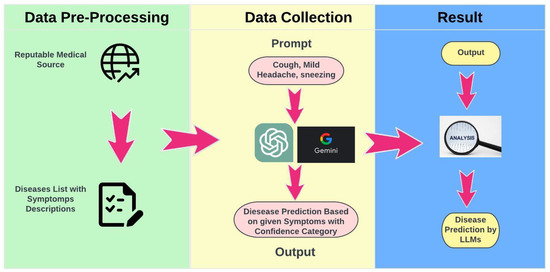
Figure 1.
Research Strategy.
Following the generation of predictions by the models, each response was systematically compared to the actual diagnoses associated with the provided symptom descriptions. This evaluation was conducted using a zero-shot diagnostic approach, wherein the models produced predictions without prior fine-tuning or domain-specific customization. Performance was assessed using key metrics, including precision, recall, and F1 scores, enabling a comprehensive evaluation of diagnostic accuracy. The systematic approach not only allowed for the identification of individual model strengths and limitations but also established a robust framework for evaluating the overall reliability and applicability of LLMs in healthcare. The consistency of this methodology minimized biases and external confounding factors, ensuring a fair and transparent evaluation of each model’s capabilities. These findings contribute to advancing the understanding of LLMs’ potential applications in medical diagnostics and provide a foundation for future research to address more complex clinical scenarios and integrate multimodal data.
3.2. Description of the LLMs Evaluated
This section introduces each of the Large Language Models (LLMs) used in the study—Gemini, GPT-3.5, GPT-4, o1 Preview, and GPT-4o. Each model demonstrates strengths in generating clinically relevant information and performing tasks such as clinical prediction, diagnosis, and providing data-driven insights to support health maintenance and recovery. These LLMs are widely accessible, commonly used by the general public, and are essential for rapid evaluation to determine their suitability and effectiveness in clinical applications and healthcare research. Each model possesses unique computational strengths for working with clinical datasets, contributing to enhanced diagnostic reliability in healthcare settings [12].
Gemini: Gemini marks a notable advancement in LLM technology, particularly with its specialized design for domain-specific applications, including healthcare. The architecture of Gemini is crafted to enable nuanced understanding and response generation in specialized fields, making it an invaluable tool for tasks such as medical diagnostics and healthcare inquiries, where precision and accuracy are critical. Gemini’s ability to integrate and reason across multimodal inputs underscores its potential to transform how medical information is processed, establishing a new standard for AI in healthcare [7].
GPT-3.5: As a foundational model, GPT-3.5, the predecessor to GPT-4, offers robust capabilities in language understanding and generation, though with slightly less proficiency compared to its successor. GPT-3.5 serves as a comparative benchmark to measure progress in LLMs and their applicability in medical diagnostics. Despite its earlier status, GPT-3.5 achieved a notable accuracy rate of 53 percent on the Medical Knowledge Self-Assessment Program, demonstrating its capability to understand and respond to clinical and healthcare-related inquiries, and illustrating its value in diagnostic tasks [5].
GPT-4: Developed by OpenAI, GPT-4 stands at the forefront of language understanding and generation capabilities, designed to interpret complex queries. Its architecture is aligned specifically for assessing diagnostic accuracy based on symptom descriptions. GPT-4 achieved a 75 percent accuracy rate on the Medical Knowledge Self-Assessment Program, highlighting its advanced understanding of complex medical questions and emphasizing its role in refining diagnostic accuracy from symptom narratives [5].
o1 Preview: As an iteration within the GPT series, o1 Preview prioritizes high diagnostic accuracy alongside real-time performance for medical applications. Its architecture is optimized for handling complex inquiries efficiently, making it particularly useful in settings that require timely diagnostic support. Built on the strengths of the GPT-4 architecture, o1 Preview is intended for seamless integration in fast-paced healthcare environments, supporting clinicians in making quick and precise diagnostic decisions, which positions it as a practical tool in modern medical practice [5].
GPT-4o: Designed as a specialized variant of GPT-4, GPT-4o emphasizes balanced, real-time diagnostic performance, especially in clinical settings. This model builds upon GPT-4’s capabilities, adapting them to workflows that demand a prompt, accurate analysis of patient symptoms. OpenAI developed GPT-4o with a focus on diagnostic accuracy while ensuring smooth integration into healthcare systems that require swift decision-making. By prioritizing real-time applicability, GPT-4o aids healthcare professionals in making well-informed diagnoses both efficiently and effectively [5].
3.3. Data Collection Methods
The dataset for this study was constructed using 50 distinct diseases, each evaluated by Large Language Models (LLMs): Gemini, GPT-3.5, GPT-4, GPT-4o, and o1 preview model. This approach resulted in a total of 250 individual evaluations (50 diseases × 5 models) as shown in Table 1.

Table 1.
Distribution of dataset evaluations across models.
Source and selection of diseases: The symptom data for the selected diseases were gathered from reputable medical sources, including the Centers for Disease Control and Prevention (CDC) [39], World Health Organization (WHO) [40], Mayo Clinic [41], Cleveland Clinic [42], and Johns Hopkins Hospital [43]. For each disease, a detailed list of symptoms was compiled, which formed the basis for creating diagnostic prompts.
Dataset and disease selection: The dataset encompassed 50 common diseases (Figure 2), such as seasonal allergies, the common cold, and food-related illnesses, which are frequently encountered in everyday medical practice. Each disease was tested against all models (Gemini, GPT-3.5, GPT-4, o1 Preview and GPT-4o), ensuring consistency in model comparison and yielding 250 total data points for the analysis. This dataset focused on widely recognized symptoms to evaluate the models’ ability to provide accurate diagnostic insights. The dataset can be accessed on GitHub: https://github.com/gkgupta11k/Health_LLM_Research_Dataset (accessed on 13 January 2025).

Figure 2.
Word cloud visualization of the diseases included in the dataset.
While the dataset included some diversity in symptom presentation, it did not incorporate extensive demographic variation such as age, gender, or ethnicity, as the study primarily focused on the conditions themselves rather than patient-specific variations. The current dataset allowed for an initial exploration of LLMs in medical diagnostics, but future studies will seek to expand this dataset to include a wider range of diseases and incorporate more diverse patient demographics, which will help evaluate the models’ generalizability across different populations.
Diagnostic procedure: For each selected disease, a comprehensive list of symptoms was compiled, forming the basis for diagnostic prompts. These prompts asked the LLMs to predict the disease based on the symptoms and to provide a confidence score for each prediction. To ensure consistency and comparability, the same prompts were applied uniformly across all models. After receiving the models’ predictions, the results were manually verified to assess the accuracy and reliability of the diagnoses, providing the foundation for the study’s findings.
This methodology underscores the study’s goal of exploring the potential of LLMs as tools for recognizing common health conditions. By focusing on frequently encountered diseases, this research aims to provide valuable insights into the capabilities and limitations of AI technologies in everyday health applications, while also laying the groundwork for future studies involving more complex and diverse cases.
In this study, each model was presented with 250 individual prompts (one for each combination of the 50 diseases and 5 LLMs), amounting to a total of 250 prompt iterations. Each prompt provided the model with a list of symptoms and requested a single-word diagnosis with a confidence level, without requiring additional explanations. The prompt design aligned with the zero-shot approach, where models are required to make diagnostic predictions based solely on the provided symptoms without any prior examples or step-by-step reasoning. This setup assessed each model’s inherent capability to interpret symptoms and identify the most likely disease independently, simulating a straightforward diagnostic task typical in clinical settings where minimal additional context is provided.
Prompt for models:
The following dialogue presents a prompt used to test the diagnostic capabilities of various language models.

3.4. Evaluation Metrics for Diagnosing Diseases Through LLMs
The efficacy of Language Learning Models (LLMs) in diagnosing diseases from descriptions of medical symptoms was evaluated using a detailed, multi-step process. This approach incorporated the use of precision, recall, and the F1 score—metrics renowned for their ability to provide a rounded perspective on the accuracy of predictive models in identifying correct diagnoses and highlighting the omission of relevant diagnoses.
In our study, we evaluated the LLMs’ outputs for each dataset entry, systematically classifying each response based on its diagnostic accuracy. The classifications were as follows:
- True positive (TP): instances where the LLM correctly identified the disease, showcasing the model’s ability to accurately match symptom descriptions with the correct disease diagnosis.
- False positive (FP): instances where the LLM incorrectly identified a disease, attributing a condition to the symptom descriptions that did not align with the actual disease present, thereby overestimating the model’s diagnostic accuracy.
- False negative (FN):instances where the LLM either attributed a different disease than the one actually present based on the symptom descriptions or failed to recognize the presence of a disease altogether, thereby underestimating the model’s diagnostic sensitivity.
We then proceeded to compute the following metrics based on the points assigned to each prediction. If the model’s prediction matched the actual disease, it was assigned 1 point in the true positive TP category. If the model predicted a disease that did not match the actual disease (an incorrect prediction), it was assigned 1 point in the false positive (FP) category. Finally, if the model failed to predict the correct disease, either by predicting an incorrect disease or by failing to predict any disease at all, it was assigned 1 point in the false negative (FN) category. After the evaluation was completed for each model, we then totaled the TP, FP, and FN points, and these totals were used in the following equations to calculate the precision, recall, and F1 score.
- Precision: this metric evaluates the exactness of the model’s positive predictions (i.e., the proportion of TP observations among all positive diagnoses made by the model), offering insight into the accuracy of the model’s disease identification.
- Recall: this metric assesses the model’s ability to identify all pertinent instances (i.e., the ratio of TP observations to all actual positives within the dataset), providing a measure of the model’s comprehensiveness in disease detection.
- F1 Score: This metric serves as a balanced measure of both precision and recall, particularly valuable when the contributions of both metrics are of equal importance. It is calculated as the harmonic mean of precision and recall, furnishing a singular measure of the model’s overall diagnostic performance.
Employing these metrics enabled a comprehensive evaluation of the LLMs’ diagnostic capabilities, providing nuanced insights into the precision of correct diagnoses and the models’ overall efficacy in disease identification.
4. Results
4.1. Overview of Findings
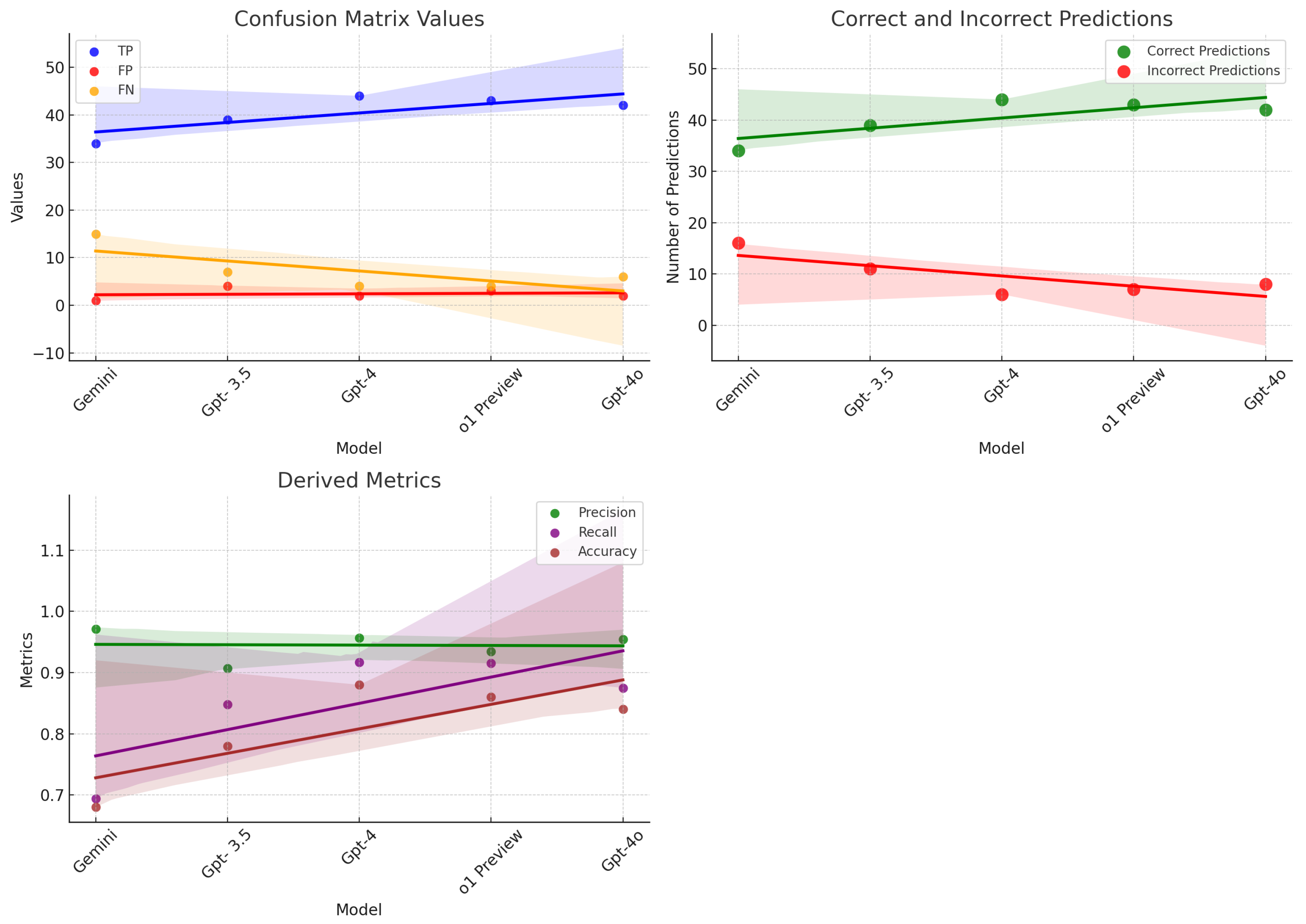
Our comprehensive evaluation delved into the diagnostic abilities of five state-of-the-art Language Learning Models (LLMs)—Gemini, GPT-3.5, GPT-4, o1 Preview, and GPT-4o. The goal was to assess how effectively these models could analyze and diagnose medical conditions based on detailed descriptions of symptoms. The findings, illustrated in Figure 3, revealed significant differences in the diagnostic accuracy and capabilities of each model, shedding light on their potential utility in clinical settings. GPT-4 emerged as the standout performer in our study, demonstrating exceptional diagnostic accuracy. This model’s success is attributable to its extensive training on a vast array of medical literature and patient data, which has equipped it with a profound understanding of medical symptomatology. GPT-4’s ability to consistently and accurately identify diseases from symptom descriptions showcases its advanced algorithmic structure and sophisticated data processing capabilities. It sets a benchmark in the realm of AI-driven medical diagnostics, proving to be a robust tool that could revolutionize how healthcare providers approach diagnosis and treatment planning. Close behind in terms of performance, o1 Preview also demonstrated impressive diagnostic abilities, achieving a precision of 0.93 and a recall of 0.91. This performance made it highly comparable to GPT-4, with slight variations in its ability to recall certain diseases. GPT-4o, another high-performing model, displayed a balanced diagnostic capability with a precision of 0.95 and a recall of 0.88, reinforcing its reliability in medical diagnostic tasks. GPT-3.5 displayed robust diagnostic skills as well. Although it did not surpass GPT-4, its effectiveness in converting complex symptom data into accurate health assessments makes it a valuable asset in the medical field. GPT-3.5 supports clinical decision-making by providing reliable interpretations of medical contexts, which can greatly aid physicians in diagnosing and understanding patient conditions more effectively. Its solid performance underlines the reliability of well-trained LLMs in handling medical diagnostic tasks, highlighting the potential for AI to assist significantly in everyday healthcare operations. Gemini, although it did not achieve as many correct diagnoses as its counterparts, was noted for its extraordinary diagnostic precision. This model adopts a conservative approach, prioritizing accuracy over quantity in its outputs. Such high-confidence predictions make Gemini especially suitable for use in clinical scenarios where accuracy is critical, such as in the diagnosis of complex or rare conditions where the cost of a misdiagnosis can be particularly high. Gemini’s approach to minimizing false positives is vital in clinical practices where precision is paramount, and the margin for error is minimal.
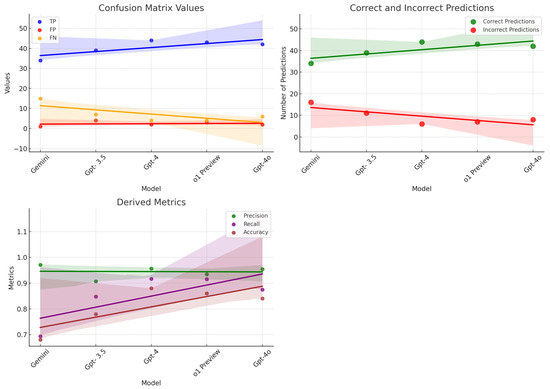
Figure 3.
Performance metrics and predictions of models: scatter plots showcasing confusion matrix values, derived metrics, and the correctness of model predictions.
The collective performance of these LLMs paints an optimistic picture of the role of AI in enhancing medical diagnostics. The integration of these advanced models into healthcare could lead to faster, more accurate, and highly reliable diagnostic processes. GPT-4’s broad diagnostic capabilities, GPT-3.5’s dependable performance, and Gemini’s meticulous precision collectively embody the advancement of artificial intelligence within the healthcare sector. The addition of o1 Preview and GPT-4o further expands the versatility of these models in medical diagnostics. These findings from our study not only underscore the substantial progress that AI technology has made in unraveling and understanding the complexities of human health but also pave the way for their future applications in medical practice. By enhancing the efficiency and accuracy of diagnostics, these models could serve as invaluable tools for medical practitioners, enabling better patient outcomes and transforming the landscape of healthcare delivery. As we continue to explore and refine these technologies, their integration into clinical workflows holds the promise of making healthcare more effective, personalized, and accessible to all.
4.2. Comparative Analysis
The comparative evaluation of Gemini, GPT-3.5, GPT-4, o1 Preview, and GPT-4o through our study provides an illuminating overview of their diagnostic abilities, each underscored by unique strengths as revealed by the performance metrics summarized in Table 2 and visually presented in Figure 3.

Table 2.
Comparative performance of LLMs in digital diagnostics.
GPT-4, with its outstanding number of correct answers, stood as a testament to its comprehensive training regimen, encompassing a wide array of medical data. This extensive preparation was reflected in its superior F1 score, indicative of the model’s proficiency in deciphering complex medical language and accurately mapping symptoms to diagnoses.
Figure 3 graphically displays the comparative accuracy of the models, highlighting GPT-4’s dominance in correctly answering questions, thereby showcasing its exceptional capability to navigate the complexities inherent in the symptom–diagnosis correlation. o1 Preview closely followed, with high accuracy in predicting diseases, while GPT-4o also displayed strong capabilities across multiple diagnostic tasks.
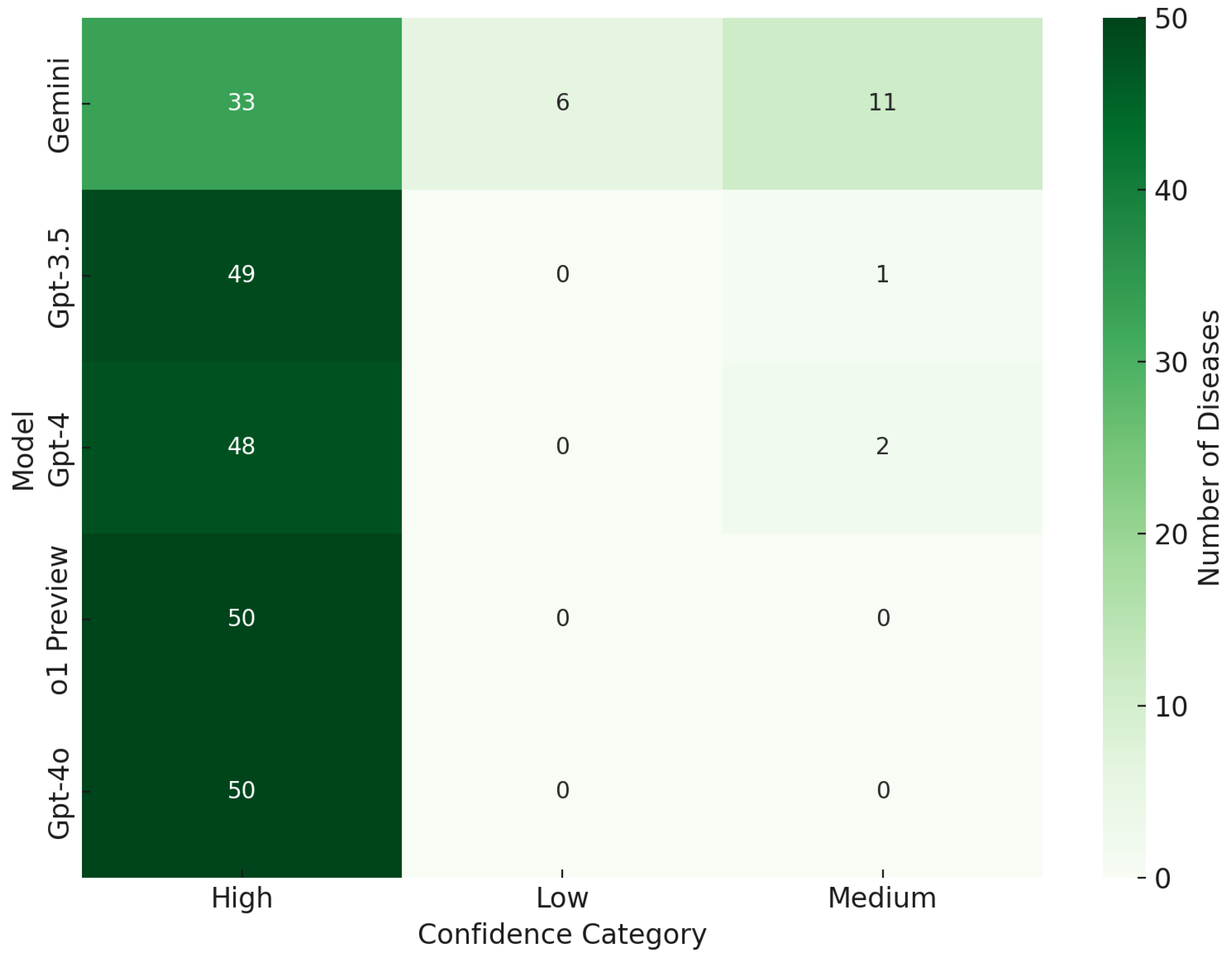
Further enhancing our analytical perspective, Figure 4 delves into the confidence levels associated with each model’s predictions. Here, the confidence distributions of GPT-4, GPT-4o, o1 Preview, and GPT-3.5 were primarily classified under the “High” category, underscoring their robust assertiveness and reliability in diagnostic conclusions. o1 Preview, similar to GPT-4, exhibited strong confidence in its predictions, maintaining a perfect record in the “High” confidence category with no low or medium confidence classifications, further highlighting its reliability for critical diagnostic tasks. On the other hand, Gemini’s inclination towards high-confidence responses, despite a lower overall number of predictions, spotlighted its unparalleled precision. This trait is particularly crucial in healthcare contexts where the stakes of a misdiagnosis are high, emphasizing the need for accuracy and high confidence in diagnostics.
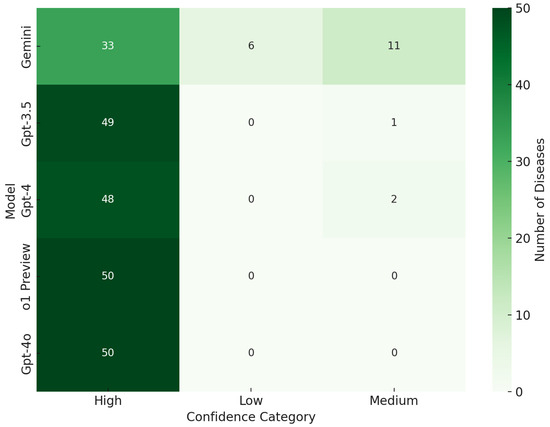
Figure 4.
Model confidence category.
However, Gemini’s impressive precision came at the cost of recall, as evidenced by its performance metrics. This suggests a cautious approach to diagnosis, where the model opts for certainty over breadth, potentially overlooking certain conditions in the process. Meanwhile, GPT-3.5, although not as advanced as GPT-4, demonstrated significant diagnostic utility, balancing precision and recall effectively. Its solid performance affirmed its value in scenarios where cutting-edge models like GPT-4 might not be available or necessary.
Integrating these insights, the study underscored the multifaceted diagnostic capabilities inherent in these LLMs. GPT-4 emerged as a versatile asset across various medical domains, whereas Gemini’s precision earmarked it as an invaluable resource for confirming diagnoses with high confidence. On the other hand, GPT-3.5’s reliable performance ensured its continued relevance in the evolving landscape of AI in healthcare. The addition of o1 Preview and GPT-4o to this analysis emphasized the growing role of LLMs in providing scalable and reliable diagnostic support in clinical environments.
The findings from our study advocate for a strategic incorporation of LLMs within healthcare settings, leveraging each model’s distinct strengths. Such an approach not only augments the diagnostic process but also enhances the overall quality of care, paving the way for an AI-integrated healthcare ecosystem that prioritizes accuracy, efficiency, and patient safety.
5. Discussion
5.1. Interpretation of Results
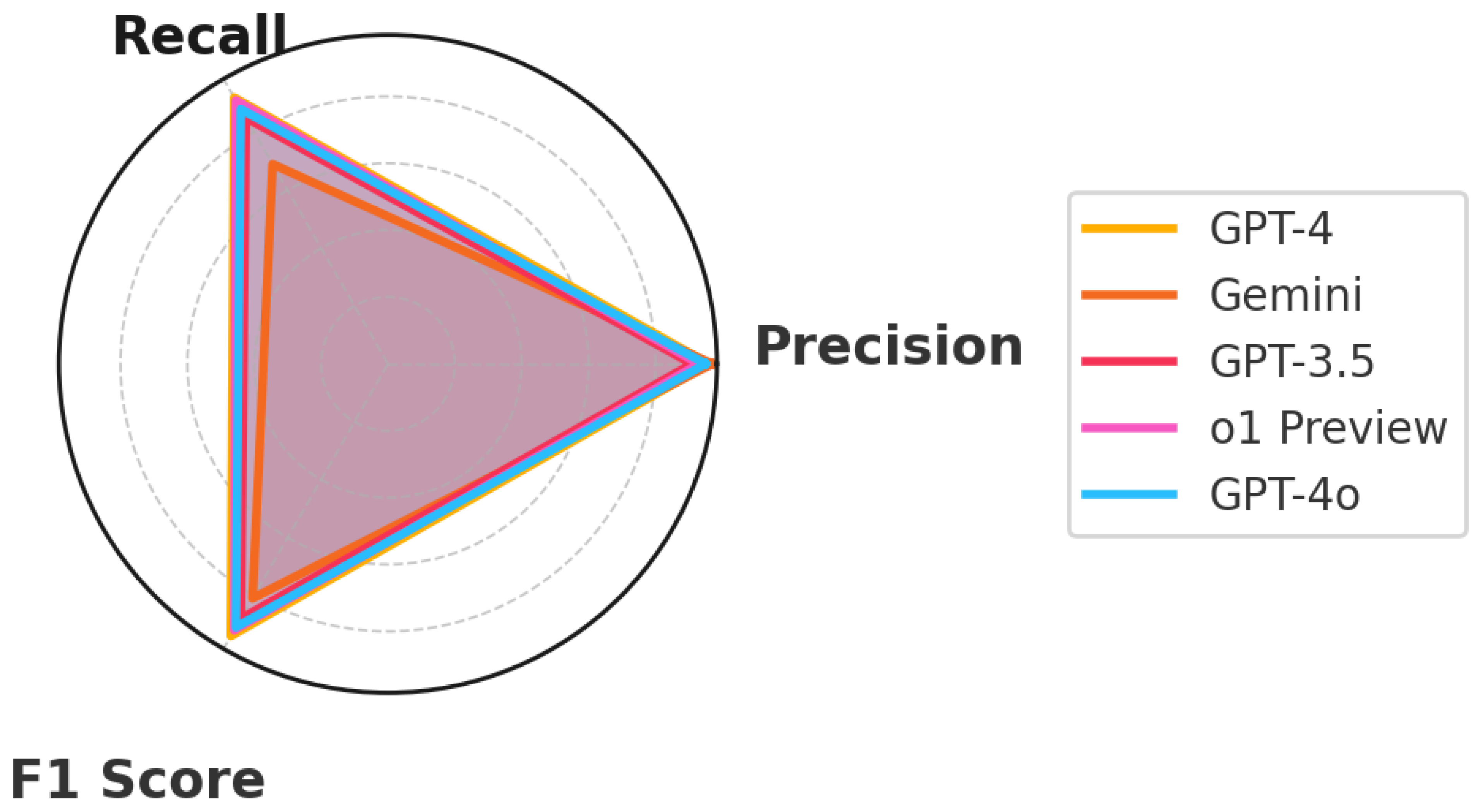
This study carefully examined the capabilities of five advanced Language Learning Models (LLMs) diagnosing everyday illnesses based on symptom descriptions (Figure 5). The goal was to assess how these models could assist in medical diagnostics by analyzing symptom descriptions and mapping them to possible conditions. The standout performer, GPT-4, showcased the potential of AI in medical diagnostics through its ability to understand and process complex medical data. This model’s effectiveness highlighted its extensive training across diverse medical scenarios, making it exceptionally good at matching symptoms with the correct medical conditions. o1 Preview, closely following GPT-4, demonstrated impressive diagnostic capabilities, with a strong balance of precision and recall, making it a reliable alternative for clinical applications. GPT-4o, also highly accurate, displayed balanced diagnostic abilities, emphasizing its potential in real-time medical settings. Although not as advanced as GPT-4, GPT-3.5 still demonstrated significant capability in making accurate medical diagnoses. Its ability to deliver reliable assessments makes it a useful tool in healthcare, particularly where newer technologies might not yet be available. Gemini, known for its high precision, focused on accuracy in its predictions, which is especially important in medical settings where mistakes can have serious consequences.
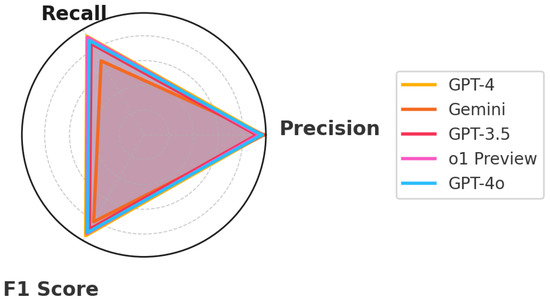
Figure 5.
Performance metrics.
5.2. Enhancing Diagnostic Processes with Large Language Models
The introduction of Large Language Models (LLMs) into the medical context is likely to change the way healthcare is provided at the very first stages of contact between a patient and a healthcare professional. In the future, LLMs will likely improve the speed and quality of first medical consultations, allowing more rapid assessments of patient information, thereby taking much pressure off medical experts or allowing other professionals to fill the role. In settings where resources are scarce, especially when access to first human consultations is unfeasible, LLMs could quickly analyze the data provided by symptoms and tell the patient what they might be suffering from [30,44].
In triaging these cases, LLMs could help to prioritize patients based on urgency and route them to the appropriate level of care. All of this could lead to improved patient outcomes, such as faster interventions, fewer missed appointments, and reduced wait times before treatment. Patient education is also one of the benefits of using these models. LLMs can also be used to provide patients with additional information about their symptoms and potential diagnoses—additional knowledge that empowers people to better understand their health [12].
Yet, such incorporation of AI-enabled tools in healthcare systems is bound to be fraught with hurdles, and we shall have to navigate several hefty issues. For example, adherence to healthcare regulations, such as the Health Insurance Portability and Accountability Act (HIPAA) in the United States, comprehensive federal legislation that sets metalegal standards and outlines the protection and secure transfer of patients’ data in healthcare systems [14]. All the information gathered by LLMs during healthcare interactions must be strictly HIPAA-compliant; they should master the vital data and conversation with full security and confidentiality.
In addition, LLMs might prove valuable, but if integrated, their use must complement, rather than substitute, the human side of delivering care, augmenting physician sensibilities of the human system over replacing the physician. Error rates of LLMs for complex diagnoses need to be constantly validated against medical benchmarks [28]. Deviating from the norm could result in a misdiagnosis. Moreover, the models also need to be kept up-to-date through regular checking and flagging.
That commitment to multidisciplinary work among technologists, clinicians, and regulators is also crucial to using LLMs in healthcare effectively. From conceiving AI tools that are both architecture- and design-wise, solid, and clinically useful to providing right-size validation and ensuring ethical use, stringent workflows across disciplines are necessary. Testing each model goes beyond mere optimization of technical performance to confirm that the tool is capable of effectively helping clinicians in actual practice [45].
In conclusion, incorporating LLMs into healthcare is a promising avenue for improving diagnosis and patient care. However, to address these challenges, we must implement careful planning and robust processes. Doing so will help to make the most of AI in healthcare and improve outcomes for patients and the medical system.
5.3. Limitations of the Study
The LLMs in this study were primarily evaluated on common, less complex illnesses, which do not fully represent the broader, more challenging aspects of diagnosing chronic or severe conditions. Chronic diseases often involve complex symptoms that are difficult to interpret without a comprehensive understanding of an individual’s medical history and additional diagnostic tests. Moreover, the reliance on text analysis in our study ignores the multimodal nature of traditional medical diagnostics, which often include visual elements like scans, detailed patient histories, and physical examinations. Future improvements in AI models for healthcare should aim to incorporate these various data types to provide more accurate and holistic diagnoses.
5.4. Future Research Directions
Future research should focus on creating LLMs that can analyze various types of medical information beyond just text. This includes integrating visual data from scans and other medical images to create more comprehensive diagnostic tools. Also, developing AI systems that can understand and process information across different languages and cultural contexts will be crucial for their global applicability. This study used a proof-of-concept dataset, and we plan to explore this further on a larger-scale dataset in the future to better evaluate the models’ performance across more complex medical conditions. Expanding the dataset will allow for deeper insights into the LLMs’ capabilities and limitations in diagnosing a wider range of health issues.
Furthermore, it is crucial to ensure that these systems are developed with strict adherence to ethical standards, particularly regarding patient privacy. Exploring ways to securely integrate AI into healthcare while respecting patient confidentiality will be essential. Additionally, putting these AI models into real-world clinical settings to assess their performance and impact on healthcare efficiency and patient outcomes will provide critical insights into their practical value and limitations.
This research demonstrates the significant potential and challenges of using advanced AI models in healthcare diagnostics. With careful development and ethical considerations, these tools could greatly enhance the ability to diagnose and treat patients more efficiently and accurately. Continued exploration and improvement of these technologies could lead to their successful integration into everyday medical practice, benefiting both healthcare providers and patients.
6. Conclusions
The exploration into the capabilities of Large Language Models (LLMs) like Gemini, GPT-3.5, GPT-4, o1 Preview, and GPT-4o, culminated in a comprehensive understanding of their potential to enhance digital diagnostics in healthcare. This study rigorously evaluated the performance of these models, shedding light on their strengths in identifying symptoms of common illnesses and their limitations. The core findings demonstrated that LLMs like GPT-4 offered considerable promise in processing medical language with high accuracy. They heralded a significant step forward in providing immediate, accessible healthcare guidance. o1 Preview and GPT-4o also performed impressively, showcasing strong diagnostic capabilities with balanced precision and recall, making them reliable tools for real-time medical applications. The specialized Gemini model’s remarkable precision pointed towards the feasibility of creating niche, domain-focused LLMs that could provide precise diagnostic support. GPT-3.5, while slightly overshadowed by its successors, still displayed commendable capabilities, indicative of the rapid advancements within the field of AI in healthcare. These results reinforce the transformative potential of LLMs in digital diagnostics, suggesting that they can complement conventional diagnostic methods, enhancing the quality and accessibility of patient care. However, the journey from potential to actualized utility in a clinical setting will require overcoming substantial hurdles, including the integration of multimodal data, ethical considerations, and ensuring adherence to stringent healthcare standards. The limitations of this study, predominantly its scope restricted to textual analysis and the manual evaluation process, set a clear directive for future research. The next phase should aim to expand the capabilities of LLMs beyond text, incorporating visual and empirical data to align closely with comprehensive clinical diagnostics. Moreover, the study emphasizes the necessity to build ethically aligned, culturally sensitive, and linguistically diverse LLMs to serve global healthcare needs effectively. In the quest to harness AI for healthcare, LLMs emerge not as standalone solutions but as part of a collaborative toolset augmenting the expertise of medical professionals. As research progresses, it will be paramount to embed these models within real-world clinical workflows to fully assess their practicality and reliability. With continued development and responsible implementation, LLMs are poised to play a pivotal role in shaping the future of healthcare, making diagnostics more accessible, accurate, and patient-centric.
Author Contributions
Conceptualization, G.K.G. and A.S.; methodology, G.K.G.; software, G.K.G.; validation, G.K.G., A.S. and S.V.M.; formal analysis, G.K.G.; investigation, G.K.G.; resources, G.K.G.; data curation, G.K.G.; writing—original draft preparation, G.K.G.; writing—review and editing, G.K.G., A.S., S.V.M. and A.E.; visualization, G.K.G.; supervision, A.S.; project administration, G.K.G. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data are available on GitHub at https://github.com/gkgupta11k/Health_LLM_Research_Dataset (accessed on 13 January 2025).
Conflicts of Interest
Author Abul Ehtesham was employed by the Davey Tree Expert Company. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| LLM | Large Language Model |
| GPT | Generative Pre-trained Transformer |
| HIPAA | Health Insurance Portability and Accountability Act |
| NLP | Natural Language Processing |
| AI | Artificial Intelligence |
| CDC | Centers for Disease Control and Prevention |
| WHO | World Health Organization |
| EHR | Electronic Health Records |
| FP | False positive |
| TP | True positive |
| FN | False negative |
References
- Minaee, S.; Mikolov, T.; Nikzad, N.; Chenaghlu, M.; Socher, R.; Amatriain, X.; Gao, J. Large Language Models: A Survey. arXiv 2024, arXiv:2402.06196. [Google Scholar]
- Zhao, W.X.; Zhou, K.; Li, J.; Tang, T.; Wang, X.; Hou, Y.; Min, Y.; Zhang, B.; Zhang, J.; Dong, Z.; et al. A Survey of Large Language Models. arXiv 2023, arXiv:2303.18223. [Google Scholar]
- Singh, A. Exploring Language Models: A Comprehensive Survey and Analysis. In Proceedings of the 2023 International Conference on Research Methodologies in Knowledge Management, Artificial Intelligence and Telecommunication Engineering (RMKMATE), Chennai, India, 1–2 November 2023; pp. 1–4. [Google Scholar] [CrossRef]
- Hadi, M.U.; Al Tashi, Q.; Qureshi, R.; Shah, A.; Muneer, A.; Irfan, M.; Zafar, A.; Shaikh, M.B.; Akhtar, N.; Hassan, S.Z.; et al. Large Language Models: A Comprehensive Survey of its Applications, Challenges, Limitations, and Future Prospects. TechRxiv 2024. [Google Scholar] [CrossRef]
- OpenAI. GPT-4 Technical Report. arXiv 2023, arXiv:2303.08774. [Google Scholar]
- Choudhury, A.; Chaudhry, Z. Large Language Models and User Trust: Focus on Healthcare. J. Med. Internet Res. 2024, 26, e56764. [Google Scholar] [CrossRef]
- Team, G. Gemini: A Family of Highly Capable Multimodal Models. arXiv 2024, arXiv:2312.11805. [Google Scholar]
- Webster, P. Six Ways Large Language Models are Changing Healthcare. Nat. Med. 2023, 29, 2969–2971. [Google Scholar] [CrossRef]
- Peng, W.; Feng, Y.; Yao, C.; Zhang, S.; Zhuo, H.; Qiu, T.; Zhang, Y.; Tang, J.; Gu, Y.; Sun, Y. Evaluating AI in Medicine: A Comparative Analysis of Expert and ChatGPT Responses to Colorectal Cancer Questions. Sci. Rep. 2024, 14, 2840. [Google Scholar] [CrossRef]
- Cui, H.; Fang, X.; Xu, R.; Kan, X.; Ho, J.C.; Yang, C. Multimodal Fusion of EHR in Structures and Semantics: Integrating Clinical Records and Notes with Hypergraph and LLM. arXiv 2024, arXiv:2403.08818. [Google Scholar]
- Montagna, S.; Ferretti, S.; Klopfenstein, L.C.; Florio, A.; Pengo, M.F. Data Decentralisation of LLM-Based Chatbot Systems in Chronic Disease Self-Management. In Proceedings of the 2023 ACM Conference on Information Technology for Social Good, Lisbon, Portugal, 6–8 September 2023; pp. 205–212. [Google Scholar] [CrossRef]
- Kusa, W.; Mosca, E.; Lipani, A. “Dr LLM, what do I have?”: The impact of user beliefs and prompt formulation on health diagnoses. In Proceedings of the Third Workshop on NLP for Medical Conversations; Khosla, S., Ed.; Association for Computational Linguistics: Bali, Indonesia, 2023; pp. 13–19. [Google Scholar] [CrossRef]
- Lai, T.; Shi, Y.; Du, Z.; Wu, J.; Fu, K.; Dou, Y.; Wang, Z. Psy-LLM: Scaling up Global Mental Health Psychological Services with AI-based Large Language Models. arXiv 2023, arXiv:2307.11991. [Google Scholar]
- Humphrey, B.A. Data Privacy vs. Innovation: A Quantitative Analysis of Artificial Intelligence in Healthcare and Its Impact on HIPAA regarding the Privacy and Security of Protected Health Information. Ph.D. Thesis, Robert Morris University, Moon Twp, PA, USA, 2021. [Google Scholar]
- Dhakal, U.; Singh, A.K.; Devkota, S.; Sapkota, Y.; Lamichhane, B.; Paudyal, S.; Dhakal, C. GPT-4’s assessment of its performance in a USMLE-based case study. arXiv 2024, arXiv:2402.09654. [Google Scholar]
- Denecke, K.; May, R.; Rivera Romero, O. Potential of Large Language Models in Health Care: Delphi Study. J. Med. Internet Res. 2024, 26, e52399. [Google Scholar] [CrossRef]
- Meskó, B.; Hetényi, G.; Győrffy, Z. The role of artificial intelligence in precision medicine. Expert Rev. Precis. Med. Drug Dev. 2017, 2, 239–241. [Google Scholar] [CrossRef]
- Singh, A.; Ehtesham, A.; Mahmud, S.; Kim, J.H. Revolutionizing Mental Health Care through LangChain: A Journey with a Large Language Model. In Proceedings of the 2024 IEEE 14th Annual Computing and Communication Workshop and Conference (CCWC), Las Vegas, NV, USA, 8–10 January 2024; pp. 73–78. [Google Scholar] [CrossRef]
- de Curtò, J.; de Zarzà, I.; Roig, G.; Cano, J.C.; Manzoni, P.; Calafate, C.T. LLM-Informed Multi-Armed Bandit Strategies for Non-Stationary Environments. Electronics 2023, 12, 2814. [Google Scholar] [CrossRef]
- Chiu, Y.Y.; Sharma, A.; Lin, I.W.; Althoff, T. A Computational Framework for Behavioral Assessment of LLM Therapists. arXiv 2024, arXiv:2401.00820. [Google Scholar]
- Batsis, J.A.; Mackenzie, T.A.; Emeny, R.T.; Lopez-Jimenez, F.; Bartels, S.J. Low Lean Mass With and Without Obesity, and Mortality: Results From the 1999–2004 National Health and Nutrition Examination Survey. J. Gerontol. Ser. A 2017, 72, 1445–1451. [Google Scholar] [CrossRef]
- Baharudin, N.; Mohamed-Yassin, M.S.; Daher, A.M.; Ramli, A.S.; Khan, N.M.N.; Abdul-Razak, S. Prevalence and factors associated with lipid-lowering medications use for primary and secondary prevention of cardiovascular diseases among Malaysians: The REDISCOVER study. BMC Public Health 2022, 22, 228. [Google Scholar] [CrossRef]
- Abbasian, M.; Azimi, I.; Rahmani, A.M.; Jain, R. Conversational Health Agents: A Personalized LLM-Powered Agent Framework. arXiv 2024, arXiv:2310.02374. [Google Scholar]
- Meng, X.; Yan, X.; Zhang, K.; Liu, D.; Cui, X.; Yang, Y.; Zhang, M.; Cao, C.; Wang, J.; Wang, X.; et al. The application of large language models in medicine: A scoping review. iScience 2024, 27, 109713. [Google Scholar] [CrossRef]
- Clusmann, J.; Kolbinger, F.R.; Muti, H.S.; Carrero, Z.I.; Eckardt, J.N.; Laleh, N.G.; Löffler, C.M.L.; Schwarzkopf, S.C.; Unger, M.; Veldhuizen, G.P.; et al. The future landscape of large language models in medicine. Commun. Med. 2023, 3, 141. [Google Scholar] [CrossRef] [PubMed]
- Reese, J.T.; Danis, D.; Caufield, J.H.; Groza, T.; Casiraghi, E.; Valentini, G.; Mungall, C.J.; Robinson, P.N. On the limitations of large language models in clinical diagnosis. medRxiv 2023. [Google Scholar] [CrossRef]
- Yu, K.; Beam, A.L.; Kohane, I.S. Artificial intelligence in healthcare: A critical analysis of the legal and ethical implications. Int. J. Med. Inform. 2020, 141, 104431. [Google Scholar]
- Singh, A.K.; Lamichhane, B.; Devkota, S.; Dhakal, U.; Dhakal, C. Do Large Language Models Show Human-like Biases? Exploring Confidence—Competence Gap in AI. Information 2024, 15, 92. [Google Scholar] [CrossRef]
- Ullah, E.; Parwani, A.; Baig, M.M.; Singh, R. Challenges and Barriers of Using Large Language Models (LLM) Such as ChatGPT for Diagnostic Medicine with a Focus on Digital Pathology—A Recent Scoping Review. Diagn. Pathol. 2024, 19, 43. [Google Scholar] [CrossRef]
- Jo, E.; Epstein, D.A.; Jung, H.; Kim, Y.H. Understanding the Benefits and Challenges of Deploying Conversational AI Leveraging Large Language Models for Public Health Intervention. In Proceedings of the 2023 CHI Conference on Human Factors in Computing Systems, Hamburg, Germany, 23–28 April 2023; Association for Computing Machinery: New York, NY, USA, 2023. Article 18. pp. 1–16. [Google Scholar] [CrossRef]
- Baric-Parker, J.; Anderson, E. Patient Data-Sharing for AI: Ethical Challenges, Catholic Solutions. Linacre Q. 2020, 87, 471–481. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Tang, R.; Jiang, X.; Hu, X. LLM for Patient-Trial Matching: Privacy-Aware Data Augmentation Towards Better Performance and Generalizability. Am. Med. Inform. Assoc. (AMIA) Annu. Symp. 2024. Available online: https://par.nsf.gov/biblio/10448809 (accessed on 13 January 2025).
- Jin, M.; Yu, Q.; Shu, D.; Zhang, C.; Fan, L.; Hua, W.; Zhu, S.; Meng, Y.; Wang, Z.; Du, M.; et al. Health-LLM: Personalized Retrieval-Augmented Disease Prediction System. arXiv 2024, arXiv:2402.00746. [Google Scholar]
- Kim, Y.; Xu, X.; McDuff, D.; Breazeal, C.; Park, H.W. Health-LLM: Large Language Models for Health Prediction via Wearable Sensor Data. arXiv 2024, arXiv:2401.06866. [Google Scholar]
- Xu, X.; Yao, B.; Dong, Y.; Gabriel, S.; Yu, H.; Hendler, J.; Ghassemi, M.; Dey, A.K.; Wang, D. Mental-LLM: Leveraging Large Language Models for Mental Health Prediction via Online Text Data. Proc. ACM Interact. Mob. Wearable Ubiquitous Technol. 2024, 8, 32. [Google Scholar] [CrossRef]
- Ghosh, A.; Acharya, A.; Jain, R.; Saha, S.; Chadha, A.; Sinha, S. CLIPSyntel: CLIP and LLM Synergy for Multimodal Question Summarization in Healthcare. arXiv 2023, arXiv:2312.11541. [Google Scholar] [CrossRef]
- Shieh, A.; Tran, B.; He, G.; Kumar, M.; Freed, J.A.; Majety, P. Assessing ChatGPT 4.0’s test performance and clinical diagnostic accuracy on USMLE STEP 2 CK and clinical case reports. Sci. Rep. 2024, 14, 9330. [Google Scholar] [CrossRef]
- Yuan, J.; Tang, R.; Jiang, X.; Hu, X. Large Language Models for Healthcare Data Augmentation: An Example on Patient-Trial Matching. AMIA Annu. Symp. Proc. 2024, 2023, 1324–1333. [Google Scholar]
- Centers for Disease Control and Prevention. Symptoms of Diseases. Available online: https://www.cdc.gov (accessed on 9 October 2024).
- World Health Organization. Disease Symptoms and Information. Available online: https://www.who.int (accessed on 9 October 2024).
- Mayo Clinic. Symptoms of Common Diseases. Available online: https://www.mayoclinic.org (accessed on 9 October 2024).
- Cleveland Clinic. Disease Symptoms and Conditions. Available online: https://my.clevelandclinic.org (accessed on 9 October 2024).
- Johns Hopkins Hospital. Symptoms and Disease Information. Available online: https://www.hopkinsmedicine.org (accessed on 9 October 2024).
- Sallam, M. ChatGPT Utility in Healthcare Education, Research, and Practice: Systematic Review on the Promising Perspectives and Valid Concerns. Healthcare 2023, 11, 887. [Google Scholar] [CrossRef] [PubMed]
- Frantzidis, C.A.; Bamidis, P.D. Description and Future Trends of ICT Solutions Offered Towards Independent Living: The Case of LLM Project. In Proceedings of the 2nd International Conference on PErvasive Technologies Related to Assistive Environments, Corfu, Greece, 9–13 June 2009; p. 59. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).