1. Introduction
The tumor suppressor gene
TP53 is the most commonly mutated gene in human cancer [
1]. In breast cancer, mutation of
TP53 occurs at a frequency of 25–35% and is often associated with an earlier age at onset, a poor prognosis and a triple-negative molecular subtype [
2,
3,
4,
5]. p53 is a transcription factor and maintains genomic integrity primarily by regulating downstream target genes to induce apoptosis, cell cycle arrest, senescence, or other stress responses to halt the progression of cancer [
6,
7]. In addition to mutation, decreasing the level of wildtype
TP53 is also a major mechanism of functional inactivation of p53 [
8]. MDM2 (Mouse Double Minute 2 homolog) is a key negative regulator of p53, being both a target gene of p53 and an E3 ubiquitin ligase that targets p53 for proteasomal degradation, which maintains low p53 protein levels in the absence of stress [
8,
9,
10].
Mdm2 knockout in mice leads to early embryonic lethality, whereas mice with a deletion of both
Mdm2 and
p53 are viable and develop normally, demonstrating that the negative regulation of p53 is a critical role of MDM2 [
11,
12]. Accordingly,
MDM2 amplification and/or overexpression may be an important mechanism of aberrant p53 inactivation, and has been observed in many tumors, often mutually exclusive with
TP53 mutation [
10,
13].
SNP309T>G (rs2279744) is a sequence variant in the
MDM2 intronic promoter P2, and its minor G-allele was found to increase binding of transcription factor SP1, P2 promoter activity, and
MDM2 expression [
14]. In the initial study, the SNP309G-allele was found associated with a younger age at onset among individuals with Li-Fraumeni syndrome [
14]. However, most subsequent studies of various types of sporadic cancer did not find such a premature onset associated with the G-allele, and revealed inconsistent results with respect to cancer risk. Overall, most studies in Asian populations showed an association of the SNP309GG genotype with an increased cancer risk, whereas studies in Caucasians predominantly found no association [
15,
16,
17,
18,
19].
A potential mechanistic explanation for these discordant results was provided by the more recent discovery of another SNP in the
MDM2 P2 promoter, SNP285G>C (rs117039649) [
20,
21]. SNP285 is only 24 bp upstream of SNP309 and in complete linkage disequilibrium with it. The minor SNP285C allele was found exclusively in Caucasians, and population genetics has revealed that it has arisen on a pre-existing SNP309G allele after the separation between Caucasians and modern day East Asians [
22]. Like SNP309, SNP285 is also located at the edge of a (different) SP1 binding site, however, whereas the SNP309G allele enhances binding of SP1, the SNP285C allele reduces binding, and might thus antagonize the molecular effects of SNP309G [
20]. In addition, SNP285 is located within an estrogen receptor binding site (ERE) that overlaps with the SP1 site [
23]. Only a few studies, mostly in Northwestern European populations, have analyzed the association of SNP285 with cancer risk, and revealed a trend for SNP285C to be associated with a reduced risk of gynecological cancers, but not other cancer types [
20,
21,
23,
24,
25,
26,
27,
28,
29]. Some of these studies have found a decreased breast cancer risk associated with the SNP285C allele among carriers of the SNP309GG genotype, but not among carriers of the SNP309TG genotype, and not among subjects unselected for the SNP309 genotype [
20,
21,
24,
25,
29,
30].
The aim of our study was to determine the distribution and allele frequencies of MDM2 SNP309 and SNP285 in a Central European (Austrian) study population. This closes an important geographical gap between the many Northern and several Southern European populations studied thus far, particularly in light of their divergent SNP285 allele frequencies. We further sought to analyze the association of SNP285 and SNP309 with breast cancer risk, with the age at breast cancer onset, with key clinical and histopathological parameters of breast cancer, in particular the p53 status, and with breast cancer survival and prognosis. By studying the potential mutual interference between SNP309 and SNP285 with respect to these associations, we aimed to address the hypothesis that SNP285 antagonizes the effects of SNP309, thus potentially confounding the many studies of SNP309 in Caucasians that have not taken SNP285 into account.
3. Discussion
The discovery of SNP285 in
MDM2 promoter P2 has led to an elegant hypothesis potentially solving a long-standing conundrum: Why is the SNP309G-allele associated with an increased cancer risk in Asian, but not in Caucasian populations [
16,
17,
18,
19,
20,
21,
27,
29]? According to this hypothesis, SNP309 likely is associated with cancer risk, consistent with the finding that genetically engineered SNP309GG mice are more tumor-prone than SNP309TT mice [
47]. However, this association is apparently confounded in Europeans by the adjacent SNP285, which also affects SP1 binding to another site, antagonizes the effects of SNP309, and is present at a low frequency in Europeans, but absent in Asians [
20,
22]. On the other hand, the present study, together with previous ones, found no evidence to support this hypothesis. Like many previous studies in European populations, we found that SNP309 is not significantly associated with breast cancer risk, although a trend for an increased risk associated with the GG genotype in the recessive inheritance model was observed [
16,
18,
19]. However, when we corrected for the potential confounding effect of SNP285, we still did not observe any significant associations. In fact, the observed odds ratios associated with SNP309 genotypes hardly changed upon exclusion of carriers of the SNP285C allele or adjustment for SNP285, consistent with previous reports [
25,
32]. We conclude that the discordant results between European and Asian populations with respect to the association of SNP309 with cancer risk must be due to differences other than the presence vs. absence of the variant C-allele of SNP285.
In the initial study of SNP309, the GG genotype was found to be associated with a significantly younger age at cancer onset in Li-Fraumeni patients [
14]. This finding was confirmed in most subsequent studies in Li-Fraumeni patients [
36,
37,
38,
39,
40]. Interestingly, an association of the SNP309GG genotype with an earlier onset was also observed in several studies of familial breast cancer, particularly in
BRCA1 mutation carriers [
17,
27,
48,
49]. However, most studies of sporadic breast cancer did not find such a premature onset associated with the SNP309G-allele [
18,
21,
50,
51,
52,
53]. Here we report an effect opposite to that in familial cancer syndromes: In our study population, the SNP309TT genotype is associated with a younger age at breast cancer onset, consistent with a previous report [
17]. Like our study, Lum et al. reported an association of the SNP309GG genotype with an increased breast cancer risk, but a delayed onset [
17]. This is unexpected given the proposed molecular function of the SNP309G-allele to increase the expression of
MDM2 and hence to attenuate the p53-pathway. We hypothesize that this seemingly paradoxical association may be due to the differential rate of p53 positivity found by us and others, which is low in SNP309GG, but high in TT-tumors (see discussion below). It has been convincingly established that
TP53 mutant patients have a much younger age at onset [
2,
3,
4]. Accordingly, we propose that the SNP309TT genotype leads to a higher selective pressure to mutate
TP53 and hence to a much higher rate of these mutations, which then causes the earlier mean age at onset associated with that genotype. This effect of differential somatic
TP53 mutation rate obviously does not exist in Li-Fraumeni patients who have a germline mutation in
TP53, which may explain why they differ from sporadic cancer cases with respect to the association of SNP309 with the age at cancer onset. Consistently, we found an association of the age at breast cancer onset with
MDM2 mRNA expression. Significantly elevated mean
MDM2 mRNA levels were observed in patients with an age ≥55 years, as well as in post-menopausal patients (H. Miedl and M. Schreiber, unpublished data).
We found that the SNP309GG genotype is significantly associated with an increased breast cancer risk in p53 negative patients, who harbour a functional p53 protein (OR, 1.82; 95% CI; 1.09–3.03; and
p = 0.02), but not in p53 positive or unselected patients (
Table 1 and
Table 3), consistent with a previous report [
50]. Thus, the impact of SNP309 on breast cancer risk appears to be limited to tumors in which p53 is functional, which is biologically plausible since the key function of
MDM2 to negatively regulate p53 becomes irrelevant if p53 is inactivated by mutation. Moreover, we found evidence for an inverse correlation of
TP53 mutation with the presence of the variant SNP309G-allele.
TP53 was positive in 31% of SNP309TT patients, 26% of TG patients, but only 13% of GG patients (
p < 0.04;
Table 4). Since the SN309G allele leads to a higher expression of
MDM2 and consequently lower levels of p53, presence of the SNP309TG and particularly the GG genotype may, at least in part, functionally substitute the inactivation of
TP53 by mutation. Thus, if p53 tumor suppressor activity is already attenuated by overexpression of
MDM2, the selective pressure to additionally mutate
TP53 may be reduced in the presence of the SNP309G allele, and this may be reflected in the lower rates of p53 positivity observed in carriers of the SNP309GG and TG genotypes. Interestingly, amplification of
MDM2, another clinically relevant mechanism in carcinogenesis to increase
MDM2 and decrease p53 expression, was also found to be associated with a reduced rate of
TP53 mutation [
10,
13,
54]. Like in the present study, a reduced rate of
TP53 mutations in carriers of the SNP309G allele has been reported for non-small cell lung cancer [
55], hepatocellular carcinoma [
35], and bladder cancer [
56]. In contrast, no significant association was found in oral cancer, pancreatic cancer, and two studies of colorectal cancer [
57,
58,
59,
60]. A lower frequency of p53 positivity was also found in breast cancer patients with the SNP309GG genotype compared to those with the SNP309TT genotype [
50]. An analogous association of a germline polymorphism with a differential mutation rate of
TP53 was also found for the R72P coding SNP (rs1042522) in
TP53 itself [
46,
61].
We found no significant association of SNP309 genotype with survival, consistent with most reports in breast cancer and other tumor types [
50,
62,
63,
64,
65]. However, the TT genotype tended to be associated with a slightly poorer metastasis-free survival, and this association was significant in multivariable analysis (
Table 6). Since the SNP309T allele is thought to lead to lower MDM2 and hence higher p53 levels [
14], we expected it to rather be associated with an improved survival. However, the TT genotype has been found associated with a poor prognosis previously, including in a breast cancer study in BRCA1/2 mutation carriers [
66,
67,
68]. We hypothesize that this counter-intuitive association is due to the much higher rate of
TP53 mutation in SNP309TT patients. To the best of our knowledge, we here report the first analysis of the association of SNP285 with breast cancer survival. We found the SNP285GC genotype non-significantly associated with a poorer disease-free and metastasis-free survival in univariable and multivariable analyses (
Table 5 and
Table 6). However, this result is based on the analysis of only four patients with the GC genotype and 128 with the SNP285GG genotype and should be interpreted with caution. The association of SNP285 with survival has previously been analyzed in two lung cancer studies. One study found no association, whereas the other study found an association of the SNP285GC genotype with a worse progression-free survival compared to the GG genotype, but no association with the overall survival [
32,
55].
5. Conclusions
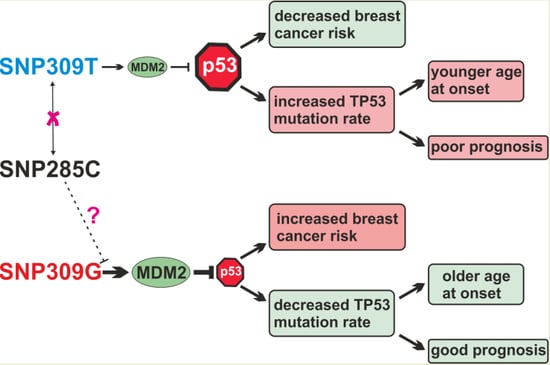
We found that the SNP309GG genotype was associated with an older age at breast cancer onset, a good prognosis in multivariable analysis of the metastasis-free survival and a trend in the same direction in related survival analyses, an increased breast cancer risk in specific subgroups, and a rate of p53 positivity less than half compared to the SNP309TT genotype. These findings are consistent with a model in which the p53-response is attenuated in the presence of the SNP309GG genotype. A direct consequence of this attenuation may be that the risk of breast cancer and other cancers is increased, but that the selective pressure to mutate
TP53 is reduced. An attenuated p53-response is presumably still functional to some extent as opposed to one that is completely inactivated by mutation, and consequently leads to a later onset of breast cancer and a trend towards a good prognosis. None of the observed associations of SNP285 were significant in our study, which may in part be due to the very low frequency of the variant C-allele. However, the observed trends towards an increased breast cancer risk and a poor survival are at odds with the proposed molecular function of the SNP285C-allele to reduce binding of SP1 and expression of
MDM2. Accordingly, we conclude that these associations of SNP285C are bystander effects of its complete linkage disequilibrium with SNP309G. The odds ratios in subgroups according to SNP309 genotype (
Table 2), and in clinical subgroups (
Table 3) support this conclusion. For example, the odds ratios associated with both the SNP309GG and the SNP285GC genotype are higher in the Ki67-high than in the Ki67-low subgroup (
Table 3). This is consistent with the molecular function of SNP309G to attenuate the anti-proliferative p53-response, but not with that of SNP285C to enhance it. These bystander effects may be more pronounced in our study population compared to previous ones due to the lower frequency of the SNP285GC genotype and the complete absence of SNP285CC subjects. Finally, the fact that SNP309 is associated with an increased cancer risk in most Asian but not European studies has been suggested to be due to the antagonistic molecular function and consequential confounding effect of SNP285, which is present in Europeans, but absent in Asians. However, the results of the present study do not support this hypothesis. Specifically, when we excluded SNP285GC individuals from our analyses, thus mimicking the situation in Asian populations, the observed associations (ORs) were still not significant, and remained essentially unaltered (
Table 1). In conclusion, we report here the first analysis of SNP285 in a Central European study population, the first analysis of the association of SNP285 with breast cancer survival, the first analysis of SNP285 in clinically relevant subgroups of breast cancer, and the first analysis of potential confounding effects of SNP285 on the association of SNP309 with the age at onset and survival.