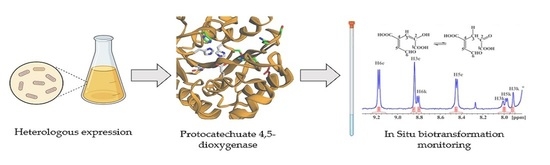
Characterization of Protocatechuate 4,5-Dioxygenase from Pseudarthrobacter phenanthrenivorans Sphe3 and In Situ Reaction Monitoring in the NMR Tube
Abstract
:1. Introduction
2. Results
2.1. PCA 4,5-Dioxygenase Activity and Transcriptional Analysis with RT-PCR and RT-qPCR
2.2. Heterologous Expression and Catalytic Properties of the Recombinant PcaA
2.3. Substrate Specificity
2.4. In Situ Monitoring of the PcaA Enzymatic Reaction Using PCA or Gallate as Substrates in the NMR Tube Bioreactor
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Growth Conditions
4.2. Preparation of Sphe3 Cell Extracts
4.3. Enzyme Assays
4.4. Substrate Specificity Assessment
4.5. Cloning and Heterologous Expression of pcaA Gene
4.6. Purification of Recombinant PCA 4,5-Dioxygenase
4.7. In situ Biotransfomation Monitoring in the NMR Tube
4.8. Transcriptional Analysis with RT-PCR and Quantitative Real-Time PCR (RT-qPCR)
4.9. 3D Model Construction of PcaA
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kamimura, N.; Masai, E. The protocatechuate 4, 5-cleavage pathway: Overview and new findings. In Biodegradative Bacteria; Nojiri, H., Tsuda, M., Fukuda, M., Kamagata, Y., Eds.; Springer: Tokyo, Japan, 2014; pp. 207–226. [Google Scholar] [CrossRef]
- Crawford, R.L. Novel pathway for degradation of protocatechuic acid in Bacillus species. J. Bacteriol. 1975, 121, 531–536. [Google Scholar] [CrossRef] [Green Version]
- Kasai, D.; Fujinami, T.; Abe, T.; Mase, K.; Katayama, Y.; Fukuda, M.; Masai, E. Uncovering the protocatechuate 2, 3-cleavage pathway genes. J. Bacteriol. 2009, 191, 6758–6768. [Google Scholar] [CrossRef] [Green Version]
- Harwood, C.S.; Parales, R.E. The β-ketoadipate pathway and the biology of self-identity. Ann. Rev. Microbiol. 1996, 50, 553–590. [Google Scholar] [CrossRef]
- Kamimura, N.; Aoyama, T.; Yoshida, R.; Takahashi, K.; Kasai, D.; Abe, T.; Mase, K.; Katayama, Y.; Fukuda, M.; Masai, E. Characterization of the protocatechuate 4, 5-cleavage pathway operon in Comamonas sp. strain E6 and discovery of a novel pathway gene. Appl. Environ. Microbiol. 2010, 76, 8093–8101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bugg, T.D.; Ahmad, M.; Hardiman, E.M.; Rahmanpour, R. Pathways for degradation of lignin in bacteria and fungi. Nat. Prod. Rep. 2011, 28, 1883–1896. [Google Scholar] [CrossRef]
- Sim, H.W.; Jung, M.; Cho, Y.K. Purification and characterization of protocatechuate 3, 4-dioxygenase from Pseudomonas pseudoalcaligenes KF707. J. Korean Soc. Appl. Biol. Chem. 2013, 56, 401–408. [Google Scholar] [CrossRef]
- Ni, B.; Zhang, Y.; Chen, D.W.; Wang, B.J.; Liu, S.J. Assimilation of aromatic compounds by Comamonas testosteroni: Characterization and spreadability of protocatechuate 4, 5-cleavage pathway in bacteria. Appl. Microbiol. Biotechnol. 2013, 97, 6031–6041. [Google Scholar] [CrossRef] [PubMed]
- Bugg, T.D.; Ahmad, M.; Hardiman, E.M.; Singh, R. The emerging role for bacteria in lignin degradationand bioproduct formation. Curr. Opin. Biotechnol. 2011, 22, 394–400. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pantoja, D.; Donoso, R.; Junca, H.; González, B.; Pieper, D.H. Phylogenomics of aerobic bacterial degradation of aromatics. In Aerobic Utilization of Hydrocarbons, Oils and Lipids; Rojo, F., Ed.; Springer: Cham, Switzerland, 2017; pp. 1–48. [Google Scholar] [CrossRef]
- Chen, Y.P.; Lovell, C.R. Purification and properties of a homodimeric protocatechuate 4, 5-dioxygenase from Rhizobium leguminosarum. Arch. Microbiol. 1994, 161, 191–195. [Google Scholar] [CrossRef]
- Barry, K.P.; Taylor, E.A. Characterizing the promiscuity of LigAB, a lignin catabolite degrading extradiol dioxygenase from Sphingomonas paucimobilis SYK-6. Biochemistry 2013, 52, 6724–6736. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vaillancourt, F.H.; Bolin, J.T.; Eltis, L.D. The ins and outs of ring-cleaving dioxygenases. Crit. Rev. Biochem. Mol. Biol. 2006, 41, 241–267. [Google Scholar] [CrossRef] [PubMed]
- Burroughs, A.M.; Glasner, M.E.; Barry, K.P.; Taylor, E.A.; Aravind, L. Oxidative opening of the aromatic ring: Tracing the natural history of a large superfamily of dioxygenase domains and their relatives. J. Biol. Chem. 2019, 294, 10211–10235. [Google Scholar] [CrossRef] [PubMed]
- Mampel, J.; Providenti, M.A.; Cook, A.M. Protocatechuate 4, 5-dioxygenase from Comamonas testosteroni T-2: Biochemical and molecular properties of a new subgroup within class III of extradiol dioxygenases. Arch. Microbiol. 2005, 183, 130–139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Providenti, M.A.; Mampel, J.; MacSween, S.; Cook, A.M.; Wyndham, R.C. Comamonas testosteroni BR6020 possesses a single genetic locus for extradiol cleavage of protocatechuate. Microbiology 2001, 147, 2157–2167. [Google Scholar] [CrossRef] [Green Version]
- Maruyama, K.; Shibayama, T.; Ichikawa, A.; Sakou, Y.; Yamada, S.; Sugisaki, H. Cloning and characterization of the genes encoding enzymes for the protocatechuate meta-degradation pathway of Pseudomonas ochraceae NGJ1. Biosc. Biotechnol. Biochem. 2004, 68, 1434–1441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Masai, E.; Shinohara, S.; Hara, H.; Nishikawa, S.; Katayama, Y.; Fukuda, M. Genetic and biochemical characterization of a 2-pyrone-4, 6-dicarboxylic acid hydrolase involved in the protocatechuate 4, 5-cleavage pathway of Sphingomonas paucimobilis SYK-6. J. Bacteriol. 1999, 181, 55–62. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wattiau, P.; Bastiaens, L.; van Herwijnen, R.; Daal, L.; Parsons, J.R.; Renard, M.E.; Springael, D.; Cornelis, G.R. Fluorene degradation by Sphingomonas sp. LB126 proceeds through protocatechuic acid: A genetic analysis. Res. Microbiol. 2001, 152, 861–872. [Google Scholar] [CrossRef]
- Noda, Y.; Nishikawa, S.; Shiozuka, K.; Kadokura, H.; Nakajima, H.; Yoda, K.; Katayama, Y.; Morohoshi, N.; Haraguchi, T.; Yamasaki, M. Molecular cloning of the protocatechuate 4, 5-dioxygenase genes of Pseudomonas paucimobilis. J. Bacteriol. 1990, 172, 2704–2709. [Google Scholar] [CrossRef] [Green Version]
- Patil, N.K.; Kundapur, R.; Shouche, Y.S.; Karegoudar, T. Degradation of a plasticizer, di-n-butylphthalate by Delftia sp. TBKNP-05. Curr. Microbiol. 2006, 52, 225–230. [Google Scholar] [CrossRef]
- Busse, H.J. Review of the taxonomy of the genus Arthrobacter, emendation of the genus Arthrobacter sensu lato, proposal to reclassify selected species of the genus Arthrobacter in the novel genera Glutamicibacter gen. nov., Paeniglutamicibacter gen. nov., Pseudoglutamicibacter gen. nov., Paenarthrobacter gen. nov. and Pseudarthrobacter gen. nov., and emended description of Arthrobacter roseus. Int. J. Syst. Evol. Microbiol. 2016, 66, 9–37. [Google Scholar] [CrossRef] [Green Version]
- Kallimanis, A.; Kavakiotis, K.; Perisynakis, A.; Spröer, C.; Pukall, R.; Drainas, C.; Koukkou, A. Arthrobacter phenanthrenivorans sp. nov., to accommodate the phenanthrene-degrading bacterium Arthrobacter sp. strain Sphe3. Inter. J. Syst. Evol. Microbiol. 2009, 59, 275–279. [Google Scholar] [CrossRef]
- Kallimanis, A.; Frillingos, S.; Drainas, C.; Koukkou, A.I. Taxonomic identification, phenanthrene uptake activity, and membrane lipid alterations of the PAH degrading Arthrobacter sp. strain Sphe3. App. Microbiol. Biotechnol. 2007, 76, 709–717. [Google Scholar] [CrossRef] [PubMed]
- Kallimanis, A.; LaButti, K.M.; Lapidus, A.; Clum, A.; Lykidis, A.; Mavromatis, K.; Pagani, I.; Liolios, K.; Ivanova, N.; Goodwin, L. Complete genome sequence of Arthrobacter phenanthrenivorans type strain (Sphe3). Stand. Genomic. Sci. 2011, 4, 123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chatterjee, S.; Dutta, T.K. Complete degradation of butyl benzyl phthalate by a defined bacterial consortium: Role of individual isolates in the assimilation pathway. Chemosphere 2008, 70, 933–941. [Google Scholar] [CrossRef]
- Niewerth, H.; Schuldes, J.; Parschat, K.; Kiefer, P.; Vorholt, J.A.; Daniel, R.; Fetzner, S. Complete genome sequence and metabolic potential of the quinaldine-degrading bacterium Arthrobacter sp. Rue61a. BMC Genom. 2012, 13, 534. [Google Scholar] [CrossRef] [Green Version]
- Pau, M.Y.; Davis, M.I.; Orville, A.M.; Lipscomb, J.D.; Solomon, E.I. Spectroscopic and electronic structure study of the enzyme−substrate complex of intradiol dioxygenases: Substrate activation by a high-spin ferric non-heme iron site. J. Am. Chem. Soc. 2007, 129, 1944–1958. [Google Scholar] [CrossRef] [Green Version]
- Vandera, E.; Samiotaki, M.; Parapouli, M.; Panayotou, G.; Koukkou, A.I. Comparative proteomic analysis of Arthrobacter phenanthrenivorans Sphe3 on phenanthrene, phthalate and glucose. J. Proteom. 2015, 113, 73–89. [Google Scholar] [CrossRef] [PubMed]
- Eaton, R.W. Plasmid-encoded phthalate catabolic pathway in Arthrobacter keyseri 12B. J. Bacteriol. 2001, 183, 3689–3703. [Google Scholar] [CrossRef] [Green Version]
- Plotnikova, E.; Solyanikova, I.; Egorova, D.; Shumkova, E.; Golovleva, L. Degradation of 4-chlorobiphenyl and 4-chlorobenzoic acid by the strain Rhodococcus ruber P25. Microbiology 2012, 81, 143–153. [Google Scholar] [CrossRef]
- Ono, K.; Nozaki, M.; Hayaishi, O. Purification and some properties of protocatechuate 4, 5-dioxygenase. Biochim. Biophys. Acta 1970, 220, 224–238. [Google Scholar] [CrossRef]
- Chatzikonstantinou, A.V.; Tsiailanis, A.D.; Gerothanassis, I.P.; Stamatis, H.; Ravera, E.; Fragai, M.; Luchinat, C.; Parigi, G.; Tzakos, A.G. The NMR tube bioreactor. Methods Enzymol. 2020, 633, 71–101. [Google Scholar] [CrossRef]
- Chatzikonstantinou, A.V.; Chatziathanasiadou, M.V.; Ravera, E.; Fragai, M.; Parigi, G.; Gerothanassis, I.P.; Luchinat, C.; Stamatis, H.; Tzakos, A.G. Enriching the biological space of natural products and charting drug metabolites, through real time biotransformation monitoring: The NMR tube bioreactor. Biochim. Biophys. Acta Gen. Subj. 2018, 1862, 1–8. [Google Scholar] [CrossRef]
- Primikyri, A.; Sayyad, N.; Quilici, G.; Vrettos, E.I.; Lim, K.; Chi, S.W.; Musco, G.; Gerothanassis, I.P.; Tzakos, A.G. Probing the interaction of a quercetin bioconjugate with Bcl-2 in living human cancer cells with in-cell NMR spectroscopy. FEBS Lett. 2018, 592, 3367–3379. [Google Scholar] [CrossRef] [Green Version]
- Primikyri, A.; Mazzone, G.; Lekka, C.; Tzakos, A.G.; Russo, N.; Gerothanassis, I.P. Understanding zinc(II) chelation with quercetin and luteolin: A combined NMR and theoretical study. J. Phys. Chem. B 2015, 119, 83–95. [Google Scholar] [CrossRef]
- Troganis, A.N.; Sicilia, E.; Barbarossou, K.; Gerothanassis, I.P.; Russo, N. Solvation properties of N-substituted cis and trans amides are not identical: Significant enthalpy and entropy changes are revealed by the use of variable temperature 1H NMR in aqueous and chloroform solutions and ab initio calculations. J. Phys. Chem. A 2005, 109, 11878–11884. [Google Scholar] [CrossRef]
- Kamimura, N.; Takamura, K.; Hara, H.; Kasai, D.; Natsume, R.; Senda, T.; Katayama, Y.; Fukuda, M.; Masai, E. Regulatory system of the protocatechuate 4, 5-cleavage pathway genes essential for lignin downstream catabolism. J. Bacteriol. 2010, 192, 3394–3405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sugimoto, K.; Senda, T.; Aoshima, H.; Masai, E.; Fukuda, M.; Mitsui, Y. Crystal structure of an aromatic ring opening dioxygenase LigAB, a protocatechuate 4, 5-dioxygenase, under aerobic conditions. Structure 1999, 7, 953–965. [Google Scholar] [CrossRef] [Green Version]
- Webb, B.; Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinform. 2016, 54, 5–6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sugimoto, K.; Senda, M.; Kasai, D.; Fukuda, M.; Masai, E.; Senda, T. Molecular mechanism of strict substrate specificity of an extradiol dioxygenase, DesB, derived from Sphingobium sp. SYK-6. PLoS ONE 2014, 9, e92249. [Google Scholar] [CrossRef]
- Eltis, L.D.; Bolin, J.T. Evolutionary relationships among extradiol dioxygenases. J. Bacteriol. 1996, 178, 5930–5937. [Google Scholar] [CrossRef] [Green Version]
- Peng, X.; Egashira, T.; Hanashiro, K.; Masai, E.; Nishikawa, S.; Katayama, Y.; Kimbara, K.; Fukuda, M. Cloning of a Sphingomonas paucimobilis SYK-6 gene encoding a novel oxygenase that cleaves lignin-related biphenyl and characterization of the enzyme. Appl. Environ. Microbiol. 1998, 64, 2520–2527. [Google Scholar] [CrossRef] [Green Version]
- Pérez-Pantoja, D.; Donoso, R.; Agulló, L.; Córdova, M.; Seeger, M.; Pieper, D.H.; González, B. Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ. Microbiol. 2012, 14, 1091–1117. [Google Scholar] [CrossRef] [PubMed]
- Duetz, W.A.; Marqués, S.; Wind, B.; Ramos, J.L.; van Andel, J. Catabolite repression of the toluene degradation pathway in Pseudomonas putida harboring pWW0 under various conditions of nutrient limitation in chemostat culture. Appl. Environ. Microbiol. 1996, 62, 601–606. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blencke, H.M.; Homuth, G.; Ludwig, H.; Mäder, U.; Hecker, M.; Stülke, J. Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: Regulation of the central metabolic pathways. Metab. Eng. 2003, 5, 133–149. [Google Scholar] [CrossRef]
- Tropel, D.; van der Meer, J.R. Bacterial transcriptional regulators for degradation pathways of aromatic compounds. Microbiol. Mol. Biol. Rev. 2004, 68, 474–500. [Google Scholar] [CrossRef] [Green Version]
- Jiménez, J.I.; Miñambres, B.; García, J.L.; Díaz, E. Genomic analysis of the aromatic catabolic pathways from Pseudomonas putida KT2440. Environ. Microbiol. 2002, 4, 824–841. [Google Scholar] [CrossRef]
- Romero-Silva, M.J.; Mendez, V.; Agullo, L.; Seeger, M. Genomic and functional analyses of the gentisate and protocatechuate ring-cleavage pathways and related 3-hydroxybenzoate and 4-hydroxybenzoate peripheral pathways in Burkholderia xenovorans LB400. PLoS ONE 2013, 8, e56038. [Google Scholar] [CrossRef] [Green Version]
- Dagley, S.; Geary, P.; Wood, J. The metabolism of protocatechuate by Pseudomonas testosteroni. Biochem. J. 1968, 109, 559. [Google Scholar] [CrossRef] [Green Version]
- Nogales, J.; Canales, Á.; Jiménez-Barbero, J.; García, J.L.; Díaz, E. Molecular characterization of the gallate dioxygenase from Pseudomonas putida KT2440 the prototype of a new subgroup of extradiol dioxygenases. J. Biol. Chem. 2005, 280, 35382–35390. [Google Scholar] [CrossRef] [Green Version]
- Whiting, A.K.; Boldt, Y.R.; Hendrich, M.P.; Wackett, L.P.; Que, L. Manganese (II)-dependent extradiol-cleaving catechol dioxygenase from Arthrobacter globiformis CM-2. Biochemistry 1996, 35, 160–170. [Google Scholar] [CrossRef]
- Fielding, A.J.; Kovaleva, E.G.; Farquhar, E.R.; Lipscomb, J.D.; Que, L. A hyperactive cobalt-substituted extradiol-cleaving catechol dioxygenase. J. Biol. Inor. Chem. 2011, 16, 341–355. [Google Scholar] [CrossRef] [Green Version]
- Yun, S.H.; Yun, C.Y.; Kim, S.I. Characterization of protocatechuate 4, 5-dioxygenase induced from p-hydroxybenzoate-cultured Pseudomonas sp. K82. J. Microbiol. 2004, 42, 152–155. [Google Scholar]
- Zabinski, R.; Münck, E.; Champion, P.; Wood, J. Kinetic and Mössbauer studies on the mechanism of protocatechuic acid 4, 5-oxygenase. Biochemistry 1972, 11, 3212–3219. [Google Scholar] [CrossRef] [PubMed]
- Ribbons, D.; Evans, W. Oxidative metabolism of protocatechuic acid by certain soil pseudomonads: A new ring-fission mechanism. Biochem. J. 1962, 83, 482. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Masai, E.; Momose, K.; Hara, H.; Nishikawa, S.; Katayama, Y.; Fukuda, M. Genetic and Biochemical characterization of 4-Carboxy-2-Hydroxymuconate-6-Semialdehyde dehydrogenase and its role in the protocatechuate 4, 5-cleavage pathway in Sphingomonas paucimobilis SYK-6. J. Bacteriol. 2000, 182, 6651–6658. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chakraborty, I.; Bodurtha, K.J.; Heeder, N.J.; Godfrin, M.P.; Tripathi, A.; Hurt, R.H.; Shukla, A.; Bose, A. Massive electrical conductivity enhancement of multilayer graphene/polystyrene composites using a nonconductive filler. ACS Appl. Mater. Interfaces 2014, 6, 16472–16475. [Google Scholar] [CrossRef]
- Nogales, J.; Canales, Á.; Jiménez-Barbero, J.; Serra, B.; Pingarrón, J.M.; García, J.L.; Díaz, E. Unravelling the gallic acid degradation pathway in bacteria: The gal cluster from Pseudomonas putida. Mol. Microbiol. 2011, 79, 359–374. [Google Scholar] [CrossRef]
- Brecker, L.; Ribbons, D.W. Biotransformations monitored in situ by proton nuclear magnetic resonance spectroscopy. Trends Biotechnol. 2000, 18, 197–202. [Google Scholar] [CrossRef]
- Yang, T.; Bar-Peled, M. Identification of a novel UDP-sugar pyrophosphorylase with a broad substrate specificity in Trypanosoma cruzi. Biochem. J. 2010, 429, 533–543. [Google Scholar] [CrossRef] [Green Version]
- Kopp, D.; Willows, R.D.; Sunna, A. Cell-free enzymatic conversion of spent coffee grounds into the platform chemical lactic acid. Front. Bioeng Biotechnol. 2019, 7, 389. [Google Scholar] [CrossRef] [Green Version]
- Dalvit, C. Ligand-and substrate-based 19F NMR screening: Principles and applications to drug discovery. Progr. NMR Spectrosc. 2007, 51, 243–271. [Google Scholar] [CrossRef]
- Werner, R.M.; Johnson, A. 31 P NMR of the pyruvate kinase reaction: An undergraduate experiment in enzyme kinetics. Biochem. Mol. Biol. Educ. 2017, 45, 509–514. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Escobedo-Hinojosa, W.; Wissner, J.L.; Hauer, B. A real-time 31P-NMR-based approach for the assessment of glycerol kinase catalyzed monophosphorylations. Methods X 2021, 8, 101285. [Google Scholar] [CrossRef]
- Limtiaco, J.F.; Beni, S.; Jones, C.J.; Langeslay, D.J.; Larive, C.K. NMR methods to monitor the enzymatic depolymerization of heparin. Anal. Bioanal. Chem. 2011, 399, 593–603. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Korchak, S.; Jagtap, A.P.; Glöggler, S. Signal-enhanced real-time magnetic resonance of enzymatic reactions at millitesla fields. Chem. Sci. 2021, 12, 314–319. [Google Scholar] [CrossRef]
- Vandera, E.; Kavakiotis, K.; Kallimanis, A.; Kyrpides, N.C.; Drainas, C.; Koukkou, A.I. Heterologous expression and characterization of two 1-hydroxy-2-naphthoic acid dioxygenases from Arthrobacter phenanthrenivorans. Appl. Environ. Microbiol. 2012, 78, 621–627. [Google Scholar] [CrossRef] [Green Version]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Deschenes, L.A.; David, A. Origin 6.0: Scientific Data Analysis and Graphing Software Origin Lab Corporation (Formerly Microcal Software, Inv.); BoutUniversity of Texas, Austin; Commercial price: 595; Academic price: 446; 2000. pp. 9567–9568. Available online: www.originlab.com (accessed on 18 March 2017).
- William, S.; Feil, H.; Copeland, A. Bacterial Genomic DNA Isolation using CTAB. DOE Joint Genome Institute, Walnut Creek, CA 2012. Available online: https://jgi.doe.gov/wp-content/uploads/2014/02/JGI-Bcterial-DNA-isolation-CTAB-Protocol-2012.pdf (accessed on 18 March 2017).
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 1983, 166, 557–580. [Google Scholar] [CrossRef]
- Chung, C.; Miller, R.H. A rapid and convenient method for the preparation and storage of competent bacterial cells. Nucleic Acids Res. 1988, 16, 3580. [Google Scholar] [CrossRef] [Green Version]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [Green Version]
- Sambrook, J.; Fritsch, E.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Corbella, M.; Puyet, A. Real-time reverse transcription-PCR analysis of expression of halobenzoate and salicylate catabolism-associated operons in two strains of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2003, 69, 2269–2275. [Google Scholar] [CrossRef] [Green Version]
- Applied Biosystems. Relative Quantifcation of Gene Expression in User Bulletin 2 ABI PRISM 7700 Sequence Detection System; Applied Biosystems: Waltham, MA, USA, 1999; pp. 1–36. [Google Scholar]
- Reynolds, C.R.; Islam, S.A.; Sternberg, M.J. EzMol: A web server wizard for the rapid visualization and image production of protein and nucleic acid structures. J. Mol. Biol. 2018, 430, 2244–2248. [Google Scholar] [CrossRef] [PubMed]
- Sehnal, D.; Bittrich, S.; Deshpande, M.; Svobodová, R.; Berka, K.; Bazgier, V.; Velankar, S.; Burley, S.K.; Koča, J.; Rose, A.S. Mol* Viewer: Modern web app for 3D visualization and analysis of large biomolecular structures. Nucleic Acids Res. 2021. [Google Scholar] [CrossRef]







Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsagogiannis, E.; Vandera, E.; Primikyri, A.; Asimakoula, S.; Tzakos, A.G.; Gerothanassis, I.P.; Koukkou, A.-I. Characterization of Protocatechuate 4,5-Dioxygenase from Pseudarthrobacter phenanthrenivorans Sphe3 and In Situ Reaction Monitoring in the NMR Tube. Int. J. Mol. Sci. 2021, 22, 9647. https://doi.org/10.3390/ijms22179647
Tsagogiannis E, Vandera E, Primikyri A, Asimakoula S, Tzakos AG, Gerothanassis IP, Koukkou A-I. Characterization of Protocatechuate 4,5-Dioxygenase from Pseudarthrobacter phenanthrenivorans Sphe3 and In Situ Reaction Monitoring in the NMR Tube. International Journal of Molecular Sciences. 2021; 22(17):9647. https://doi.org/10.3390/ijms22179647
Chicago/Turabian StyleTsagogiannis, Epameinondas, Elpiniki Vandera, Alexandra Primikyri, Stamatia Asimakoula, Andreas G. Tzakos, Ioannis P. Gerothanassis, and Anna-Irini Koukkou. 2021. "Characterization of Protocatechuate 4,5-Dioxygenase from Pseudarthrobacter phenanthrenivorans Sphe3 and In Situ Reaction Monitoring in the NMR Tube" International Journal of Molecular Sciences 22, no. 17: 9647. https://doi.org/10.3390/ijms22179647