Abstract
Auxin response factors (ARFs) are important plant transcription factors that are differentially expressed in response to auxin and various abiotic stresses. ARFs play important roles in mediating plant growth and stress responses; however, these factors have not been studied in eggplants. In this study, genome–wide identification and the functional analysis of the ARF gene family in eggplants (Solanum melongena L.) were performed. A total of 20 ARF (SmARF) genes were identified and phylogenetically classified into three groups. Our analysis revealed four functional domains and 10 motifs in these proteins. Subcellular localization showed that the SmARFs localized in the nucleus. To investigate the biological functions of the SmARFs under 2,4–D and salt stress treatments, quantitative real–time RT–PCR (qRT–PCR) was conducted. Most SmARF genes exhibited changes in expression in response to 2,4–D treatments in the flowers, especially SmARF4 and 7B. All SmARF genes quickly responded to salt stress, except SmARF17 and 19 in leaves, SmARF1A and 7B in roots, and SmARF2A, SmARF7B, and SmARF16B in stems. These results helped to elucidate the role of ARFs in auxin signaling under 2,4–D and salt stress in eggplants.
1. Introduction
Eggplant (Solanum melongena L.) is an excellent source of fiber, vitamins, minerals, and antioxidant polyphenols, making it a popular food and an economically important vegetable crop species that is cultivated worldwide [1,2]. This plant species originates in Africa and is widely cultivated in Asia, Africa, Europe, and the Near East [3]. China is the largest producer of eggplant, accounting for 68% of global production in 2020 [4]. Flower abscission is a serious constraint to the yield and quality of the plant [5,6]. During planting, 2,4–D (2,4–dichlorophenoxyacetic acid), a synthetic auxin plant growth regulator, can be sprayed on flower buds to mitigate flower abscission and increase fruit set [7].
Auxin is a plant hormone that is strongly involved in several biological processes, including organ formation and development [8,9,10,11,12]. Two transcription factor families are associated with the auxin signal transduction pathway: the auxin response factor (ARF) and auxin/indole acetic acid (Aux/IAA) [13]. ARFs have a modular domain structure with three major domains. The N–terminal B3–like DNA–binding domain (DBD) is highly conserved and binds to auxin response elements (AuxREs) in the promoters of auxin–responsive genes [13,14]. The middle region activates or inhibits target genes, depending on their sequence [13,14]. The C–terminal dimerization domain (CTD) contains the motifs III and IV, which are also found in the Aux/IAA family.
The functions of ARFs have been studied in Arabidopsis thaliana, tomato (Solanum lycopersicum), papaya (Carica papaya L.), sugar beet, peach, and other plant species [13,15,16,17,18,19,20,21,22,23,24,25]. AtARF 2, 3, 5, 6, 7, 8, 17, and 19 regulate apical bud formation, pollen wall synthesis, vascular bundle development, hypocotyl tropic movement, and adventitious root formation [26,27,28,29,30]. For example, the phenotypes of arf2 T–DNA insertion mutations indicated that ARF2 regulates leaf senescence [31] and floral organ abscission [32]. SlARF6A, 8A, 8B, and 24 regulate leaf shape development [33]. SlARF 3, 4, 5 act in tomato development and can stimulate the formation of epidermal cells and trichomes, improve plant resistance to water deficit, and regulate fruit set and development [34,35,36]. However, the biological functions of the SmARF gene family members in response to auxin in eggplants are unknown.
Soil salinization is one of the most crucial abiotic stresses that affects the growth of plant leaves, roots, and stems [37]. Various ARF genes act in response to salinity stress in some plants, such as sugar beet (Beta vulgaris L.), tomato (Solanum lycopersicum), rice (Oryza sativa), sweet potato, and Tamarix chinensis [19,38,39,40,41,42,43]. It has been reported that many SlARFs were differentially expressed in tomato leaves and roots under salt stress conditions, and all SlARF genes were down–regulated in leaves except SlARF1, SlARF4, and SlARF19, whose expression was significantly induced [38]. SlARF8A and SlARF10A were significantly up–regulated by salt and drought stresses [38], while the down–regulation and loss function of SlARF4 increased tomato tolerance to salinity and drought stress [39]. In rice, both OsARF11 and OsARF15 showed differential expression under salt stress conditions [40]. Most BvARF genes in sugar beet were up–regulated or down–regulated to varying degrees by salt stress, and with significant changes in nine genes [19]. In Tamarix chinensis, the expression profiling of TcARFs showed that only TcARF6 responds to salt stress [42].These reports suggest that ARFs respond to salt stress and participate in the crosstalk between auxin and salt stress signaling.
The newest eggplant genome [44] obtained from the scaffold of v3.0 using 3D chromosome conformation capture (Hi–C) information was used to identify the members of SmARF family using known SlARF protein sequences from tomato. We found 20 potential ARF (SmARF) genes and investigated the phylogenetics, gene structure, conserved motifs, and putative cis–acting regulatory elements (CAREs). We observed the subcellular location of 10 SmARFs, and gene expression patterns in various tissues and in response to hormones and salt stress were also determined. This is the first identification and characterization of SmARFs in eggplant, and the results should facilitate future work to explore the involvement of SmARFs in auxin–mediated responses, as well as in salt stress–tolerant eggplant varieties.
2. Results
2.1. Identification and Sequence Analysis of ARF Genes in Eggplant
Full–length putative ARF protein sequences were obtained by querying tomato ARFs [17] against the eggplant genome reported by Barchi et al. (2021) [44] using BLASTP. A total of 20 candidate genes were identified, and the presence of SmARF domains was evaluated via the comparison of these sequences to other ARFs in the Pfam databases. To achieve the consistent nomenclature of ARF genes across species, eggplant members of this gene family were named based on their phylogenetic relationship, and by the numbering of the closest tomato homologs. All SmARF proteins were found to contain an ARF domain (Table 1), as revealed through the Pfam analysis tool http://pfam.sanger.ac.uk/ (accessed on 15 December 2021). Of the identified genes, SmARF19 and SmARF16A proteins contain the longest (1423 aa) and shortest (379 aa) amino acid sequences, with a mean of 824.85 aa. The molecular weight of the SmARF proteins ranged from 43.09 (SmARF6A) to 158.50 (SmARF19) kDa, with a mean of 90.46 kDa. The pI varied from 5.44 (SmARF8A) to 8.06 (SmARF6A), with a mean of 6.41. The localization of all SmARF proteins were predicted in the nucleus (Table 1).

Table 1.
Characteristics of putative genes encoding auxin response factors in Solanum melongena.
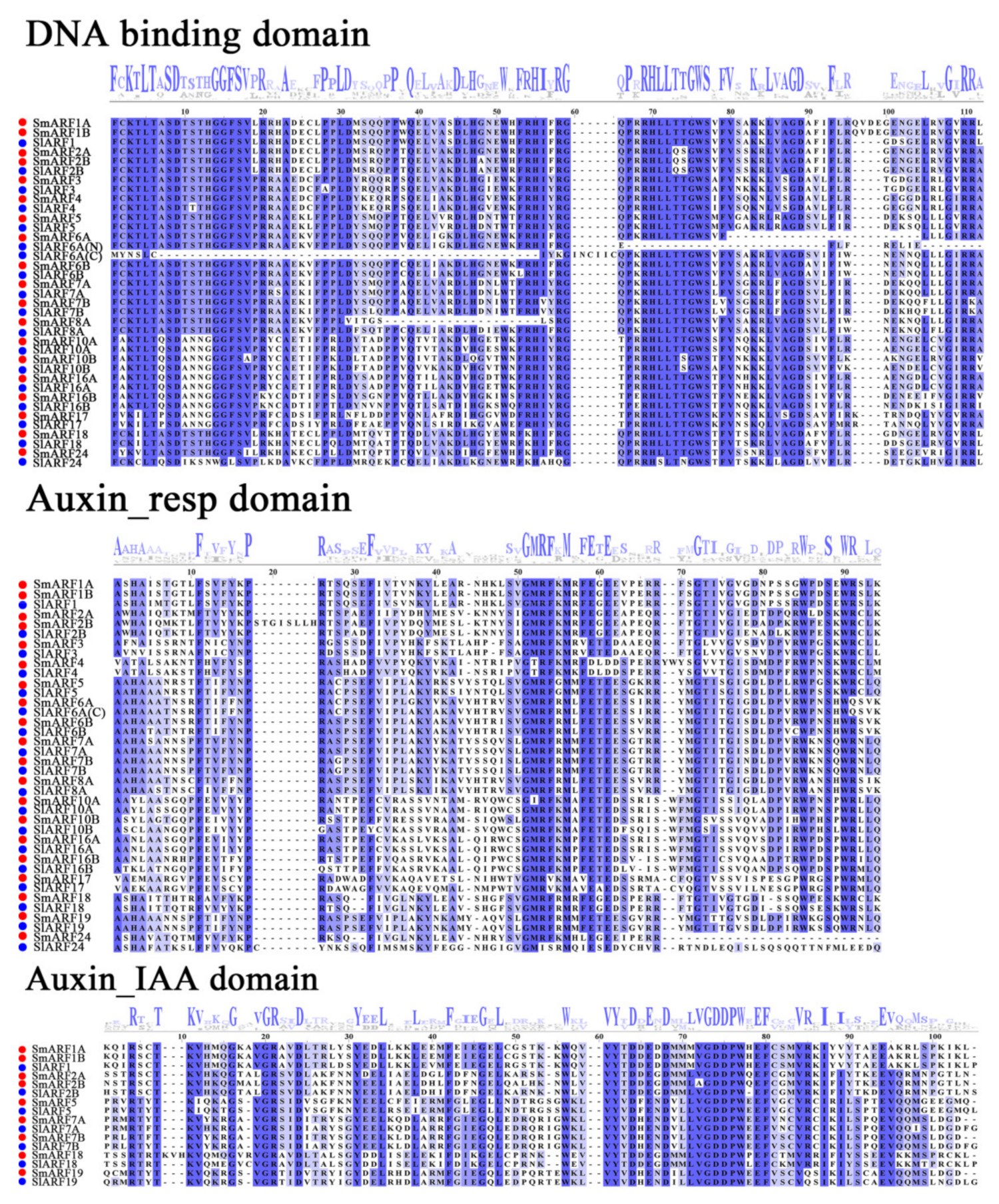
An alignment of eggplant ARFs was made against tomato homolog ARFs protein sequences, with the three highly conserved domains (B3, auxin_resp, and Aux/IAA–binding domain) aligned in multiple (Figure 1). Of the 20 sequences, all but SmARF19 included a typical DBD domain (B3), with a high conservation of the N–terminal DBDs in these proteins. All of the putative SmARFs contained an auxin_resp domain. However, only nine SmARF proteins (SmARF1A, 1B, 2A, 2B, 5, 7A, 7B, 18, and 19) contained an Aux/IAA–binding domain (Figure 1).

Figure 1.
Multiple alignment profiles of the B3 (DNA binding domain), auxin_resp, and CTD (Aux/IAA–binding) domains of the SmARF proteins obtained with the Jalview program. Red and blue codes were used to distinguish the eggplant and tomato sequences. All sequences show high levels of amino acid conservation in blue. The logo at the top shows amino acids, and large letters represent greater conservation.
2.2. Phylogenetic Tree Analysis and Grouping of Auxin Response Factor (ARF) Genes in Eggplant
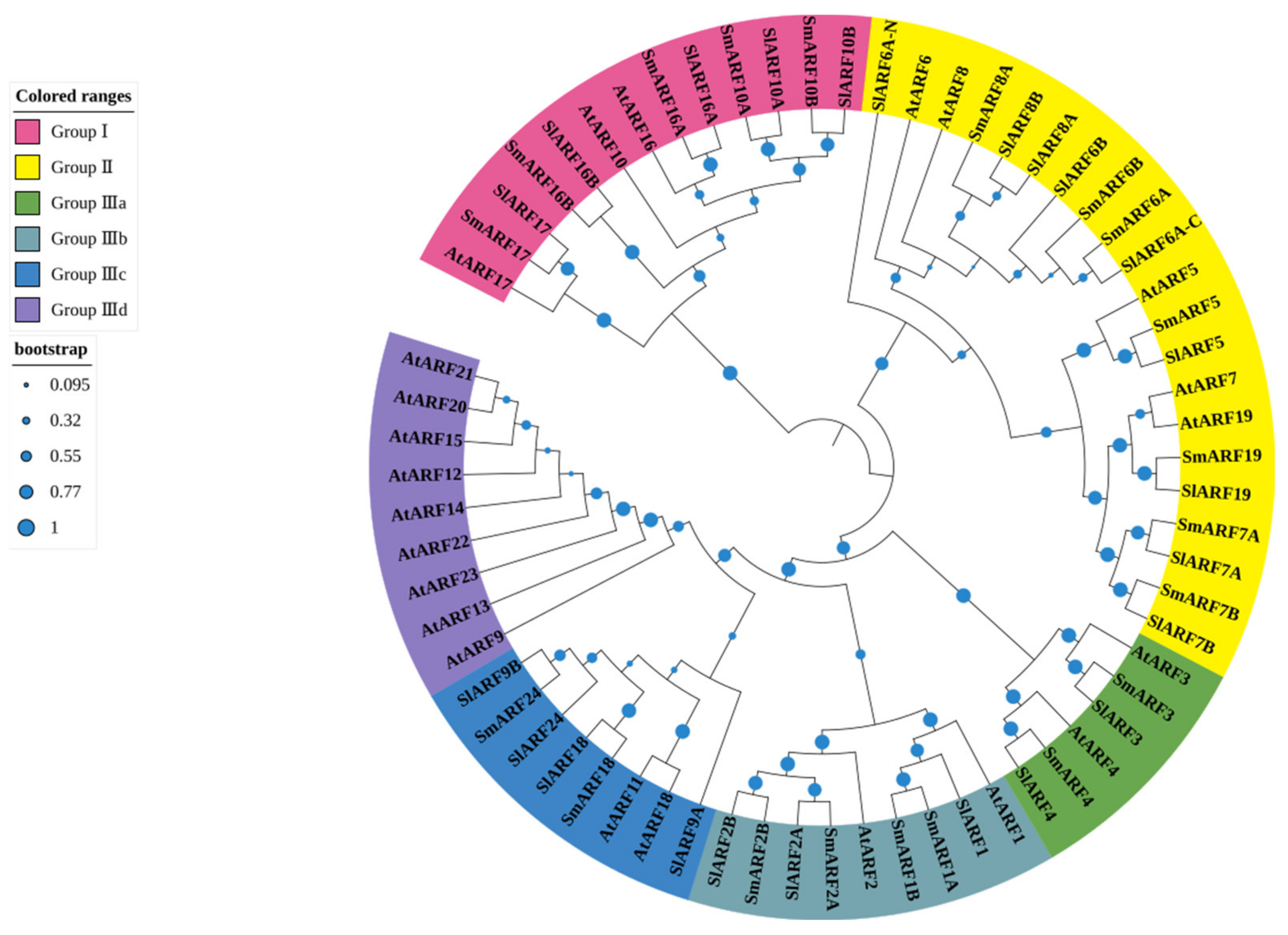
To analyze the evolutionary relationship between candidate SmARF genes, a phylogenetic tree was constructed using ARF protein sequences from eggplant (20), Arabidopsis (23) [13], and tomato (22) [17] (Figure 2). In total, the 67 sequences included in the phylogenetic tree were grouped into three major clades (I, II, and III; Figure 2). Group I included five SmARF proteins, five SlARF proteins, and three AtARF proteins. Group II had seven SmARFs, nine SlARFs, and five AtARFs. Group III was divided into four subgroups, designated Group IIIa, b, c, and d. Group IIIa included two SmARFs, two SlARFs, and two AtARFs. Group IIIb contained four SmARFs, three SlARFs, and two AtARFs. Group IIIc had two SmARFs, four SlARFs, and two AtARFs. Subclass IIId contained no SmARF or SlARF sequences, but included nine AtARFs (Figure 2). A total of 20 sister pairs were formed in the combined phylogenetic tree, with sixteen SmARF–SlARF pairs, one SmARF–SmARF pair, one SlARF–SlARF pair, and two AtARF–AtARF pairs. Groups I, II, IIIa, IIIb, IIIc, and IIId all included at least one member from the SmARFs, SlARFs, and AtARFs, but group IIId contained only AtARFs (Figure 2).
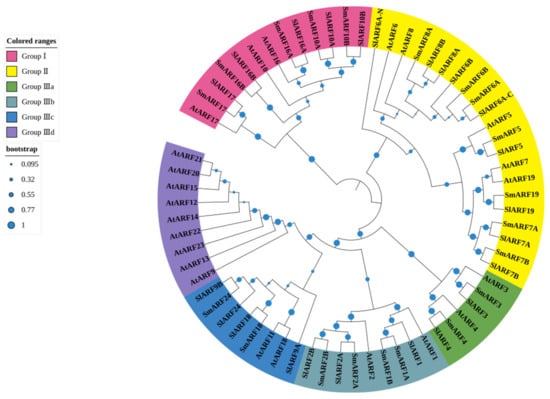
Figure 2.
Phylogenetic analysis of genes encoding auxin response factors (ARFs) in Arabidopsis, eggplant, and tomato. A phylogenetic tree was constructed using MEGA 7 software through the neighbor–joining method, with 1000 bootstrap replicates using ARF protein sequences from eggplant (20), Arabidopsis (23), and tomato (22).
2.3. Motif Analysis
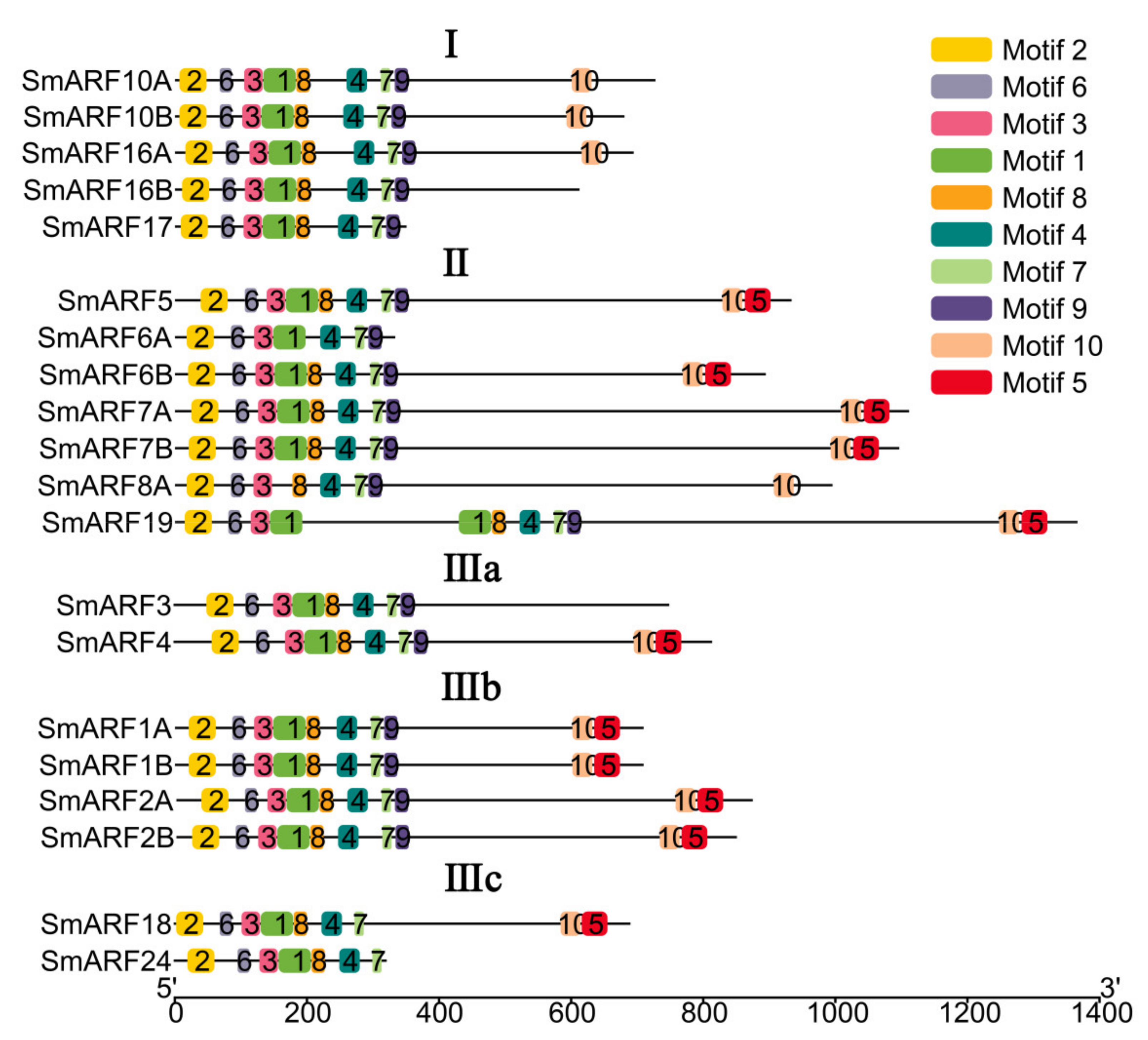
The structural features of the SmARF proteins were analyzed using MEME, and 10 conserved motifs were identified (Figure 3). Motifs 2, 3, 4, and 6 were detected in most ARF proteins (Figure 3). All of the motifs common or specific to particular groups are represented in the phylogenetic tree (Figure 3). Motif 5 was not found in any genes in group I. In group II, all of the genes contained motifs 1–10, except SmARF6A, which lacked motif 5, 8, and 10, and SmARF8A, which lacked motif 1 and motif 5. Group IIIa, b, and c also contained all motifs, in addition to SmARF3, which lacked motifs 5 and 10, SmARF18 which lacked motif 9, and SmARF24, which lacked motifs 5, 9, and 10.

Figure 3.
Schematic representation of conserved motifs in auxin response factor proteins in eggplant (Solanum melongena L.). The horizontal scale indicates protein length (number of amino acids).
2.4. Prediction of Cis–Acting Elements in Promoters
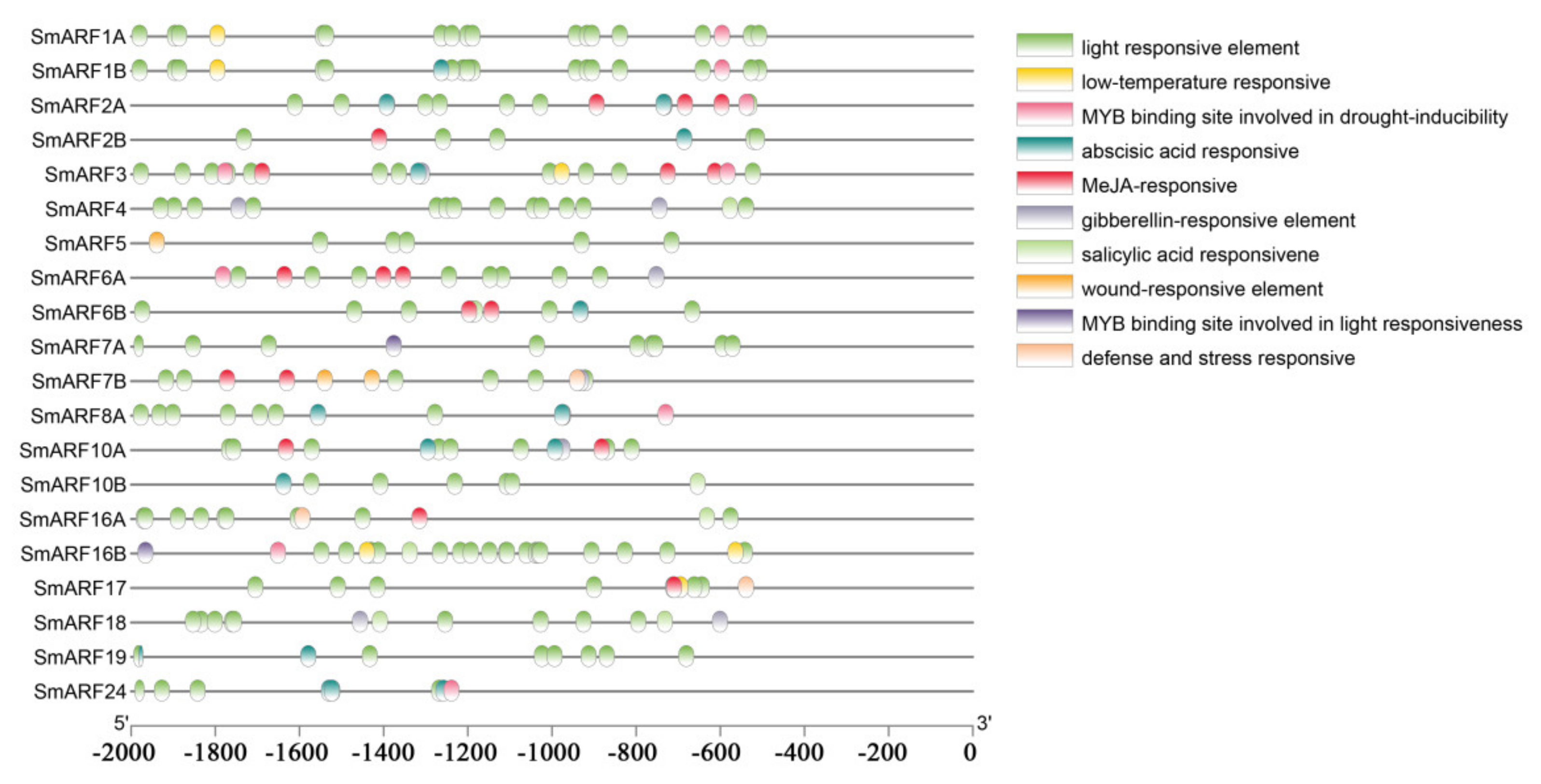
Promoters control gene expression, and transcription factors can regulate gene expression by binding to cis–acting elements. Various cis–acting elements either induce or inhibit gene expression in response to different biotic or abiotic stress signals. We predicted and analyzed potential cis–acting elements in the promoters of all the identified SmARFs using the PlantCARE tool. Several well–characterized elements were identified in the promoter regions of SmARFs, including light–responsive, low–temperature responsive, gibberellin–responsive, methyl jasmonate (MeJA)–responsive, abscisic acid–responsive, defense–responsive, stress–responsive, salicylic acid–responsive, and wound–responsive elements, in addition to MYB–binding sites (Figure 4). Multiple elements were present in each promoter. The identified light–responsive elements included TCT–motif, LAMP–element, I–box, GT1–motif, G–box, GATT–motif, GA–motif, Box 4, ATCT–motif, and AE–box.
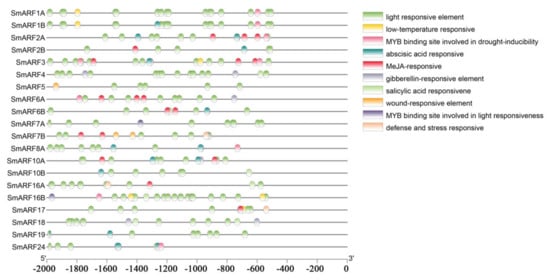
Figure 4.
Cis–acting regulatory elements upstream of auxin response factor genes in eggplant (Solanum melongena). The presence of cis–acting elements in a 2000–bp sequence upstream of the ATG start codons was assessed using PlantCARE. The horizontal scale indicates promoter length.
2.5. Subcellular Localization of SmARFs
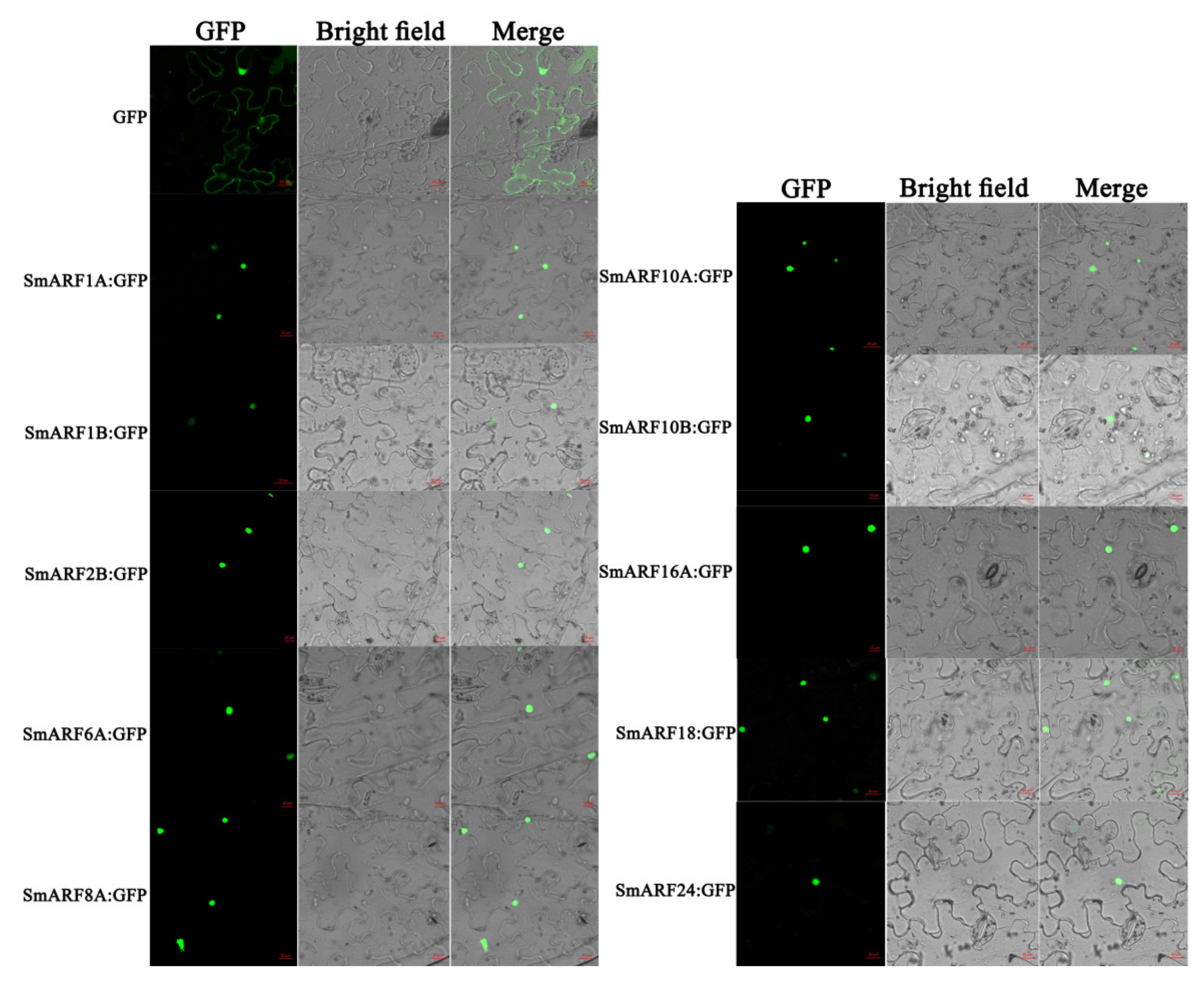
Ten SmARFs (SmARF1A, SmARF6A, SmARF2B, SmARF1B, SmARF18, SmARF8A, SmARF10B, SmARF24, SmARF16A, and SmARF10A) from each of the three clades were selected for the analysis of subcellular localization. To perform this, SmARF:GFP dual–expression vectors were constructed for each SmARF, and a GFP construct was used as a positive control. Fluorescence microscopy revealed that the GFP signals of all 10 fusion proteins were detected in the nucleus (Figure 5).

Figure 5.
Subcellular localization of SmARFs (bars = 20 μm). SmARF:GFP fusion proteins were transiently expressed in tobacco leaves, and their localization was determined using confocal microscopy. The green dots correspond to the nucleus.
2.6. Expression Patterns of SmARF Genes in Eggplant
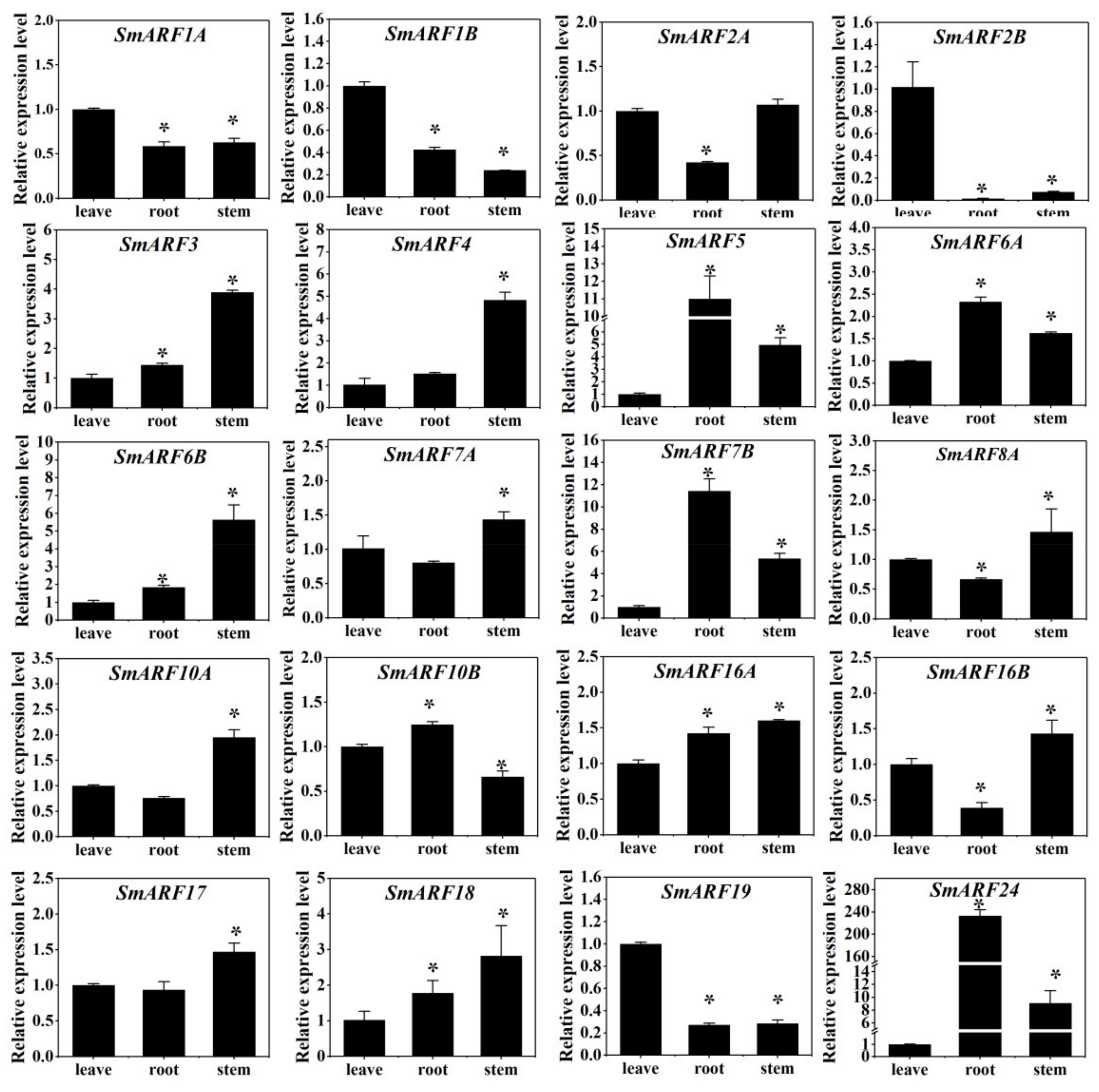
To explore the function of the SmARFs throughout the plant, the expression patterns of the SmARF genes were determined in eggplant leaves, roots, and stems using qPCR (Figure 6). Some SmARFs were differentially expressed in leaves (SmARF1A, 1B, 2B, and 19), roots (SmARF5, 6A, 7B, and 24), and stems (SmARF3, 4, 6B, 7A, 8A, 10A, 16B, 17, 18).

Figure 6.
The expression profiles of SmARF genes in the leaves, roots, and stems of 3–week–old eggplants. The relative expression level was calculated using the method of 2−△△Ct. The relative mRNA levels of the leaves were used for the reference. The values are means ± SD (n = 3). * represents significance at p < 0.05 comparing with reference.
2.7. Expression Analysis of SmARF Genes in Response to 2,4–D
ARFs are sensitive to auxin, and they are strong regulators of auxin signal transduction pathways. A synthetic growth regulator, 2,4–D, is usually sprayed on the flower stalk to reduce flower abscission and increase fruit set. To determine how ARFs are involved in the response to auxin, the 3 month–old eggplant flower buds were treated before blooming with 2,4–D for 0 h, 2 h, 4 h, and 6 h and then gene expression was analyzed using qPCR (Figure 7).

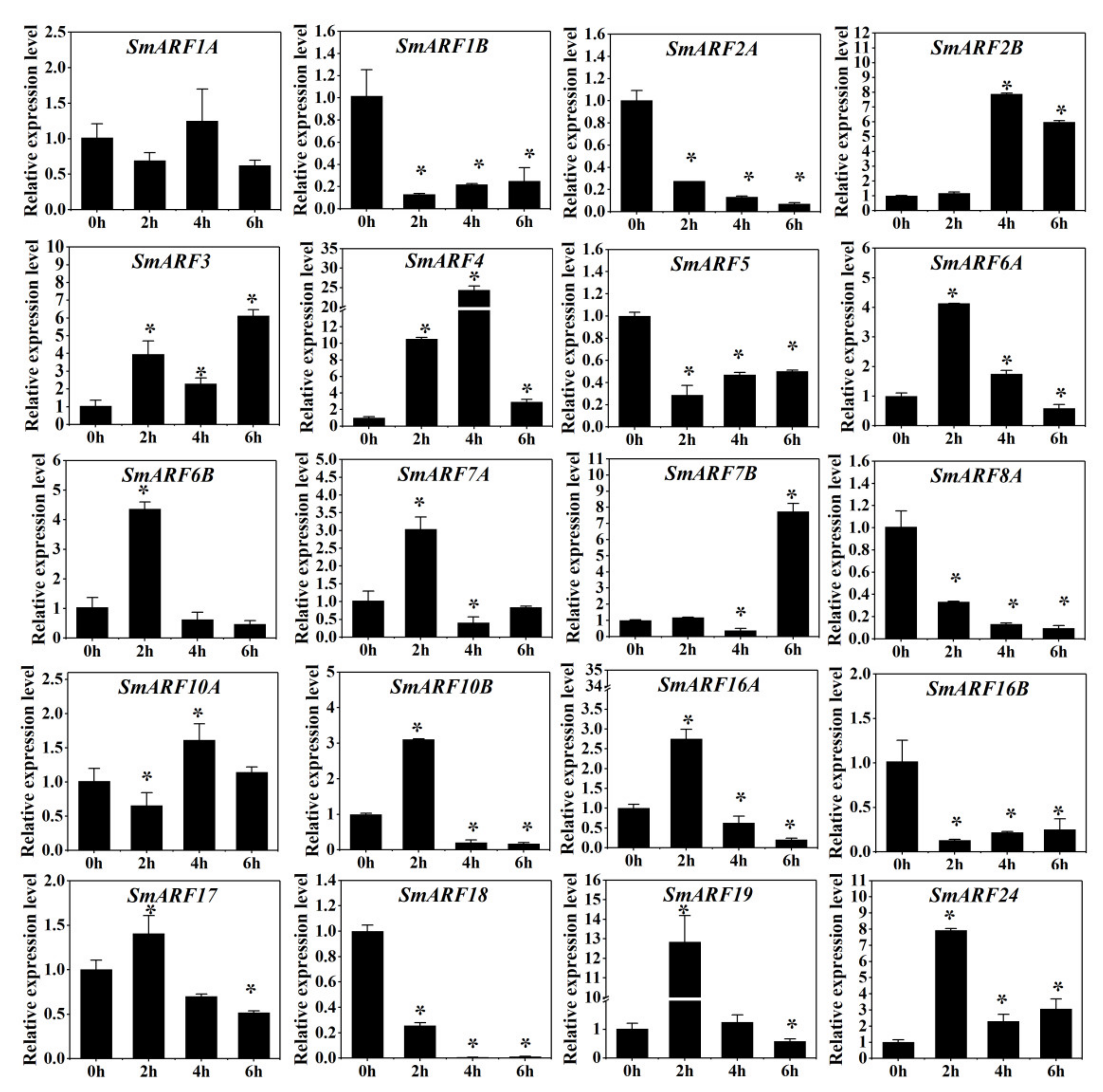
Figure 7.
Expression analysis of SmARFs after 2,4–D treatment. The relative mRNA levels of the group at 0 h were used for the reference. The relative expression was calculated using the of 2−△△Ct method. Values are means ± SD (n = 3). * represents significance at p < 0.05 compared with the reference.
A two–hour 2,4–D treatment increased the expression of SmARF2B, SmARF6A, SmARF6B, SmARF7A, SmARF10B, SmARF16A, SmARF19, and SmARF24 by more than 2–fold, and decreased the expression of SmARF1B, SmARF2A, SmARF5, SmARF8A, SmARF16B, and SmARF18 by more than 2–fold. However, after four hours of exposure, the expression levels of SmARF3, SmARF6A, SmARF6B, SmARF7A, SmARF10B, SmARF16A, SmARF19, and SmARF24 decreased to lower levels than without treatment. A six–hour treatment increased the expression of SmARF3, and SmARF17.
2.8. Expression Analysis of SmARF Genes in Response to Salt Stress
During cultivation, soil secondary salinization is one of the main stresses that can limit the growth of eggplants. Here, we treated eggplant seedlings with 250 mm NaCl for 0 h, 2 h, and 24 h, and then analyzed the expression of the 20 SmARF genes using qRT–PCR (Figure 8).

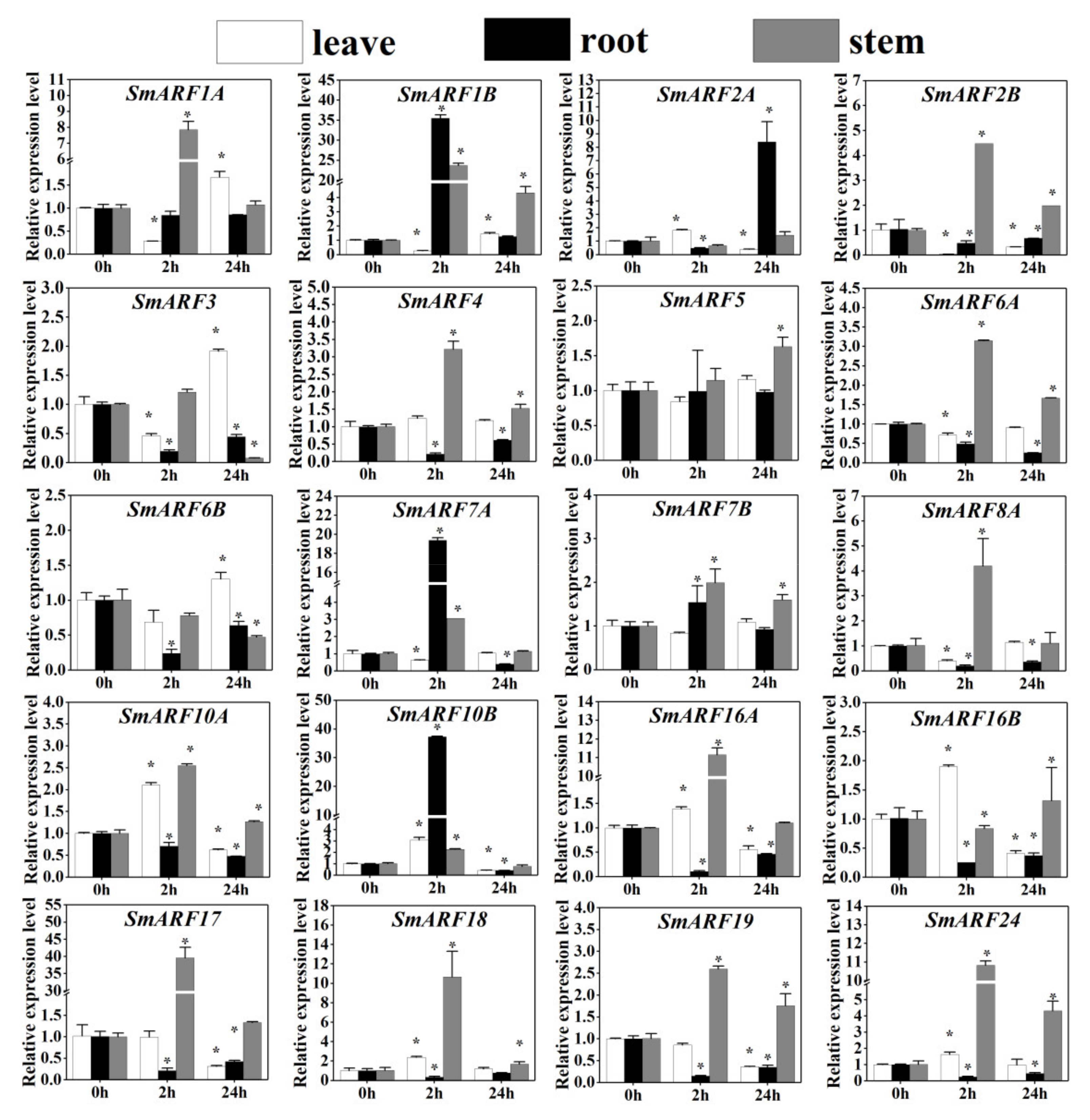
Figure 8.
The expression profiles of auxin response factor (ARF) genes in eggplant (Solanum melongena L.) in response to NaCl treatment. The relative expression levels of 20 ARF genes in leaves, roots, and stems of 3 week–old eggplants were measured at 0, 2, and 24 h of treatment. The relative mRNA levels of the group at 0 h were used as for reference. The relative expression was calculated using the method of 2−△△Ct. Values are means ± SD (n = 3). * represents significance at p < 0.05 comparing with the references.
All of the SmARFs exhibited rapid responses to salt stress in leaves except SmARF4/5/7B/17/19, in roots except SmARF1A/5/7B, and in stems except SmARF5. A two–hour salt treatment increased the expression of seven genes (SmARF2A, SmARF10A, SmARF10B, SmARF16A, SmARF16B, SmARF18, and SmARF16) in leaves, three genes (SmARF1B, SmARF7A, and SmARF10B) in roots, and 15 genes (SmARF1A, SmARF1B, SmARF2B, SmARF3, SmARF4, SmARF6A, SmARF7A, SmARF7B, SmARF8A, SmARF10B, SmARF16A, SmARF17, SmARF18, SmARF19, and SmARF24) in stems, suggesting an involvement of the ARF genes in salt stress responses in eggplant.
3. Discussion
Multiple ARF transcription factors have been reported for Arabidopsis thaliana [13], tomato (Solanum lycopersicum) [17], papaya (Carica papaya L.) [18], longan (Dimocarpus longan L.) [23], apple (Malus domestica) [25], and rice (Oryza sativa) [45], with 23, 22, 11, 17, 31, and 25 genes, respectively. In this study, we identified and characterized 20 SmARF transcription factors in eggplant. They were unevenly distributed on 12 chromosomes and have highly similar domains. Each ARF generally consists of three conservative domains, B3, auxin_resp, and Aux/IAA–binding [46]. Among the 20 SmARFs, all but SmARF19 included a typical DBD domain (B3). The lack of a B3 domain suggests that the protein encoded by this gene is not able to recognize and bind to the auxin response element in the promoter sequences of target genes [45]. As transcription factors, ARF proteins generally act in the nucleus. Consistent with the subcellular location of ARF in other species, all SmARF members are predicted to be in the nucleus, consistent with their function as transcription factors [17,47]. Our results with fluorescent reporters confirmed nuclear localization for 10 of these proteins (Figure 5).
A phylogenetic tree was constructed to analyze relationships between ARF family genes in eggplant, A. thaliana, and tomato (Figure 2). The results showed that most sister gene pairs with high bootstrap values (≥99%) were identified between eggplant and tomato. The absence of any SmARF gene in subclass IIId suggested that the nine duplicated AtARF genes of this group were originally derived from a single AtARF gene. This result was consistent with the tomato ARFs [25], suggesting that ARFs in eggplant were highly homologous to those in tomato. Conserved motifs for transcription factors correlate with protein interactions, transcriptional activity, and DNA–binding [24]. Ten conserved motifs were identified in this study (Figure 2). All the motifs common or specific to particular groups were represented in the phylogenetic tree (Figure 3). Although the number of members in each phylogenetic group varied, there was a strong conservation of the patterns of motifs within a group. The comparison of sequences of novel functional domains/motifs across multiple orthologous proteins is an approach that is widely used to predict protein functions based on evolutionary conservation. Although most motifs in SmARFs are conserved, other motifs may be associated with novel functions in plants and should be further investigated. Defense–responsive, stress–responsive, light–responsive, low temperature–responsive, drought–responsive, gibberellin–responsive, MeJA–responsive, abscisic acid–responsive, salicylic acid–responsive, and wound–responsive elements, as well as MYB–binding sites, were identified upstream of SmARF genes (Figure 4).
To identify the functions of SmARFs in eggplant, we analyzed their expression in leaves, roots, and stems. The data indicated that the expression of SmARFs was ubiquitous in all tissues. Among these genes, SmARF2B was significantly highly expressed in leaves. Similarly, the expressions of its tomato homolog, SlARF2B, were also significantly higher in leaves [25]. In Arabidopsis thaliana, AtARF2 could regulate leaf senescence [31]. Thus, we speculated that SmARF2B might play a similar role in eggplant leaves. However not all cases were the same. For example, SmARF10A was significantly high expressed in stems, while SlARF10A has been reported to be involved in the spatial restriction of the auxin response that drive leaf blade outgrowth in tomato [38]. Further work will investigate the function of SmARFs on growth and development of eggplants.
As transcription factor, ARFs participate in signal pathways related to auxin response and regulate the flower–to–fruit transition [38,39,40,41,42,43,48]. The growth regulator 2,4–D was used to mitigate flower abscission and increase fruit set. In this study, the expression of SmARF2B was significantly increased in flower buds after 2,4–D treatment (Figure 7). The expression of its tomato homolog, SlARF2, was clearly responded to auxin. In addition, SlARF2 could regulate lateral root formation and flower organ senescence [49]. The silenced AtARF2/3/4 line leads to abnormal morphology of pollen grains [32]. The expression of SmARF7A was significantly increased after 2,4–D treatment for 2 h. The highly expression level of SlARF7 in placental tissues of the mature flower could activate the auxin response, attenuating genes that might repress the auxin response and prevent fruit set [50]. We also found that SmARF10A was significantly increased after 2,4–D treatment for 4 h. It has been proven that SlARF10A, the target of sly–miR160, regulated auxin–mediated ovary patterning, as well as floral organ abscission and lateral organ lamina outgrowth [51]. Thus, we speculated that SmARFs can exhibit many different functions in different plant species.
Transcription factors (TFs) coordinate gene expression by activating or inhibiting transcription in response to various abiotic stress signals [30,38,39,40,41,42,44,45,46,47,48,49,50,51,52]. In this study, the expression patterns of SmARF genes in response to NaCl treatment indicated that they might have crucial roles in eggplant leaves, roots, and stems (Figure 8). For example, the expression of SmARF2A was significantly decreased in 2 h and increased 24 h after salt treatment. Previous studies have reported that the up–regulation of SlARF2 results in various asexual reproduction growth phenotypes, including increased lateral root formation [49]. The inhibition of AtARF2, AtARF3, and AtARF4 expression via tasiRNAs may release the repression of Arabidopsis lateral root growth [48]. In addition, SmARF7A was significantly repressed or induced, respectively, in leaves, roots, and stems under salt stress, which indicates that SmARF7A might be involved in the stress response within eggplant. Similar expression profiles werealso found in SlARF7A, which is phylogenetically close to SmARF7A from tomato [38]. It had shown that AtARF7 is involved in the control of lateral root formation in Arabidopsis [30]. The other ARF transcription factors also played an essential role in the response to phytohormones and abiotic stress. For example, the IbARF5 gene from sweet potato (Ipomoea batatas) could increase its tolerance to salt and drought stress in transgenic Arabidopsis [41]. In Tamarix chinensis, TcARF6 was rapidly expressed in response to salt stress, but it was significantly downregulated specifically in the roots [42].
4. Materials and Methods
4.1. Identification of ARFs in the Eggplant Genome
SmARF gene sequences were obtained from the Eggplant Genome database https://solgenomics.net/organism/Solanum_melongena/genome (accessed on 18 December 2021) [44], the S. lycopersicum database https://solgenomics.net/organism/Solanum_lycopersicum/genome (accessed on 18 December 2021) [53], and the Arabidopsis thaliana database https://www.arabidopsis.org (accessed on 18 December 2021) [54].
To identify ARFs and to remove redundant sequences, “auxin response factor” was input as keywords to search the database and genome websites. The number of amino acids, molecular weight, and theoretical isoelectric point (pI) of putative ARF sequences were calculated using EXPASy https://web.expasy.org/protparam (accessed on 25 December 2021). Subcellular localization was predicted using the BUSCA tool.
4.2. Phylogenetic Analysis of ARF Genes
Full–length protein sequences were used for phylogenetic analysis. The protein sequences were aligned using ClustalX software with default parameters [55]. Phylogenetic trees were generated using MEGA7 software, with a bootstrap test of 1000 replicates [56]. The final tree was viewed and modified using Interactive Tree Of Life (iTOL) software https://itol.embl.de/personal_page.cgi (accessed on 3 January 2022) [57].
4.3. Structural Characterization
The subcellular location and intron numbers of ARFs were obtained from genome databases. Conserved motifs were identified using MEME software version 5.0.5 http://meme–suite.org/tools/meme (accessed on 5 January 2022) with the following parameters: any number of repetitions, a maximum of 10 misfits, and an optimum motif width of 6–200 amino acid residues [58]. The upstream sequences (2 kb) of SmARFs were also retrieved from the genome database, and regulatory elements were identified by analysis using the PlantCARE database http://bioinformatics.psb.ugent.be/webtools/plantcare/html (accessed on 7 January 2022) [59]. Next, the structure and composition of SmARF genes were investigated using TBtools software [60]. The conserved motifs senquences were listed in Table S2 and the Cis-acting elements on promoters of SmARFs were listed in Table S3.
4.4. Subcellular Localization
The complete coding sequences of 10 SmARFs (SmARF1A, SmARF6A, SmARF2B, SmARF1B, SmARF18, SmARF8A, SmARF10B, SmARF24, SmARF16A, and SmARF10A) were obtained from the Eggplant Genome database [44]. HindIII and BamHI enzymes were used to linearize the CaMV35S promoter vector with GFP. Sequences of 10 SmARFs were combined into the vector using In–Fusion Snap Assembly cloning kits (Takara, Dalian, China) according to the manual. The fusion construct was transferred into Agrobacterium tumefaciens strain GV3101, and subcellular localization assays were performed as previously reported [61]. The protein localization was determined under 20× confocal microscopy. The primer sequences were listed in Table S1.
4.5. Growth Conditions and Treatments
Seeds of the high generation inbred line No. 108 [62] were sterilized for 10 min in 50% sodium hypochlorite, rinsed four times with sterile distilled water, and sown in pots containing peat. Seeds were grown at 25 ± 2 °C under a light regimen of 16 h light and 8 h dark at a temperature of 16 ± 2 °C for 40 days. The plants were then transferred to a greenhouse and grown under the same temperature and photoperiod.
For chemical treatment, the herbicide 2,4–dichlorophenoxyacetic acid (2,4–D; BDH Chemical) was dissolved in water according to the manual, and the pH was adjusted to 7.0. Flower buds with no obvious pests, diseases, or mechanical damage were soaked in 1 mL of 1.0 mM 2,4–D for 0, 2, 4, and 6 h.
For salt stress treatment, plants with three true leaves were treated with 250 mM NaCl solution (salt stress group). Leaves, stems, and roots samples were harvested after 0, 2, and 24 h treatment.
4.6. RNA Extraction and qRT–PCR Analysis
All samples were frozen in liquid nitrogen and stored at −80 °C for RNA extraction and other analyses. Total RNA was extracted from flower buds, leaves, roots, and stems using the RNA Isolation Kit (Ambion, Thermo Fisher Scientific, Waltham, MA, USA) following the manufacturer’s protocol, and stored at −80 °C. RNA integrity was evaluated using the Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA).
First–strand cDNA was synthesized from 1 μg of total RNA using a Prime Script RT Reagent Kit (Takara, Dalian, China). PCR reactions were performed using the ABI 7500 Fast Real–Time PCR system (Applied Biosystems, Foster City, California, USA) and the QuantiFast SYBR Green PCR Kit (Qiagen, Duesseldorf, Germany). The amplification parameters were 95 °C for 5 min, followed by 45 cycles at 95 °C for 10 s, 60 °C for 10 s, and 72 °C for 10 s. The mRNA expression levels were normalized to the level of SmActin expression using the 2−ΔΔCt method [49]. Each experiment included three biological replicates. The primer sequences are listed in Table S1.
4.7. Statistical Analysis
In all the presented figures, error bars indicated standard deviation. SPSS 23.0 (SPSS, Inc., Chicago, IL, USA) was used to assess statistical significance using one–way ANOVA and Duncan’s New Multiple Range test (p < 0.05).
5. Conclusions
This is the first study of the evolution, expression profiles, and putative functions of ARF genes in eggplants. In the current study, 20 SmARF members were identified from the eggplant genome, and divided into three clades. Their gene structures were similar, and most members have a conserved ARF domain. Fluorescent reporters confirmed nuclear localization for 10 of these proteins. These members were highly expressed in leaves (SmARF1A, 1B, 2B, and 19), roots (SmARF5, 6A, 7B, and 24), and stems (SmARF3, 4, 6B, 7A, 8A, 10A, 16B, 17, and 18). These genes may play important roles in regulating eggplant development. The similar expression profiles of SmARF2B/2A/7A/10A to SlARF2/7/10A when under 2,4–D or salt treatment showed that these genes may play a similar role in regulating root formation and flower organ senescence, fruit set and lateral root formation, and floral organ abscission and lateral organ lamina outgrowth. These data will help to improve genotype selection, gene function analysis, and the improvement of agronomic traits in eggplants.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms23116219/s1.
Author Contributions
Data curation, J.C. and S.W.; Investigation, J.C.; Methodology, J.C.; Project administration, J.C. and F.W.; Writing—original draft, J.C.; Writing—review and editing, F.Y., J.L. and M.W. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the National Natural Sciences Foundations of China (grant number: 32172556), Youth Project of Shandong Provincial Natural Science Foundation (grant number: ZR2020QC155), Key Project of Shandong Provincial Natural Science Foundation (ZR2020KC039), and Special Project Protected Horticulture Advantage Team of Shandong Agricultural University “Double First–Class” Science and Technology Team (grant number: SYL2017YSTD07).
Institutional Review Board Statement
The study was conducted in accordance with the Declaration of Helsinki. This study did not include human samples and it did not require the approval of the Institutional Review Board.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Kaushik, P. Genetic Analysis for fruit phenolics content, flesh color, and browning related traits in eggplant (Solanum melongena). Int. J. Mol. Sci. 2019, 20, 2990. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mishiba, K.I.; Nishida, K.; Inoue, N.; Fujiwara, T.; Teranishi, S.; Iwata, Y.; Takeda, S.; Koizumi, N. Genetic engineering of eggplant accumulating β–carotene in fruit. Plant Cell Rep. 2020, 39, 1029–1039. [Google Scholar] [CrossRef] [PubMed]
- Wei, Q.; Wang, W.; Hu, T.; Hu, H.; Wang, J.; Bao, C. Construction of a SNP–based genetic map using SLAF–Seq and QTL analysis of morphological traits in eggplant. Front. Genet. 2020, 11, 178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Food and Agricultural Organization of the United Nations. Available online: http://www.fao.org/faostat/en/#rankings/countries_by_commodity (accessed on 28 December 2021).
- Olsson, V.; Butenko, M.A. Abscission in plants. Curr. Biol. 2018, 28, R338–R339. [Google Scholar] [CrossRef] [Green Version]
- Liljegren, S.J. Organ abscission: Exit strategies require signals and moving traffic. Curr. Opin. Plant Biol. 2012, 15, 670–676. [Google Scholar] [CrossRef]
- Cooper, W.C.; Rasmussen, G.K.; Rogers, B.J.; Reece, P.C.; Henry, W.H. Control of abscission in agricultural crops and its physiological basis. Plant Physiol. 1968, 43, 1560–1576. [Google Scholar]
- Benková, E.; Michniewicz, M.; Sauer, M.; Teichmann, T.; Seifertová, D.; Jürgens, G.; Friml, J. Local, efflux–dependent auxin gradients as a common module for plant organ formation. Cell 2003, 115, 591–602. [Google Scholar] [CrossRef] [Green Version]
- Ljung, K. Auxin metabolism and homeostasis during plant development. Development 2013, 140, 943–950. [Google Scholar] [CrossRef] [Green Version]
- Woodward, A.W.; Bartel, B. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 707–735. [Google Scholar] [CrossRef] [Green Version]
- Goldental–Cohen, S.; Israeli, A.; Ori, N.; Yasuor, H. Auxin Response Dynamics During Wild–Type and entire Flower Development in Tomato. Plant Cell Physiol. 2017, 58, 1661–1672. [Google Scholar] [CrossRef] [Green Version]
- Nemhauser, J.L.; Zambryski, P.C.; Roe, J.L. Auxin signaling in Arabidopsis flower development? Curr. Opin. Plant Biol. 1998, 1, 531–535. [Google Scholar] [CrossRef]
- Guilfoyle, T.J.; Hagen, G. Auxin response factors. Curr. Opin. Plant Biol. 2007, 10, 453–460. [Google Scholar] [CrossRef]
- Chandler, J.W. Auxin response factors. Plant Cell Environ. 2016, 39, 1014–1028. [Google Scholar] [CrossRef] [Green Version]
- Audran–Delalande, C.; Bassa, C.; Mila, I.; Regad, F.; Zouine, M.; Bouzayen, M. Genome–wide identification, functional analysis and expression profiling of the Aux/IAA gene family in tomato. Plant Cell Physiol. 2012, 53, 659–672. [Google Scholar] [CrossRef] [Green Version]
- Kumar, R.; Tyagi, A.K.; Sharma, A.K. Genome–wide analysis of auxin response factor (ARF) gene family from tomato and analysis of their role in flower and fruit development. Mol. Genet. Genom. MGG 2011, 285, 245–260. [Google Scholar] [CrossRef]
- Zouine, M.; Fu, Y.; Chateigner–Boutin, A.L.; Mila, I.; Frasse, P.; Wang, H.; Audran, C.; Roustan, J.P.; Bouzayen, M. Characterization of the tomato ARF gene family uncovers a multi–levels post–transcriptional regulation including alternative splicing. PLoS ONE 2014, 9, e84203. [Google Scholar] [CrossRef]
- Liu, K.; Yuan, C.; Li, H.; Lin, W.; Yang, Y.; Shen, C.; Zheng, X. Genome–wide identification and characterization of auxin response factor (ARF) family genes related to flower and fruit development in papaya (Carica papaya L.). BMC Genom. 2015, 16, 901. [Google Scholar] [CrossRef] [Green Version]
- Cui, J.; Li, X.; Li, J.; Wang, C.; Cheng, D.; Dai, C. Genome–wide sequence identification and expression analysis of ARF family in sugar beet (Beta vulgaris L.) under salinity stresses. PeerJ 2020, 8, e9131. [Google Scholar] [CrossRef]
- Diao, D.; Hu, X.; Guan, D.; Wang, W.; Yang, H.; Liu, Y. Genome–wide identification of the ARF (auxin response factor) gene family in peach and their expression analysis. Mol. Biol. Rep. 2020, 47, 4331–4344. [Google Scholar] [CrossRef]
- Li, W.; Chen, F.; Wang, Y.; Zheng, H.; Yi, Q.; Ren, Y.; Gao, J. Genome–wide identification and functional analysis of ARF transcription factors in Brassica juncea var. tumida. PLoS ONE 2020, 15, e0232039. [Google Scholar] [CrossRef] [Green Version]
- Ma, F.; Huang, J.; Yang, J.; Zhou, J.; Sun, Q.; Sun, J. Identification, expression and miRNA targeting of auxin response factor genes related to phyllody in the witches’ broom disease of jujube. Gene 2020, 746, 144656. [Google Scholar] [CrossRef]
- Peng, Y.; Fang, T.; Zhang, Y.; Zhang, M.; Zeng, L. Genome–wide identification and expression analysis of auxin response factor (ARF) gene family in Longan (Dimocarpus longan L.). Plants 2020, 9, 221. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Luo, W.; Sun, Y.; Chang, H.; Ma, K.; Zhao, Z.; Lu, L. Identification and expression analysis of miR160 and their target genes in cucumber. Biochem. Genet. 2021, 60, 127–152. [Google Scholar] [CrossRef]
- Luo, X.C.; Sun, M.H.; Xu, R.R.; Shu, H.R.; Wang, J.W.; Zhang, S.Z. Genomewide identification and expression analysis of the ARF gene family in apple. J. Genet. 2014, 93, 785–797. [Google Scholar] [CrossRef]
- Hardtke, C.S.; Ckurshumova, W.; Vidaurre, D.P.; Singh, S.A.; Stamatiou, G.; Tiwari, S.B.; Hagen, G.; Guilfoyle, T.J.; Berleth, T. Overlapping and non–redundant functions of the Arabidopsis auxin response factors MONOPTEROS and NONPHOTOTROPIC HYPOCOTYL 4. Development 2004, 131, 1089–1100. [Google Scholar] [CrossRef] [Green Version]
- Mallory, A.C.; Bartel, D.P.; Bartel, B. MicroRNA–directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 2005, 17, 1360–1375. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Tiwari, S.B.; Hagen, G.; Guilfoyle, T.J. AUXIN RESPONSE FACTOR7 restores the expression of auxin–responsive genes in mutant Arabidopsis leaf mesophyll protoplasts. Plant Cell 2005, 17, 1979–1993. [Google Scholar] [CrossRef] [Green Version]
- Schruff, M.C.; Spielman, M.; Tiwari, S.; Adams, S.; Fenby, N.; Scott, R.J. The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 2006, 133, 251–261. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Okushima, Y.; Fukaki, H.; Onoda, M.; Theologis, A.; Tasaka, M. ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 2007, 19, 118–130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, P.O.; Lee, I.C.; Kim, J.; Kim, H.J.; Ryu, J.S.; Woo, H.R.; Nam, H.G. Auxin response factor 2 (ARF2) plays a major role in regulating auxin–mediated leaf longevity. J. Exp. Bot. 2010, 61, 1419–1430. [Google Scholar] [CrossRef] [PubMed]
- Ellis, C.M.; Nagpal, P.; Young, J.C.; Hagen, G.; Guilfoyle, T.J.; Reed, J.W. AUXIN RESPONSE FACTOR1 and AUXIN RESPONSE FACTOR2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 2005, 132, 4563–4574. [Google Scholar] [CrossRef] [Green Version]
- Wu, L.; Tian, Z.; Zhang, J. Functional Dissection of Auxin Response Factors in Regulating Tomato Leaf Shape Development. Front. Plant Sci. 2018, 9, 957. [Google Scholar] [CrossRef] [Green Version]
- Chen, M.; Zhu, X.; Liu, X.; Wu, C.; Yu, C.; Hu, G.; Chen, L.; Chen, R.; Bouzayen, M.; Zouine, M.; et al. Knockout of auxin response factor SlARF4 improves tomato resistance to water deficit. Int. J. Mol. Sci. 2021, 22, 3347. [Google Scholar] [CrossRef]
- Liu, S.; Zhang, Y.; Feng, Q.; Qin, L.; Pan, C.; Lamin–Samu, A.T.; Lu, G. Tomato AUXIN RESPONSE FACTOR 5 regulates fruit set and development via the mediation of auxin and gibberellin signaling. Sci. Rep. 2018, 8, 2971. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Yan, F.; Tang, Y.; Yuan, Y.; Deng, W.; Li, Z. Auxin response gene SlARF3 plays multiple roles in tomato development and is involved in the formation of epidermal cells and trichomes. Plant Cell Physiol. 2015, 56, 2110–2124. [Google Scholar] [CrossRef] [Green Version]
- Munns, R.; Passioura, J.B.; Colmer, T.D.; Byrt, C.S. Osmotic adjustment and energy limitations to plant growth in saline soil. New Phytol. 2020, 225, 1091–1096. [Google Scholar] [CrossRef] [Green Version]
- Bouzroud, S.; Gouiaa, S.; Hu, N.; Bernadac, A.; Mila, I.; Bendaou, N.; Smouni, A.; Bouzayen, M.; Zouine, M. Auxin Response Factors (ARFs) are potential mediators of auxin action in tomato response to biotic and abiotic stress (Solanum lycopersicum). PLoS ONE 2018, 13, e0193517. [Google Scholar] [CrossRef] [Green Version]
- Bouzroud, S.; Gasparini, K.; Hu, G.; Barbosa, M.; Rosa, B.L.; Fahr, M.; Bendaou, N.; Bouzayen, M.; Zsögön, A.; Smouni, A.; et al. Down Regulation and Loss of Auxin Response Factor 4 Function Using CRISPR/Cas9 Alters Plant Growth, Stomatal Function and Improves Tomato Tolerance to Salinity and Osmotic Stress. Genes 2020, 11, 272. [Google Scholar] [CrossRef] [Green Version]
- Jain, M.; Khurana, J.P. Transcript profiling reveals diverse roles of auxin–responsive genes during reproductive development and abiotic stress in rice. FEBS J. 2009, 276, 3148–3162. [Google Scholar] [CrossRef]
- Kang, C.; He, S.; Zhai, H.; Li, R.; Zhao, N.; Liu, Q. A Sweetpotato Auxin Response Factor Gene (IbARF5) is involved in carotenoid biosynthesis and salt and drought tolerance in transgenic Arabidopsis. Front. Plant Sci. 2018, 9, 1307. [Google Scholar] [CrossRef]
- Ye, Y.; Wang, J.; Wang, W.; Xu, L.A. ARF family identification in Tamarix chinensis reveals the salt responsive expression of TcARF6 targeted by miR167. PeerJ 2020, 8, e8829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Agharbaoui, Z.; Leclercq, M.; Remita, M.A.; Badawi, M.A.; Lord, E.; Houde, M.; Danyluk, J.; Diallo, A.B.; Sarhan, F. An integrative approach to identify hexaploid wheat miRNAome associated with development and tolerance to abiotic stress. BMC Genom. 2015, 16, 339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barchi, L.; Rabanus–Wallace, M.T.; Prohens, J.; Toppino, L.; Padmarasu, S.; Portis, E.; Rotino, G.L.; Stein, N.; Lanteri, S.; Giuliano, G. Improved genome assembly and pan–genome provide key insights into eggplant domestication and breeding. Plant J. Cell Mol. Biol. 2021, 107, 579–596. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Pei, K.; Fu, Y.; Sun, Z.; Li, S.; Liu, H.; Tang, K.; Han, B.; Tao, Y. Genome–wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 2007, 394, 13–24. [Google Scholar] [CrossRef]
- Korasick, D.A.; Jez, J.M.; Strader, L.C. Refining the nuclear auxin response pathway through structural biology. Curr. Opin. Plant Biol. 2015, 27, 22–28. [Google Scholar] [CrossRef] [Green Version]
- Xing, H.; Pudake, R.N.; Guo, G.; Xing, G.; Hu, Z.; Zhang, Y.; Sun, Q.; Ni, Z. Genome–wide identification and expression profiling of auxin response factor (ARF) gene family in maize. BMC Genom. 2011, 12, 178. [Google Scholar] [CrossRef] [Green Version]
- Marin, E.; Jouannet, V.; Herz, A.; Lokerse, A.S.; Weijers, D.; Vaucheret, H.; Nussaume, L.; Crespi, M.D.; Maizel, A. miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 2010, 22, 1104–1117. [Google Scholar] [CrossRef] [Green Version]
- Ren, Z.; Liu, R.; Gu, W.; Dong, X. The Solanum lycopersicum auxin response factor SlARF2 participates in regulating lateral root formation and flower organ senescence. Plant Sci. Int. J. Exp. Plant Biol. 2017, 256, 103–111. [Google Scholar] [CrossRef]
- de Jong, M.; Mariani, C.; Vriezen, W.H. The role of auxin and gibberellin in tomato fruit set. J. Exp. Bot. 2009, 60, 1523–1532. [Google Scholar] [CrossRef] [Green Version]
- Damodharan, S.; Zhao, D.; Arazi, T. A common miRNA160–based mechanism regulates ovary patterning, floral organ abscission and lamina outgrowth in tomato. Plant J. Cell Mol. Biol. 2016, 86, 458–471. [Google Scholar] [CrossRef] [Green Version]
- Hou, Q.; Qiu, Z.; Wen, Z.; Zhang, H.; Li, Z.; Hong, Y.; Qiao, G.; Wen, X. Genome–Wide identification of ARF gene family suggests a functional expression pattern during fruitlet abscission in Prunus avium L. Int. J. Mol. Sci. 2021, 22, 1968. [Google Scholar] [CrossRef]
- Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635–641. [Google Scholar] [CrossRef] [Green Version]
- Berardini, T.Z.; Reiser, L.; Li, D.; Mezheritsky, Y.; Muller, R.; Strait, E.; Huala, E. The Arabidopsis information resource: Making and mining the “gold standard” annotated reference plant genome. Genesis 2015, 53, 474–485. [Google Scholar] [CrossRef] [Green Version]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL): An online tool for phylogenetic tree display and annotation. Bioinformatics 2007, 23, 127–128. [Google Scholar] [CrossRef] [Green Version]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis–acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Sparkes, I.A.; Runions, J.; Kearns, A.; Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat. Protoc. 2006, 1, 2019–2025. [Google Scholar] [CrossRef]
- Duan, Z.; Tian, S.; Yang, G.; Wei, M.; Li, J.; Yang, F. The Basic Helix–Loop–Helix Transcription Factor SmbHLH1 Represses Anthocyanin Biosynthesis in Eggplant. Front. Plant Sci. 2021, 12, 757936. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).