Exploring the Potential of Metatranscriptomics to Describe Microbial Communities and Their Effects in Molluscs
Abstract
:1. Introduction
2. Results
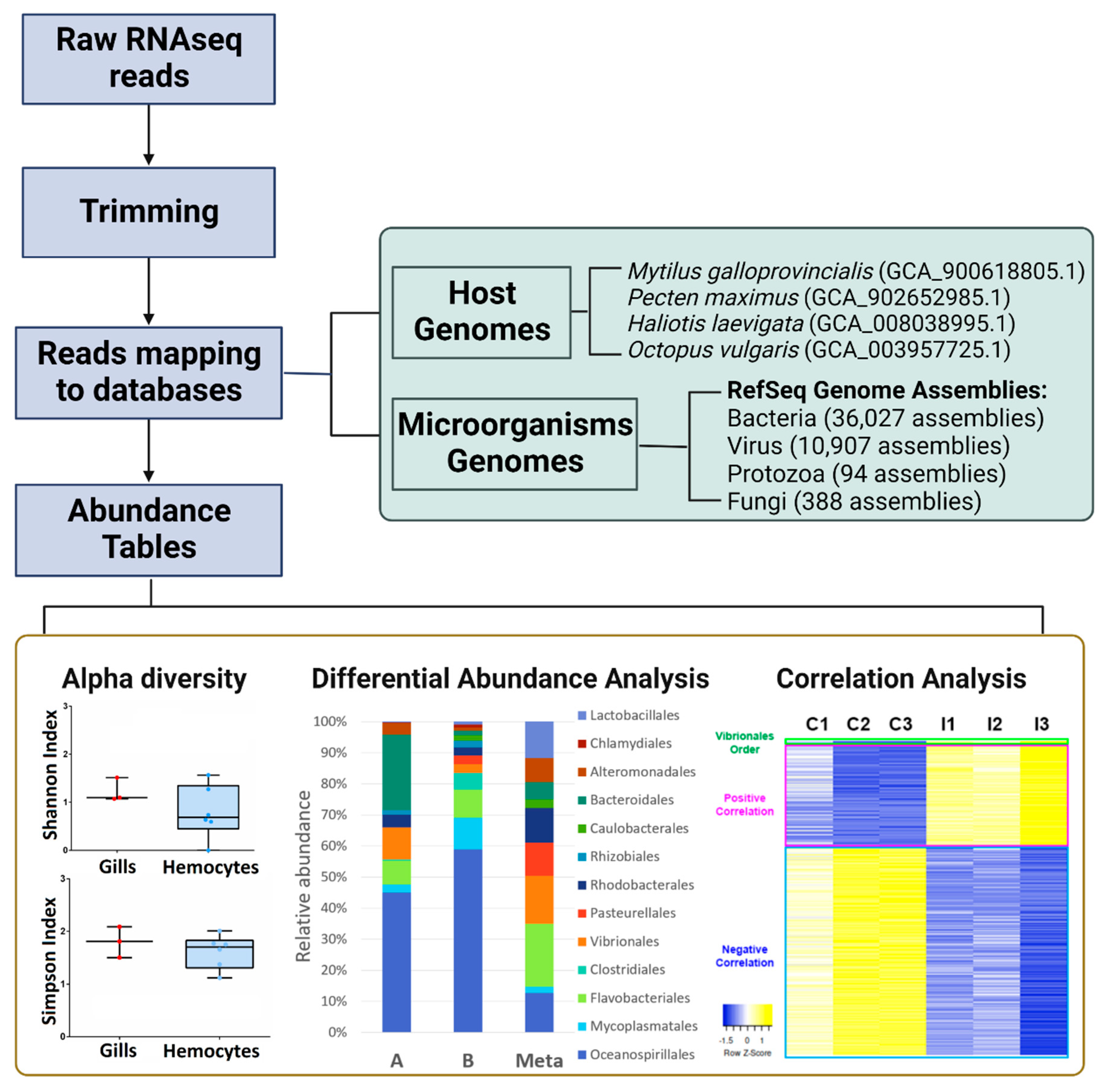
2.1. Comparison between 16S rRNA Sequencing and RNA-Seq Taxonomic Profiling
2.2. Microbiome Comparison between Gills and Haemocytes of Mytilus galloprovincialis
2.3. Effect of a Pathogenic Stimulus on the Microbiome Composition of M. galloprovincialis and Gene Expression
2.4. Haemocyte Microbiome of Molluscs
3. Discussion
4. Materials and Methods
4.1. 16S rRNA Sequencing
4.2. Taxonomic Profiling from RNA-Seq Data
4.3. Differential Abundance Analysis and Alpha Diversity
4.4. Correlation Microbiome/Transcriptome Results
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Suttle, C.A. Marine viruses–major players in the global ecosystem. Nat. Rev. Microbiol. 2007, 5, 801–812. [Google Scholar] [CrossRef]
- Azam, F.; Malfatti, F. Microbial structuring of marine ecosystems. Nat. Rev. Microbiol. 2007, 5, 782–791. [Google Scholar] [CrossRef]
- Rosani, U.; Gerdol, M. A bioinformatics approach reveals seven nearly-complete RNA-virus genomes in bivalve RNA-seq data. Virus Res. 2017, 239, 33–42. [Google Scholar] [CrossRef]
- Offret, C.; Jégou, C.; Mounier, J.; Fleury, Y.; Le Chevalier, P. New insights into the haemo- and coelo-microbiota with antimicrobial activities from Echinodermata and Mollusca. J. Appl. Microbiol. 2019, 126, 1023–1031. [Google Scholar] [CrossRef]
- Vezzulli, L.; Stagnaro, L.; Grande, C.; Tassistro, G.; Canesi, L.; Pruzzo, C. Comparative 16SrDNA Gene-Based Microbiota Profiles of the Pacific Oyster (Crassostrea gigas) and the Mediterranean Mussel (Mytilus galloprovincialis) from a Shellfish Farm (Ligurian Sea, Italy). Microb. Ecol. 2018, 75, 495–504. [Google Scholar] [CrossRef]
- Rosani, U.; Shapiro, M.; Venier, P.; Allam, B. A Needle in A Haystack: Tracing Bivalve-Associated Viruses in High-Throughput Transcriptomic Data. Viruses 2019, 11, 205. [Google Scholar] [CrossRef] [Green Version]
- Gantt, S.E.; López-Legentil, S.; Erwin, P.M. Stable microbial communities in the sponge Crambe crambe from inside and outside a polluted Mediterranean harbor. FEMS Microbiol. Lett. 2017, 364, 1–7. [Google Scholar] [CrossRef]
- Rosado Rodríguez, G.; Otero Morales, E. Assessment of heavy metal contamination at Tallaboa Bay (Puerto Rico) by marine sponges’ bioaccumulation and fungal community composition. Mar. Pollut. Bull. 2020, 161, 111803. [Google Scholar] [CrossRef]
- Lloyd-Price, J.; Abu-Ali, G.; Huttenhower, C. The healthy human microbiome. Genome Med. 2016, 8, 51. [Google Scholar] [CrossRef] [Green Version]
- Saltzman, E.T.; Thomsen, M.; Hall, S.; Vitetta, L. Perna canaliculus and the Intestinal Microbiome. Mar. Drugs 2017, 15, 207. [Google Scholar] [CrossRef]
- Douglas, A.E. Simple animal models for microbiome research. Nat. Rev. Microbiol. 2019, 17, 764–775. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Romero, A.; Costa M d Forn-Cuni, G.; Balseiro, P.; Chamorro, R.; Dios, S.; Figueras, A.; Novoa, B. Occurrence, seasonality and infectivity of Vibrio strains in natural populations of mussels Mytilus galloprovincialis. Dis. Aquat. Organ. 2014, 108, 149–163. [Google Scholar] [CrossRef] [PubMed]
- Sanschagrin, S.; Yergeau, E. Next-generation sequencing of 16S ribosomal RNA gene amplicons. J. Vis. Exp. 2014, 90, 51709. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Popovic, A.; Parkinson, J. Characterization of Eukaryotic Microbiome Using 18S Amplicon Sequencing. Methods Mol. Biol. 2018, 1849, 29–48. [Google Scholar] [CrossRef] [PubMed]
- Auguste, M.; Lasa, A.; Pallavicini, A.; Gualdi, S.; Vezzulli, L.; Canesi, L. Exposure to TiO2 nanoparticles induces shifts in the microbiota composition of Mytilus galloprovincialis hemolymph. Sci. Total Environ. 2019, 670, 129–137. [Google Scholar] [CrossRef]
- Auguste, M.; Lasa, A.; Balbi, T.; Pallavicini, A.; Vezzulli, L.; Canesi, L. Impact of nanoplastics on hemolymph immune parameters and microbiota composition in Mytilus galloprovincialis. Mar. Environ. Res. 2020, 159, 105017. [Google Scholar] [CrossRef]
- Li, L.L.; Amara, R.; Souissi, S.; Dehaut, A.; Duflos, G.; Monchy, S. Impacts of microplastics exposure on mussel (Mytilus edulis) gut microbiota. Sci. Total Environ. 2020, 745, 141018. [Google Scholar] [CrossRef]
- Li, Y.F.; Xu, J.K.; Chen, Y.W.; Ding, W.Y.; Shao, A.Q.; Liang, X.; Zhu, Y.T.; Yang, J.L. Characterization of Gut Microbiome in the Mussel Mytilus galloprovincialis in Response to Thermal Stress. Front. Physiol. 2019, 10, 1086. [Google Scholar] [CrossRef] [Green Version]
- Iori, S.; Rovere, G.D.; Ezzat, L.; Smits, M.; Ferraresso, S.S.; Babbucci, M.; Marin, M.G.; Masiero, L.; Fabrello, J.; Garro, E.; et al. The effects of glyphosate and AMPA on the mediterranean mussel Mytilus galloprovincialis and its microbiota. Environ. Res. 2020, 182, 108984. [Google Scholar] [CrossRef]
- Saco, A.; Rey-Campos, M.; Novoa, B.; Figueras, A. Transcriptomic Response of Mussel Gills After a Vibrio splendidus Infection Demonstrates Their Role in the Immune Response. Front. Immunol. 2020, 11, 615580. [Google Scholar] [CrossRef]
- Rey-Campos, M.; Moreira, R.; Gerdol, M.; Pallavicini, A.; Novoa, B.; Figueras, A. Immune Tolerance in Mytilus galloprovincialis Hemocytes After Repeated Contact with Vibrio splendidus. Front. Immunol. 2019, 10, 1894. [Google Scholar] [CrossRef] [Green Version]
- Pauletto, M.; Milan, M.; Moreira, R.; Novoa, B.; Figueras, A.; Babbucci, M.; Patarnello, T.; Bargelloni, L. Deep transcriptome sequencing of Pecten maximus hemocytes: A genomic resource for bivalve immunology. Fish Shellfish Immunol. 2014, 37, 154–165. [Google Scholar] [CrossRef]
- Zhang, X.; Shi, J.; Sun, Y.; Habib, Y.J.; Yang, H.; Zhang, Z.; Wang, Y. Integrative transcriptome analysis and discovery of genes involving in immune response of hypoxia/thermal challenges in the small abalone Haliotis diversicolor. Fish Shellfish Immunol. 2019, 84, 609–626. [Google Scholar] [CrossRef]
- Castellanos-Martínez, S.; Arteta, D.; Catarino, S.; Gestal, C. De novo transcriptome sequencing of the Octopus vulgaris hemocytes using Illumina RNA-Seq technology: Response to the infection by the gastrointestinal parasite Aggregata octopiana. PLoS ONE 2014, 9, e107873. [Google Scholar] [CrossRef]
- Li, Y.F.; Chen, Y.W.; Xu, J.K.; Ding, W.Y.; Shao, A.Q.; Zhu, Y.T.; Wang, C.; Liang, X.; Yang, J.L. Temperature elevation and Vibrio cyclitrophicus infection reduce the diversity of haemolymph microbiome of the mussel Mytilus Coruscus. Sci. Rep. 2019, 9, 16391. [Google Scholar] [CrossRef] [Green Version]
- Unzueta-Martínez, A.; Scanes, E.; Parker, L.M.; Ross, P.M.; O’Connor, W.; Bowen, J.L. Microbiomes of the Sydney Rock Oyster are acquired through both vertical and horizontal transmission. Anim. Microbiome 2022, 4, 32. [Google Scholar] [CrossRef]
- Razzauti, M.; Galan, M.; Bernard, M.; Maman, S.; Klopp, C.; Charbonnel, N.; Vayssier-Taussat, M.; Eloit, M.; Cosson, J.F. A Comparison between Transcriptome Sequencing and 16S Metagenomics for Detection of Bacterial Pathogens in Wildlife. PLoS Negl. Trop. Dis. 2015, 9, e0003929. [Google Scholar] [CrossRef] [Green Version]
- Durazzi, F.; Sala, C.; Castellani, G.; Manfreda, G.; Remondini, D.; De Cesare, A. Comparison between 16S rRNA and shotgun sequencing data for the taxonomic characterization of the gut microbiota. Sci. Rep. 2021, 11, 3030. [Google Scholar] [CrossRef]
- Musella, M.; Wathsala, R.; Tavella, T.; Rampelli, S.; Barone, M.; Palladino, G.; Biagi, E.; Brigidi, P.; Turroni, S.; Franzellitti, S.; et al. Tissue-scale microbiota of the Mediterranean mussel (Mytilus galloprovincialis) and its relationship with the environment. Sci. Total Environ. 2020, 717, 137209. [Google Scholar] [CrossRef]
- Wang, Z.; Lu, X.; Liang, Y.; Zheng, Z. A Marteilia-like parasite in blue mussels Mytilus edulis in China. J. Aquat. Anim. Health 2012, 24, 161–164. [Google Scholar] [CrossRef]
- Kerr, R.; Ward, G.M.; Stentiford, G.D.; Alfjorden, A.; Mortensen, S.; Bignell, J.P.; Feist, S.W.; Villalba, A.; Carballal, M.J.; Cao, A.; et al. Marteilia refringens and Marteilia pararefringens sp. nov. are distinct parasites of bivalves and have different European distributions. Parasitology 2018, 145, 1483–1492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peeler, E.J.; Reese, R.A.; Cheslett, D.L.; Geoghegan, F.; Power, A.; Thrush, M.A. Investigation of mortality in Pacific oysters associated with Ostreid herpesvirus-1 μVar in the Republic of Ireland in 2009. Prev. Vet. Med. 2012, 105, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Jouaux, A.; Lafont, M.; Blin, J.L.; Houssin, M.; Mathieu, M.; Lelong, C. Physiological change under OsHV-1 contamination in Pacific oyster Crassostrea gigas through massive mortality events on fields. BMC Genom. 2013, 14, 590. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burioli, E.; Prearo, M.; Houssin, M. Complete genome sequence of Ostreid herpesvirus type 1 µVar isolated during mortality events in the Pacific oyster Crassostrea gigas in France and Ireland. Virology 2017, 509, 239–251. [Google Scholar] [CrossRef] [PubMed]
- Walker, P.J.; Winton, J.R. Emerging viral diseases of fish and shrimp. Vet. Res. 2010, 41, 51. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Crane, M.; Hyatt, A. Viruses of fish: An overview of significant pathogens. Viruses 2011, 3, 2025–2046. [Google Scholar] [CrossRef] [Green Version]
- Mugimba, K.K.; Byarugaba, D.K.; Mutoloki, S.; Evensen, Ø.; Munang’andu, H.M. Challenges and Solutions to Viral Diseases of Finfish in Marine Aquaculture. Pathogens 2021, 10, 673. [Google Scholar] [CrossRef]
- Rosani, U. Tracing RNA viruses associated with Nudibranchia gastropods. PeerJ 2022, 10, e13410. [Google Scholar] [CrossRef]
- Rey-Campos, M.; González, L.D.; Novoa, B.; Figueras, A. Metatranscriptomics as a Tool to Unmask Mollusca Dark Virome. 2022; in preparation. [Google Scholar]
- Bettencourt, R.; Rodrigues, M.; Barros, I.; Cerqueira, T.; Freitas, C.; Costa, V.; Pinheiro, M.; Egas, C.; Santos, R.S. Site-related differences in gene expression and bacterial densities in the mussel Bathymodiolus azoricus from the Menez Gwen and Lucky Strike deep-sea hydrothermal vent sites. Fish Shellfish Immunol. 2014, 39, 343–353. [Google Scholar] [CrossRef]
- Li, Y.F.; Yang, N.; Liang, X.; Yoshida, A.; Osatomi, K.; Power, D.; Batista, F.M.; Yang, J.L. Elevated Seawater Temperatures Decrease Microbial Diversity in the Gut of Mytilus coruscus. Front. Physiol. 2018, 9, 839. [Google Scholar] [CrossRef]
- Richards, A.L.; Muehlbauer, A.L.; Alazizi, A.; Burns, M.B.; Findley, A.; Messina, F.; Gould, T.J.; Cascardo, C.; Pique-Regi, R.; Blekhman, R.; et al. Gut Microbiota Has a Widespread and Modifiable Effect on Host Gene Regulation. mSystems 2019, 4, e00323-18. [Google Scholar] [CrossRef] [Green Version]
- Grieneisen, L.; Muehlbauer, A.L.; Blekhman, R. Microbial control of host gene regulation and the evolution of host-microbiome interactions in primates. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2020, 375, 20190598. [Google Scholar] [CrossRef]
- Sielemann, K.; Hafner, A.; Puker, B. The reuse of public datasets in the life sciences: Potential risks and rewards. PeerJ 2020, 8, e9954. [Google Scholar] [CrossRef]
- Roche, D.G.; Kruuk, L.E.; Lanfear, R.; Binning, S.A. Public Data Archiving in Ecology and Evolution: How Well Are We Doing? PLoS Biol. 2015, 13, e1002295. [Google Scholar] [CrossRef] [Green Version]
- Lasa, A.; di Cesare, A.; Tassistro, G.; Borello, A.; Gualdi, S.; Furones, D.; Carrasco, N.; Cheslett, D.; Brechon, A.; Paillard, C.; et al. Dynamics of the Pacific oyster pathobiota during mortality episodes in Europe assessed by 16S rRNA gene profiling and a new target enrichment next-generation sequencing strategy. Environ. Microbiol. 2019, 21, 4548–4562. [Google Scholar] [CrossRef] [Green Version]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids. Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Yilmaz, P.; Parfrey, L.W.; Yarza, P.; Gerken, J.; Pruesse, E.; Quast, C.; Schweer, T.; Peplies, J.; Ludwig, W.; Glöckner, F.O. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res. 2014, 42, D643–D648. [Google Scholar] [CrossRef] [Green Version]
- Gerdol, M.; Moreira, R.; Cruz, F.; Gómez-Garrido, J.; Vlasova, A.; Rosani, U.; Venier, P.; Naranjo-Ortiz, M.A.; Murgarella, M.; Greco, S.; et al. Massive gene presence-absence variation shapes an open pan-genome in the Mediterranean mussel. Genome Biol. 2020, 21, 275. [Google Scholar] [CrossRef]
- Kenny, N.J.; McCarthy, S.A.; Dudchenko, O.; James, K.; Betteridge, E.; Corton, C.; Dolucan, J.; Mead, D.; Oliver, K.; Omer, A.D.; et al. The gene-rich genome of the scallop Pecten maximus. GigaScience 2020, 9, giaa037. [Google Scholar] [CrossRef]
- Botwright, N.A.; Zhao, M.; Wang, T.; McWilliam, S.; Colgrave, M.L.; Hlinka, O.; Li, S.; Suwansa-Ard, S.; Subramanian, S.; McPherson, L.; et al. Greenlip Abalone (Haliotis laevigata) Genome and Protein Analysis Provides Insights into Maturation and Spawning. G3 (Bethesda) 2019, 9, 3067–3078. [Google Scholar] [CrossRef]
- Zarrella, I.; Herten, K.; Maes, G.E.; Tai, S.; Yang, M.; Seuntjens, E.; Ritschard, E.A.; Zach, M.; Styfhals, R.; Sanges, R.; et al. The survey and reference assisted assembly of the Octopus vulgaris genome. Sci. Data 2019, 6, 13. [Google Scholar] [CrossRef]






| Project | Species | Tissue | Sample Features | Reference |
|---|---|---|---|---|
| PRJNA858246 PRJNA858248 | Mytilus galloprovincialis | Whole body | 16S rRNA sequencing Ría de Vigo | Current work |
| PRJNA638821 | Mytilus galloprovincialis | Gills | Mussel gills after a V. splendidus infection by bath | [20] |
| PRJNA466718 | Mytilus galloprovincialis | Haemocytes | Mussel haemocytes after a V. splendidus injection in the adductor muscle | [21] |
| PRJNA222492 | Pecten maximus | Haemocytes | Control from Vigo | [22] |
| PRJNA481417 | Haliotis diversicolor | Haemocytes | Control from China | [23] |
| PRJNA253995 | Octopus vulgaris | Haemocytes | Control from Vigo | [24] |
| Gillbath | |||
| ID | Pearson | Uniprot Annotation | Function |
| C_8095 | −0.993 | Regulator of MON1-CCZ1 complex | Endosomal/autophagic flux |
| C_35153 | −0.991 | Beta-galactosidase-1-like protein | Probable glycosyl hydrolase |
| C_53367 | −0.991 | Zinc finger protein 233 | Transcriptional regulation |
| C_83652 | −0.990 | tRNA-splicing endonuclease subunit Sen54 | Responsible for identification and cleavage of the splice sites in pre-tRNA |
| C_51191 | 0.989 | AP-5 complex subunit mu-1 | May be involved in endosomal transport |
| C_36214 | 0.989 | NA- and Cl-dependent neutral and basic amino acid transporter B(0+) | Uptake of a broad range of amino acids in a Na+/Cl--dependent manner |
| C_35181 | 0.989 | Pre-mRNA-splicing factor ISY1 homolog | Processing of microRNAs during embryonic stem cell differentiation |
| C_55942 | −0.987 | Protein mab-21-like 3 | Neural development |
| C_63893 | −0.986 | E3 ubiquitin-protein ligase TRIM56 | Role in innate antiviral immunity |
| C_28519 | −0.985 | Metabotropic glutamate receptor 3 | Signalling via G proteins |
| C_32331 | −0.985 | 15-hydroxyprostaglandin dehydrogenase | Conversion of hydroxylated arachidonic acid to oxidised metabolites |
| C_111951 | 0.985 | Retrovirus-related Pol polyprotein from type-2 | - |
| C_94710 | −0.985 | Chromatin assembly factor 1 subunit A | Chromatin assembly in DNA replication and DNA repair |
| C_92181 | −0.985 | Embryonic stem cell-specific 5-hydroxymethylcytosine-binding protein | - |
| C_17718 | −0.984 | Ankyrin-1 | Attaches integral membrane proteins to cytoskeletal elements |
| C_21601 | −0.984 | Sialoadhesin | Acts as an endocytic receptor-mediating clathrin-dependent endocytosis |
| C_9272 | −0.983 | Neuronal acetylcholine receptor subunit alpha-10 | Ionotropic receptor with a role in the modulation of auditory stimuli |
| C_36504 | −0.983 | Innexin unc-9 | Plays a role in maintaining gap junction activity to promote locomotion |
| C_48464 | 0.983 | Neurensin-1 | Neural organelle transport |
| C_22050 | −0.983 | Cytochrome P450 10 | Functions as monooxygenases |
| Haemocytes injection | |||
| ID | Pearson | Uniprot Annotation | Function |
| C_7894 | 0.997 | GTPase IMAP family member 4 | Regulation of apoptosis |
| C_15253 | −0.997 | Nuclear pore complex protein Nup54 | Required for the trafficking across the nuclear membrane |
| C_13141 | −0.997 | Bardet–Biedl syndrome 7 protein homolog | Brain development and ciliary trafficking |
| C_58173 | −0.996 | Long-chain fatty acid transport protein 3 | Long-chain fatty acids translocation at the plasma membrane |
| C_15154 | −0.996 | tRNA (uracil-5-)-methyltransferase homolog A | May be involved in nucleic acid metabolism and/or modifications |
| C_49199 | −0.995 | Choline dehydrogenase, mitochondrial | Dehydrogenation of choline to betaine aldehyde in mitochondria |
| C_76285 | −0.995 | Kelch-like protein 25 | Homeostatic mechanism, translational regulation |
| C_30541 | −0.994 | 4-aminobutyrate aminotransferase, mitochondrial | Catalyses neurotransmitter conversions |
| C_6950 | −0.994 | C-Myc-binding protein | Control transcriptional activity of MYC, involved in cell cycle and apoptosis |
| C_98220 | −0.994 | UNC93-like protein | Involved in innate and adaptive immune response by regulating TLR signalling |
| C_52106 | −0.991 | Programmed cell death protein 2 | May play an important role in cell death and/or in regulation of cell proliferation |
| C_7630 | −0.990 | GPI inositol-deacylase | Important for efficient transport of GPI-anchored proteins from the ER to the Golgi |
| C_37669 | 0.990 | Torsin-1A-interacting protein 2 | Required for endoplasmic reticulum integrity |
| C_79722 | 0.990 | Collagen alpha-4(VI) chain | Cell-binding protein |
| C_18830 | −0.990 | Copper chaperone for superoxide dismutase | Metalloprotein responsible for the delivery of Cu to SOD1. Antioxidant |
| C_120023 | −0.990 | Zinc finger protein 45 | May be involved in transcriptional regulation |
| C_67714 | −0.990 | Leucine-rich repeat-containing protein 63 | Associated with innate immunity. Specially pathogen recognition |
| C_5776 | 0.990 | Rab9 effector protein with Kelch motifs | Rab9 effector required for endosome to trans-Golgi network (TGN) transport |
| C_69835 | −0.989 | TATA box-binding protein-associated factor RNA polymerase I subunit C | Involved in the transcription initiation |
| C_482 | −0.989 | Ribonuclease Oy | Releases mononucleotides from RNA in the order of 3’-GMP, 3’-AMP and 3’-UMP |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rey-Campos, M.; Ríos-Castro, R.; Gallardo-Escárate, C.; Novoa, B.; Figueras, A. Exploring the Potential of Metatranscriptomics to Describe Microbial Communities and Their Effects in Molluscs. Int. J. Mol. Sci. 2022, 23, 16029. https://doi.org/10.3390/ijms232416029
Rey-Campos M, Ríos-Castro R, Gallardo-Escárate C, Novoa B, Figueras A. Exploring the Potential of Metatranscriptomics to Describe Microbial Communities and Their Effects in Molluscs. International Journal of Molecular Sciences. 2022; 23(24):16029. https://doi.org/10.3390/ijms232416029
Chicago/Turabian StyleRey-Campos, Magalí, Raquel Ríos-Castro, Cristian Gallardo-Escárate, Beatriz Novoa, and Antonio Figueras. 2022. "Exploring the Potential of Metatranscriptomics to Describe Microbial Communities and Their Effects in Molluscs" International Journal of Molecular Sciences 23, no. 24: 16029. https://doi.org/10.3390/ijms232416029





