Characterization of a Vibriophage Infecting Pathogenic Vibrio harveyi
Abstract
1. Introduction
2. Results
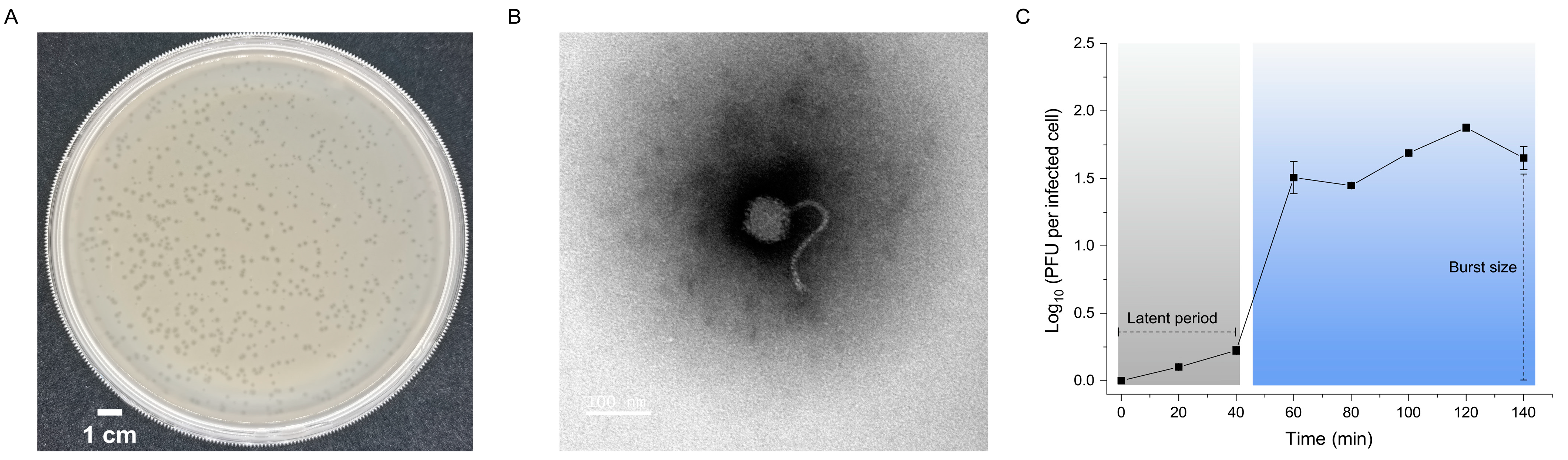
2.1. Biological Characterization of R21Y
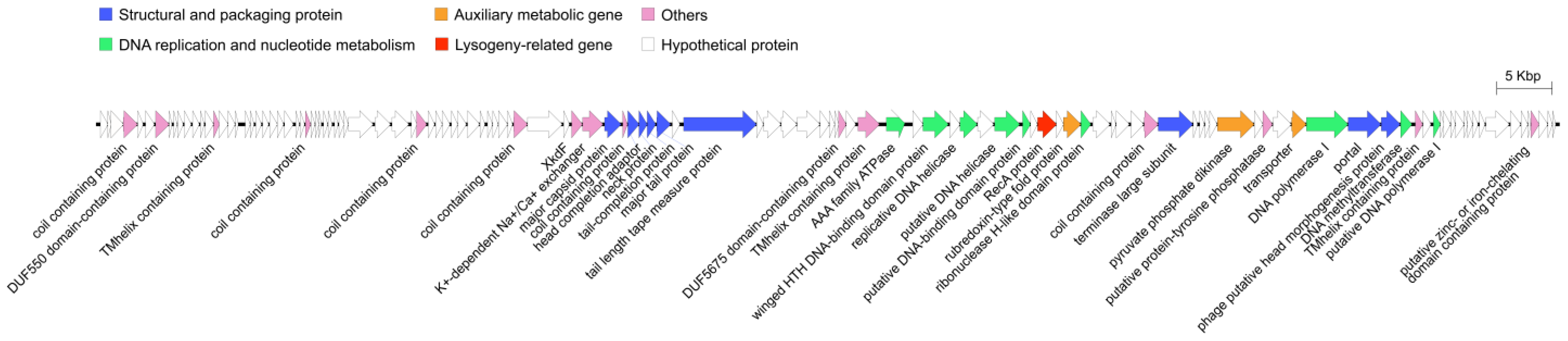
2.2. Genomic Features of R21Y
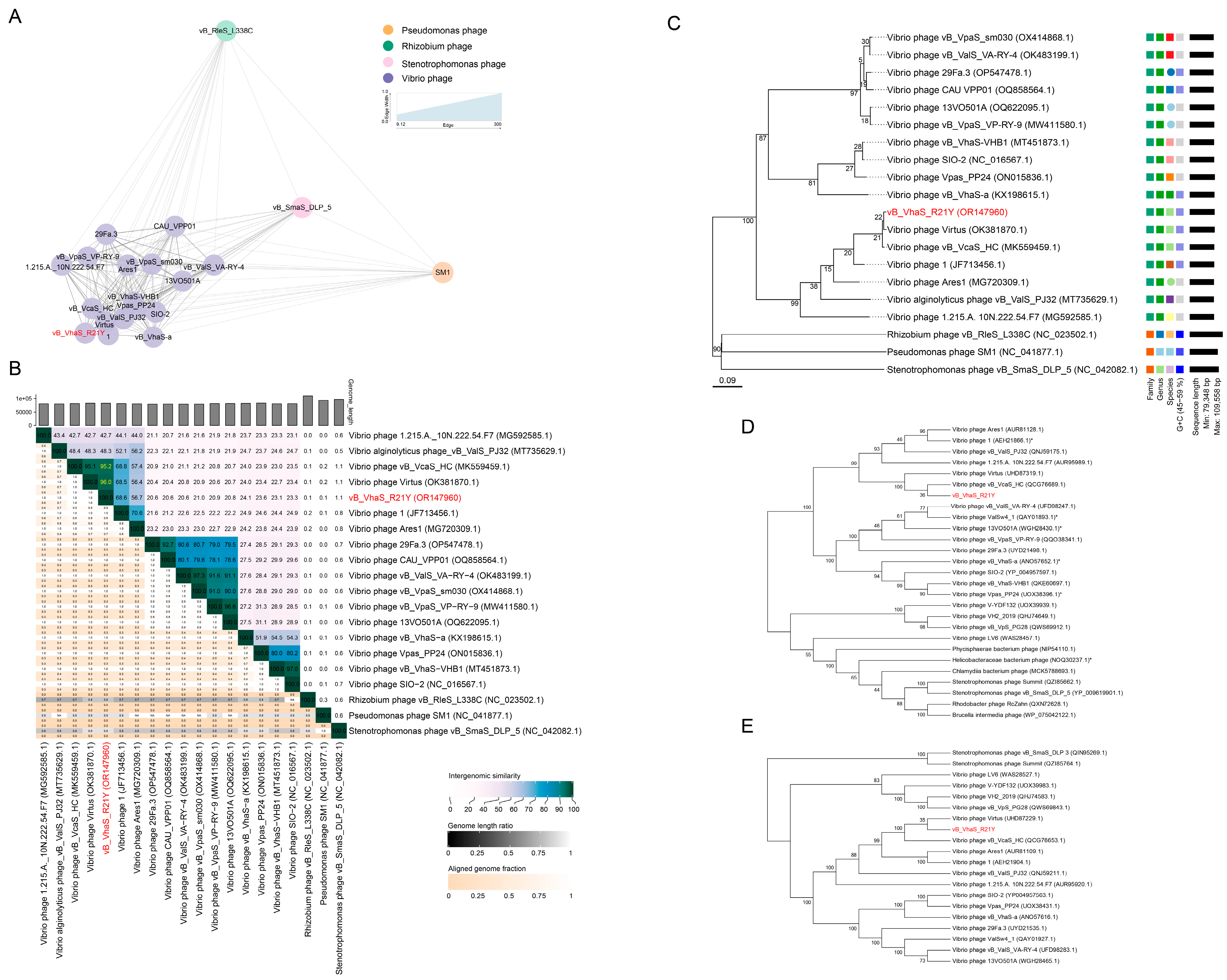
2.3. Taxonomy and Phylogeny of R21Y
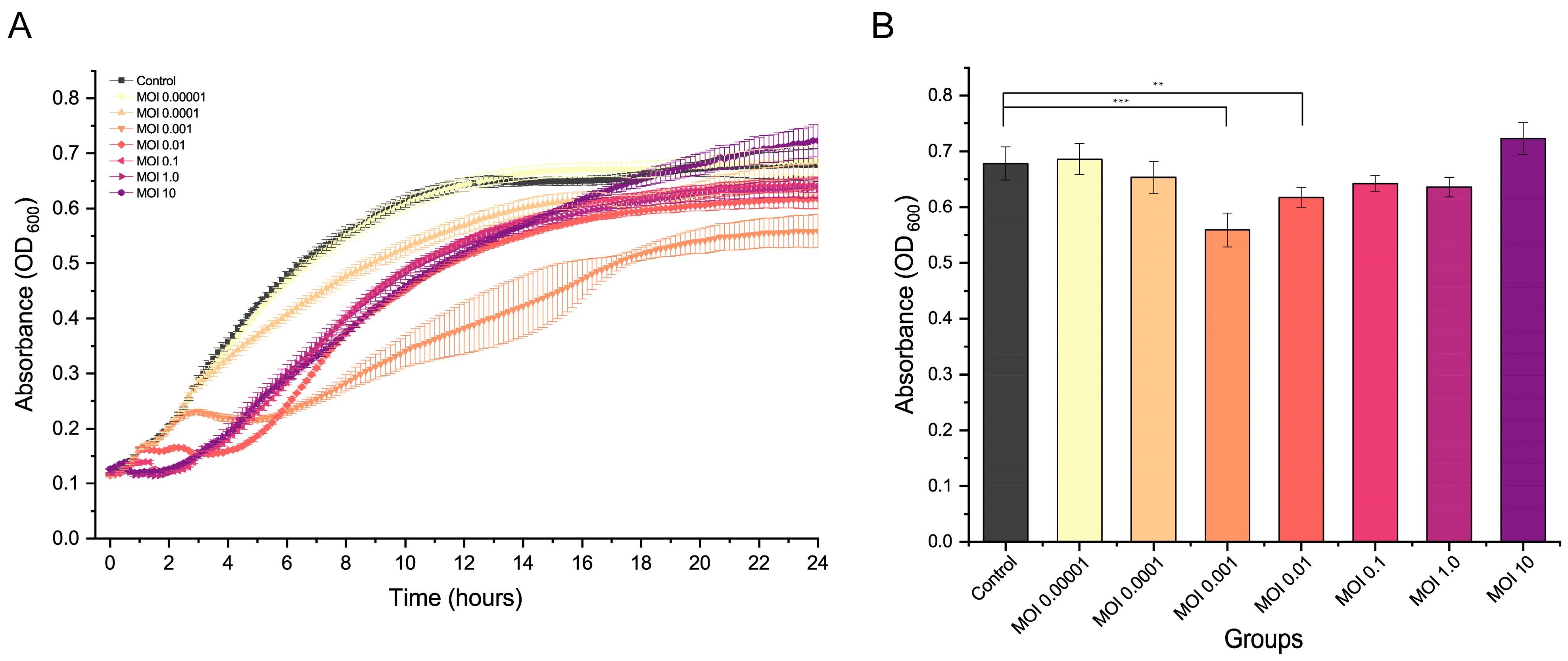
2.4. Lytic Ability of R21Y
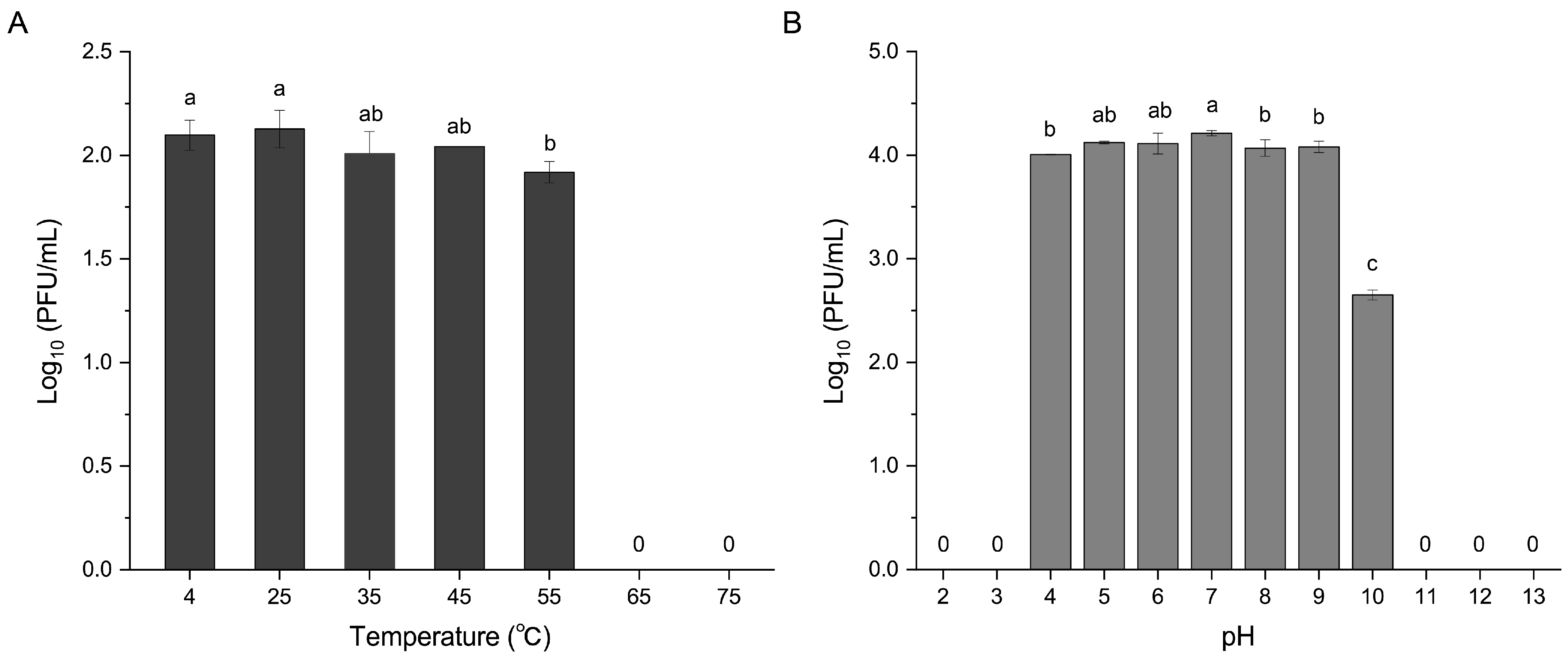
2.5. Thermal and pH Stability of R21Y
3. Discussion
4. Materials and Methods
4.1. Phage Isolation and Purification
4.2. Preparation of High-Titer Phage Suspensions
4.3. Transmission Electron Microscopy
4.4. One-Step Growth Curve
4.5. Host Range
4.6. Lysogeny Test
4.7. Lytic Efficacy
4.8. Thermal and pH Stability
4.9. DNA Extraction
4.10. Genome Sequencing and Annotation
4.11. Taxonomic Network and Phylogenetic Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Johnson, C.N. Fitness factors in vibrios: A mini-review. Microb. Ecol. 2013, 65, 826–851. [Google Scholar] [CrossRef]
- Akolkar, J.K.; Matson, J.S. Stress responses in pathogenic Vibrios and their role in host and environmental survival. In Vibrio spp. Infections; Almagro-Moreno, S., Pukatzki, S., Eds.; Springer International Publishing: Cham, Germany, 2023; pp. 213–232. [Google Scholar]
- Cardinaud, M.; Barbou, A.; Capitaine, C.; Bidault, A.; Dujon, A.M.; Moraga, D.; Paillard, C. Vibrio harveyi adheres to and penetrates tissues of the European abalone Haliotis tuberculata within the first hours of contact. Appl. Environ. Microbiol. 2014, 80, 6328–6333. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.H.; He, X.; Austin, B. Vibrio harveyi: A serious pathogen of fish and invertebrates in mariculture. Mar. Life Sci. Technol. 2020, 2, 231–245. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Xu, L.; Liu, S.; Wang, Q.; Guo, Z.; Chen, C.; Feng, J. What drives changes in the virulence and antibiotic resistance of Vibrio harveyi in the South China Sea? J. Fish Dis. 2020, 43, 853–862. [Google Scholar] [CrossRef] [PubMed]
- Cabello, F.C. Heavy use of prophylactic antibiotics in aquaculture: A growing problem for human and animal health and for the environment. Environ. Microbiol. 2006, 8, 1137–1144. [Google Scholar] [CrossRef]
- Defoirdt, T.; Boon, N.; Sorgeloos, P.; Verstraete, W.; Bossier, P. Alternatives to antibiotics to control bacterial infections: Luminescent vibriosis in aquaculture as an example. Trends Biotechnol. 2007, 25, 472–479. [Google Scholar] [CrossRef]
- Defoirdt, T.; Sorgeloos, P.; Bossier, P. Alternatives to antibiotics for the control of bacterial disease in aquaculture. Curr. Opin. Microbiol. 2011, 14, 251–258. [Google Scholar] [CrossRef]
- Golkar, Z.; Bagasra, O.; Pace, D.G. Bacteriophage therapy: A potential solution for the antibiotic resistance crisis. J. Infect. Dev. Countries 2014, 8, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, C.; Sarkar, P.; Issa, R.; Haldar, J. Alternatives to conventional antibiotics in the era of antimicrobial resistance. Trends Microbiol. 2019, 27, 323–338. [Google Scholar] [CrossRef] [PubMed]
- Dion, M.B.; Oechslin, F.; Moineau, S. Phage diversity, genomics and phylogeny. Nat. Rev. Microbiol. 2020, 18, 125–138. [Google Scholar] [CrossRef]
- Correa, A.M.S.; Howard-Varona, C.; Coy, S.R.; Buchan, A.; Sullivan, M.B.; Weitz, J.S. Revisiting the rules of life for viruses of microorganisms. Nat. Rev. Microbiol. 2021, 19, 501–513. [Google Scholar] [CrossRef]
- Zhang, M.; Zhang, T.; Yu, M.; Chen, Y.L.; Jin, M. The life cycle transitions of temperate phages: Regulating factors and potential ecological implications. Viruses 2022, 14, 1904. [Google Scholar] [CrossRef]
- Richards, G.P. Bacteriophage remediation of bacterial pathogens in aquaculture: A review of the technology. Bacteriophage 2014, 4, e975540. [Google Scholar] [CrossRef]
- Oakey, H.J.; Owens, L. A new bacteriophage, VHML, isolated from a toxin-producing strain of Vibrio harveyi in tropical Australia. J. Appl. Microbiol. 2000, 89, 702–709. [Google Scholar] [CrossRef]
- Briers, Y. Phage lytic enzymes. Viruses 2019, 11, 113. [Google Scholar] [CrossRef] [PubMed]
- Zampara, A.; Sørensen, M.C.H.; Grimon, D.; Antenucci, F.; Vitt, A.R.; Bortolaia, V.; Briers, Y.; Brøndsted, L. Exploiting phage receptor binding proteins to enable endolysins to kill Gram-negative bacteria. Sci. Rep. 2020, 10, 12087. [Google Scholar] [CrossRef] [PubMed]
- Hendrix, R.W. Bacteriophages: Evolution of the majority. Theor. Popul. Biol. 2002, 61, 471–480. [Google Scholar] [CrossRef] [PubMed]
- Pasharawipas, T.; Thaikua, S.; Sriurairatana, S.; Ruangpan, L.; Direkbusarakum, S.; Manopvisetcharean, J.; Flegel, T.W. Partial characterization of a novel bacteriophage of Vibrio harveyi isolated from shrimp culture ponds in Thailand. Virus Res. 2005, 114, 63–69. [Google Scholar] [CrossRef]
- Patil, R.J.; Desai, S.N.; Roy, P.; Durgaiah, M.; Saravanan, R.S.; Vipra, A. Simulated hatchery system to assess bacteriophage efficacy against Vibrio harveyi. Dis. Aquat. Org. 2014, 112, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Misol, G.N., Jr.; Kokkari, C.; Katharios, P. Biological and genomic characterization of a novel jumbo bacteriophage, vB_VhaM_pir03 with broad host lytic activity against Vibrio harveyi. Pathogens 2020, 9, 1051. [Google Scholar] [CrossRef]
- Cui, H.; Cong, C.; Wang, L.; Li, X.; Li, J.; Yang, H.; Li, S.; Xu, Y. Protective effectiveness of feeding phage cocktails in controlling Vibrio harveyi infection of turbot Scophthalmus maximus. Aquaculture 2021, 535, 736390. [Google Scholar] [CrossRef]
- Luo, Z.H.; Ye, D.Z.; Yun, Y.P. Vibrio harveyi Giant Phage VP4B and Application Thereof. Patent CN103555671A, 5 February 2014. [Google Scholar]
- Du, S.Y.; Jiang, S.J.; Li, X.; Ma, R.J.; Li, X.S.; Zhao, D.D.; Luo, C.S.; Zhang, Q.; Liu, Y.Q.; Gai, C.Y.; et al. Lytic vibrio harveyi Phage RDP-VP-19012 and Application Thereof. Patent CN112063595B, 11 December 2020. [Google Scholar]
- Pan, Q.; Ren, H.Y.; Sun, H.Z.; Yan, Y.X.; Qin, M.R. Vibrio harveyi Phage vB _ KaS _ PK22, Phage Composition and Application Thereof. Patent CN111705041B, 2 July 2021. [Google Scholar]
- Pan, Q.; Ren, H.Y.; Sun, H.Z.; Yan, Y.X.; Wang, C.Y. Vibrio harveyi Phage, Phage Composition and Application Thereof. Patent CN111676197B, 14 December 2021. [Google Scholar]
- Xu, L.H.; Li, D.F.; Qin, W.N.; Tong, Y.G.; Lin, W.; Sun, Z.T. Vibrio harveyi High-Efficiency Lytic Phage vB _ Vhas-yong3 and Application Thereof. Patent CN112391355A, 23 February 2021. [Google Scholar]
- Xu, L.H.; Li, D.F.; Qin, W.N.; Tong, Y.G.; Lin, W.; Sun, Z.T.; Wang, C.L.; Mu, C.K.; Han, S.M. Vibrio harveyi High-Efficiency Lytic Phage vB-Vhas-yong1 and Application Thereof. Patent CN112442487A, 5 March 2021. [Google Scholar]
- Xu, L.H.; Li, D.F.; Qin, W.N.; Tong, Y.G.; Lin, W.; Sun, Z.T.; Wang, C.L.; Wu, C.K. Vibrio harveyi High-Efficiency Lytic Phage vB _ Vhas-yong2 and Application Thereof. Patent CN112391356A, 23 February 2021. [Google Scholar]
- Zhang, H.; Wang, Y.; Pang, M.D.; Sun, L.C. Vibrio Bacteriophage and Preparation Method and Application of Bactericidal Composition Thereof. Patent CN109207440B, 27 August 2021. [Google Scholar]
- Wei, J.G.; Kang, S.Z.; Qin, Q.W.; Zhang, L.H.; Liao, J.M.; Zhang, D.Z.; Mu, G.H.; Chen, J.; Lin, D.R.; Yang, A.B. Vibrio harveyi Phage V-YDF132 and Application Thereof. Patent CN115505576A, 23 December 2022. [Google Scholar]
- Vidgen, M.; Carson, J.; Higgins, M.; Owens, L. Changes to the phenotypic profile of Vibrio harveyi when infected with the Vibrio harveyi myovirus-like (VHML) bacteriophage. J. Appl. Microbiol. 2006, 100, 481–487. [Google Scholar] [CrossRef] [PubMed]
- Vinod, M.G.; Shivu, M.M.; Umesha, K.R.; Rajeeva, B.C.; Krohne, G.; Karunasagar, I.; Karunasagar, I. Isolation of Vibrio harveyi bacteriophage with a potential for biocontrol of luminous vibriosis in hatchery environments. Aquaculture 2006, 255, 117–124. [Google Scholar] [CrossRef]
- Karunasagar, I.; Shivu, M.M.; Girisha, S.K.; Krohne, G.; Karunasagar, I. Biocontrol of pathogens in shrimp hatcheries using bacteriophages. Aquaculture 2007, 268, 288–292. [Google Scholar] [CrossRef]
- Shivu, M.M.; Rajeeva, B.C.; Girisha, S.K.; Karunasagar, I.; Krohne, G.; Karunasagar, I. Molecular characterization of Vibrio harveyi bacteriophages isolated from aquaculture environments along the coast of India. Environ. Microbiol. 2007, 9, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Crothers-Stomps, C.; Hoj, L.; Bourne, D.G.; Hall, M.R.; Owens, L. Isolation of lytic bacteriophage against Vibrio harveyi. J. Appl. Microbiol. 2010, 108, 1744–1750. [Google Scholar] [CrossRef]
- Phumkhachorn, P.; Rattanachaikunsopon, P. Isolation and partial characterization of a bacteriophage infecting the shrimp pathogen Vibrio harveyi. Afr. J. Microbiol. Res. 2010, 4, 1794–1800. Available online: https://academicjournals.org/journal/AJMR/article-full-text-pdf/AD36E5414186 (accessed on 29 July 2023).
- Pasharawipas, T.; Manopvisetcharean, J.; Flegel, T.W. Phage treatment of Vibrio harveyi: A general concept of protection against bacterial infection. Res. J. Microbiol. 2011, 6, 560–567. Available online: https://scialert.net/abstract/?doi=jm.2011.560.567 (accessed on 29 July 2023). [CrossRef]
- Thiyagarajan, S.; Chrisolite, B.; Alavandi, S.V.; Poornima, M.; Kalaimani, N.; Santiago, T.C. Characterization of four lytic transducing bacteriophages of luminescent Vibrio harveyi isolated from shrimp (Penaeus monodon) hatcheries. FEMS Microbiol. Lett. 2011, 325, 85–91. [Google Scholar] [CrossRef][Green Version]
- Baudoux, A.C.; Hendrix, R.W.; Lander, G.C.; Bailly, X.; Podell, S.; Paillard, C.; Johnson, J.E.; Potter, C.S.; Carragher, B.; Azam, F. Genomic and functional analysis of Vibrio phage SIO-2 reveals novel insights into ecology and evolution of marine siphoviruses. Environ. Microbiol. 2012, 14, 2071–2086. [Google Scholar] [CrossRef]
- Khemayan, K.; Prachumwat, A.; Sonthayanon, B.; Intaraprasong, A.; Sriurairatana, S.; Flegel, T.W. Complete genome sequence of virulence-enhancing siphophage VHS1 from Vibrio harveyi. Appl. Environ. Microbiol. 2012, 78, 2790–2796. [Google Scholar] [CrossRef]
- Surekhamol, I.S.; Deepa, G.D.; Somnath Pai, S.; Sreelakshmi, B.; Varghese, S.; Bright Singh, I.S. Isolation and characterization of broad spectrum bacteriophages lytic to Vibrio harveyi from shrimp farms of Kerala, India. Lett. Appl. Microbiol. 2014, 58, 197–204. [Google Scholar] [CrossRef]
- Luo, Z.H.; Yu, Y.P.; Jost, G.; Xu, W.; Huang, X.L. Complete genome sequence of a giant Vibrio bacteriophage VH7D. Mar. Genomics 2015, 24 Pt 3, 293–295. [Google Scholar] [CrossRef] [PubMed]
- Lal, T.M.; Sano, M.; Ransangan, J. Isolation and characterization of large marine bacteriophage (Myoviridae), VhKM4 infecting Vibrio harveyi. J. Aquat. Anim. Health 2017, 29, 26–30. [Google Scholar] [CrossRef] [PubMed]
- Stalin, N.; Srinivasan, P. Efficacy of potential phage cocktails against Vibrio harveyi and closely related Vibrio species isolated from shrimp aquaculture environment in the south east coast of India. Vet. Microbiol. 2017, 207, 83–96. [Google Scholar] [CrossRef]
- Wang, Y.; Barton, M.; Elliott, L.; Li, X.; Abraham, S.; O’Dea, M.; Munro, J. Bacteriophage therapy for the control of Vibrio harveyi in greenlip abalone (Haliotis laevigata). Aquaculture 2017, 473, 251–258. [Google Scholar] [CrossRef]
- Wu, L.; Tian, Y.; Pang, M.; Yang, Z.; Bao, H.; Zhou, Y.; Sun, L.; Wang, R.; Zhang, H. A novel vibriophage vB_VhaS_PcB-1G capable of inhibiting virulent Vibrio harveyi pathogen. Aquaculture 2021, 542, 736854. [Google Scholar] [CrossRef]
- Droubogiannis, S.; Katharios, P. Genomic and biological profile of a novel bacteriophage, Vibrio phage Virtus, which improves survival of Sparus aurata larvae challenged with Vibrio harveyi. Pathogens 2022, 11, 630. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Zhang, L.; Liao, J.; Zhang, D.; Wu, S.; Zhang, X.; Qin, Q.; Wei, J. Isolation and characterization of a newly discovered phage, V-YDF132, for lysing Vibrio harveyi. Viruses 2022, 14, 1802. [Google Scholar] [CrossRef]
- Kar, P.; Das, T.K.; Ghosh, S.; Pradhan, S.; Chakrabarti, S.; Mondal, K.C.; Ghosh, K. Characterization of a Vibrio-infecting bacteriophage, VPMCC5, and proposal of its incorporation as a new genus in the Zobellviridae family. Virus Res. 2022, 321, 198904. [Google Scholar] [CrossRef]
- Lelin, C.; Thirumalaikumar, E.; Uma, G.; Babu, M.M.; Ajan, C.; Vimal, S.; Citarasu, T. Isolation and partial characterization of bacteriophages infecting Vibrio harveyi from shrimp farm effluent water. Aquacult. Int. 2022, 30, 2081–2094. [Google Scholar] [CrossRef]
- Benala, M.; Vaiyapuri, M.; Sivam, V.; Raveendran, K.; Mothadaka, M.P.; Badireddy, M.R. Genome characterization and infectivity potential of vibriophage-varphiLV6 with Lytic activity against luminescent Vibrios of Penaeus vannamei shrimp aquaculture. Viruses 2023, 15, 868. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Tian, Y.; Li, Z.; Yang, Y.; Ai, C.; Zhang, R. A broad-host-range lytic phage vB_VhaS-R18L as a candidate against vibriosis. Front. Microbiol. 2023, 14, 1191157. [Google Scholar] [CrossRef]
- Droubogiannis, S.; Pavlidi, L.; Skliros, D.; Flemetakis, E.; Katharios, P. Comprehensive characterization of a novel bacteriophage, vB_VhaS_MAG7 against a fish pathogenic strain of Vibrio harveyi and its in vivo efficacy in phage therapy trials. Int. J. Mol. Sci. 2023, 24, 8200. [Google Scholar] [CrossRef] [PubMed]
- Tian, F.; Li, J.; Hu, Y.; Zhao, F.; Ren, H.; Pan, Q.; Nazir, A.; Li, F.; Tong, Y. Characterization and complete genome sequence analysis of a newly isolatedphage against Vibrio parahaemolyticus from sick shrimp in Qingdao, China. PLoS ONE 2022, 17, e0266683. [Google Scholar] [CrossRef]
- Gallet, R.; Kannoly, S.; Wang, I.-N. Effects of bacteriophage traits on plaque formation. BMC Microbiol. 2011, 11, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, Z.; Zhao, J.; Wang, L.; Xie, G.; Huang, J.; Zhang, Y. A novel vibriophage vB_VcaS_HC containing lysogeny-related gene has strong lytic ability against pathogenic bacteria. Virol. Sin. 2021, 36, 281–290. [Google Scholar] [CrossRef]
- Pan, L.; Li, D.; Sun, Z.; Lin, W.; Hong, B.; Qin, W.; Xu, L.; Liu, W.; Zhou, Q.; Wang, F.; et al. First characterization of a Hafnia phage reveals extraordinarily large burst size and unusual plaque polymorphism. Front. Microbiol. 2021, 12, 754331. [Google Scholar] [CrossRef]
- Sui, B.; Qi, X.; Wang, X.; Ren, H.; Liu, W.; Zhang, C. Characterization of a novel bacteriophage swi2 harboring two lysins can naturally lyse Escherichia coli. Front. Microbiol. 2021, 12, 670799. [Google Scholar] [CrossRef]
- Chang, Y.T.; Lin, C.Y.; Chen, Y.H.; Hsueh, P.R. Update on infections caused by Stenotrophomonas maltophilia with particular attention to resistance mechanisms and therapeutic options. Front. Microbiol. 2015, 6, 893. [Google Scholar] [CrossRef]
- Taslem Mourosi, J.; Awe, A.; Guo, W.; Batra, H.; Ganesh, H.; Wu, X.; Zhu, J. Understanding bacteriophage tail fiber interaction with host surface receptor: The key “blueprint” for reprogramming phage host range. Int. J. Mol. Sci. 2022, 23, 12146. [Google Scholar] [CrossRef] [PubMed]
- Dabrowska, K. Phage therapy: What factors shape phage pharmacokinetics and bioavailability? systematic and critical review. Med. Res. Rev. 2019, 39, 2000–2025. [Google Scholar] [CrossRef] [PubMed]
- Wildschut, J.D.; Lang, R.M.; Voordouw, J.K.; Voordouw, G. Rubredoxin: Oxygen oxidoreductase enhances survival of Desulfovibrio vulgaris hildenborough under microaerophilic conditions. J. Bacteriol. 2006, 188, 6253–6260. [Google Scholar] [CrossRef] [PubMed]
- Livingstone, D.R. Oxidative stress in aquatic organisms in relation to pollution and aquaculture. Revue Médecine Vétérinaire 2003, 154, 427–430. Available online: https://api.semanticscholar.org/CorpusID:131783646 (accessed on 27 September 2023).
- Baum, L.; Nguyen, M.; Jia, Y.; Biazik, J.; Thomas, T. Characterization of a novel roseophage and the morphological and transcriptional response of the sponge symbiont Ruegeria AU67 to infection. Environ. Microbiol. 2021, 23, 2532–2549. [Google Scholar] [CrossRef]
- Al-Khannaq, M.; Lytton, J. Regulation of K+-dependent Na+/Ca2+-exchangers (NCKX). Int. J. Mol. Sci. 2022, 24, 598. [Google Scholar] [CrossRef]
- Guan, K.L.; Dixon, J.E. Bacterial and viral protein tyrosine phosphatases. Semin. Cell Biol. 1993, 4, 389–396. [Google Scholar] [CrossRef]
- Morelli, M.J.; Ten Wolde, P.R.; Allen, R.J. DNA looping provides stability and robustness to the bacteriophage lambda switch. Proc. Natl. Acad. Sci. USA 2009, 106, 8101–8106. [Google Scholar] [CrossRef]
- Clokie, M.R.J.; Kropinski, A.M. Bacteriophages; Springer: Berlin/Heidelberg, Germany, 2009; Volume 1. [Google Scholar]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Chen, Y.; Li, W.; Shi, K.; Fang, Z.; Yang, Y.; Zhang, R. Isolation and characterization of a novel phage belonging to a new genus against Vibrio parahaemolyticus. Virol. J. 2023, 20, 1–11. [Google Scholar] [CrossRef]
- Hammerl, J.A.; Gollner, C.; Al Dahouk, S.; Nockler, K.; Reetz, J.; Hertwig, S. Analysis of the first temperate broad host range brucellaphage (BiPBO1) isolated from B. inopinata. Front. Microbiol. 2016, 7, 24. [Google Scholar] [CrossRef]
- Ma, R.; Chen, X.; Li, Y.; Jiao, N.; Zhang, R. Diversity, evolution and life strategies of CbK-like phages. Environ. Microbiol. 2023, 25, 1250–1264. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.G.; Jun, J.W.; Giri, S.S.; Yun, S.; Kim, H.J.; Kim, S.W.; Kang, J.W.; Han, S.J.; Jeong, D.; Park, S.C. Isolation and characterisation of pVa-21, a giant bacteriophage with anti-biofilm potential against Vibrio alginolyticus. Sci. Rep. 2019, 9, 6284. [Google Scholar] [CrossRef] [PubMed]
- Ma, R.; Lai, J.; Chen, X.; Wang, L.; Yang, Y.; Wei, S.; Jiao, N.; Zhang, R.; Ellermeier, C.D. A novel phage infecting Alteromonas represents a distinct group of siphophages infecting diverse aquatic copiotrophs. mSphere 2021, 6, e00454-21. [Google Scholar] [CrossRef]
- Zerbino, D.R.; Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 18, 821–829. [Google Scholar] [CrossRef] [PubMed]
- Garneau, J.R.; Depardieu, F.; Fortier, L.-C.; Bikard, D.; Monot, M. PhageTerm: A tool for fast and accurate determination of phage termini and packaging mechanism using next-generation sequencing data. Sci. Rep. 2017, 7, 8292. [Google Scholar] [CrossRef]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef]
- Chan, P.P.; Lin, B.Y.; Mak, A.J.; Lowe, T.M. tRNAscan-SE 2.0: Improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 2021, 49, 9077–9096. [Google Scholar] [CrossRef]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Bryant, S.H. CD-Search: Protein domain annotations on the fly. Nucleic Acids Res. 2004, 32, W327-31. [Google Scholar] [CrossRef] [PubMed]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Zheng, D.; Zhou, S.; Chen, L.; Yang, J. VFDB 2022: A general classification scheme for bacterial virulence factors. Nucleic Acids Res. 2022, 50, D912–D917. [Google Scholar] [CrossRef]
- Alcock, B.P.; Huynh, W.; Chalil, R.; Smith, K.W.; Raphenya, A.R.; Wlodarski, M.A.; Edalatmand, A.; Petkau, A.; Syed, S.A.; Tsang, K.K.; et al. CARD 2023: Expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2023, 51, D690–D699. [Google Scholar] [CrossRef]
- Bin Jang, H.; Bolduc, B.; Zablocki, O.; Kuhn, J.H.; Roux, S.; Adriaenssens, E.M.; Brister, J.R.; Kropinski, A.M.; Krupovic, M.; Lavigne, R.; et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 2019, 37, 632–639. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Moraru, C.; Varsani, A.; Kropinski, A.M. VIRIDIC—A novel tool to calculate the intergenomic similarities of prokaryote-infecting viruses. Viruses 2020, 12, 1268. [Google Scholar] [CrossRef] [PubMed]
- Meier-Kolthoff, J.P.; Göker, M.; Kelso, J. VICTOR: Genome-based phylogeny and classification of prokaryotic viruses. Bioinformatics 2017, 33, 3396–3404. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
| Species | Strain | Strain Type | Infectivity |
|---|---|---|---|
| V. harveyi | BVH1 | Pathogenic | + |
| 20150916-2 | Pathogenic | − | |
| Chan01 | Nonpathogenic | − | |
| CGMCC 1.1593 T | Nonpathogenic | − | |
| LMG 4044 T | Nonpathogenic | − | |
| V. alginolyticus | ATCC 17749 T | Pathogenic | − |
| HJ-1 | Nonpathogenic | − | |
| JL2674 | Nonpathogenic | − | |
| ZWCM4003 | Nonpathogenic | − | |
| V. azureus | JL1214 | Nonpathogenic | − |
| JL3560 | Nonpathogenic | − | |
| JL3577 | Nonpathogenic | − | |
| V. campbellii | HJ-2 | Nonpathogenic | − |
| JL2671 | Nonpathogenic | − | |
| JL3506 | Nonpathogenic | − | |
| JL3507 | Nonpathogenic | − | |
| V. chagasic | JL3518 | Nonpathogenic | − |
| V. diazotrophicus | JCM 21185 | Pathogenic | − |
| V. fortis | JL3515 | Nonpathogenic | − |
| JL3565 | Nonpathogenic | − | |
| JL3678 | Nonpathogenic | − | |
| V. hyugaensis | JL3710 | Nonpathogenic | − |
| V. inhibens | JL3707 | Nonpathogenic | − |
| V. mytili | JL3548 | Nonpathogenic | − |
| V. natriegens | WPAGA4 | Nonpathogenic | − |
| V. neocaledonicus | JL3539 | Nonpathogenic | − |
| JL3558 | Nonpathogenic | − | |
| JL3563 | Nonpathogenic | − | |
| JL3703 | Nonpathogenic | − | |
| V. owens | JL2663 | Nonpathogenic | − |
| JL3650 | Nonpathogenic | − | |
| JL2918 | Nonpathogenic | − | |
| V. parahemolyticus | Chan02 | Nonpathogenic | − |
| ATCC 17802 T | Pathogenic | − | |
| V. plantisponsor | DSM 21026 | Pathogenic | − |
| V. ponticus | JL4287 | Nonpathogenic | − |
| V. rotiferianus | JL3557 | Nonpathogenic | − |
| V. tubiashii | JL3833 | Nonpathogenic | − |
| V. variabilis | JL3468 | Nonpathogenic | − |
| V. xiii | JL2919 | Nonpathogenic | − |
| V. ziniensis | ZWAL4003 | Nonpathogenic | − |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Yun, H.; Chen, R.; Jiao, N.; Zheng, Q.; Yang, Y.; Zhang, R. Characterization of a Vibriophage Infecting Pathogenic Vibrio harveyi. Int. J. Mol. Sci. 2023, 24, 16202. https://doi.org/10.3390/ijms242216202
Li Y, Yun H, Chen R, Jiao N, Zheng Q, Yang Y, Zhang R. Characterization of a Vibriophage Infecting Pathogenic Vibrio harveyi. International Journal of Molecular Sciences. 2023; 24(22):16202. https://doi.org/10.3390/ijms242216202
Chicago/Turabian StyleLi, Yingying, Huayi Yun, Ruo Chen, Nianzhi Jiao, Qiang Zheng, Yunlan Yang, and Rui Zhang. 2023. "Characterization of a Vibriophage Infecting Pathogenic Vibrio harveyi" International Journal of Molecular Sciences 24, no. 22: 16202. https://doi.org/10.3390/ijms242216202
APA StyleLi, Y., Yun, H., Chen, R., Jiao, N., Zheng, Q., Yang, Y., & Zhang, R. (2023). Characterization of a Vibriophage Infecting Pathogenic Vibrio harveyi. International Journal of Molecular Sciences, 24(22), 16202. https://doi.org/10.3390/ijms242216202







