Abstract
The flavin monooxygenase (FMO) enzyme was discovered in mammalian liver cells that convert a carcinogenic compound, N-N′-dimethylaniline, into a non-carcinogenic compound, N-oxide. Since then, many FMOs have been reported in animal systems for their primary role in the detoxification of xenobiotic compounds. In plants, this family has diverged to perform varied functions like pathogen defense, auxin biosynthesis, and S-oxygenation of compounds. Only a few members of this family, primarily those involved in auxin biosynthesis, have been functionally characterized in plant species. Thus, the present study aims to identify all the members of the FMO family in 10 different wild and cultivated Oryza species. Genome-wide analysis of the FMO family in different Oryza species reveals that each species has multiple FMO members in its genome and that this family is conserved throughout evolution. Taking clues from its role in pathogen defense and its possible function in ROS scavenging, we have also assessed the involvement of this family in abiotic stresses. A detailed in silico expression analysis of the FMO family in Oryza sativa subsp. japonica revealed that only a subset of genes responds to different abiotic stresses. This is supported by the experimental validation of a few selected genes using qRT-PCR in stress-sensitive Oryza sativa subsp. indica and stress-sensitive wild rice Oryza nivara. The identification and comprehensive in silico analysis of FMO genes from different Oryza species carried out in this study will serve as the foundation for further structural and functional studies of FMO genes in rice as well as other crop types.
1. Introduction
The discovery of flavin monooxygenases (FMOs) traces back to the 1960s in mammalian hepatic microsomes, where it was needed to convert N-N′-dimethylaniline into N-oxide [1]). N-N′-dimethylaniline is a tertiary amine compound that has potential carcinogenic activity. Therefore, it requires detoxification by the liver cell [2]. FMOs belong to one of the protein families needed for detoxification of the carcinogens in the cell [3,4,5,6,7,8].
FMOs are found in all domains of life and metabolize several xenobiotic compounds like toxins, several drugs, and pesticides. They need cofactors flavin adenine dinucleotide (FAD), nicotinamide adenine dinucleotide phosphate (NADPH), and dioxygen for their activity [7,9,10,11,12,13]. FMO catalyzes the transfer of hydroxyl groups to small, nucleophilic substrates with heteroatoms like sulfur, nitrogen, iodine, and selenium, thus making them polar and therefore easily excretable from the cell [7,14]. Yeast has a single FMO [15], and animals have five FMOs [16,17,18,19]. The first plant FMO, YUCCA, was reported in Arabidopsis in 2001, after almost 40 years of its discovery [20]. YUCCA catalyzes the rate-limiting step in auxin biosynthesis. Previous reports have shown that plants harbour multiple FMO proteins, for example, Arabidopsis has 29 putative FMOs, and cultivated rice, Oryza sativa subsp. Japonica, has around 20 putative FMOs, thereby indicating that this family might have evolved in plants to perform diverse functions [7,14]
Evolutionary analysis of plant and animal FMOs revealed plant FMOs to belong to three major clades—clade I, clade II and clade III—based on their homology with animal FMOs [21]. Arabidopsis, Poplar, and rice FMOs were then assigned to these clades based on their putative functions like pathogen defence, auxin biosynthesis, and S-oxygenation [7,14,22,23,24,25,26,27]. Several studies have been carried out to identify and/or characterize the FMO genes belonging to the auxin biosynthetic clade (named YUCCAs) in plants [26,27,28,29,30,31,32,33,34,35,36,37,38]. However, only very few reports are available for the other two clades [21,39,40,41,42,43,44]. In addition to the previously reported roles, studies indicate the role of FMOs in abiotic stresses as well. YUCCA6 overexpression was found to improve drought tolerance and delayed leaf senescence in Arabidopsis plants [45,46]. Furthermore, some wheat FMOs (TaYUCCA1-A, TaYUCCA2-B, TaYUCCA3-A, TaYUCCA10.1, TaYUCCA8-B and TaYUCCA10.2) were involved in heat and drought stress [44]. Despite playing an important role in hormone metabolism, pathogen resistance, signalling, and chemical defence, the identification of FMOs and the novel reactions that they perform, as well as further knowledge of FMO functionality, has remained relatively limited over the last decade and is restricted to a small number of plant species.
Crop wild relatives (CWRs) have the potential to provide a useful pool of genetic material to plant breeders that can help in crop improvement. During the first half of the 20th century, gene pools from wild relatives were used for improvements in sugarcane by introgression. This produced a few disease-resistant “wonder canes” with an increase in cane and sugar yields, improved ratooning ability, and adaptability to various abiotic stresses [47,48,49,50]. In the 1940s and 1950s, CWRs were used in breeding programs for major food crops [51]. Besides trait introgression from wild relatives to cultivated species, research on the genomes of wild relatives has resulted in a process known as neodomestication. Neodomestication is a novel approach for crop improvement that allows the creation of a domesticated crop from a non-domesticated species by manipulating critical domestication genes [52]. The neodomestication of a wild relative of rice, reported in 2021, is the first example of a newly domesticated cereal crop with a polyploid genome. Researchers have used the CRISPR genome editing tool to target the traits controlling seed shattering, plant height, grain size, and flowering time in a wild species, Oryza alta, which is salinity stress-tolerant and pest-resistant. As a result, a shorter plant with longer grains and reduced flowering time was developed. These characteristics make O. alta suitable for grain production in temperate climates [53,54]. This opens the door to the rapid neodomestication of the wild relatives carrying resilience and adaptation traits for further crop improvement. Thus, the efficient use of genetic diversity from wild relatives would help in new gene discoveries and can be used for crop improvement [55,56,57,58,59,60,61,62].
Rice is of significant value to humans as it is a major staple food for more than half of the world population. The genus Oryza has been domesticated independently in Asia (10,000 years ago), which is Oryza sativa, and Africa (3000 years ago), which is Oryza glaberrima. The global population is expected to increase to 9 billion by 2050. Therefore, rice breeders need to develop new and improved cultivars of rice that have higher yields and adaptation to different biogeographic regions [63,64,65,66]. CWRs of rice can provide genetic resources for these important traits as they are adapted to wide geographic ranges and are tolerant to many biotic and abiotic stresses [63,67,68]. A total of 27 species of the genus Oryza are present after 15 million years of evolution, with 11 genome types. Some 6 of them were diploid (n = 12, AA, BB, CC, EE, FF, GG) and 5 were polyploid (n = 24, BBCC, CCDD, HHJJ, HHKK, KKLL). Cultivated rice is diploid, having an AA genome. For trait introgression from CWRs or their neodomestication, the first step is to gather information about genomic differences and similarities between the wild and cultivated species [52,63]. This can also speed up the discovery of new candidate genes for stress tolerance in plants.
Our lab has been working on identification and characterization of several gene families that contribute to tolerance towards abiotic stresses [64,69,70,71,72,73]. In this study, we have explored the presence of the FMO family of genes in 10 genomes of rice (both wild and cultivated) to understand the evolution, conservation and functional diversification of the FMO family. We have also investigated the possible involvement of this family in providing tolerance to different abiotic stresses using publicly available expression datasets and further experimental validation.
2. Results
2.1. The Number of FMO Genes Varies in Wild and Cultivated Species of Rice
The hidden Markov model (HMM) profile (PF00743) search was done against the Gramene [74] database for different Oryza species (Oryza brachyantha, Oryza punctata, Oryza meridionalis, Oryza glumaepatula, Oryza glaberrima, Oryza barthii, Oryza nivara, Oryza sativa subsp. indica, Oryza rufipogon and Oryza sativa subsp. japonica). We found that the number of FMO genes, having conserved binding motifs for FAD and NADPH, varies in cultivated and wild rice species. There are 25 different FMO members in O. brachyantha, 22 in O. punctata, 27 in O. meridionalis, 30 in O. glumaepatula, 24 in O. glaberrima, 25 in O. barthii, 24 in O. nivara, 29 in O. sativa subsp. indica, 24 in O. rufipogon and 28 in O. sativa subsp. japonica. Table 1 lists the FMO genes found in various species and these species have been arranged according to their evolution. Among these species, O. brachyantha is the most distant, while O. sativa subsp. indica is most recent one. The nomenclature of various genes in different Oryza species has been assigned according to their appearance on the chromosomes in ascending order, prefixed by the genus and species name.

Table 1.
Identification and nomenclature of FMO encoding genes in wild and cultivated rice. The names of the FMO encoding genes identified, from the Ensembl Gramene database, have been assigned according to increasing order of chromosomal distribution. Suffixes of OsI, OsJ, Osgl have been apportioned to the cultivated varieties indica, japonica and glaberrima respectively and Oba, Obr, On, Om, Or and Op have been allocated to O. barthii, O. brachyantha, O. nivara, O. meridionalis, O. rufipogon and O. punctata, respectively. (A) shows nomenclature for O. brachyantha, O. punctata, O. meridionalis, O. glumaepatula, O. glaberrima and (B) show nomenclature for O. barthii, O. nivara, O. indica, O. rufipogon and O. japonica. In the table, each rice chromosome is indicated by a different colored row.
2.2. The Distribution of FMO Genes on Chromosomes Reveals the Presence of Gene Clusters
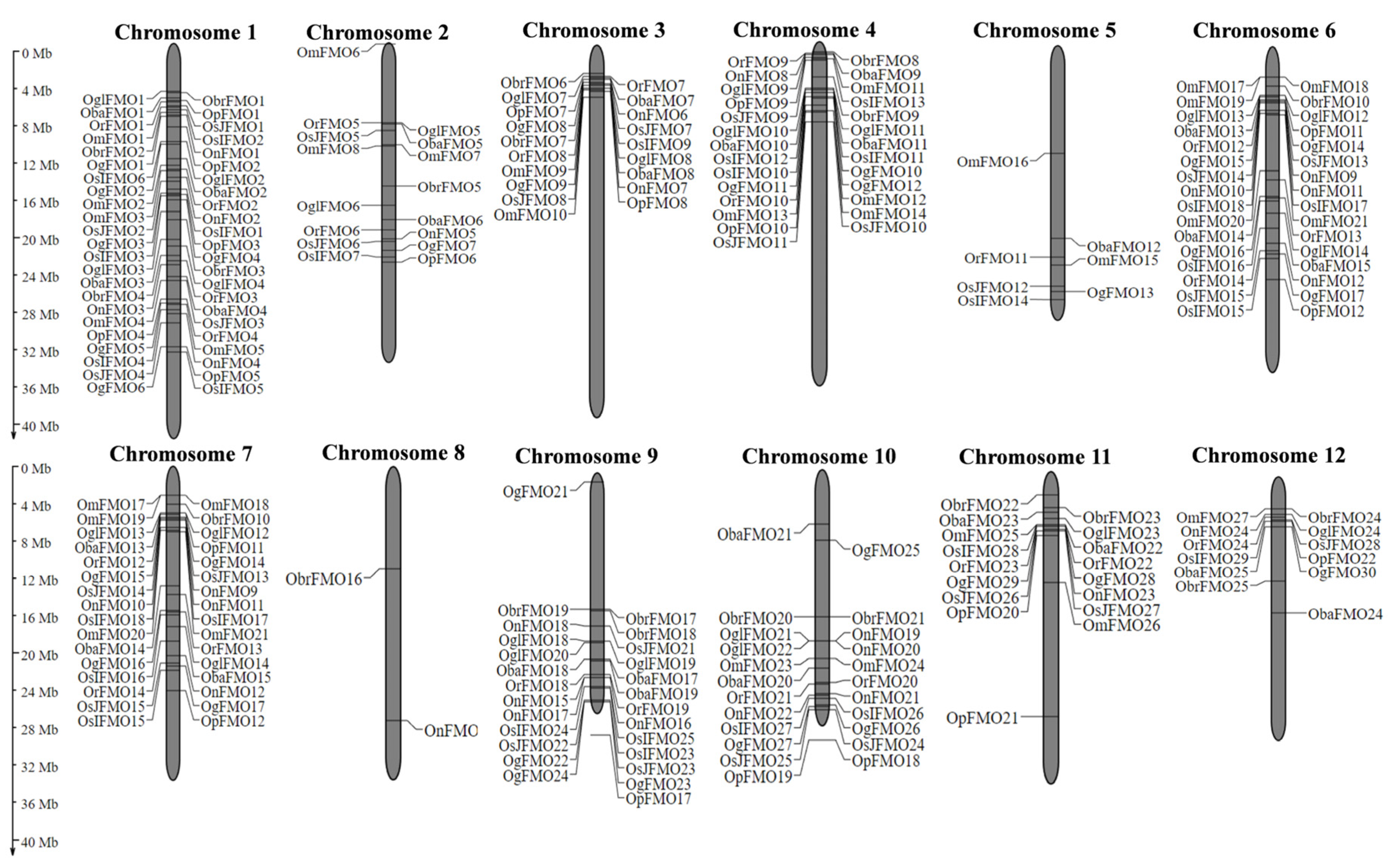
Information regarding the coordinates of FMO genes belonging to different Oryza species was obtained from the Gramene database and genes from all the species were mapped on the rice chromosomes using MapGene2Chromosome v2 [75]. We found that FMO genes are mainly distributed on 11 out of 12 rice chromosomes. Unlike other chromosomes, Chromosome (Chr) number 8 was the least populated, consisting of single FMO gene from O. brachyantha (ObrFMO18) and O. nivara (OnFMO14) (Figure 1). Among these 12 chromosomes, the maximum members of a single species are present on Chr 1 and Chr 4 and the lowest are found on Chr 5 and Chr 8. Furthermore, careful observation of chromosomal distribution of FMO genes revealed that the majority of the orthologous FMO genes in different Oryza species were found in the same order on a particular chromosome. Also, it is noteworthy that some FMO genes form clusters as they are adjacent to each other on the chromosomes. For example, in Oryza sativa subsp. japonica OsJFMO7 and OsJFMO8 forms a cluster on Chr 3, OsJFMO13 and OsJFMO14 on Chr 6, OsJFMO16, OsJFMO17, OsJFMO18, and OsJFMO19 on Chr 7, OsJFMO22 and OsJFMO23 on Chr 9, OsJFMO24 and OsJFMO25 on Chr 10. Further, OsJFMO26 and OsJFMO27 forms a cluster on Chr 11.
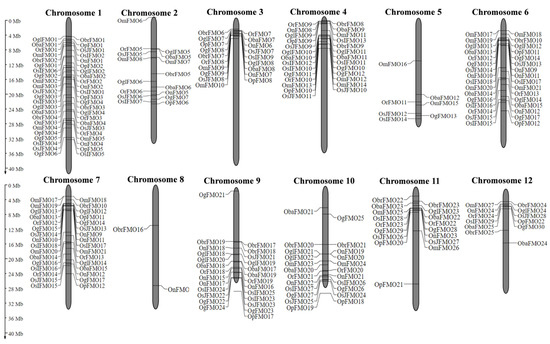
Figure 1.
Chromosomal distribution of FMO genes in wild and cultivated rice. The distribution of the FMO genes on the twelve chromosomes of rice was determined using MapGene2Chromosome tool. The chromosome numbers are depicted on top of each chromosome. The position of each gene on the respective chromosome has been depicted in terms of megabase pairs by numbers beside each gene.
2.3. Phylogenetic Analysis Shows Conservation among FMO Family and the Majority of Them Belongs to the S-Oxygenating Clade in Different Oryza Species
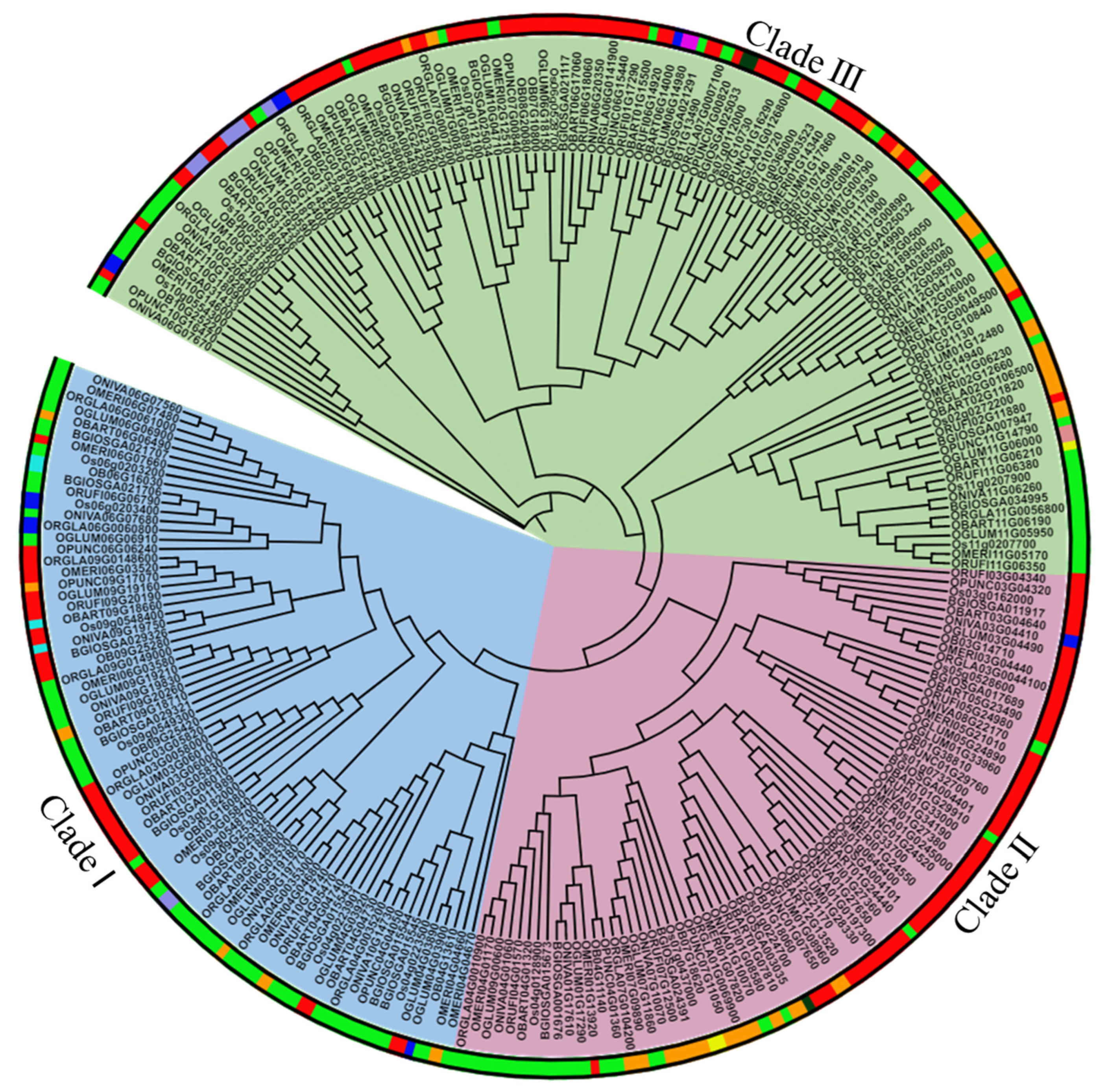
To determine the evolutionary and phylogenetic relationships among FMO proteins in different Oryza species, a rooted tree was constructed using MEGA7 software [76] and the proteins were grouped into three different clades based on their putative functions (Figure 2). Clade I was assigned as a pathogen defense clade, Clade II as an auxin biosynthesis clade, and Clade III as an S-oxygenating clade based on its constituents’ homology to known and characterized Arabidopsis members [14,21]. Cultivated rice (O. sativa subsp. indica, O. sativa subsp. japonica, O. glaberrima) has 8 members in Clade I, 6–7 in Clade II and 10–14 members in Clade III. In wild rice species (O. brachyantha, O. punctata, O. meridionalis, O. glumaepatula, O. barthii, O. nivara and O. rufipogon), there are 4–9 members in clade I, 5–8 members in clade II and 8–13 members in clade III. Most family members belong to the S-oxygenating clade. We also noticed that OnFMO10 is the outlier in the phylogenetic tree (Figure 2). This might be because of its lower homology to all the other protein sequences of the Oryza species.
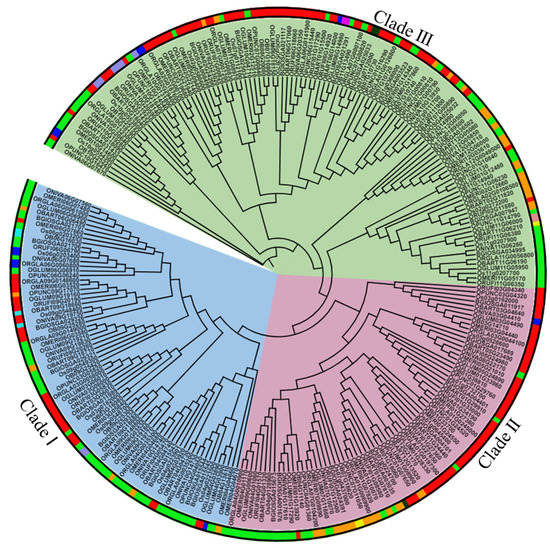
Figure 2.
Evolutionary and sub-cellular localization analysis of FMO genes in wild and cultivated rice. A rooted circular phylogenetic tree, depicting evolutionary connection between the FMO encoding genes in wild and cultivated rice, was determined by maximum likelihood method with 1000 bootstrap replicates using MEGA 7.0 and visualised using iTOL. The subcellular localization of the genes has been indicated by a color strip around the tree. Red—cytoplasm, parrot green—chloroplast, dark green—endoplasmic reticulum, orange—plastid, blue—nucleus, cyan—vacuole, pink—golgi, peach—cytoskeleton, yellow—extracellular space. The branch lengths indicate the evolutionary time between the two nodes.
The localization of FMO proteins was predicted using WolfPsort software [77]. A maximum number of proteins were found to be localized in the, cytoplasm followed by chloroplast (Figure 2). In Oryza sativa subsp. japonica, 11 proteins were found to be localized in the chloroplast (light green strip), 10 in the cytoplasm (red strip), 4 in the plastid (orange strip), 1 in the nucleus (blue strip), 1 in the peroxisome (purple strip), and 1 in the endoplasmic reticulum (dark green strip). However, in O. sativa subsp. indica, 13 proteins were found in the chloroplast, 11 in the cytoplasm, 4 in the plastid and 1 in the ER. In wild rice species, besides these organelles, very few FMO proteins were also found to be localized in the vacuole (cyan strip), cytoskeleton (peach strip) and extracellular compartment (yellow strip). Based on the results of multiple sequence alignment by ClustalW and the phylogenetic tree, orthologs from different species were identified. These are represented in Table 2.

Table 2.
Distribution of FMO genes in 10 genomes of rice. The chromosome-wise distribution of the identified FMO family members in the genomes of ten Oryza species, arranged according to their evolutionary history. O. brachyantha is the most distant and O. sativa subp. japonica is the most recent among these species. Each row contains list of genes that are orthologs of each other in different Oryza species. The blank space denotes the absence of the corresponding ortholog in the respective species. * Indicates the presence of an ortholog that belongs to some different chromosome in a particular row.
2.4. Protein Motif Analysis Reveals Five Conserved Motifs in FMO Members from Different Oryza Species
Following the phylogenetic analysis of FMO members in different Oryza species, we analyzed the conservation of different motifs, 5 of which are signatory conserved motifs of FMOs. It has been reported that FMO binds the adenine motif of FAD with a conserved GxGxxG sequence which is the FAD-binding motif found at the N terminus of the protein [78]. An identical (GxGxxG) but less conserved motif, the NADPH-binding motif is located at the centre of the protein [79]. Another motif, ATG containing motif (ATGY) is present at the C-terminus and occurs in proteins that carry out N-oxidation. Finally, a conserved motif FxGxxHxxY/F, FMO-identifying sequence is found in all known plant FMOs. It can be distributed anywhere in the protein [79]. We found that orthologs from different Oryza species exhibit high conservation in protein motifs (Figure 3). Variation in protein motif conservation in some orthologs from Oryza brachyantha has been observed as it is the most distant ancestor among these species. We also checked the presence of signatory conserved motifs of the FMO family.

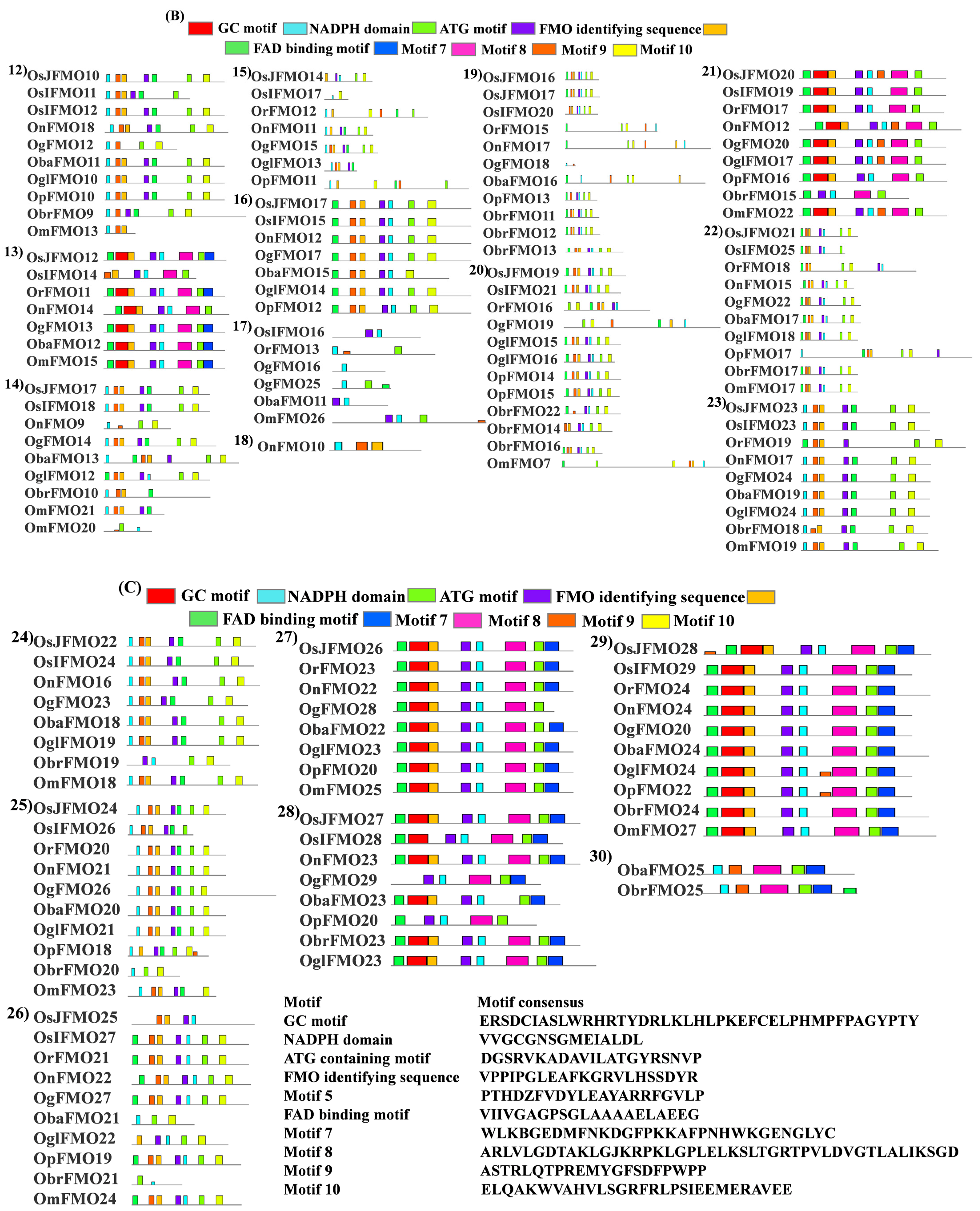
Figure 3.
Schematic representation of the conserved FMO-identifying motifs in the FMO proteins in wild and cultivated rice. FMO-identifying motifs previously reported in literature were searched for in the FMO protein sequences of the cultivated Oryza sativa subsp. japonica and wild species, Oryza nivara. The motif sequences have been marked by colours representing each of the conserved motifs, bright green: FAD binding domain; red: GC motif, violet: FMO-identifying sequence; cyan: NADPH domain and light olive green: ATG-containing motif.
In our analysis, we found that almost all of these motifs were present in the protein sequences of different Oryza species. However, few of them lack these motifs. The FAD-binding motif was mostly present at the N terminus of the protein, while the ATG-containing motif was present at the C terminus. Also, it was observed that ATG-binding motif was present twice in some of the proteins. Besides these, 5 more conserved motifs were seen in these sequences, which are named motif 5, motif 7, motif 8, motif 9, and motif 10 in this study. Again, OnFMO10 (Figure 4B(18)) shows the difference in protein motif arrangement from the rest of the members, with only 3 motifs present in total, NADPH-containing motif, motif 5, and motif 9. OgFMO16 (Figure 4B(17)) shows the presence of only 1 motif, the NADPH-binding motif, and OsIFMO16 shows the presence of two motifs, NADPH and FMO-identifying motifs (Figure 4A,B). This is probably because this gene has not been sequenced properly and the available sequence in the Gramene database has missing 5′ bases, which are denoted as a stretch of N in the database. In the 5th and 30th sets (Figure 4A(5),C(30) of orthologs that have only wild rice members, the FMO-identifying sequence was absent while 5–6 other conserved motifs were present.

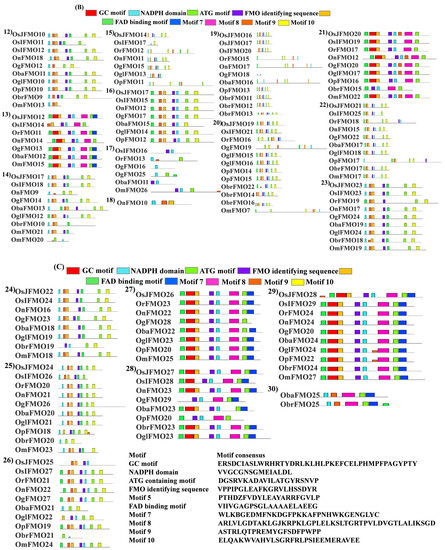
Figure 4.
Protein motif analysis in FMO genes in wild and cultivated rice. Conserved motifs, in addition to the FMO-identifying motifs, were analyzed using the MEME suite. The sequences of all the analyzed motifs have been depicted in the box. The FMO protein motifs in the wild and cultivated rice have been represented in sets (A) 1–11 (B) 12–23 and (C) 24–30. Wild rice FMO orthologs have been grouped together in each set. Similar motif organization was observed in the different sets.
2.5. Analysis of Gene Architecture Depicts Conservation among Orthologs from Different Oryza Species
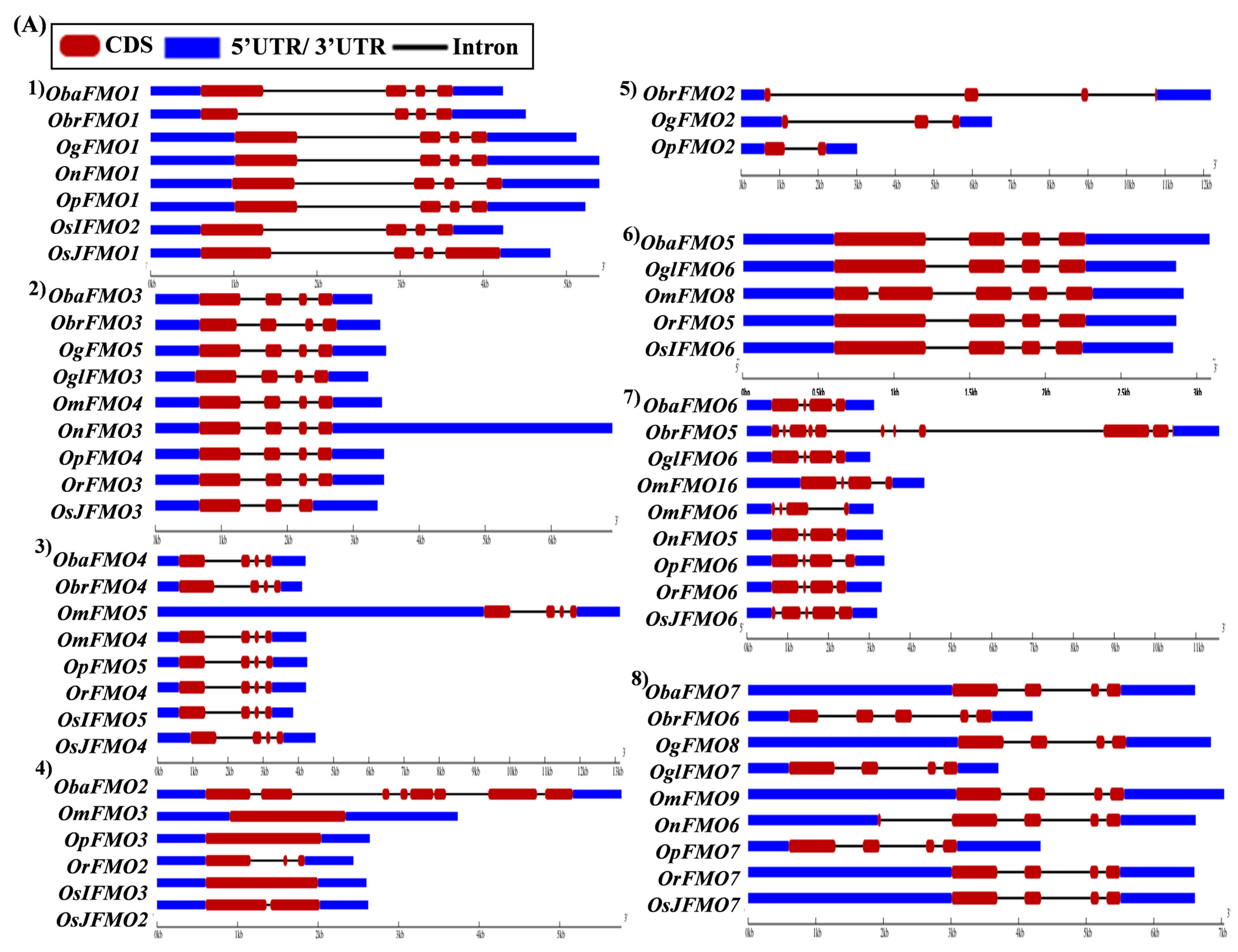
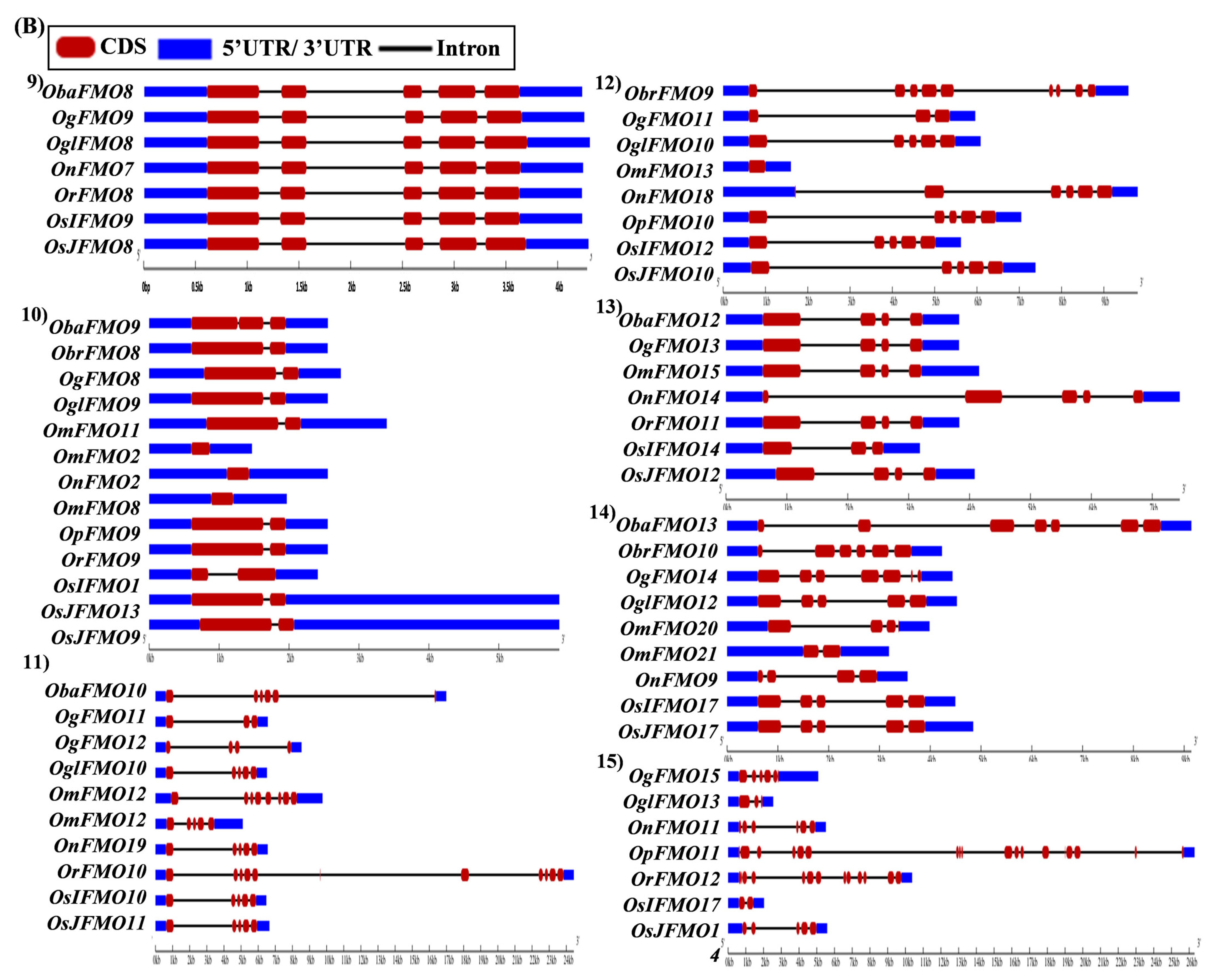
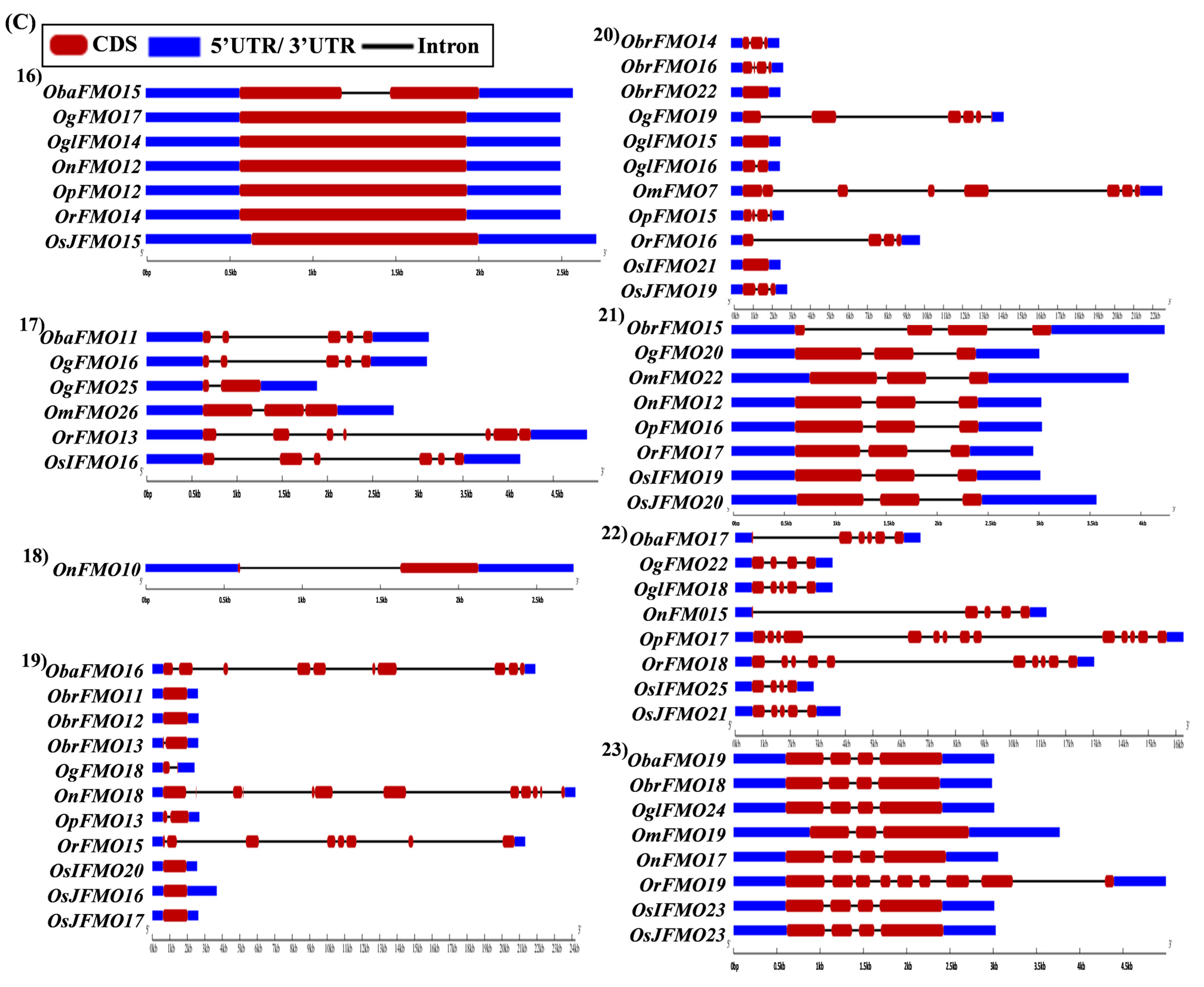
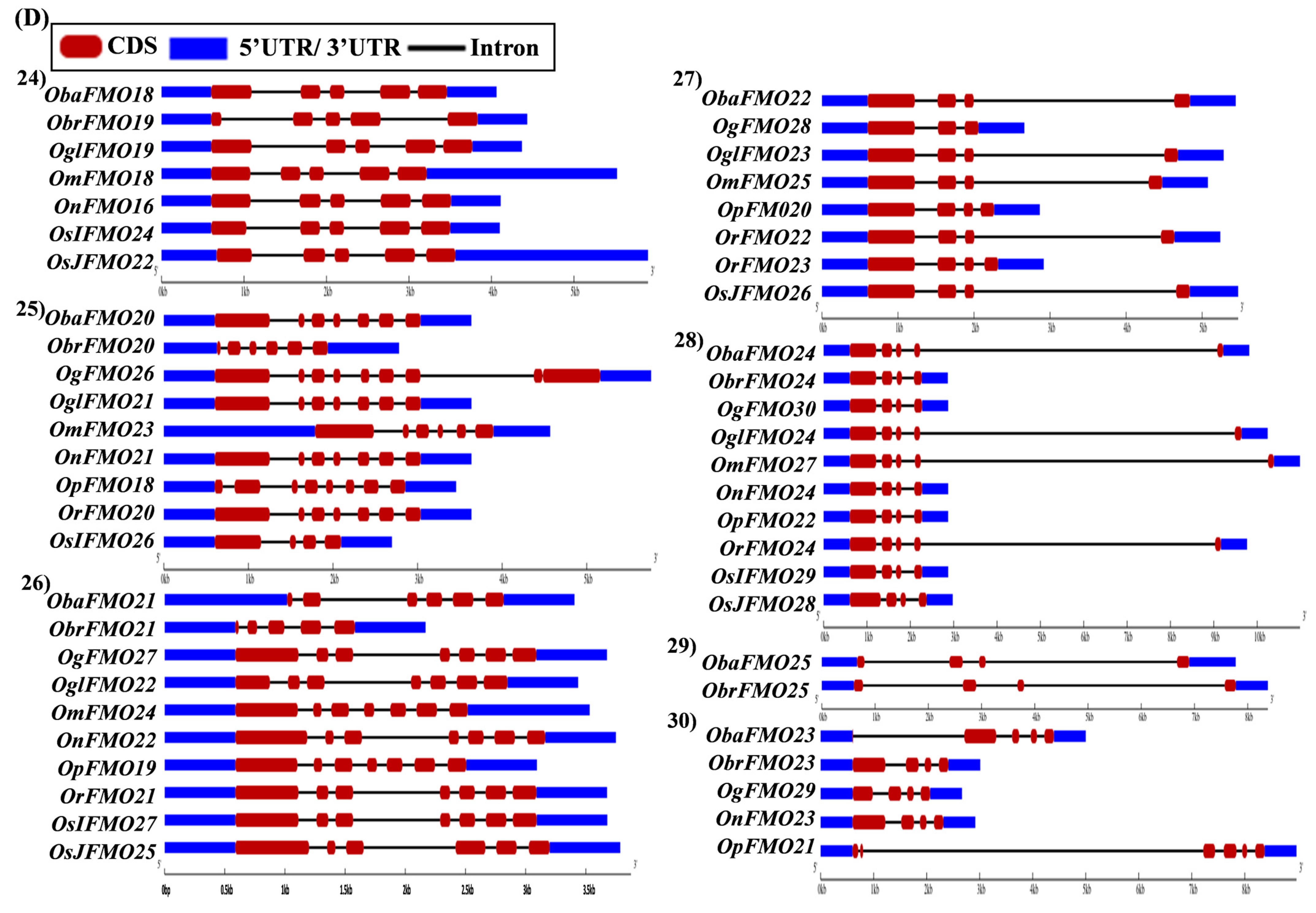
The exon–intron arrangement for the genes encoding the FMO proteins in different Oryza species was evaluated using the Gene Structure Display Server tool [80]. The gene structures of the orthologous genes have been represented together in Figure 5. The orthologs were largely found to share similar exon–intron arrangements, with a few exceptions. OnFMO3 possessed an exceptionally long 3′ UTR, while OmFMO5 had a long 5′ UTR (Figure 5A(2,3)). The exon–intron arrangement of ObaFMO2 was found to be completely different from that of the other members in the set, consisting of eight exons, while the rest comprised one to a maximum of three exons (Figure 5A(4)). In the fifth set, ObrFMO2 had longer introns as compared to OgFMO2 and OpFMO2 (Figure 5A(5)) A similar observation was also made for ObrFMO5, ObaFMO10, OpFMO11 and OrFMO15 and (Figure 5A(7),B(11,15),C(19)).
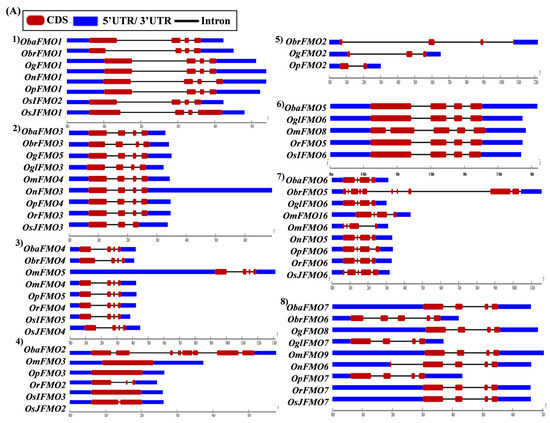
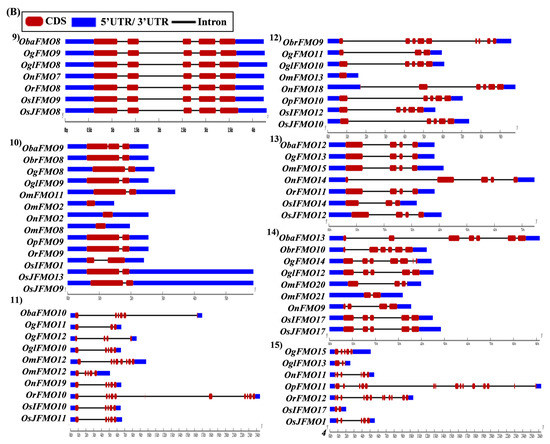
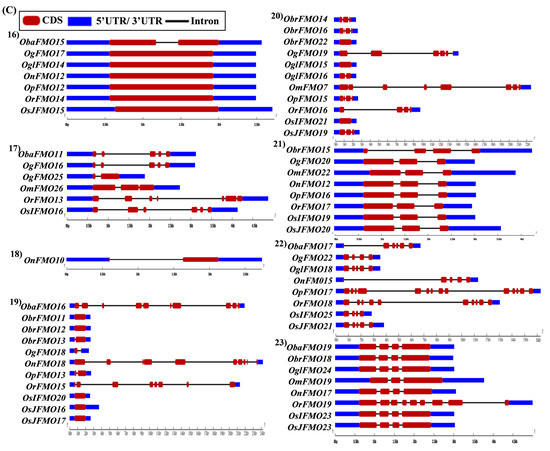
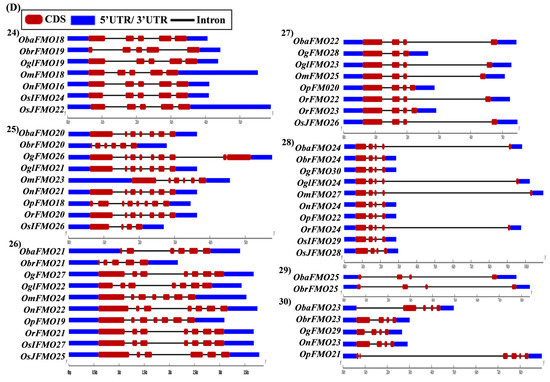
Figure 5.
Gene architecture of the FMO gene family in cultivated and wild relatives of rice. The exon–intron arrangement of the FMO gene family from the different Oryza species, (A) 1–8, (B) 9–15, (C) 16–23 and (D) 24–30 was predicted using the Gene Structure Display Server 2.0. The length of UTR, exon, and intron has been depicted in proportion to the actual sizes, which is also indicated using a scale at the top, blue: 5′ UTR/3′ UTR region; red: CDS; solid line: intron. Different sets show conserved gene structures.
In the nineteenth set, ObaFMO16, OnFMO18, and OrFMO15 consisted of 8–10 exons, while the rest were either intronless or consisted of a maximum of two exons. Another example of this arrangement would be the twentieth set, wherein OgFMO19, OmFMO7 and OrFMO16 contained varying numbers of exons ranging from 4–8, while others consisted of two exons or no introns at all (Figure 5C(19,20)). In the sixteenth set, all the genes were intronless except ObaFMO15 (Figure 5C(16)). In the twenty-eight set of orthologs, ObaFMO24, OmFMO27, OrFMO24 and OglFMO24 consisted of longer introns and five exons, one more than the other genes in the same set (Figure 5D(28)). Overall, it can be seen that the FMO gene family has variable sizes of exon, intron, and UTRs.
2.6. Domain Assessment Reveals Loss of Extra Copies of FAD Domain during the Course of Domestication
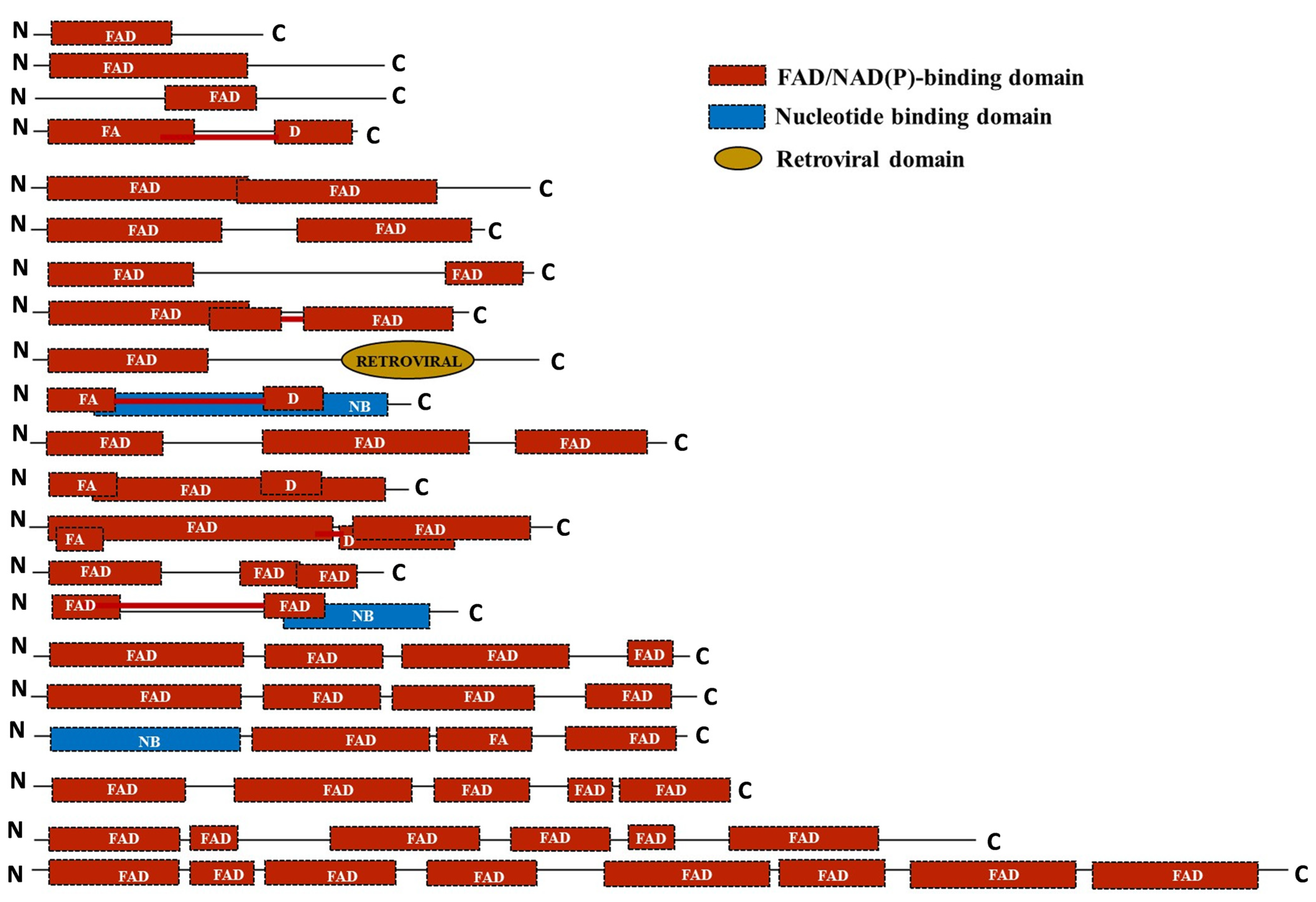
To assess the domain architecture of the identified putative FMO proteins, analysis was done using the SUPERFAMILY database [81]. The protein sequences were found to harbour single-to-multiple copies of the FAD/NADP-binding domain. Additionally, a nucleotide-binding domain and retroviral domains were also found in a few of the sequences like OgFMO19, OgFMO2, ObrFMO25, and ObaFMO2, respectively. While all the FMO family proteins of the cultivated rice varieties, both indica and japonica, consisted of single-to-double repeats of the domain, wild relatives of rice mostly contained two-to-five repeats of the domain such as OpFMO11, OpFMO17, OrFMO15, OrFMO18, OnFMO18, OgFMO19. OnFMO18, and OgFMO19. In fact, OmFMO7 was found to harbor eight copies of the domain, the most among all the proteins in wild vs cultivated varieties. The presence of multiple domains also correlated with the sequence length of the proteins. Interestingly, we also found a few of the domains to be arranged in such a way that they overlapped with the adjacent domain repeat, while others were split because they were linked together by a linker or spacer region (Figure 6).
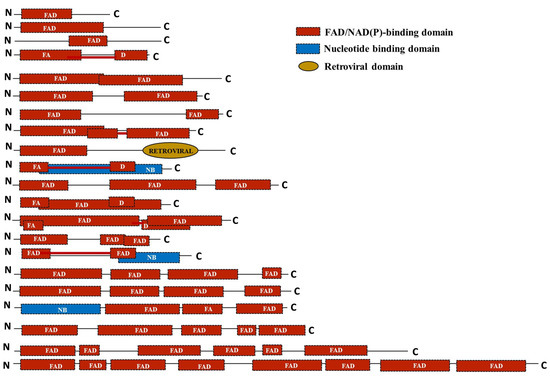
Figure 6.
Schematic representation of the domain architecture of the FMO family of proteins in the cultivated and wild relatives of rice. The domain organization of the FMO members in different Oryza species was predicted by the SUPERFAMILY database. FAD; flavin adenine dinucleotide domain, NB; nucleotide-binding domain; retroviral domain. The length of the domains has been represented in proportion to their actual sizes and are not to scale. Domains connected by a red line mean they are split.
2.7. Developmental and Stress-Mediated Expression Profiling of FMO Genes Reveals Tissue Specificity and Perturbations in a Subset of Genes in Different Abiotic Stresses
Next, we analyzed the expression profiling of the genes encoding the FMO family of proteins in the cultivated variety Oryza sativa subsp. japonica. Normalized and curated transcript abundance data were retrieved from the publicly available Genevestigator database. Of the 28 genes, expression data for 6 genes viz, OsJFMO2 (Os01g0368000), OsJFMO12 (Os05g0528600), OsJFMO17 (Os07g0111900), OsJFMO18 (Os07g0112000), OsJFMO26 (Os11g0207700), OsJFMO27 (Os11g0207900) could not be retrieved due to data unavailability.
The spatial expression profiling revealed differential expression of the genes in the different tissues, namely endosperm, embryo, seedling, culm, leaf, flag leaf, panicle, spikelet, and root. OsJFMO6 and OsJFMO28 were found to be highly expressed only in the endosperm. Most of the other genes such as OsJFMO5, OsJFMO8, OsJFMO14, OsJFMO19, OsJFMO22, OsJFMO23, OsJFMO3, OsJFMO4, OsJFMO20, and OsJFMO7 showed low to medium level of expression in all the tissues. Interestingly, the transcript abundance of only one gene, OsJFMO24, was maintained across tissues. OsJFMO21 and OsJFMO9 showed medium-to-high levels of expression in all the tissues. In fact, OsFMO9 showed maximum expression (14.36-fold change) in roots as compared to other genes. OsJFMO9 and OsJFMO10 were found to be highly expressed in the panicle (11.66- and 10.82-fold, respectively) and spikelet (11.95- and 11-fold, respectively). OsJFMO9 was highly induced (13.33-fold change) in the roots of the seedling. OsJFMO25 and OsJFMO9 showed higher expression (11.26-, 11.04-, and 10.79-fold) in the embryo. (Figure 7A).
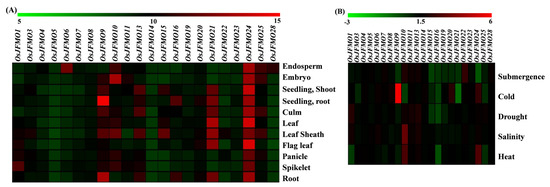
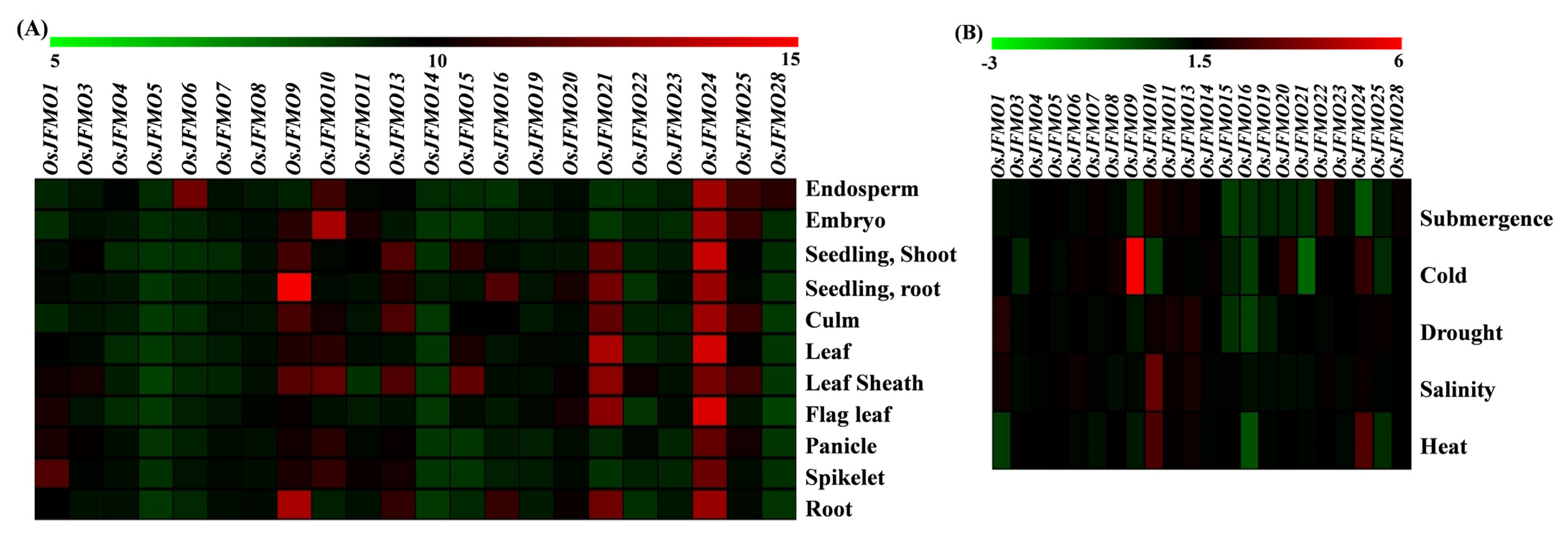
Figure 7.
Spatial and stress-mediated expression profiling of different FMO genes cultivated variety O. sativa subsp. japonica. Expression data were retrieved from the publicly available Affymetrix as well as RNA-seq datasets from the Genevestigator for (A) different tissues during development and (B) under different abiotic stress conditions namely heat, salinity, cold, drought and submergence. All expression values with p value < 0.05 have been depicted in log2 fold change. The heatmaps have been generated using MeV software. The color scale below each heatmap shows the level of expression, with red color showing the highest expression and green showing the lowest expression.
Further, we explored the expression profiling of the FMO genes under various stress conditions such as salinity, heat, cold, drought, and submergence. All expression values have been denoted as log2 fold change. Out of 28, 10 genes show little or no perturbations under various stresses examined. These are OsJFMO4, OsJFMO5, OsJFMO6, OsJFMO7, OsJFMO8, OsJFMO11, OsJFMO13, OsJFMO14, OsJFMO23, and OsJFMO28. However, 11 genes, OsJFMO1, OsJFMO3, OsJFMO9, OsJFMO10, OsJFMO15, OsJFMO16, OsJFMO19, OsJFMO20, OsJFMO21, OsJFMO24, and OsJFMO25 were found to be significantly repressed in one or the other stress. Only a few of them were found to be upregulated. For example, OsJFMO10 upregulation was highly induced under salinity (2.44-fold) stress. OsJFMO1 is the only gene found to be upregulated under drought stress. OsJFMO9 was found to be 5–6-fold upregulated under cold stress, the highest level among all the genes (Figure 7B). Overall, it seems that stresses impose perturbation in a subset of genes among this family. The expression data of each gene has been added in the Supplementary Materials (Table S1).
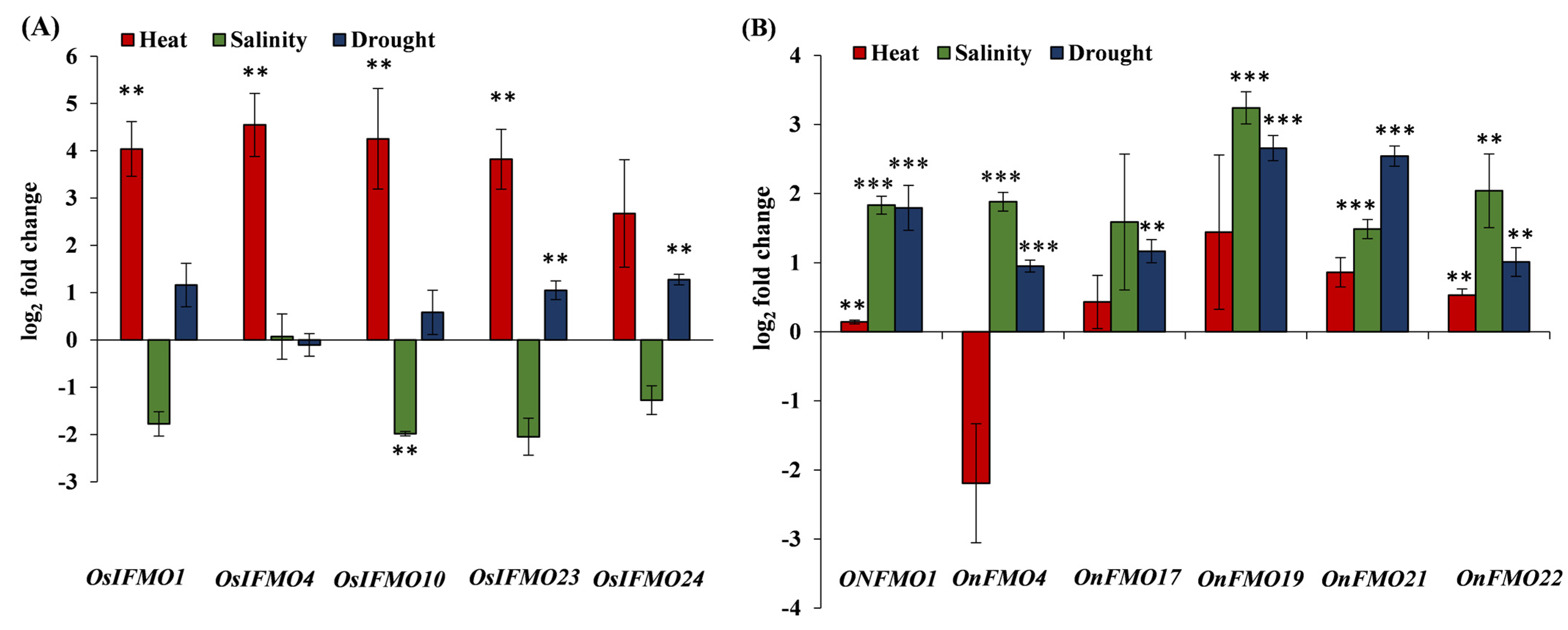
Next, we validated some of the genes encoding for the FMO family of proteins based on the publicly available stress-mediated expression profiling data. The genes selected belong to the three different clades. Leaves cut from one month old plants of the cultivated species Oryza sativa subsp. indica and its closest wild rice relative, Oryza nivara, were subjected to different stress conditions such as high temperature, salinity, and drought. In Oryza sativa subsp. indica, OsIFMO1, OsIFMO4, OsIFMO10, OsIFMO23 and OsIFMO24, five genes were found to be significantly upregulated under high temperature stress. However, none of the genes show significant upregulation under drought stress. A contrasting trend was observed in salinity stress where OsIFMO1, OsIFMO10, OsIFMO24 and OsIFMO10 shows downregulation (0.07 to −2.04-fold) (Figure 8A). Under drought stress, all of the genes, OnFMO1, OnFMO17, OnFMO21, OnFMO4, and OnFMO20, were significantly up regulated in the wild relative O. nivara. In heat stress, four of them, OnFMO1, OnFMO17, OnFMO19, OnFMO21 and OnFMO22 shows upregulation. In salinity stress, OnFMO19 (log 2-fold change, 3.24) and OnFMO22 (log 2-fold change, 2.04) shows perturbation in a significant range (Figure 8B). The ortholog of OnFMO22 i.e., OsIFMO27, was not detected in any of the stresses, thus indicating null expression of the gene at this developmental stage. This is the first report on experimentally validated expression profiling of FMO encoding genes from the wild rice O. nivara. Our expression profiling data was mostly, but not entirely, correlated with publicly available expression data of Oryza sativa subsp. japonica. OsIFMO10 and OsIFMO24 show a similar trend in heat and drought stress but an opposite trend in salt stress. Similarly, OsIFMO1 and OsIFMO4 show similar trends in drought and salinity, but opposite trends in heat stress. Thus, it can be concluded that, while FMO encoding genes in indica variety are induced by heat, their orthologs in O. nivara are induced by salinity and drought.
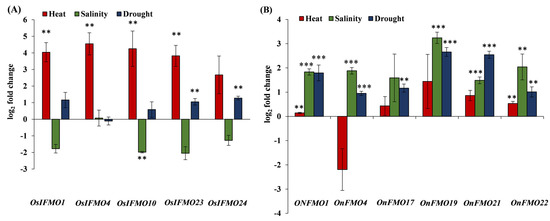
Figure 8.
Real-Time expression analysis of FMO genes in wild and cultivated rice. Expression profiling of selected FMO members from one month old leaves of (A) Oryza sativa subsp. indica (PB1 variety) and (B) Oryza nivara using qRT-PCR in response to heat stress (42 °C), salinity stress (200 mM NaCl) and drought stress (water withheld) for 24 h. Mean fold change (log2) is depicted, and the expression data is plotted against the untreated samples. The error bar represents the standard deviation where n = 6. *** signifies p value < 0.05 upto four or more decimal places and ** signifies p value < 0.05 for two decimal places.
3. Discussion
Plant FMOs are currently a significantly under-utilized class of enzymes, but studies in animal systems show that they have a rich potential for the discovery of new activities and novel bioactive metabolites. The evolutionary diversity of plant FMOs can provide a remarkable untapped resource of “green” biocatalysts [7]. Plant FMOs could provide unique breeding targets for subsistence agriculture based on their known important functions in plant metabolism. The identification of the entire complement of FMOs from Oryza species used in this study is a significant step forward in FMO research. The comparison of FMO genes from different Oryza species also provides useful information on evolutionary aspects of this gene family and shows conservation which clearly indicates the mandatory roles of FMOs, which cell cannot afford to change. However, all the substrates and reactions catalyzed by FMOs are yet to be identified.
The difference in the number of FMO genes in different Oryza species indicates the gain or loss of FMOs during the evolution of individual species. The size of gene families can increase by single gene duplications or duplication of a segment of chromosome i.e., segmental duplication. Gene family size can reduce by single gene deletion or deletion within a segmental piece [82,83]. The most distant ancestor among these species, O. brachyantha, has 25 FMO members, O. punctata has 22, and cultivated species O. sativa subsp. indica and O. sativa subsp. japonica have 28 and 29 genes, respectively. This increase in number in domesticated rice clearly indicates gene duplication events in their genome. There are a few non-functional FMOs in animal systems, such as human FMO6, which is a pseudogene [19,84]. Pseudogenes have missing promoters/deleted sequences/%frameshifts/a lesser number of introns or premature stop codons [85,86]. It is possible that a few of them are non-functional in rice as well. Across different Oryza species, a variable number of gene members of the jumonji C domain-containing protein family and the CBS-domain-containing protein family have been observed, implying that change in gene family size is very common in evolution [64,87,88].
Tandem duplications, which are duplication of adjacent identical chromosome segments (exon(s)/gene) resulting in the formation of another gene or specific exon, occur frequently in the plant genome [89]. The presence of FMO gene clusters on different chromosomes might be a result of tandem duplication. Similar clusters were also observed in FMOs belonging to clade II in wheat [90]. The Arabidopsis receptor-like kinase (RLKs) family has been expanded via tandem duplication [91]. Furthermore, it has been reported that tandem duplication has resulted in genome size expansion in rice, which may have occurred specifically in the rice lineage as a response to different stresses [92]. These duplications within the gene families may benefit the species in terms of functional diversity and response. Also, the majority of the orthologous FMO gene family members are found on the same order on chromosomes, indicating that these orthologous genes are present as syntenic blocks in chromosomal regions.
Phylogenetic analysis reveals that the gene family is conserved across these species due to sequence similarity. Also, the presence of maximum FMO members in clade III indicates their role in S-oxygenation. In 2007, the first plant FMO belonging to this clade was isolated and characterised from Arabidopsis [21]. FMOGS-OX is one out of seven FMOs present in Arabidopsis from this clade [41]. This FMO catalyzes the S-oxygenation of methionine-derived glucosinolates (GSLs) like methylthioalkyl. GSLs are plant secondary metabolites that are present in cruciferous plants, especially brassicales. GSLs and their hydrolytic products are involved in various plant functions like defense against pathogens and herbivores [93]. FMOs belonging to rice and other plant species that do not produce GSLs, on the other hand, may be involved in the S-oxygenation of a wide range of sulfur-containing compounds which are widespread in the plant kingdom [39]. However, the actual substrates of these FMOs are not known. The localization of FMOs in different intracellular compartments might provide functional diversity to this gene family by simultaneous physical segregation and the operation of various metabolic processes in the same cell.
The majority of FMO proteins from these species have conserved motifs that have been reported in the literature. The absence of some conserved motifs in a few FMO members again raises the possibility that these are non-functional pseudogenes but are related to other members of this family due to significant sequence similarity. Based on the absence of few conserved motifs, recent identification of NRL [NONPHOTOTROPIC HYPOCOTYL 3/ROOT PHOTOTROPISM 2-like (NPH3/RPT2-Like)] genes from rice have revealed the presence of two pseudogenes among the 27 that are present in the genome [94]. In fact, rice has many pseudogenes that belongs to different gene families [95]. However, the presence of these non-functional proteins in the FMO family can only be confirmed through functional studies.
Most orthologs from different Oryza species have similar exon–intron arrangements, again demonstrating the conservation of this gene family. However, the presence of an exceptionally long 3′ UTR in OnFMO3 and 5′ UTR in OmFMO5 indicate its role in specific developmental processes, as longer 5′ UTRs are generally found in genes involved in regulating growth processes in a tissue-specific manner [96]. The FMO-encoding genes with longer introns may be highly expressed as compared to those with shorter or no introns since, in plants, generally, genes with longer intronic regions and UTRs have high transcript abundance [97,98,99]. Also, it has been shown that the presence of both longer 3′ UTRs and introns act as cis elements to promote non-sense mediated decay (NMD) which is a quality check mechanism in eukaryotes including plants to eliminate identified aberrant mRNAs with a premature termination codon [100]. Thus, it is possible that OnFMO3, OmFMO5, and ObrFMO2 may be subjected to NMD owing to the presence of longer UTRs and introns in its gene architecture. Different gene structures in some sets of orthologs also indicate that gene duplication in this family has been followed by subsequent diversification [101,102] which can help these FMOs in performing various functions and/or in spatio-temporal regulation. The variable sizes of exon, intron, and UTRs in the FMO gene family implies that genes have undergone extensive shuffling during evolution [103,104].
From domain analysis, we have found that wild rice strains have mostly 2–5 FAD domains, with few members having more than five domains as well, whereas cultivated rice strains have only 1–2 FAD domains. There are three possibilities here. First, all the FAD domains in wild rice species work equally, and to reduce the genetic load on the cell, some of them have been deleted and the complete function has been assigned to a lesser number of domains in the cultivated rice. Second, only 1 or 2 domains are actually functional and are sufficient for the function of an FMO, and so the extra copies of this domain have been deleted. Another possibility is that these domains may be split in wild rice and together, they make functional domains during protein folding. A recent study has shown that domain gains and losses occur frequently during proteome evolution and concurrently with the evolution of cells. Fascinatingly, the number of gains tends to exceed the losses in the proteome, which could be to redefine the survival strategy of an organism [105]. In fact, it has been shown that higher eukaryotes trade-off the organismal budget for possessing the unique number of genes and domain architecture (economy), with the ability to resist damage and adaptability to environmental change [106]. It seems possible that a sharp reduction in the genetic size, together with a tendency to retain flexibility and robustness during domestication, has resulted in the subsequent loss of the extra repeats of the FAD domain of the FMO family in the cultivated varieties. Furthermore, the presence of nucleotide-binding domains in some FMO proteins clearly indicates that some members of this family play a role in pathogen defense, as this domain aids in pathogen recognition and signalling [107,108,109,110].
From the in silico expression analysis of available FMO genes of O. sativa subsp. japonica in different tissues, few genes were found to be highly expressed in specific tissues such as endosperm, root, panicle, spikelet, seedling root and embryo. Expression in a particular tissue indicates their functional specificity. The majority of the genes’ expression, however, was consistent across all tissues. Therefore, it can be inferred that these genes might be playing a significant role across all the developmental processes. Expression analysis of FMO genes under different abiotic stresses, as well as validation by qRT-PCR, reveals that the entire family does not respond to these stresses where only a subset of genes get downregulated and some were upregulated under these stresses. Some of the FMO genes belonging to Clade II have been found to be downregulated in drought and heat stress in wheat also [90]. In our study, we also found that few genes are downregulated under salinity stress, thus indicating the possibility that some of the members of the FMO family are not needed in these stresses and probably that its silencing could play a significant role in imparting stress tolerance in rice like SlMYB50 and SlMYB55 genes, the silencing of which enhanced salinity and drought tolerance in tomato [111,112]. However, this data is very preliminary, and detailed spatio-temporal and multiple stress-mediated expression profiling would help us understand the trend better. We believe the findings of this study will aid in the functional characterization of this gene family in rice, opening up new avenues for future research.
4. Materials and Methods
4.1. Identification of the FMO Family Members in Rice
The HMM (hidden Markov model) search of the FMO domain (PF00743) was retrieved from the Pfam database, (http://pfam.xfam.org/; EMBL-EBI, UK; accessed on 12 July 2022), which was used to identify the full-length protein sequences of FMO in the Gramene [74] database (https://www.gramene.org, accessed on 12 July 2022) for the identification of homologous sequences in 10 Oryza species: Oryza sativa subsp. japonica, O. sativa subsp. indica, O. barthi, O. brachyantha, O. glumaepatula, O. glaberrima, O. meridionalis, O. nivara, O. punctata, and O. rufipogon. All the retrieved sequences were analyzed using the Pfam 34.0 database (http://pfam.xfam.org/, accessed on 13 July 2022) for the presence of FAD domain in the FMO gene family. The developers and publishers of the Pfam database have chosen to shut down the website and reroute traffic to (InterPro (ebi.ac.uk)), another web-based resource that, like Pfam, analyses protein sequences functionally by grouping them into families and predicting the presence of domains and important sites. The data from the two databases are correlated.
4.2. Phylogenetic Analysis
Multiple sequence alignment of the FMO proteins from different Oryza species was performed using Clustal Ω and subsequently, phylogenetic tree was constructed using MEGA 7.0 [76] by the maximum likelihood method with 1000 bootstrap replicates. Tree was visualized using iTOL (https://itol.embl.de/upload.cgi, accessed on 14 July 2022). For subcellular localization, protein sequences were analyzed using the Wolf Psort: Protein subcellular localization predictor (https://wolfpsort.hgc.jp/, accessed on 12 July 2022).
4.3. Motif Analysis
The identified FMO sequences were analyzed for the presence of conserved motifs using the multiple expectation maximization for motif elicitation (MEME) program (http://meme-suite.org/, accessed on 27 July 2022), with the default parameters and the maximum number of motifs set to 10.
4.4. Evaluation of Domain Architecture of the FMO Family Proteins
The proteins sequences of the genes encoding the FMO family of proteins were analyzed using the SUPERFAMILY database (SUPERFAMILY database of structural and functional protein annotations for all completely sequenced organisms (supfam.org), accessed on 28 July 2022) for SCOP assessment using the default parameters.
4.5. Evaluation of the Gene Structure
The architecture of the genes encoding FMO proteins was determined and visualized using the Gene Structure Display Server tool, using genome sequence and CDS as query sequence (Gene Structure Display Server 2.0 (gao-lab.org), accessed on 29 July 2022).
4.6. Developmental and Stress-Mediated Expression Profiling of the FMO Encoding Genes
Normalized curated expression profiling data of the FMO encoding genes under different stresses and in the various anatomical parts under different developmental stages were retrieved from the Affymetrix, as well as RNA-seq datasets of the publicly available Genevestigator database (https://genevestigator.com; NEBION, Switzerland; accessed on 4 August 2022). The heatmaps were generated using MeV 4.9.0 application tool.
For qRT-PCR-based expression profiling of selected FMO genes, leaves cut from one month old plants of Oryza sativa subsp. indica and its closest wild relative Oryza nivara, growing under controlled greenhouse conditions, were subjected to salt stress (200 mM NaCl), drought (water withheld) and high temperature (42 °C) for 24 h. Total RNA was isolated using Trizol reagent (ThermoFisher Scientific, Waltham, MA, USA) and first-strand cDNA synthesis done using the RevertAid first-strand cDNA synthesis kit (ThermoFisher Scientific, Waltham, MA, USA). Real-time PCR was performed to the Applied Biosystems 7500 Step-One Instrument (Applied Biosystems 7500, Foster City, CA, USA). The eukaryotic Elongation Factor-1α (eEF-1α) was used as an internal control for normalization in the qRT-PCR. The log2 expression values of each gene under salinity, heat and drought conditions have been calculated with respect to the untreated control (having log2 value 0) [113].
5. Conclusions
Rice genome sequencing has substantially sped up the discovery and characterization of breeder’s important genes, as well as a new understanding of their evolutionary history. In our study, we have identified the entire complement of the FMO family from 10 different Oryza species including both wild and cultivated. Wild and cultivated rice species have different numbers of FMO genes. Phylogenetic, motif, and gene structure analysis indicate that this family is conserved across these species. The phylogenetic analysis of FMO orthologues in cultivated rice and its wild relatives also reveals that nearly all of the orthologs that shared comparable structural organizations were clustered together with a convincing bootstrap value (1000). Some of the genes from O. brachyantha show a difference in motif and gene structure from other orthologs because it has FF genome and it is the most distant ancestor. Unlike, O. brachyantha, O. punctata which has the BB genome, is different from the rest of the Oryza species with an AA genome and did not show any difference among its orthologs. From domain analysis, we have found that wild rice species have multiple domains which have been subsequently lost during evolution of our cultivated species. Also, the expression of this gene family in different rice tissues is reported in this study. From the in silico expression analysis and validation of selected genes by qRT-PCR in cultivated rice, O. sativa subsp. indica and wild rice O. nivara under different abiotic stresses, it can be concluded that the entire family does not respond to different stresses, implying that they perform diverse functions in the cell. Altogether, this study provides the total number of FMO genes present in rice, which has not been reported till now, and also assesses the similarities and differences in this family across different Oryza species. This information can help researchers in the functional characterization of rice FMOs. The stress responsiveness of some FMOs reported in this study further encourages the researcher to use them for improving stress tolerance in the cultivated rice varieties or neodomestication of CWRs.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms24044190/s1.
Author Contributions
Conceptualization, S.L.S.-P.; Data curation, Y.G. and B.B.; Formal analysis, Y.G.; Investigation, S.L.S.-P.; Resources, S.L.S.-P.; Software, B.B.; Supervision, S.L.S.-P.; Validation, B.B.; Visualization, A.P.; Writing—original draft, Y.G.; Writing—review & editing, A.P. and S.L.S.-P. All authors have read and agreed to the published version of the manuscript.
Funding
Funded by ICGEB.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The datasets supporting the conclusions of this article are included within the article and its additional files. The sequence data for all the Oryza species were obtained from Gramene data resource (https://gramene.org (accessed on 12 July 2022). For O. sativa subsp. japonica, the sequences were also retrieved from the RGAP (http://rice.plantbiology.msu.edu/ (accessed on 15 September 2021)).
Acknowledgments
S.L.S.-P. acknowledges ICGEB for core grant support. Y.G. and B.B. acknowledges Council of Scientific and Industrial Research (CSIR), Government of India, for providing research fellowship during their research work.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Pettit, F.H.; Ziegler, D.M. The catalytic demethylation of N, N-dimethylaniline-N-oxide by liver microsomes. Biochem. Biophys. Res. Commun. 1963, 13, 193–197. [Google Scholar] [CrossRef]
- Bickel, M.H. The pharmacology and biochemistry of N-oxides. Pharmacol. Rev. 1969, 21, 325–355. [Google Scholar] [PubMed]
- Jakoby, W.B.; Ziegler, D.M. The enzymes of detoxication. J. Biol. Chem. 1990, 265, 20715–20718. [Google Scholar] [CrossRef] [PubMed]
- Huijbers, M.M.E.; Montersino, S.; Westphal, A.H.; Tischler, D.; Van-Berkel, W.J.H. Flavin dependent monooxygenases. Arch. Biochem. Biophys. 2014, 544, 2–17. [Google Scholar] [CrossRef]
- Mascotti, M.L.; Lapadula, W.J.; Ayub, M.J. The origin and evolution of Baeyer—Villiger Monooxygenases (BVMOs): An ancestral family of flavin monooxygenases. PLoS ONE 2015, 10, e0132689. [Google Scholar] [CrossRef]
- Nicoll, C.R.; Bailleul, G.; Fiorentini, F.; Mascotti, M.L.; Fraaije, M.W.; Mattevi, A. Ancestral-sequence reconstruction unveils the structural basis of function in mammalian FMOs. Nat. Struct. Mol. Biol. 2020, 1, 14–24. [Google Scholar] [CrossRef]
- Thodberg, S.; Jakobsen Neilson, E.H. The “Green” FMOs: Diversity, Functionality and Application of Plant Flavoproteins. Catalysts 2020, 10, 329. [Google Scholar] [CrossRef]
- Zhang, J.; Chen, J.; Wu, Y.F.; Wang, Z.P.; Qiu, J.G.; Li, X.L.; Cai, F.; Xiao, K.Q.; Sun, X.X.; Rosen, B.P.; et al. Oxidation of organoarsenicals and antimonite by a novel flavin monooxygenase widely present in soil bacteria. Environ. Microbiol. 2022, 24, 752–761. [Google Scholar] [CrossRef]
- Ziegler, D.M.; Pettit, F.H. Formation of an intermediate N oxide in the oxidative demethylation of N, N-dimethylaniline catalyzed by liver microsomes. Biochem. Biophys. Res. Commun. 1964, 15, 188–193. [Google Scholar] [CrossRef]
- Cashman, J.R.; Zhang, J. Human flavin-containing monooxygenases. Annu. Rev. Pharmacol. Toxicol. 2006, 46, 65–100. [Google Scholar] [CrossRef]
- Mascotti, M.L.; Juri-Ayub, M.; Furnham, N.; Tornton, J.M.; Laskowski, R.A. Chopping and changing: The evolution of the flavin-dependent monooxygenases. J. Mol. Biol. 2016, 428, 3131–3146. [Google Scholar] [CrossRef] [PubMed]
- Rendić, S.P.; Crouch, R.D.; Guengerich, F.P. Roles of selected non-P450 human oxidoreductase enzymes in protective and toxic effects of chemicals: Review and compilation of reactions. Arch. Toxicol. 2022, 96, 2145–2246. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Zhou, Q.; Wu, Y.; Chen, X.; Zhong, F. Properties and mechanisms of flavin-dependent monooxygenases and their applications in natural product synthesis. Int. J. Mol. Sci. 2022, 23, 2622. [Google Scholar] [CrossRef] [PubMed]
- Schlaich, N.L. Flavin-containing monooxygenases in plants: Looking beyond detox. Trends Plant Sci. 2007, 12, 412–418. [Google Scholar] [CrossRef] [PubMed]
- Suh, J.K.; Poulson, L.L.; Ziegler, D.M.; Robertus, J.D. Molecular cloning and kinetic characterization of a flavin-containing monooxygenase from Saccharomyces cerevisiae. Arch. Biochem. Biophys. 1996, 336, 268–274. [Google Scholar] [CrossRef]
- Lawton, M.P.; Cashman, J.R.; Cresteil, T.; Dolphin, C.T.; Elfarra, A.A.; Hines, R.N.; Hodgson, E.; Kimura, T.; Ozols, J.; Phillips, I.R. A nomenclature for the mammalian flavin-containing monooxygenase gene family based on amino acid sequence identities. Arch. Biochem. Biophys. 1994, 308, 254–257. [Google Scholar] [CrossRef]
- Longin-Sauvageon, C.; Lattard, V.; Lilaz-Michel, C.; Buronfosse, T.; Benoit, E. Expression of two different FMOs in sheep liver. Drug Metab. Dispos. 1998, 26, 284–287. [Google Scholar]
- Veeravalli, S.; Phillips, I.R.; Freire, R.T.; Varshavi, D.; Everett, J.R.; Shephard, E.A. Flavin-containing monooxygenase 1 catalyzes the production of taurine from hypotaurine. Drug Metab. Dispos. 2020, 48, 378–385. [Google Scholar] [CrossRef]
- Uno, Y.; Shimizu, M.; Ogawa, Y.; Makiguchi, M.; Kawaguchi, H.; Yamato, O.; Ishizuka, M.; Yamazaki, H. Molecular and functional characterization of flavin-containing monooxygenases in pigs, dogs, and cats. Biochem. Pharmacol. 2022, 202, 115125. [Google Scholar] [CrossRef]
- Zhao, Y.; Christensen, S.K.; Fankhauser, C.; Cashman, J.R.; Cohen, J.D.; Weigel, D.; Chory, J. A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 2001, 291, 306–309. [Google Scholar] [CrossRef]
- Hansen, B.G.; Kliebenstein, D.J.; Halkier, B.A. Identification of a flavin-monooxygenase as the S-oxygenating enzyme in aliphatic glucosinolate biosynthesis in Arabidopsis. Plant J. 2007, 50, 902–910. [Google Scholar] [CrossRef] [PubMed]
- Romano, C.P.; Robson, P.R.H.; Smith, H.; Estelle, M.; Klee, H. Transgene-mediated auxin overproduction in Arabidopsis: Hypocotyl elongation phenotype and interactions with the hy6-1 hypocotyl elongation and axr1 auxin-resistant mutants. Plant Mol. Biol. 1995, 27, 1071–1083. [Google Scholar] [CrossRef] [PubMed]
- Marsch-Martinez, N.; Greco, R.; Arkel, G.V.; Herrera-Estrella, L.; Pereira, A. Activation tagging using the En-I maize transposon system in Arabidopsis. Plant Physiol. 2002, 129, 1544–1556. [Google Scholar] [CrossRef] [PubMed]
- Mishina, T.E.; Zeier, J. The Arabidopsis flavin-dependent monooxygenase FMO1 is an essential component of biologically induced systemic acquired resistance. Plant Physiol. 2006, 141, 1666–1675. [Google Scholar] [CrossRef]
- Bartsch, M.; Gobbato, E.; Bednarek, P.; Debey, S.; Schultze, J.L.; Bautor, J.; Parker, J.E. Salicylic acid-independent enhanced disease susceptibility1 signaling in Arabidopsis immunity and cell death is regulated by the monooxygenase FMO1 and the nudix hydrolase NUDT7. Plant Cell 2006, 18, 1038–1051. [Google Scholar] [CrossRef]
- Cheng, Y.; Dai, X.; Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev. 2006, 20, 1790–1799. [Google Scholar] [CrossRef]
- Cheng, Y.; Dai, X.; Zhao, Y. Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation in Arabidopsis. Plant Cell 2007, 19, 2430–2439. [Google Scholar] [CrossRef]
- Yamamoto, Y.; Kamiya, N.; Morinaka, Y.; Matsuoka, M.; Sazuka, T. Auxin biosynthesis by the YUCCA genes in rice. Plant Physiol. 2007, 143, 1362–1371. [Google Scholar] [CrossRef]
- Woo, Y.M.; Park, H.J.; Su’udi, M.; Yangm, J.I.; Park, J.J.; Back, K.; Park, Y.M.; An, G. Constitutively wilted 1, a member of the rice YUCCA gene family, is required for maintaining water homeostasis and an appropriate root to shoot ratio. Plant Mol. Biol. 2007, 65, 125–136. [Google Scholar] [CrossRef]
- Zhao, Y. Auxin biosynthesis: A simple two-step pathway converts tryptophan to indole-3-acetic acid in plants. Mol. Plant 2012, 5, 334–338. [Google Scholar] [CrossRef]
- Liu, H.; Ying, Y.Y.; Zhang, L.; Gao, Q.H.; Li, J.; Zhang, Z.; Fang, J.G.; Duan, K. Isolation and characterization of two YUCCA flavin monooxygenase genes from cultivated strawberry (Fragaria× ananassa Duch.). Plant Reprod. 2012, 31, 1425–1435. [Google Scholar] [CrossRef]
- Kim, J.I.; Baek, D.; Park, H.C.; Chun, H.J.; Oh, D.H.; Lee, M.K.; Cha, J.Y.; Kim, W.Y.; Kim, M.C.; Chung, W.S.; et al. Overexpression of Arabidopsis YUCCA6 in potato results in high-auxin developmental phenotypes and enhanced resistance to water deficit. Mol. Plant. 2013, 6, 337–349. [Google Scholar] [CrossRef] [PubMed]
- Abu-Zaitoon, Y.M. Phylogenetic analysis of putative genes involved in the tryptophan-dependent pathway of auxin biosynthesis in rice. Appl. Biochem. Biotechnol. 2014, 172, 2480–2495. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Yin, N.; Niu, Z.; Hui, W.; Song, J.; Huang, C.; Wang, H.; Kong, L.; Feng, D. Isolation and characterization of three TaYUC10 genes from wheat. Gene 2014, 546, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Ke, Q.; Wang, Z.; Ji, C.Y.; Jeong, J.C.; Lee, H.S.; Li, H.; Li, H.; Xu, B.; Deng, X.; Kwak, S.S. Transgenic poplar expressing Arabidopsis YUCCA6 exhibits auxin-overproduction phenotypes and increased tolerance to abiotic stress. Plant Physiol. Biochem. 2015, 94, 19–27. [Google Scholar] [CrossRef]
- Li, W.; Zhao, X.; Zhang, X. Genome-wide analysis and expression patterns of the YUCCA genes in maize. J. Genet. Genom. 2015, 42, 707–710. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.G.; Liu, H.H.; Wang, S.P.; Li, H.G. Genome-wide identification and expression analysis of the YUCCA gene family in soybean (Glycine max L.). J. Plant Growth Regul. 2017, 81, 265–275. [Google Scholar] [CrossRef]
- Yang, Y.L.; Li, N.; Hui, W.R.; Yuan, B.J.; Fan, P.; Liu, J.X.; Wang, H.G.; Feng, D.S. Seed-specific expression of TaYUC10 significantly increases auxin and protein content in wheat seeds. Plant Reprod. 2020, 10, 301–314. [Google Scholar] [CrossRef]
- Li, J.; Hansen, B.G.; Ober, J.A.; Kliebenstein, D.J.; Halkier, B.A. Subclade of flavin-monooxygenases involved in aliphatic glucosinolate biosynthesis. Plant Physiol. 2008, 148, 1721–1733. [Google Scholar] [CrossRef]
- Yoshimoto, N.; Onuma, M.; Mizuno, S.; Sugino, Y.; Nakabayashi, R.; Imai, S.; Tsuneyoshi, T.; Sumi, S.; Saito, K. Identification of a flavin-containing S-oxygenating monooxygenase involved in alliin biosynthesis in garlic. Plant J. 2015, 83, 941–951. [Google Scholar] [CrossRef]
- Kong, W.; Li, J.; Yu, Q.; Cang, W.; Xu, R.; Wang, Y.; Ji, W. Two novel flavin-containing monooxygenases involved in biosynthesis of aliphatic glucosinolates. Front. Plant Sci. 2016, 7, 1292. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, M.; Zeier, T.; Bernsdorff, F.; Reichel-Deland, V.; Kim, D.; Hohmann, M.; Scholten, N.; Schuck, S.; Bräutigam, A.; Hölzel, T.; et al. Flavin monooxygenase-generated N-hydroxypipecolic acid is a critical element of plant systemic immunity. Cell 2018, 173, 456–469. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, N.; Saito, K. S-Alk(en)ylcysteine sulfoxides in the genus Allium: Proposed biosynthesis, chemical conversion, and bioactivities. J. Exp. Bot. 2019, 70, 4123–4137. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, R.; Sawada, Y.; Boerzhijin, S.; Kuwahara, A.; Sato, M.; Hirai, M.Y. Abscisic acid-mediated induction of FLAVIN-CONTAINING MONOOXYGENASE 2 leads to reduced accumulation of methylthioalkyl glucosinolates in Arabidopsis thaliana. Plant Sci. 2021, 303, 110764. [Google Scholar] [CrossRef] [PubMed]
- Cha, J.Y.; Kim, W.Y.; Kang, S.B.; Kim, J.I.; Baek, D.; Jung, I.J.; Kim, M.R.; Li, N.; Kim, H.J.; Nakajima, M.; et al. A novel thiol-reductase activity of Arabidopsis YUC6 confers drought tolerance independently of auxin biosynthesis. Nat. Commun. 2015, 6, 8041. [Google Scholar] [CrossRef] [PubMed]
- Cha, J.Y.; Kim, M.R.; Jung, I.J.; Kang, S.B.; Park, H.J.; Kim, M.G.; Yun, D.J.; Kim, W.Y. The thiol reductase activity of YUCCA6 mediates delayed leaf senescence by regulating genes involved in auxin redistribution. Front. Plant Sci. 2016, 7, 626. [Google Scholar] [CrossRef]
- Bremer, G. Problems in breeding and cytology of sugar cane. Euphytica 1961, 10, 59–78. [Google Scholar] [CrossRef]
- Roach, B.T. “Nobilisation of sugarcane”. Proc. Int. Soc. Sugar Cane Technol. 1972, 14, 206–216. [Google Scholar]
- Santchurn, D.; Badaloo, M.G.H.; Zhou, M.; Labuschagne, M.T. Contribution of sugarcane crop wild relatives in the creation of improved varieties in Mauritius. Plant Genet. Resour. 2019, 17, 151–163. [Google Scholar] [CrossRef]
- Pradheep, K.; Ahlawat, S.P.; Nivedhitha, S.; Gupta, V. Crop wild relatives in India: Inventorization, prioritization and conservation. Indian J. Plant Genet. Res. 2022, 35, 124–130. [Google Scholar] [CrossRef]
- Brown, W.L. Global Crop Resources: Gene Banks and the World’s Food. Science 1987, 236, 617–618. [Google Scholar] [CrossRef] [PubMed]
- Fornasiero, A.; Wing, R.A.; Ronald, P. Rice domestication. Curr. Biol. 2022, 32, R20–R24. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Lin, T.; Meng, X.; Du, H.; Zhang, J.; Liu, G. A route to de novo domestication of wild allotetraploid rice. Cell 2021, 184, 1156–1170. [Google Scholar] [CrossRef] [PubMed]
- Gutaker, R.M.; Chater, C.C.; Brinton, J.; Castillo-Lorenzo, E.; Breman, E.; Pironon, S. Scaling up neodomestication for climate-ready crops. Curr. Opin. Plant Biol. 2022, 66, 102169. [Google Scholar] [CrossRef] [PubMed]
- Hajjar, R.; Hodgkin, T. The use of wild relatives in crop improvement: A survey of developments over the last 20 years. Euphytica 2007, 156, 1–13. [Google Scholar] [CrossRef]
- Agarwal, P.; Parida, S.K.; Raghuvanshi, S.; Kapoor, S.; Khurana, P.; Khurana, J.P.; Tyagi, A.K. Rice improvement through genome-based functional analysis and molecular breeding in India. Rice 2016, 9, 1. [Google Scholar] [CrossRef]
- Nair, K.P. Utilizing crop wild relatives to combat global warming. Adv. Agron. 2019, 153, 175–258. [Google Scholar]
- Coyne, C.J.; Kumar, S.; Wettberg, E.J.; Marques, E.; Berger, J.D.; Redden, R.J.; Noel Ellis, T.H.; Brus, J.; Zablatzká, L.; Smýkal, P. Potential and limits of exploitation of crop wild relatives for pea, lentil, and chickpea improvement. Legume Sci. 2020, 2, e36. [Google Scholar] [CrossRef]
- Das, S. Contributions of crop-wild relatives toward broadening the list of leafy vegetables. Int. J. Veg. Sci. 2022, 2022, 1–14. [Google Scholar] [CrossRef]
- Wambugu, P.W.; Henry, R. Supporting in situ conservation of the genetic diversity of crop wild relatives using genomic technologies. Mol. Ecol. 2022, 31, 2207–2222. [Google Scholar] [CrossRef]
- Renzi, J.P.; Coyne, C.J.; Berger, J.; Wettberg, E.; Nelson, M.; Ureta, S.; Hernández, F.; Smýkal, P.; Brus, J. How Could the Use of Crop Wild Relatives in Breeding Increase the Adaptation of Crops to Marginal Environments? Front. Plant Sci. 2022, 13, 886162. [Google Scholar] [CrossRef] [PubMed]
- Curtin, S.; Qi, Y.; Peres, L.E.; Fernie, A.R.; Zsögön, A. Pathways to de novo domestication of crop wild relatives. Plant Physiol. 2022, 188, 1746–1756. [Google Scholar] [CrossRef]
- Stein, J.C.; Yu, Y.; Copetti, D.; Zwickl, D.J.; Zhang, L.; Zhang, C.; Chougule, K.; Gao, D.; Iwata, A.; Goicoechea, J.L.; et al. Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover, and innovation across the genus Oryza. Nat. Genet. 2018, 50, 285–296. [Google Scholar] [CrossRef] [PubMed]
- Tomar, S.; Subba, A.; Bala, M.; Singh, A.K.; Pareek, A.; Singla-Pareek, S.L. Genetic conservation of CBS domain containing protein family in Oryza Species and their association with abiotic stress responses. Int. J. Mol. Sci. 2022, 23, 1687. [Google Scholar] [CrossRef] [PubMed]
- Gauchan, D.; Timsina, K.P.; Gairhe, S.; Timsina, J.; Joshi, K.D. Cereal demand and production projections for 2050: Opportunities for achieving food self-sufficiency in Nepal. In Agriculture, Natural Resources and Food Security; Springer: Cham, Switzerland, 2022; pp. 19–35. [Google Scholar]
- Samal, P.; Babu, S.C.; Mondal, B.; Mishra, S.N. The global rice agriculture towards 2050: An inter-continental perspective. Outlook Agric. 2022, 51, 164–172. [Google Scholar] [CrossRef]
- Satori, D.; Tovar, C.; Faruk, A.; Hammond Hunt, E.; Muller, G.; Cockel, C.; Kühn, N.; Leitch, I.J.; Lulekal, E.; Pereira, L.; et al. Prioritising crop wild relatives to enhance agricultural resilience in sub-Saharan Africa under climate change. Plants People Planet 2022, 4, 269–282. [Google Scholar] [CrossRef]
- Kashyap, A.; Garg, P.; Tanwar, K.; Sharma, J.; Gupta, N.C.; Ha, P.T.T.; Bhattacharya, R.C.; Mason, A.S.; Rao, M. Strategies for utilization of crop wild relatives in plant breeding programs. Theor. Appl. Genet. 2022, 135, 4151–4167. [Google Scholar] [CrossRef]
- Kushwaha, H.R.; Singh, A.K.; Sopory, S.K.; Singla-Pareek, S.L.; Pareek, A. Genome-wide expression analysis of CBS domain containing proteins in Arabidopsis thaliana (L.) and Oryza sativa (L.) reveals their developmental and stress regulation. BMC Genom. 2009, 10, 200. [Google Scholar] [CrossRef]
- Mustafiz, A.; Singh, A.K.; Pareek, A.; Sopory, S.K.; Singla-Pareek, S.L. Genome-wide analysis of rice and Arabidopsis identifies two glyoxalase genes that are highly expressed in abiotic stresses. Funct. Integr. Genom. 2011, 11, 293–305. [Google Scholar] [CrossRef]
- Singh, A.K.; Kumar, R.; Tripathi, A.K.; Gupta, B.K.; Pareek, A.; Singla-Pareek, S.L. Genome-wide investigation and expression analysis of Sodium/Calcium exchanger gene family in rice and Arabidopsis. Rice 2015, 8, 54. [Google Scholar] [CrossRef]
- Kushwaha, H.R.; Joshi, R.; Pareek, A.; Singla-Pareek, S.L. MATH-domain family shows response toward abiotic stress in Arabidopsis and Rice. Front. Plant Sci. 2016, 7, 923. [Google Scholar] [CrossRef] [PubMed]
- Bhowal, B.; Bhattacharjee, A.; Goswami, K.; Sanan-Mishra, N.; Singla-Pareek, S.L.; Kaur, C.; Sopory, S. Serotonin and melatonin biosynthesis in plants: Genome-wide Identification of the genes and their expression reveal a conserved role in stress and development. Int. J. Mol. Sci. 2021, 22, 11034. [Google Scholar] [CrossRef] [PubMed]
- Tello-Ruiz, M.K.; Jaiswal, P.; Ware, D. Gramene: A Resource for comparative analysis of plants genomes and pathways. Methods Mol. Biol. 2022, 2443, 101–131. [Google Scholar] [PubMed]
- Chao, J.; Li, Z.; Sun, Y.; Aluko, O.O.; Wu, X.; Wang, Q.; Liu, G. MG2C: A user-friendly online tool for drawing genetic maps. Mol. Hortic. 2021, 1, 1–4. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, 585–587. [Google Scholar] [CrossRef] [PubMed]
- Eswaramoorthy, S.; Bonanno, J.B.; Burley, S.K.; Swaminathan, S. Mechanism of action of a flavin containing monooxygenase. Proc. Natl. Acad. Sci. USA 2006, 103, 9832–9837. [Google Scholar] [CrossRef]
- Stehr, M.; Diekmann, H.; Smau, L.; Seth, O.; Ghisla, S.; Singh, M.; Macheroux, P. A hydrophobic sequence motif common to N-hydroxylating enzymes. Trends Biochem. Sci. 1998, 23, 56–57. [Google Scholar] [CrossRef]
- Bo, H.; Jinpu, J.; An-Yuan, G.; He, Z.; Jingchu, L.; Ge, G. GSDS 2.0: An upgraded gene features visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar]
- Gough, J.; Karplus, K.; Hughey, R.; Chothia, C. Assignment of homology to genome sequences using a library of Hidden Markov Models that represent all proteins of known structure. J. Mol. Biol. 2001, 313, 903–919. [Google Scholar] [CrossRef]
- Guo, Y.L. Gene family evolution in green plants with emphasis on the origination and evolution of Arabidopsis thaliana genes. Plant J. 2013, 73, 941–951. [Google Scholar] [CrossRef] [PubMed]
- Birchler, J.A.; Yang, H. The multiple fates of gene duplications: Deletion, hypofunctionalization, subfunctionalization, neofunctionalization, dosage balance constraints, and neutral variation. Plant Cell 2022, 34, 2466–2474. [Google Scholar] [CrossRef]
- Hines, R.N.; Hopp, K.A.; Franco, J.; Saeian, K.; Begun, F.P. Alternative processing of the human FMO6 gene renders transcripts incapable of encoding a functional flavin-containing monooxygenase. Mol. Pharmacol. 2002, 62, 320–325. [Google Scholar] [CrossRef] [PubMed]
- Balakirev, E.S.; Ayala, F.J. Pseudogenes: Are they “junk” or functional DNA? Annu. Rev. Genet. 2003, 37, 123–151. [Google Scholar] [CrossRef] [PubMed]
- Lees-Murdock, D.J.; McLoughlin, G.A.; McDaid, J.R.; Quinn, L.M.; O’Doherty, A.; Hiripi, L.; Hack, C.J.; Walsh, C.P. Identification of 11 pseudogenes in the DNA methyltransferase gene family in rodents and humans and implications for the functional loci. Genomics 2004, 84, 193–204. [Google Scholar] [CrossRef] [PubMed]
- Verma, G.; Dhar, Y.V.; Srivastava, D.; Kidwai, M.; Chauhan, P.S.; Bag, S.K.; Asif, M.H.; Chakrabarty, D. Genome-wide analysis of rice dehydrin gene family: Its evolutionary conservedness and expression pattern in response to PEG induced dehydration stress. PLoS ONE 2017, 12, e0176399. [Google Scholar] [CrossRef]
- Chowrasia, S.; Panda, A.K.; Rawal, H.C.; Kaur, H.; Mondal, T.K. Identification of jumonjiC domain-containing gene family among the Oryza species and their expression analysis in FL478, a salt tolerant rice genotype. Plant Physiol. Biochem. 2018, 1, 43–53. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Yang, Y.; Xu, T.; Wang, H.; Feng, D. Genome-wide identification and expression analysis of the TaYUCCA gene family in wheat. Mol. Biol. Rep. 2021, 48, 1269–1279. [Google Scholar] [CrossRef]
- Shiu, S.H.; Karlowski, W.M.; Pan, R.; Tzeng, Y.H.; Mayer, K.F.; Li, W.H. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 2004, 16, 122–1234. [Google Scholar] [CrossRef]
- Freeling, M. Bias in plant gene content following different sorts of duplication: Tandem, whole-genome, segmental, or by transposition. Annu. Rev. Plant Biol. 2009, 60, 433–453. [Google Scholar] [CrossRef] [PubMed]
- Halkier, B.A.; Gershenzon, J. Biology and biochemistry of glucosinolates. Ann. Rev. Plant Biol. 2006, 57, 303–333. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Sun, L.; Wang, D.; Zhang, D.; Cai, W. Genome-wide Identification and Bioinformatics Analysis of NRL Gene Family in Rice. Mol. Plant Breed. 2022, 13, 1–11. [Google Scholar] [CrossRef]
- Guan, J.; Zhang, J.; Gong, D.; Zhang, Z.; Yu, Y.; Luo, G.; Somta, P.; Hu, Z.; Wang, S.; Yuan, X.; et al. Genomic analyses of rice bean landraces reveal adaptation and yield related loci to accelerate breeding. Nat. Commun. 2022, 13, 5707. [Google Scholar] [CrossRef]
- Barrett, L.W.; Fletcher, S.; Wilton, S.D. Regulation of eukaryotic gene expression by the untranslated gene regions and other non-coding elements. Cell. Mol. Life Sci. 2012, 69, 3613–3634. [Google Scholar] [CrossRef]
- Ren, X.Y.; Vorst, O.; Fiers, M.W.; Stiekema, W.J.; Nap, J.P. In plants, highly expressed genes are the least compact. Trends Genet. 2006, 22, 528–532. [Google Scholar] [CrossRef]
- Chung, B.Y.; Simons, C.; Firth, A.E.; Brown, C.M.; Hellens, R.P. Effect of 5’UTR introns on gene expression in Arabidopsis thaliana. BMC Genom. 2006, 7, 120. [Google Scholar] [CrossRef]
- Bicknell, A.A.; Cenik, C.; Chua, H.N.; Roth, F.P.; Moore, M.J. Introns in UTRs: Why we should stop ignoring them. BioEssays 2012, 34, 1025–1034. [Google Scholar] [CrossRef]
- Kertész, S.; Kerényi, Z.; Mérai, Z.; Bartos, I.; Pálfy, T.; Barta, E.; Silhavy, D. Both introns and long 3’-UTRs operate as cis-acting elements to trigger nonsense-mediated decay in plants. Nucleic Acids Res. 2006, 34, 6147–6157. [Google Scholar] [CrossRef]
- Soukup, S.W. Evolution by gene duplication. S. Ohno. Springer-Verlag, New York. 160 pp. Teratology 1970, 9, 250–251. [Google Scholar] [CrossRef]
- Hidenori, T.; Tohru, K. Evolution of multigene families by gene duplication: A haploid model. Genetics 1998, 149, 2147–2158. [Google Scholar]
- Kolkman, J.A.; Stemmer, W.P. Directed evolution of proteins by exon shuffling. Nat. Biotechnol. 2001, 194, 23–28. [Google Scholar] [CrossRef]
- França, G.S.; Cancherini, D.V.; De Souza, S.J. Evolutionary history of exon shuffling. Genetica 2012, 140, 249–257. [Google Scholar] [CrossRef] [PubMed]
- Nasir, A.; Kim, K.M.; Caetano-Anollés, G. Global patterns of protein domain gain and loss in superkingdoms. PLoS Comput. Biol. 2014, 30, e1003452. [Google Scholar] [CrossRef] [PubMed]
- Yafremava, L.S.; Wielgos, M.; Thomas, S.; Nasir, A.; Wang, M.; Mittenthal, J.E.; Caetano-Anollés, G. A general framework of persistence strategies for biological systems helps explain domains of life. Front. Genet. 2013, 25, 4. [Google Scholar] [CrossRef] [PubMed]
- Rairdan, G.J.; Collier, S.M.; Sacco, M.A.; Baldwin, T.T.; Boettrich, T.; Moffett, P. The coiled-coil and nucleotide binding domains of the potato Rx disease resistance protein function in pathogen recognition and signaling. Plant Cell 2008, 20, 739–751. [Google Scholar] [CrossRef]
- Dalio, R.J.; Paschoal, D.; Arena, G.D.; Magalhaes, D.M.; Oliveira, T.S.; Merfa, M.V.; Maximo, H.J.; Machado, M.A. Hypersensitive response: From NLR pathogen recognition to cell death response. Ann. Appl. Biol. 2021, 178, 268–280. [Google Scholar] [CrossRef]
- Förderer, A.; Yu, D.; Li, E.; Chai, J. Resistosomes at the interface of pathogens and plants. Curr. Opin. Plant Biol. 2022, 1, 102212. [Google Scholar] [CrossRef]
- Parker, J.E.; Hessler, G.; Cui, H. A new biochemistry connecting pathogen detection to induced defense in plants. New Phytol. 2022, 234, 819–826. [Google Scholar] [CrossRef]
- Chen, Y.; Li, L.; Tang, B.; Wu, T.; Chen, G.; Xie, Q.; Hu, Z. Silencing of SlMYB55 affects plant flowering and enhances tolerance to drought and salt stress in tomato. Plant Sci. 2022, 316, 111166. [Google Scholar] [CrossRef]
- Chen, Y.; Feng, P.; Zhang, X.; Xie, Q.; Chen, G.; Zhou, S.; Hu, Z. Silencing of SlMYB50 affects tolerance to drought and salt stress in tomato. Plant Physiol. Biochem. 2022, 193, 139–152. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).