Novel Allelic Gene Variations in CmCLAVATA3 (CmCLV3) Were Identified in a Genetic Population of Melon (Cucumis melo L.)
Abstract
1. Introduction
2. Results
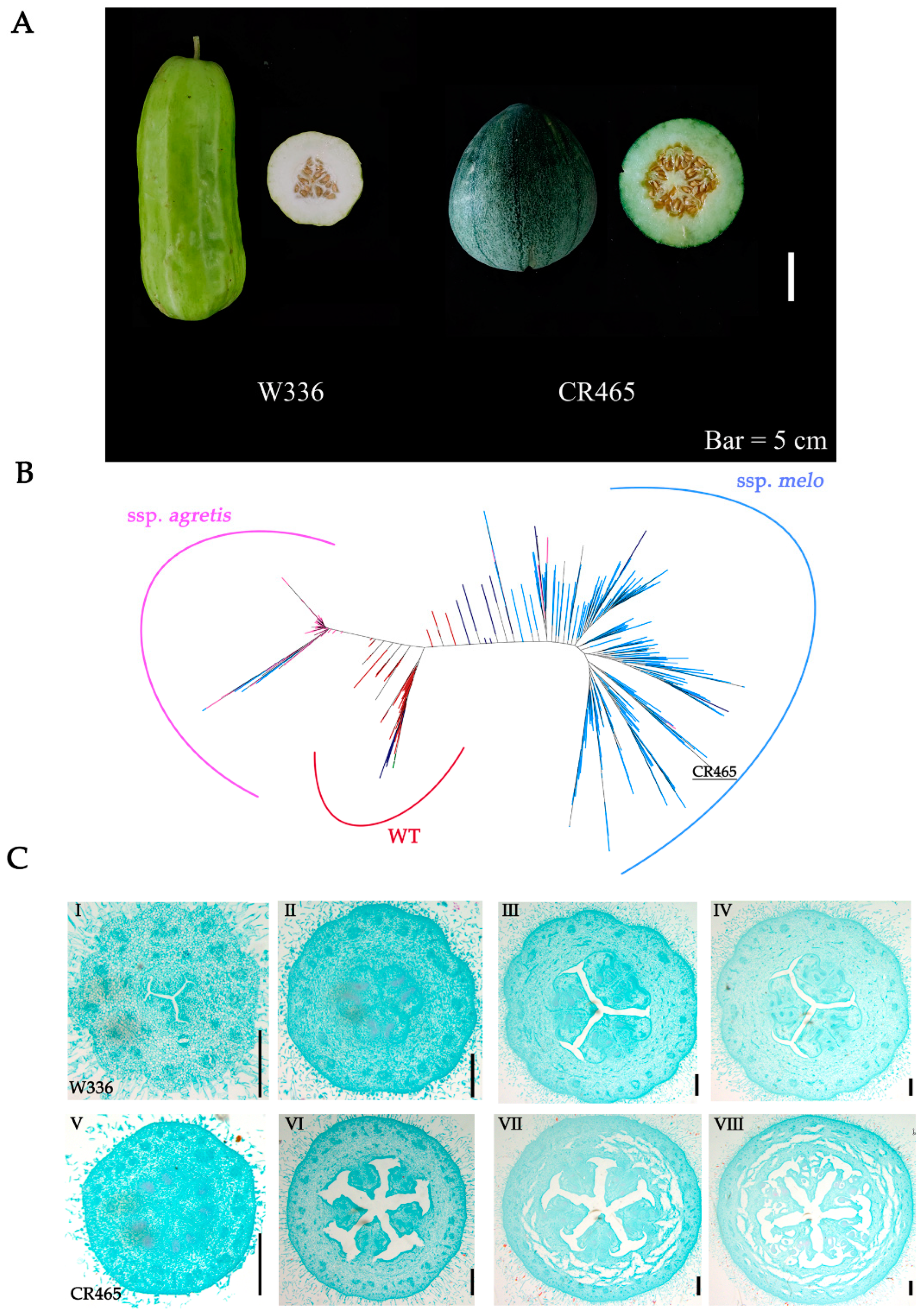
2.1. Genetic and Phenotypic Analysis of Melon Carpel Number
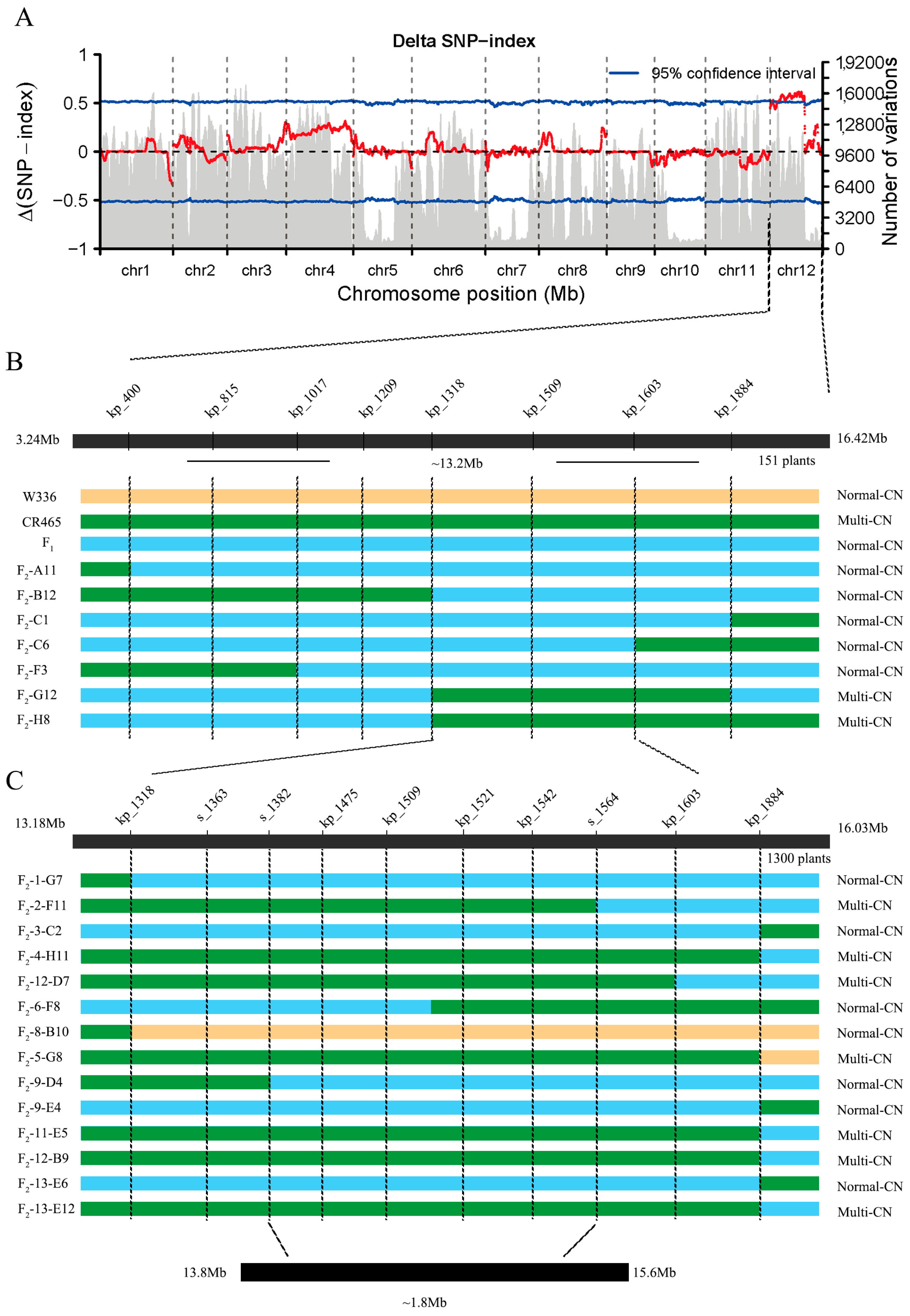
2.2. Localization of the Number of Melon Carpels Gene
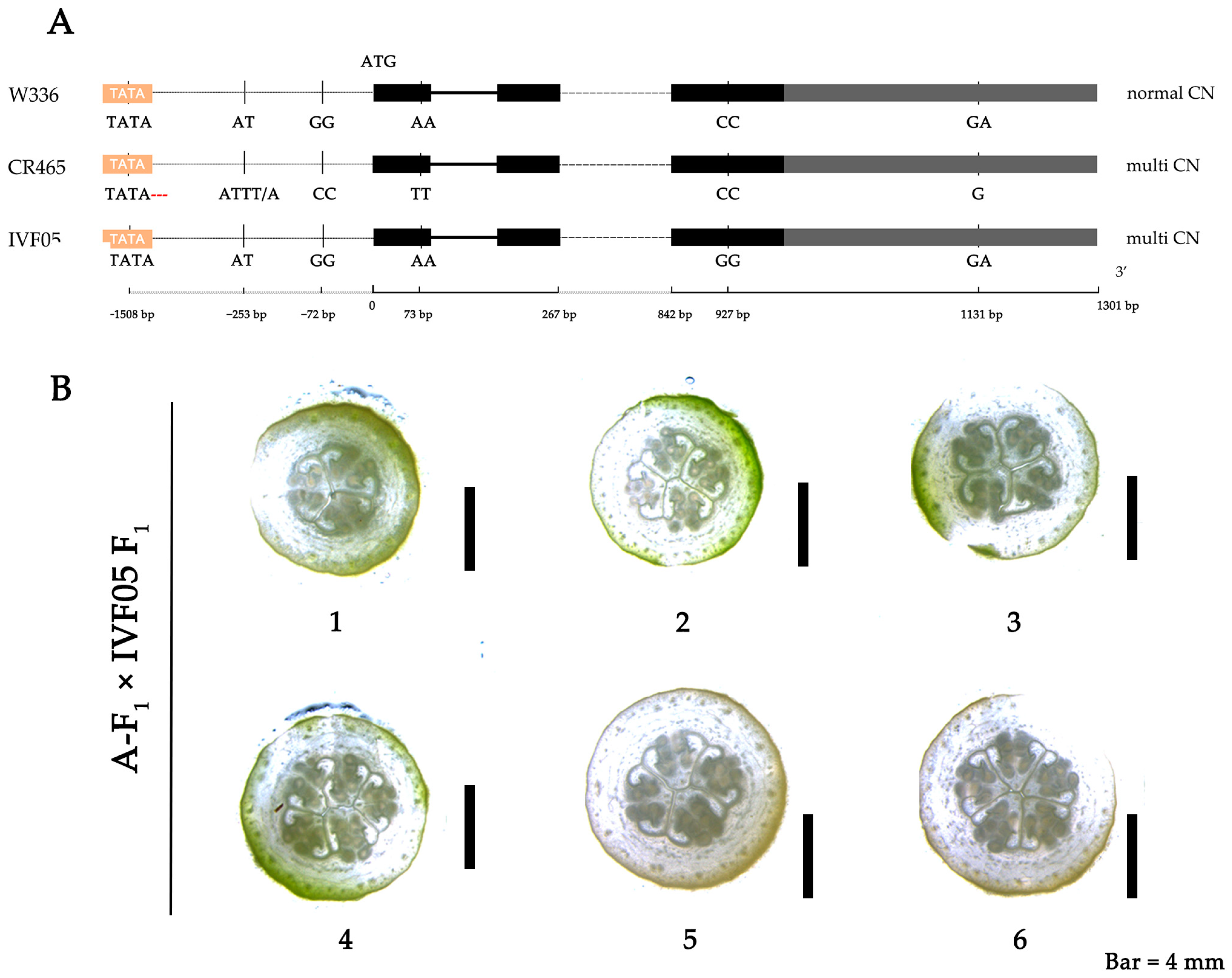
2.3. Identification of Candidate Genes and Study of Mutation Mechanism
2.4. Cis-Trans Test for Polycarpous Candidate Genes
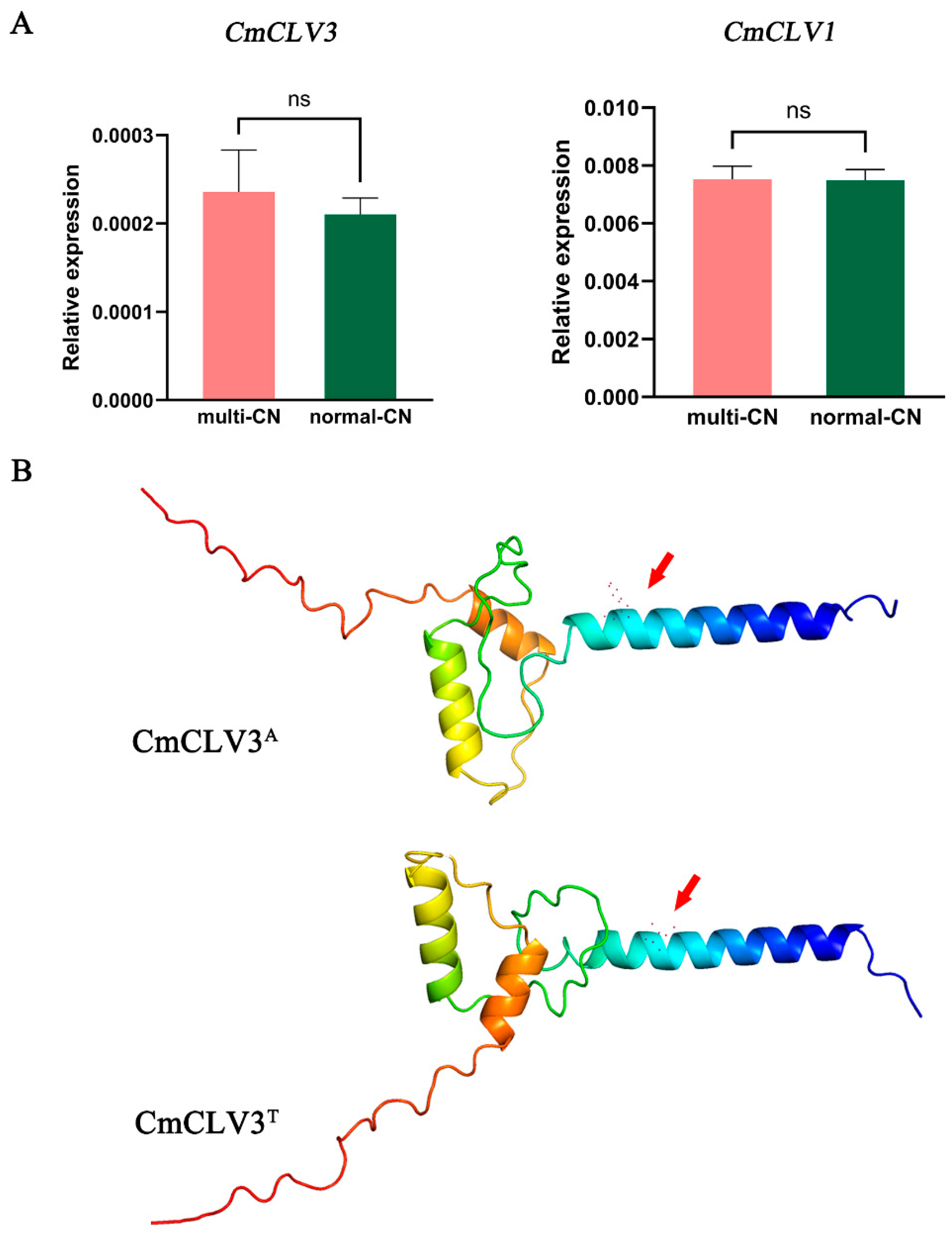
2.5. Analysis of the Expression Patterns of Candidate Genes and Prediction of Protein Structures
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Phenotypic Analysis
4.2. Constructing a Phylogenetic Tree
4.3. BSA Analysis and Initial Mapping
4.4. Fine-Mapping
4.5. Expression Analysis and Prediction of Protein Structures
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Scutt, C.P.; Vinauger-Douard, M.; Fourquin, C.; Finet, C.; Dumas, C. An evolutionary perspective on the regulation of carpel development. J. Exp. Bot. 2006, 57, 2143–2152. [Google Scholar] [CrossRef] [PubMed]
- Chanderbali, A.S.; Yoo, M.-J.; Zahn, L.M.; Brockington, S.F.; Wall, P.K.; Gitzendanner, M.A.; Albert, V.A.; Leebens-Mack, J.; Altman, N.S.; Ma, H. Conservation and canalization of gene expression during angiosperm diversification accompany the origin and evolution of the flower. Proc. Natl. Acad. Sci. USA 2010, 107, 22570–22575. [Google Scholar] [CrossRef] [PubMed]
- Scutt, C.P.; Vinauger-Douard, M.; Fourquin, C.; Ailhas, J.; Kuno, N.; Uchida, K.; Gaude, T.; Furuya, M.; Dumas, C. The identification of candidate genes for a reverse genetic analysis of development and function in the Arabidopsis gynoecium. Plant Physiol. 2003, 132, 653–665. [Google Scholar] [CrossRef] [PubMed]
- Ferrándiz, C.; Fourquin, C.; Prunet, N.; Scutt, C.P.; Sundberg, E.; Trehin, C.; Vialette-Guiraud, A.C.M. Chapter 1—Carpel Development. In Advances in Botanical Research; Kader, J.-C., Delseny, M., Eds.; Academic Press: Cambridge, MA, USA, 2010; pp. 1–73. [Google Scholar]
- Liu, H.; Li, J.; Gong, P.; He, C. The origin and evolution of carpels and fruits from an evo-devo perspective. J. Integr. Plant Biol. 2022, 65, 283–298. [Google Scholar] [CrossRef] [PubMed]
- Abiri, N.; Sinjushin, A.; Tekdal, D.; Cetiner, S. Evaluation of the possible contribution of various regulatory genes to determination of carpel number as a potential mechanism for optimal agricultural yield. Int. J. Mol. Sci. 2022, 23, 9723. [Google Scholar] [CrossRef] [PubMed]
- Tanksley, S.D. The genetic, developmental, and molecular bases of fruit size and shape variation in tomato. Plant Cell 2004, 16, S181–S189. [Google Scholar] [CrossRef] [PubMed]
- Barrero, L.S.; Cong, B.; Wu, F.; Tanksley, S.D. Developmental characterization of the fasciated locus and mapping of Arabidopsis candidate genes involved in the control of floral meristem size and carpel number in tomato. Genome 2006, 49, 991–1006. [Google Scholar] [CrossRef]
- Li, S.; Pan, Y.; Wen, C.; Li, Y.; Liu, X.; Zhang, X.; Behera, T.K.; Xing, G.; Weng, Y. Integrated analysis in bi-parental and natural populations reveals CsCLAVATA3 (CsCLV3) underlying carpel number variations in cucumber. Theor. Appl. Genet. 2016, 129, 1007–1022. [Google Scholar] [CrossRef]
- Barrero, L.S.; Tanksley, S.D. Evaluating the genetic basis of multiple-locule fruit in a broad cross section of tomato cultivars. Theor. Appl. Genet. 2004, 109, 669–679. [Google Scholar] [CrossRef]
- Soyars, C.L.; James, S.R.; Nimchuk, Z.L. Ready, aim, shoot: Stem cell regulation of the shoot apical meristem. Curr. Opin. Plant Biol. 2016, 29, 163–168. [Google Scholar] [CrossRef]
- Hu, C.; Zhu, Y.; Cui, Y.; Cheng, K.; Liang, W.; Wei, Z.; Zhu, M.; Yin, H.; Zeng, L.; Xiao, Y.; et al. A group of receptor kinases are essential for CLAVATA signalling to maintain stem cell homeostasis. Nat. Plants 2018, 4, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, J.C.; Brand, U.; Running, M.P.; Simon, R.; Meyerowitz, E.M. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science 1999, 283, 1911–1914. [Google Scholar] [CrossRef]
- Muller, R.; Borghi, L.; Kwiatkowska, D.; Laufs, P.; Simon, R. Dynamic and compensatory responses of Arabidopsis shoot and floral meristems to CLV3 signaling. Plant Cell 2006, 18, 1188–1198. [Google Scholar] [CrossRef] [PubMed]
- Ohyama, K.; Shinohara, H.; Ogawa-Ohnishi, M.; Matsubayashi, Y. A glycopeptide regulating stem cell fate in Arabidopsis thaliana. Nat. Chem. Biol. 2009, 5, 578–580. [Google Scholar] [CrossRef]
- Mayer, K.F.; Schoof, H.; Haecker, A.; Lenhard, M.; Jürgens, G.; Laux, T. Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem. Cell 1998, 95, 805–815. [Google Scholar] [CrossRef] [PubMed]
- Clark, S.E. Cell signalling at the shoot meristem. Nat. Rev. Mol. Cell Biol. 2001, 2, 276–284. [Google Scholar] [CrossRef] [PubMed]
- Yadava, S.K.; Paritosh, K.; Panjabi-Massand, P.; Gupta, V.; Chandra, A.; Sodhi, Y.; Pradhan, A.K.; Pental, D. Tetralocular ovary and high silique width in yellow sarson lines of Brassica rapa (subspecies trilocularis) are due to a mutation in Bra034340 gene, a homologue of CLAVATA3 in Arabidopsis. Theor. Appl. Genet. 2014, 127, 2359–2369. [Google Scholar] [CrossRef]
- Nimchuk, Z.L.; Zhou, Y.; Tarr, P.T.; Peterson, B.A.; Meyerowitz, E.M. Plant stem cell maintenance by transcriptional cross-regulation of related receptor kinases. Development 2015, 142, 1043–1049. [Google Scholar] [CrossRef]
- Guo, Y.; Han, L.; Hymes, M.; Denver, R.; Clark, S.E. CLAVATA2 forms a distinct CLE-binding receptor complex regulating Arabidopsis stem cell specification. Plant J. 2010, 63, 889–900. [Google Scholar] [CrossRef]
- Nimchuk, Z.L.; Tarr, P.T.; Meyerowitz, E.M. An evolutionarily conserved pseudokinase mediates stem cell production in plants. Plant Cell 2011, 23, 851–854. [Google Scholar] [CrossRef]
- Lippman, Z.; Tanksley, S.D. Dissecting the Genetic Pathway to Extreme Fruit Size in Tomato Using a Cross Between the Small-Fruited Wild Species Lycopersicon pimpinellifolium and L. esculentum var. Giant Heirloom. Genetics 2001, 158, 413–422. [Google Scholar]
- Munos, S.; Ranc, N.; Botton, E.; Berard, A.; Rolland, S.; Duffe, P.; Carretero, Y.; Le Paslier, M.C.; Delalande, C.; Bouzayen, M.; et al. Increase in tomato locule number is controlled by two single-nucleotide polymorphisms located near WUSCHEL. Plant Physiol. 2011, 156, 2244–2254. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Qi, M.; Sun, M.; Liu, Y.; Liu, Y.; Xu, T.; Li, Y.; Li, T. Tomato transcription factor SlWUS plays an important role in tomato flower and locule development. Front. Plant Sci. 2017, 8, 246807. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Gao, P.; Zhu, Q.; Zhu, Z.; Liu, H.; Wang, X.; Weng, Y.; Gao, M.; Luan, F. Resequencing of 297 melon accessions reveals the genomic history of improvement and loci related to fruit traits in melon. Plant Biotechnol. J. 2020, 18, 2545–2558. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Diao, Q.; Chen, Y.; Jin, H.; Zhang, Y.; Zhang, H. Development of KASP markers and identification of a QTL underlying powdery mildew resistance in melon (Cucumis melo L.) by bulked segregant analysis and RNA-seq. Front. Plant Sci. 2021, 11, 593207. [Google Scholar] [CrossRef] [PubMed]
- Adedze, Y.M.N.; Lu, X.; Fan, W.; Zhang, W.; Yang, X.; Deng, Z.; Alam, M.A.; Xu, G.; Zhang, L.; Li, W. Development of PCR-based markers associated with powdery mildew resistance using bulked segregant analysis (BSA-seq) in melon. Czech. J. Genet. Plant Breed. 2023, 60, 25–33. [Google Scholar] [CrossRef]
- Adedze, Y.M.N.; Lu, X.; Xia, Y.; Sun, Q.; Nchongboh, C.G.; Alam, M.A.; Liu, M.; Yang, X.; Zhang, W.; Deng, Z.; et al. Agarose-resolvable InDel markers based on whole genome re-sequencing in cucumber. Sci. Rep. 2021, 11, 3872. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, Y.; Luan, F.; Zhang, X.; Zhao, J.; Yang, Z.; Liu, S. Biparental genetic mapping reveals that CmCLAVATA3 (CmCLV3) is responsible for the variation in carpel number in melon (Cucumis melo L.). Theor. Appl. Genet. 2022, 135, 1909–1921. [Google Scholar] [CrossRef]
- Zhao, G.; Lian, Q.; Zhang, Z.; Fu, Q.; He, Y.; Ma, S.; Ruggieri, V.; Monforte, A.J.; Wang, P.; Julca, I.; et al. A comprehensive genome variation map of melon identifies multiple domestication events and loci influencing agronomic traits. Nat. Genet. 2019, 51, 1607–1615. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Yong, B.; Zhu, W.; Wei, S.; Li, B.; Wang, Y.; Xu, N.; Lu, J.; Chen, Q.; He, C. Parallel selection of loss-of-function alleles of Pdh1 orthologous genes in warm-season legumes for pod indehiscence and plasticity is related to precipitation. New Phytol. 2023, 240, 863–879. [Google Scholar] [CrossRef]
- Qiao, Q.; Edger, P.P.; Xue, L.; Qiong, L.; Lu, J.; Zhang, Y.; Cao, Q.; Yocca, A.E.; Platts, A.E.; Knapp, S.J.; et al. Evolutionary history and pan-genome dynamics of strawberry (Fragaria spp.). Proc. Natl. Acad. Sci. USA 2021, 118, e2105431118. [Google Scholar] [CrossRef] [PubMed]
- AWalker, R.; Lee, E.; Bogs, J.; McDavid, D.A.; Thomas, M.R.; Robinson, S.P. White grapes arose through the mutation of two similar and adjacent regulatory genes. Plant J. 2007, 49, 772–785. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Duan, S.; Xia, Q.; Liang, Z.; Dong, X.; Margaryan, K.; Musayev, M.; Goryslavets, S.; Zdunic, G.; Bert, P.F.; et al. Dual domestications and origin of traits in grapevine evolution. Science 2023, 379, 892–901. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4326. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Mas, J.; Benjak, A.; Sanseverino, W.; Bourgeois, M.; Mir, G.; Gonzalez, V.M.; Henaff, E.; Camara, F.; Cozzuto, L.; Lowy, E.; et al. The genome of melon (Cucumis melo L.). Proc. Natl. Acad. Sci. USA 2012, 109, 11872–11877. [Google Scholar] [CrossRef] [PubMed]
- Andrews, S.; Krueger, F.; Segonds-Pichon, A.; Biggins, L.; Krueger, C.; Wingett, S. FastQC, A quality control tool for high throughput sequence data, 2010, 370. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 2 March 2023).
- Li, H. Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 2014, 30, 2843–2851. [Google Scholar] [CrossRef] [PubMed]
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; Del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; et al. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 2013, 43, 11.10.1–11.10.33. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
| Generations | Plants | Normal-CN (Mean) | Multi-CN (Mean) | Expected Ratio | Actual Ratio | χ2 | p Value |
|---|---|---|---|---|---|---|---|
| W336(P1) | 14 | 14 (3.05) | 0 | ||||
| CR465(P2) | 33 | 0 | 33 (4.49) | ||||
| F1 | 17 | 17 (3.33) | 0 | ||||
| F2-2022 | 178 | 142 (3.18) | 41 (4.31) | 3:1 | 3.46:1 | 0.667 | 0.412 |
| Sample Name | CmCLV3H1 | CmCLV3H2 | Carpel Number (Mean) |
|---|---|---|---|
| A-F1 × ivf05 F1-1 | C/G | A/A | normal |
| A-F1 × ivf05 F1-2 | C/G | A/T | multi |
| A-F1 × ivf05 F1-3 | C/G | A/T | multi |
| A-F1 × ivf05 F1-4 | C/G | A/T | multi |
| A-F1 × ivf05 F1-5 | C/G | A/T | multi |
| A-F1 × ivf05 F1-6 | C/G | A/T | multi |
| A-F1 × ivf05 F1-7 | C/G | A/T | multi |
| B-F1 × ivf05 F1-1 | C/G | A/A | normal |
| B-F1 × ivf05 F1-2 | C/G | A/T | multi |
| B-F1 × ivf05 F1-3 | C/G | A/A | normal |
| B-F1 × ivf05 F1-4 | C/G | A/T | multi |
| B-F1 × ivf05 F1-5 | C/G | A/T | multi |
| B-F1 × ivf05 F1-6 | C/G | A/T | multi |
| B-F1 × ivf05 F1-7 | C/G | A/A | normal |
| B-F1 × ivf05 F1-8 | C/G | A/T | multi |
| B-F1 × ivf05 F1-9 | C/G | A/A | normal |
| B-F1 × ivf05 F1-10 | C/G | A/A | normal |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, H.; Jia, Y.; Chen, X.; Jiang, N.; Zhang, Z.; Chai, S. Novel Allelic Gene Variations in CmCLAVATA3 (CmCLV3) Were Identified in a Genetic Population of Melon (Cucumis melo L.). Int. J. Mol. Sci. 2024, 25, 6011. https://doi.org/10.3390/ijms25116011
Wu H, Jia Y, Chen X, Jiang N, Zhang Z, Chai S. Novel Allelic Gene Variations in CmCLAVATA3 (CmCLV3) Were Identified in a Genetic Population of Melon (Cucumis melo L.). International Journal of Molecular Sciences. 2024; 25(11):6011. https://doi.org/10.3390/ijms25116011
Chicago/Turabian StyleWu, Hangyu, Yue Jia, Xinxiu Chen, Naiyu Jiang, Zhonghua Zhang, and Sen Chai. 2024. "Novel Allelic Gene Variations in CmCLAVATA3 (CmCLV3) Were Identified in a Genetic Population of Melon (Cucumis melo L.)" International Journal of Molecular Sciences 25, no. 11: 6011. https://doi.org/10.3390/ijms25116011
APA StyleWu, H., Jia, Y., Chen, X., Jiang, N., Zhang, Z., & Chai, S. (2024). Novel Allelic Gene Variations in CmCLAVATA3 (CmCLV3) Were Identified in a Genetic Population of Melon (Cucumis melo L.). International Journal of Molecular Sciences, 25(11), 6011. https://doi.org/10.3390/ijms25116011






