Genome-Based Identification of the Dof Gene Family in Three Cymbidium Species and Their Responses to Heat Stress in Cymbidium goeringii
Abstract
1. Introduction
2. Results
2.1. Identification and Protein Features of Dofs
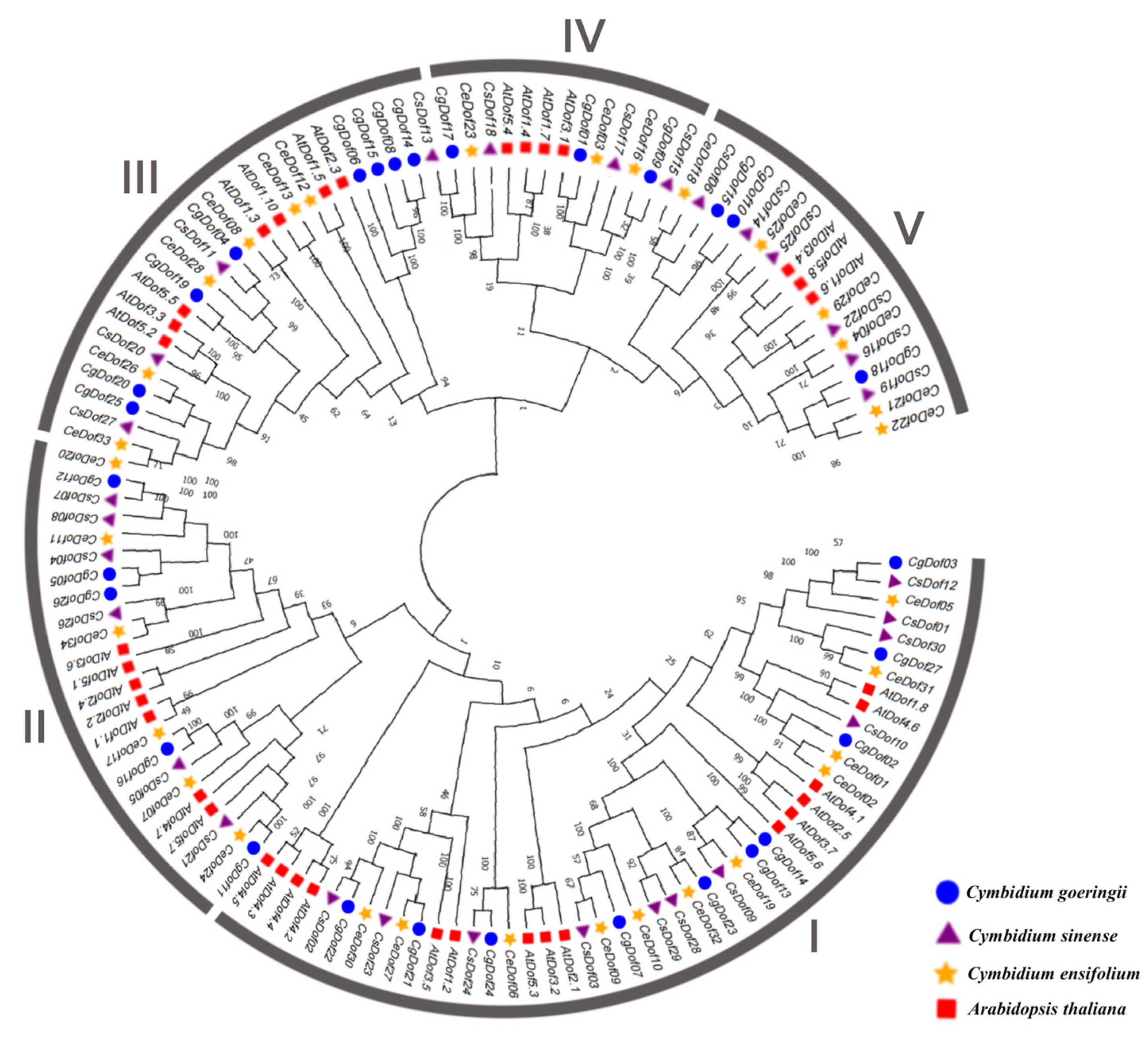
2.2. Phylogeny and Classification of Dofs
2.3. Structure and Motif Analysis of Dofs
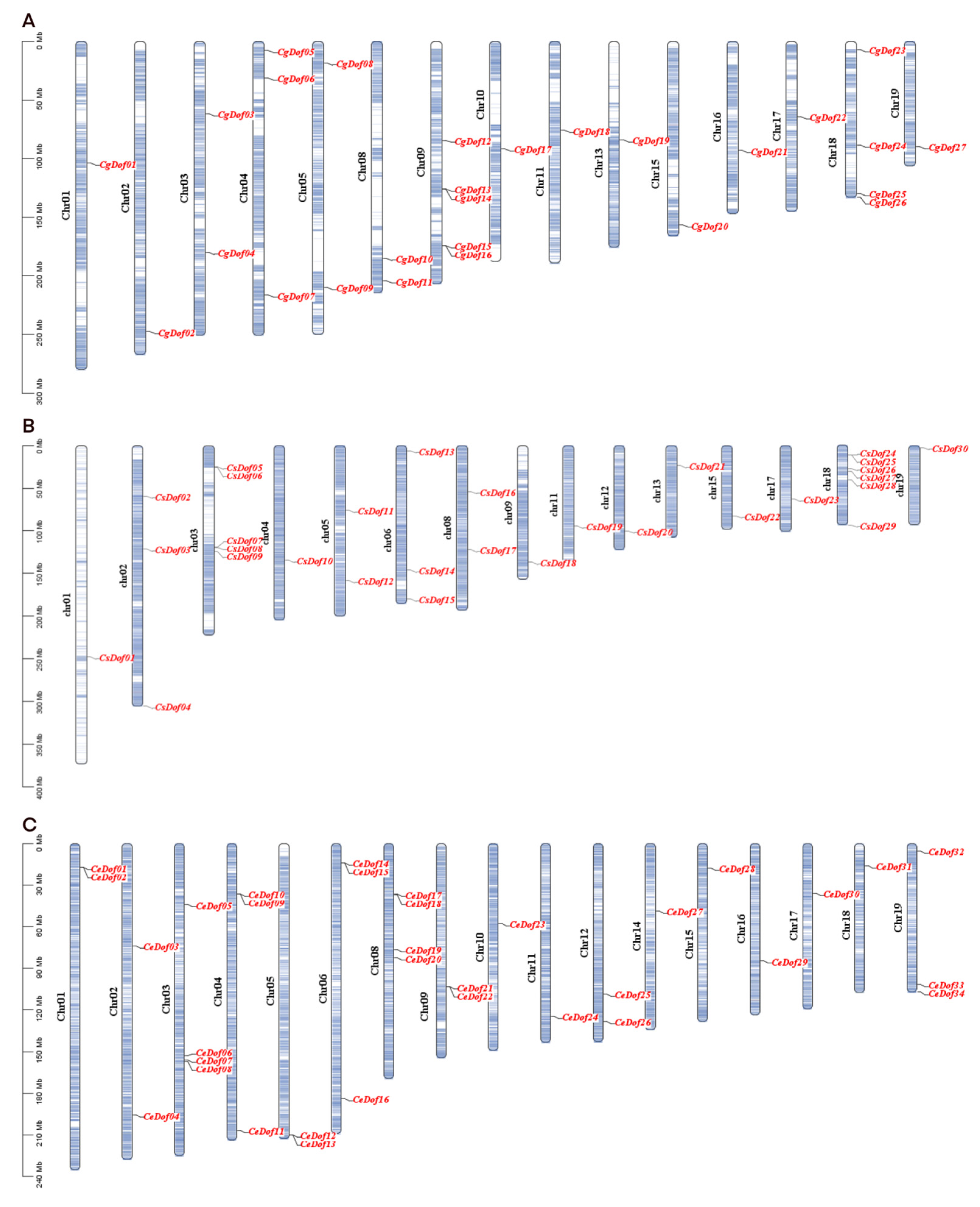
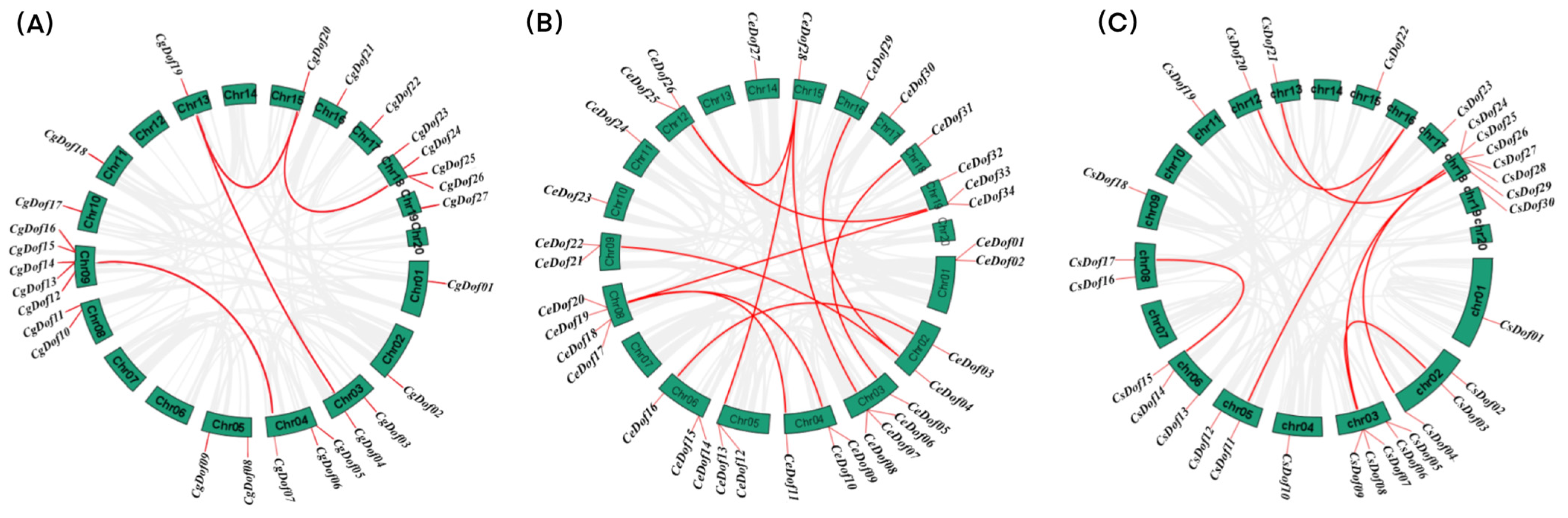
2.4. Chromosomal Localization and Collinearity of Dof Genes
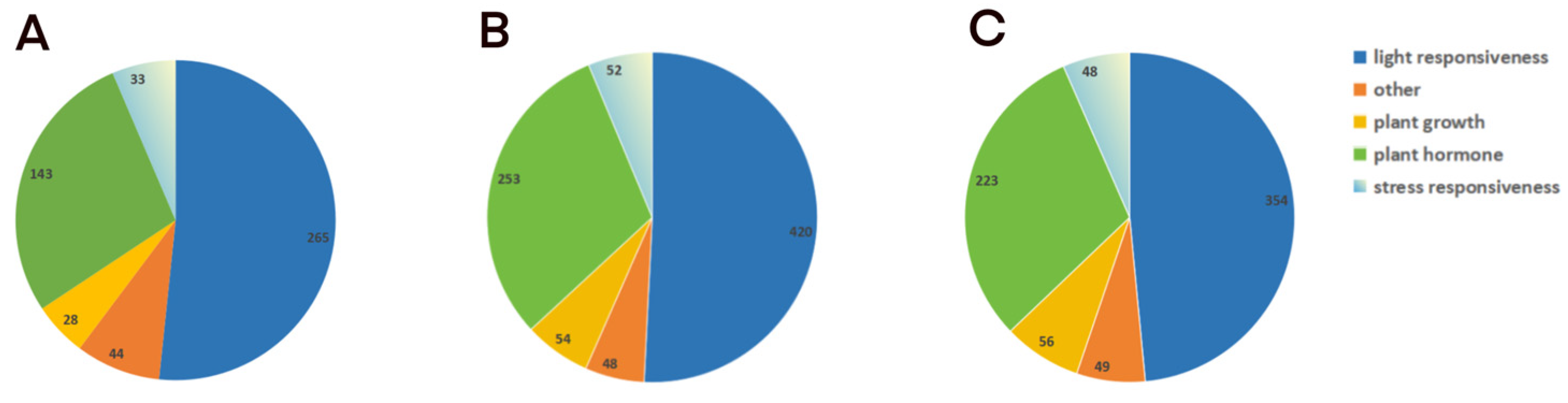
2.5. Promoter Analysis of Dofs
2.6. Prediction of Dof Protein Structure
2.7. qRT-PCR Analysis of Dof Genes
3. Discussion
4. Materials and Methods
4.1. Data Source and Plant Materials
4.2. Identification and Physicochemical Properties of Dof Genes
4.3. Phylogenetic Analysis of Dofs
4.4. Gene Structures and Conserved Motif Analysis of Dofs
4.5. Collinearity and Chromosomal Localization of Dofs
4.6. Cis-Element Analysis of Dofs
4.7. Protein Structure Prediction
4.8. Analysis of Expression and RT-qPCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Floris, M.; Mahgoub, H.; Lanet, E.; Robaglia, C.; Menand, B. Post-transcriptional regulation of gene expression in plants during abiotic stress. Int. J. Mol. Sci. 2009, 10, 3168–3185. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Malviya, N.; Kushwaha, H.; Nasim, J.; Bisht, N.C.; Singh, V.K.; Yadav, D. Insights into structural and functional diversity of Dof (DNA binding with one finger) transcription factor. Planta 2015, 241, 549–562. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S. The Dof family of plant transcription factors. Trends Plant Sci. 2002, 7, 555–560. [Google Scholar] [CrossRef] [PubMed]
- De Paolis, A.; Sabatini, S.; De Pascalis, L.; Costantino, P.; Capone, I. A rolB regulatory factor belongs to a new class of single zinc finger plant proteins. Plant J. Cell Mol. Biol. 1996, 10, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Umemura, Y.; Ishiduka, T.; Yamamoto, R.; Esaka, M. The Dof domain, a zinc finger DNA-binding domain conserved only in higher plants, truly functions as a Cys2/Cys2 Zn finger domain. Plant J. Cell Mol. Biol. 2004, 37, 741–749. [Google Scholar] [CrossRef] [PubMed]
- Imaizumi, T.; Schultz, T.F.; Harmon, F.G.; Ho, L.A.; Kay, S.A. FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science 2005, 309, 293–297. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S. A novel DNA-binding domain that may form a single zinc finger motif. Nucleic Acids Res. 1995, 23, 3403–3410. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S. Dof DNA-binding domains of plant transcription factors contribute to multiple protein-protein interactions. Eur. J. Biochem. 1997, 250, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S. Dof domain proteins: Plant-specific transcription factors associated with diverse phenomena unique to plants. Plant Cell Physiol. 2004, 45, 386–391. [Google Scholar] [CrossRef]
- Cavalar, M.; Möller, C.; Offermann, S.; Krohn, N.M.; Grasser, K.D.; Peterhänsel, C. The interaction of DOF transcription factors with nucleosomes depends on the positioning of the binding site and is facilitated by maize HMGB5. Biochemistry 2003, 42, 2149–2157. [Google Scholar] [CrossRef]
- Yanagisawa, S.; Izui, K. Molecular cloning of two DNA-binding proteins of maize that are structurally different but interact with the same sequence motif. J. Biol. Chem. 1993, 268, 16028–16036. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Risueno, M.A.; Martínez, M.; Vicente-Carbajosa, J.; Carbonero, P. The family of DOF transcription factors: From green unicellular algae to vascular plants. Mol. Genet. Genom. MGG 2007, 277, 379–390. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Li, M.Y.; Wang, F.; Tang, J.; Xiong, A.S. Genome-wide analysis of Dof family transcription factors and their responses to abiotic stresses in Chinese cabbage. BMC Genom. 2015, 16, 33. [Google Scholar] [CrossRef] [PubMed]
- Lijavetzky, D.; Carbonero, P.; Vicente-Carbajosa, J. Genome-wide comparative phylogenetic analysis of the rice and Arabidopsis Dof gene families. BMC Evol. Biol. 2003, 3, 17. [Google Scholar] [CrossRef]
- Yang, X.; Tuskan, G.A.; Cheng, M.Z. Divergence of the Dof gene families in poplar, Arabidopsis, and rice suggests multiple modes of gene evolution after duplication. Plant Physiol. 2006, 142, 820–830. [Google Scholar] [CrossRef] [PubMed]
- Cai, K.; Xie, X.; Han, L.; Chen, J.; Zhang, J.; Yuan, H.; Shen, J.; Ren, Y.; Zhao, X. Identification and functional analysis of the DOF gene family in Populus simonii: Implications for development and stress response. Front. Plant Sci. 2024, 15, 1412175. [Google Scholar] [CrossRef]
- Gu, F.; Zhang, W.; Wang, T.; He, X.; Chen, N.; Zhang, Y.; Song, C. Identification of Dof transcription factors in Dendrobium huoshanense and expression pattern under abiotic stresses. Front. Genet. 2024, 15, 1394790. [Google Scholar] [CrossRef] [PubMed]
- Alam, O.; Khan, L.U.; Khan, A.; Salmen, S.H.; Ansari, M.J.; Mehwish, F.; Ahmad, M.; Zaman, Q.U.; Wang, H.F. Functional characterisation of Dof gene family and expression analysis under abiotic stresses and melatonin-mediated tolerance in pitaya (Selenicereus undatus). Funct. Plant Biol. FPB 2024, 51, FP23269. [Google Scholar] [CrossRef]
- Jin, X.; Wang, Z.; Ai, Q.; Li, X.; Yang, J.; Zhang, N.; Si, H. DNA-Binding with One Finger (Dof) Transcription Factor Gene Family Study Reveals Differential Stress-Responsive Transcription Factors in Contrasting Drought Tolerance Potato Species. Int. J. Mol. Sci. 2024, 25, 3488. [Google Scholar] [CrossRef]
- da Silva, D.C.; da Silveira Falavigna, V.; Fasoli, M.; Buffon, V.; Porto, D.D.; Pappas, G.J., Jr.; Pezzotti, M.; Pasquali, G.; Revers, L.F. Transcriptome analyses of the Dof-like gene family in grapevine reveal its involvement in berry, flower and seed development. Hortic. Res. 2016, 3, 16042. [Google Scholar] [CrossRef]
- Cao, L.; Ye, F.; Fahim, A.M.; Ma, C.; Pang, Y.; Zhang, X.; Zhang, Q.; Lu, X. Transcription factor ZmDof22 enhances drought tolerance by regulating stomatal movement and antioxidant enzymes activities in maize (Zea mays L.). Theor. Appl. Genet. 2024, 137, 132. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Xu, Y.; Gui, J.; Huang, Y.; Ma, F.; Wu, W.; Han, T.; Qiu, W.; Yang, L.; Song, S. Characterization of Dof Transcription Factors and the Heat-Tolerant Function of PeDof-11 in Passion Fruit (Passiflora edulis). Int. J. Mol. Sci. 2023, 24, 12091. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.B.; Tang, M.; Li, H.T.; Zhang, Z.R.; Li, D.Z. Complete chloroplast genome of the genus Cymbidium: Lights into the species identification, phylogenetic implications and population genetic analyses. BMC Evol. Biol. 2013, 13, 84. [Google Scholar] [CrossRef] [PubMed]
- Li, J.W.; Chen, X.D.; Hu, X.Y.; Ma, L.; Zhang, S.B. Comparative physiological and proteomic analyses reveal different adaptive strategies by Cymbidium sinense and C. tracyanum to drought. Planta 2018, 247, 69–97. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Liu, D.K.; Wang, Q.Q.; Ke, S.; Li, Y.; Zhang, D.; Zheng, Q.; Zhang, C.; Liu, Z.J.; Lan, S. Genome-wide identification and expression analysis of the GRAS gene family in Dendrobium chrysotoxum. Front. Plant Sci. 2022, 13, 1058287. [Google Scholar] [CrossRef] [PubMed]
- Enany, S. Structural and functional analysis of hypothetical and conserved proteins of Clostridium tetani. J. Infect. Public Health 2014, 7, 296–307. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef]
- Liang, T.; Jiang, C.; Yuan, J.; Othman, Y.; Xie, X.Q.; Feng, Z. Differential performance of RoseTTAFold in antibody modeling. Brief. Bioinform. 2022, 23, bbac152. [Google Scholar] [CrossRef]
- Ai, Y.; Li, Z.; Sun, W.H.; Chen, J.; Zhang, D.; Ma, L.; Zhang, Q.H.; Chen, M.K.; Zheng, Q.D.; Liu, J.F.; et al. The Cymbidium genome reveals the evolution of unique morphological traits. Hortic. Res. 2021, 8, 255. [Google Scholar] [CrossRef]
- Liu, D.K.; Zhang, C.; Zhao, X.; Ke, S.; Li, Y.; Zhang, D.; Zheng, Q.; Li, M.H.; Lan, S.; Liu, Z.J. Genome-wide analysis of the TCP gene family and their expression pattern in Cymbidium goeringii. Front. Plant Sci. 2022, 13, 1068969. [Google Scholar] [CrossRef]
- Su, S.; Shao, X.; Zhu, C.; Xu, J.; Lu, H.; Tang, Y.; Jiao, K.; Guo, W.; Xiao, W.; Liu, Z.; et al. Transcriptome-Wide Analysis Reveals the Origin of Peloria in Chinese Cymbidium (Cymbidium sinense). Plant Cell Physiol. 2018, 59, 2064–2074. [Google Scholar] [CrossRef]
- Li, X.; Jin, F.; Jin, L.; Jackson, A.; Ma, X.; Shu, X.; Wu, D.; Jin, G. Characterization and comparative profiling of the small RNA transcriptomes in two phases of flowering in Cymbidium ensifolium. BMC Genom. 2015, 16, 622. [Google Scholar] [CrossRef]
- Hou, Q.; Yu, R.; Shang, C.; Deng, H.; Wen, Z.; Qiu, Z.; Qiao, G. Molecular characterization and evolutionary relationships of DOFs in four cherry species and functional analysis in sweet cherry. Int. J. Biol. Macromol. 2024, 263 Pt 2, 130346. [Google Scholar] [CrossRef]
- Yang, L.; Min, X.; Wei, Z.; Liu, N.; Li, J.; Zhang, Y.; Yang, Y. Genome-Wide Identification and Expression Analysis of the Dof Transcription Factor in Annual Alfalfa Medicago polymorpha. Plants 2023, 12, 1831. [Google Scholar] [CrossRef]
- Zhang, Q.; Zhong, S.; Dong, Q.; Yang, H.; Yang, H.; Tan, F.; Chen, C.; Ren, T.; Shen, J.; Cao, G.; et al. Identification of Photoperiod- and Phytohormone-Responsive DNA-Binding One Zinc Finger (Dof) Transcription Factors in Akebia trifoliata via Genome-Wide Expression Analysis. Int. J. Mol. Sci. 2023, 24, 4973. [Google Scholar] [CrossRef]
- Song, H.; Ji, X.; Wang, M.; Li, J.; Wang, X.; Meng, L.; Wei, P.; Xu, H.; Niu, T.; Liu, A. Genome-wide identification and expression analysis of the Dof gene family reveals their involvement in hormone response and abiotic stresses in sunflower (Helianthus annuus L.). Gene 2024, 910, 148336. [Google Scholar] [CrossRef] [PubMed]
- Rashid, A.; Khajuria, M.; Ali, V.; Faiz, S.; Jamwal, S.; Vyas, D. Genome-wide identification and expression profiling of WRKY family suggest their potential role in cannabinoid regulation in Cannabis sativa L. Ind. Crops Prod. 2023, 206, 117706. [Google Scholar] [CrossRef]
- Grimplet, J.; Agudelo-Romero, P.; Teixeira, R.T.; Martinez-Zapater, J.M.; Fortes, A.M. Structural and Functional Analysis of the GRAS Gene Family in Grapevine Indicates a Role of GRAS Proteins in the Control of Development and Stress Responses. Front. Plant Sci. 2016, 7, 353. [Google Scholar] [CrossRef]
- Le Hir, R.; Bellini, C. The plant-specific dof transcription factors family: New players involved in vascular system development and functioning in Arabidopsis. Front. Plant Sci. 2013, 4, 164. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, N.; Deng, X.; Liu, D.; Li, M.; Cui, D.; Hu, Y.; Yan, Y. Genome-wide analysis of wheat DNA-binding with one finger (Dof) transcription factor genes: Evolutionary characteristics and diverse abiotic stress responses. BMC Genom. 2020, 21, 276. [Google Scholar] [CrossRef]
- Yu, H.; Ma, Y.; Lu, Y.; Yue, J.; Ming, R. Expression profiling of the Dof gene family under abiotic stresses in spinach. Sci. Rep. 2021, 11, 14429. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Cai, K.; Pei, X.; Li, Y.; Hu, Y.; Meng, F.; Song, X.; Tigabu, M.; Ding, C.; Zhao, X. Genome-Wide Identification of NAC Transcription Factor Family in Juglans mandshurica and Their Expression Analysis during the Fruit Development and Ripening. Int. J. Mol. Sci. 2021, 22, 12414. [Google Scholar] [CrossRef] [PubMed]
- Koralewski, T.E.; Krutovsky, K.V. Evolution of exon-intron structure and alternative splicing. PLoS ONE 2011, 6, e18055. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Gracia, P.; Roque, E.; Medina, M.; López-Martín, M.J.; Cañas, L.A.; Beltrán, J.P.; Gómez-Mena, C. The DOF Transcription Factor SlDOF10 Regulates Vascular Tissue Formation During Ovary Development in Tomato. Front. Plant Sci. 2019, 10, 216. [Google Scholar] [CrossRef]
- Wang, H.; Zhao, S.; Gao, Y.; Yang, J. Characterization of Dof Transcription Factors and Their Responses to Osmotic Stress in Poplar (Populus trichocarpa). PLoS ONE 2017, 12, e0170210. [Google Scholar] [CrossRef]
- Wen, C.L.; Cheng, Q.; Zhao, L.; Mao, A.; Yang, J.; Yu, S.; Weng, Y.; Xu, Y. Identification and characterisation of Dof transcription factors in the cucumber genome. Sci. Rep. 2016, 6, 23072. [Google Scholar] [CrossRef]
- Malviya, N.; Gupta, S.; Singh, V.K.; Yadav, M.K.; Bisht, N.C.; Sarangi, B.K.; Yadav, D. Genome wide in silico characterization of Dof gene families of pigeonpea (Cajanus cajan (L) Millsp.). Mol. Biol. Rep. 2015, 42, 535–552. [Google Scholar] [CrossRef]
- Sang, Y.; Liu, Q.; Lee, J.; Ma, W.; McVey, D.S.; Blecha, F. Expansion of amphibian intronless interferons revises the paradigm for interferon evolution and functional diversity. Sci. Rep. 2016, 6, 29072. [Google Scholar] [CrossRef]
- Demuth, J.P.; Hahn, M.W. The life and death of gene families. BioEssays 2009, 31, 29–39. [Google Scholar] [CrossRef]
- Magadum, S.; Banerjee, U.; Murugan, P.; Gangapur, D.; Ravikesavan, R. Gene duplication as a major force in evolution. J. Genet. 2013, 92, 155–161. [Google Scholar] [CrossRef]
- Soltis, P.S.; Soltis, D.E. Ancient WGD events as drivers of key innovations in angiosperms. Curr. Opin. Plant Biol. 2016, 30, 159–165. [Google Scholar] [CrossRef]
- Conant, G.C.; Wolfe, K.H. Turning a hobby into a job: How duplicated genes find new functions. Nat. Reviews. Genet. 2008, 9, 938–950. [Google Scholar] [CrossRef]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef]
- Skirycz, A.; Reichelt, M.; Burow, M.; Birkemeyer, C.; Rolcik, J.; Kopka, J.; Zanor, M.I.; Gershenzon, J.; Strnad, M.; Szopa, J.; et al. DOF transcription factor AtDof1.1 (OBP2) is part of a regulatory network controlling glucosinolate biosynthesis in Arabidopsis. Plant J. Cell Mol. Biol. 2006, 47, 10–24. [Google Scholar] [CrossRef]
- He, L.; Su, C.; Wang, Y.; Wei, Z. ATDOF5.8 protein is the upstream regulator of ANAC069 and is responsive to abiotic stress. Biochimie 2015, 110, 17–24. [Google Scholar] [CrossRef]
- Corrales, A.R.; Carrillo, L.; Lasierra, P.; Nebauer, S.G.; Dominguez-Figueroa, J.; Renau-Morata, B.; Pollmann, S.; Granell, A.; Molina, R.V.; Vicente-Carbajosa, J.; et al. Multifaceted role of cycling DOF factor 3 (CDF3) in the regulation of flowering time and abiotic stress responses in Arabidopsis. Plant Cell Environ. 2017, 40, 748–764. [Google Scholar] [CrossRef]
- Chen, J.; Bi, Y.Y.; Wang, Q.Q.; Liu, D.K.; Zhang, D.; Ding, X.; Liu, Z.J.; Chen, S.P. Genome-wide identification and analysis of anthocyanin synthesis-related R2R3-MYB genes in Cymbidium goeringii. Front. Plant Sci. 2022, 13, 1002043. [Google Scholar] [CrossRef]
- Wang, L.; Zhao, X.; Zheng, R.; Huang, Y.; Zhang, C.; Zhang, M.M.; Lan, S.; Liu, Z.J. Genome-Wide Identification and Drought Stress Response Pattern of the NF-Y Gene Family in Cymbidium sinense. Int. J. Mol. Sci. 2024, 25, 3031. [Google Scholar] [CrossRef]
- Lamesch, P.; Berardini, T.Z.; Li, D.; Swarbreck, D.; Wilks, C.; Sasidharan, R.; Muller, R.; Dreher, K.; Alexander, D.L.; Garcia-Hernandez, M.; et al. The Arabidopsis Information Resource (TAIR): Improved gene annotation and new tools. Nucleic Acids Res. 2012, 40, D1202–D1210. [Google Scholar] [CrossRef]
- Schoch, C.L.; Ciufo, S.; Domrachev, M.; Hotton, C.L.; Kannan, S.; Khovanskaya, R.; Leipe, D.; McVeigh, R.; O’Neill, K.; Robbertse, B.; et al. NCBI Taxonomy: A comprehensive update on curation, resources and tools. Database 2020, 2020, baaa062. [Google Scholar] [CrossRef]
- Jin, J.; Tian, F.; Yang, D.C.; Meng, Y.Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2017, 45, D1040–D1045. [Google Scholar] [CrossRef]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, B.; Gao, S.; Lercher, M.J.; Hu, S.; Chen, W.H. Evolview v3: A webserver for visualization, annotation, and management of phylogenetic trees. Nucleic Acids Res. 2019, 47, W270–W275. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Geourjon, C.; Deléage, G. SOPMA: Significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Comput. Appl. Biosci. CABIOS 1995, 11, 681–684. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]






Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, X.; Zhang, M.-M.; Huang, Y.; Yu, J.; Zhao, X.; Zheng, Q.; Liu, Z.-J.; Lan, S. Genome-Based Identification of the Dof Gene Family in Three Cymbidium Species and Their Responses to Heat Stress in Cymbidium goeringii. Int. J. Mol. Sci. 2024, 25, 7662. https://doi.org/10.3390/ijms25147662
He X, Zhang M-M, Huang Y, Yu J, Zhao X, Zheng Q, Liu Z-J, Lan S. Genome-Based Identification of the Dof Gene Family in Three Cymbidium Species and Their Responses to Heat Stress in Cymbidium goeringii. International Journal of Molecular Sciences. 2024; 25(14):7662. https://doi.org/10.3390/ijms25147662
Chicago/Turabian StyleHe, Xin, Meng-Meng Zhang, Ye Huang, Jiali Yu, Xuewei Zhao, Qinyao Zheng, Zhong-Jian Liu, and Siren Lan. 2024. "Genome-Based Identification of the Dof Gene Family in Three Cymbidium Species and Their Responses to Heat Stress in Cymbidium goeringii" International Journal of Molecular Sciences 25, no. 14: 7662. https://doi.org/10.3390/ijms25147662
APA StyleHe, X., Zhang, M.-M., Huang, Y., Yu, J., Zhao, X., Zheng, Q., Liu, Z.-J., & Lan, S. (2024). Genome-Based Identification of the Dof Gene Family in Three Cymbidium Species and Their Responses to Heat Stress in Cymbidium goeringii. International Journal of Molecular Sciences, 25(14), 7662. https://doi.org/10.3390/ijms25147662





