Molecular techniques have become influential instruments in biological study, transforming our comprehension of life at the cellular and genetic levels [1]. The Special Issue "Molecular World Today and Tomorrow: Recent Trends in Biological Sciences 2.0" examines current trends in biological sciences and the impact of advanced techniques on scientific advancements. The problem highlights how molecular techniques can greatly aid in the advancement of focused medication creation and disease diagnostics. Combining molecular phylogenetic and omics approaches with expression and pathway analysis, we seek to improve our comprehension of molecular mechanisms and stress responses in biological systems [2]. This collaboration represents significant progress in genomics, proteomics, and phylogenetic methods, providing new opportunities for study and innovation.
Advancements in omics technologies, including transcriptomics, proteomics, and genomics, have significantly improved our capacity to recognize and describe biological entities [2,3]. Researchers currently use systematic methods to uncover genetic variations, forecast ecological habitats, and clarify taxonomic intricacies [4,5]. Omics data offer a thorough perspective on the molecular world, ranging from analyzing microbial populations to uncovering the genetic foundations of diseases. Integrating molecular phylogenetics with omics techniques enhances our comprehension of evolutionary connections. Researchers investigate molecular pathways across many taxonomic groups by integrating sequence data with expression profiles [6,7]. This synergy influences both evolutionary history and functional adaptations, as well as stress responses.
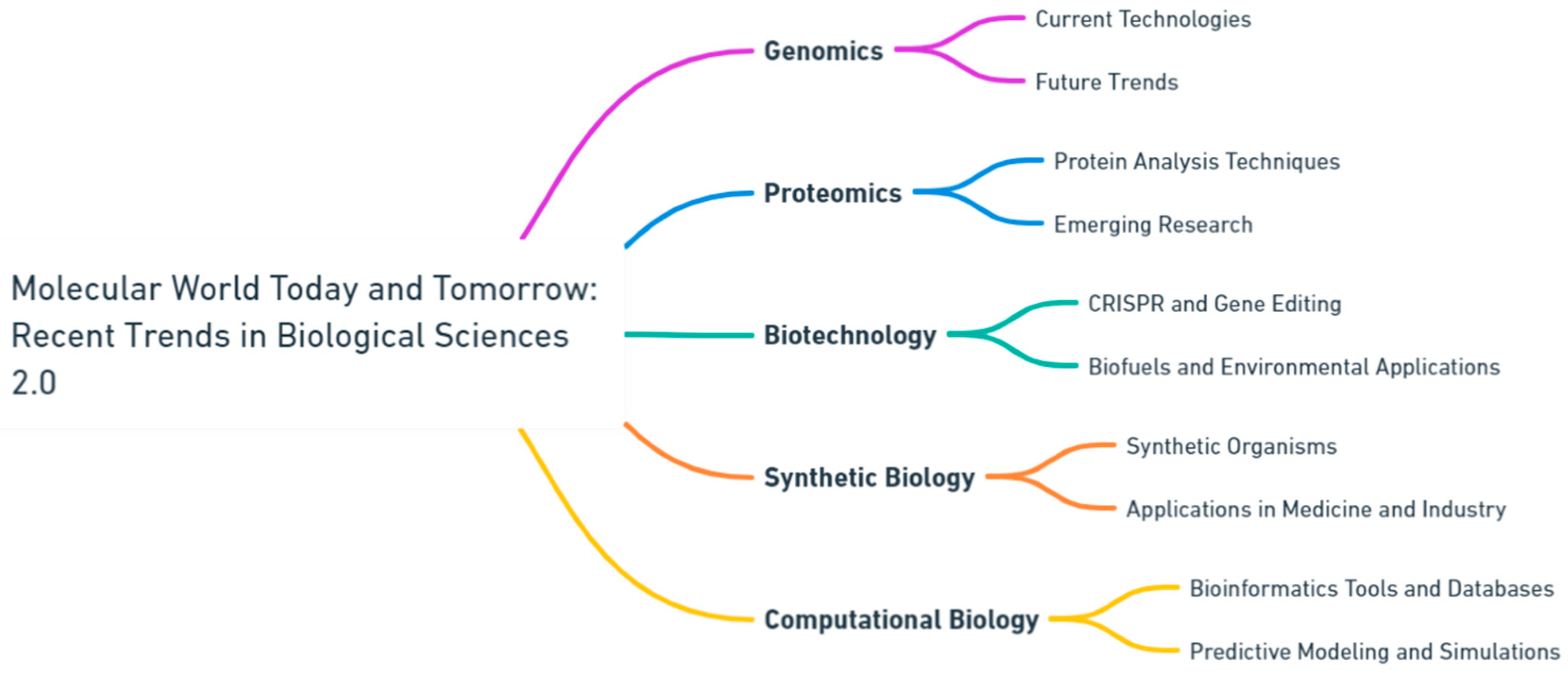
Molecular data drives precision medicine [8,9]. Genomics and proteomics play a crucial role in guiding individualized treatments by identifying therapeutic targets and predicting medication responses [10,11]. Furthermore, diagnostic tests utilizing molecular markers improve the diagnosis and prediction of diseases. As we decode the complexities of individual genomes, the future shows potential for personalized interventions. The study in this Special Issue highlights the significance of interdisciplinary approaches in addressing intricate biological issues through collaboration. It unites specialists from several disciplines, such as molecular biology, genetics, biochemistry, biophysics, and computational biology, to promote a comprehensive comprehension of life at the molecular scale. This collaborative effort is crucial for the integration of knowledge and the acceleration of scientific discoveries (Figure 1).
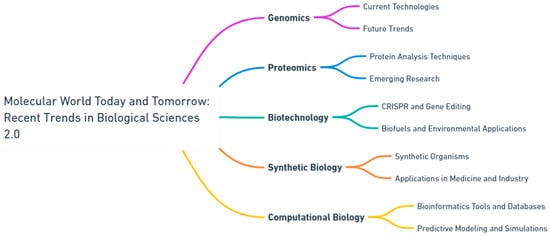
Figure 1.
Navigating the frontier: current and future perspectives in biological sciences (generated by OpenAI's Whimsical Diagrams GPT).
The edition features 20 papers, including 3 comprehensive review articles, a brief report, 1 communication, and 15 research articles. The articles in the Special Issue emphasize the importance of molecular data in promoting targeted medicines, highlighting the vital significance of molecular sciences in medicine. This interdisciplinary method highlights the significant impact of molecular approaches in research and clinical environments, presenting new opportunities for customized medicine and treatments.
In conclusions, this Special Issue demonstrates the dynamic and interdisciplinary nature of molecular sciences. It motivates researchers to expand the current knowledge and investigate new areas, assuring the ongoing development of the biological sciences field and its contribution to our understanding of life’s molecular foundation. The compilation showcases the present status of molecular sciences and paves the way for future progress, encouraging further inquiry and invention in this captivating discipline.
Conflicts of Interest
The author declare no conflicts of interest.
List of Contributions
- Arai, I.; Tsuji, M.; Takahashi, K.; Saito, S.; Takeda, H. Analyzing the Antinociceptive Effect of Interleukin-31 in Mice. Int. J. Mol. Sci. 2023, 24, 11563. https://doi.org/10.3390/ijms241411563.
- Arora, D.; Hernandez, A.G.; Walden, K.K.O.; Fields, C.J.; Yan, G. First Draft Genome Assembly of Root-Lesion Nematode Pratylenchus scribneri Generated Using Long-Read Sequencing. Int. J. Mol. Sci. 2023, 24, 7311. https://doi.org/10.3390/ijms24087311.
- Arshad, S.; Wei, M.; Ali, Q.; Mustafa, G.; Ma, Z.; Yan, Y. Paclitaxel and Caffeine–Taurine, New Colchicine Alternatives for Chromosomes Doubling in Maize Haploid Breeding. Int. J. Mol. Sci. 2023, 24, 14659. https://doi.org/10.3390/ijms241914659.
- Barreda-Castillo, J.M.; Monribot-Villanueva, J.L.; Velázquez-Rosas, N.; Bayman, P.; Guerrero-Analco, J.A.; Menchaca-García, R.A. Morphological and Physio-Chemical Responses to PEG-Induced Water Stress in Vanilla planifolia and V. pompona Hybrids. Int. J. Mol. Sci. 2023, 24, 4690. https://doi.org/10.3390/ijms24054690.
- Bhujel, B.; Oh, S.-H.; Kim, C.-M.; Yoon, Y.-J.; Kim, Y.-J.; Chung, H.-S.; Ye, E.-A.; Lee, H.; Kim, J.-Y. Mesenchymal Stem Cells and Exosomes: A Novel Therapeutic Approach for Corneal Diseases. Int. J. Mol. Sci. 2023, 24, 10917. https://doi.org/10.3390/ijms241310917.
- Costantini, M.; Esposito, R.; Ruocco, N.; Caramiello, D.; Cordella, A.; Ventola, G.M.; Zupo, V. De Novo Assembly of the Genome of the Sea Urchin Paracentrotus lividus (Lamarck 1816). Int. J. Mol. Sci. 2024, 25, 1685. https://doi.org/10.3390/ijms25031685.
- Dzidek, A.; Czerwińska-Ledwig, O.; Żychowska, M.; Pilch, W.; Piotrowska, A. The Role of Increased Expression of Sirtuin 6 in the Prevention of Premature Aging Pathomechanisms. Int. J. Mol. Sci. 2023, 24, 9655. https://doi.org/10.3390/ijms24119655.
- Félix, J.W.; Granados-Alegría, M.I.; Gómez-Tah, R.; Tzec-Simá, M.; Ruíz-May, E.; Canto-Canché, B.; Zamora-Briseño, J.A.; Bojórquez-Velázquez, E.; Oropeza-Salín, C.; Islas-Flores, I. Proteome Landscape during Ripening of Solid Endosperm from Two Different Coconut Cultivars Reveals Contrasting Carbohydrate and Fatty Acid Metabolic Pathway Modulation. Int. J. Mol. Sci. 2023, 24, 10431. https://doi.org/10.3390/ijms241310431.
- He, R.; Liu, K.; Zhang, S.; Ju, J.; Hu, Y.; Li, Y.; Liu, X.; Liu, H. Omics Analysis Unveils the Pathway Involved in the Anthocyanin Biosynthesis in Tomato Seedling and Fruits. Int. J. Mol. Sci. 2023, 24, 8690. https://doi.org/10.3390/ijms24108690.
- Jia, P.; Liu, J.; Yan, R.; Yang, K.; Dong, Q.; Luan, H.; Zhang, X.; Li, H.; Guo, S.; Qi, G. Systematical Characterization of the AT-Hook Gene Family in Juglans regia L. and the Functional Analysis of the JrAHL2 in Flower Induction and Hypocotyl Elongation. Int. J. Mol. Sci. 2023, 24, 7244. https://doi.org/10.3390/ijms24087244.
- Kim, H.-J.; Kim, K.E.; Kim, Y.J.; Kang, H.; Shin, J.W.; Kim, S.; Lee, S.H.; Jung, S.W.; Lee, T.-K. Marine Bacterioplankton Community Dynamics and Potentially Pathogenic Bacteria in Seawater around Jeju Island, South Korea, via Metabarcoding. Int. J. Mol. Sci. 2023, 24, 13561. https://doi.org/10.3390/ijms241713561.
- Kim, Y.; Wang, J.; Ma, C.; Jong, C.; Jin, M.; Cha, J.; Wang, J.; Peng, Y.; Ni, H.; Li, H.; et al. GmTCP and GmNLP Underlying Nodulation Character in Soybean Depending on Nitrogen. Int. J. Mol. Sci. 2023, 24, 7750. https://doi.org/10.3390/ijms24097750.
- Kowalski, T.W.; Feira, M.F.; Lord, V.O.; Gomes, J.D.; Giudicelli, G.C.; Fraga, L.R.; Sanseverino, M.T.; Recamonde-Mendoza, M.; Schuler-Faccini, L.; Vianna, F.S. A New Strategy for the Old Challenge of Thalidomide: Systems Biology Prioritization of Potential Immunomodulatory Drug (IMiD)-Targeted Transcription Factors. Int. J. Mol. Sci. 2023, 24, 11515. https://doi.org/10.3390/ijms241411515.
- Rocchetti, M.T.; Bellanti, F.; Zadorozhna, M.; Fiocco, D.; Mangieri, D. Multi-Faceted Role of Luteolin in Cancer Metastasis: EMT, Angiogenesis, ECM Degradation and Apoptosis. Int. J. Mol. Sci. 2023, 24, 8824. https://doi.org/10.3390/ijms24108824.
- Shi, Y.; Yu, B.; Cheng, S.; Hu, W.; Liu, F. The Change in Whole-Genome Methylation and Transcriptome Profile under Autophagy Defect and Nitrogen Starvation. Int. J. Mol. Sci. 2023, 24, 14047. https://doi.org/10.3390/ijms241814047.
- Su, H.-H.; Huang, Y.-H.; Lien, Y.; Yang, P.-C.; Huang, C.-Y. Crystal Structure of DNA Replication Protein SsbA Complexed with the Anticancer Drug 5-Fluorouracil. Int. J. Mol. Sci. 2023, 24, 14899. https://doi.org/10.3390/ijms241914899.
- Vásquez, C.; Leyton-Carcaman, B.; Cid-Alda, F.P.; Segovia, I.; Pinto, F.; Abanto, M. Physical Pretreatments Applied in Three Commercial Kits for the Extraction of High-Quality DNA from Activated Sewage Sludge. Int. J. Mol. Sci. 2023, 24, 15243. https://doi.org/10.3390/ijms242015243.
- Wang, Z.; Yadav, V.; Chen, X.; Zhang, S.; Yuan, X.; Li, H.; Ma, J.; Zhang, Y.; Yang, J.; Zhang, X.; et al. Multi-Omics Analysis Reveals Intricate Gene Networks Involved in Female Development in Melon. Int. J. Mol. Sci. 2023, 24, 16905. https://doi.org/10.3390/ijms242316905.
- Zhang, J.; Zhang, Y.; Feng, C. Genome-Wide Analysis of MYB Genes in Primulina eburnea (Hance) and Identification of Members in Response to Drought Stress. Int. J. Mol. Sci. 2024, 25, 465. https://doi.org/10.3390/ijms25010465.
- Zheng, S.; Shin, K.; Lin, W.; Wang, W.; Yang, X. Identification and Characterization of PRE Genes in Moso Bamboo (Phyllostachys edulis). Int. J. Mol. Sci. 2023, 24, 6886. https://doi.org/10.3390/ijms24086886.
References
- Hartwell, L.H.; Hopfield, J.J.; Leibler, S.; Murray, A.W. From molecular to modular cell biology. Nature 1999, 402, C47–C52. [Google Scholar] [CrossRef] [PubMed]
- Misra, B.B.; Langefeld, C.; Olivier, M.; Cox, L.A. Integrated omics: Tools, advances and future approaches. J. Mol. Endocrinol. 2019, 62, R21–R45. [Google Scholar] [CrossRef] [PubMed]
- Veenstra, T.D. Omics in systems biology: Current progress and future outlook. Proteomics 2021, 21, 2000235. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.D.; Freeland, J.R.; Parducci, L.; Evans, D.M.; Meyer, R.S.; Molano-Flores, B.; Davis, M.A. Environmental DNA as an emerging tool in botanical research. Am. J. Bot. 2023, 110, e16120. [Google Scholar] [CrossRef] [PubMed]
- Waldvogel, A.-M.; Feldmeyer, B.; Rolshausen, G.; Exposito-Alonso, M.; Rellstab, C.; Kofler, R.; Mock, T.; Schmid, K.; Schmitt, I.; Bataillon, T. Evolutionary genomics can improve prediction of species’ responses to climate change. Evol. Lett. 2020, 4, 4–18. [Google Scholar] [CrossRef] [PubMed]
- Borisov, N.; Sorokin, M.; Garazha, A.; Buzdin, A. Quantitation of molecular pathway activation using RNA sequencing data. In Nucleic Acid Detection and Structural Investigations: Methods and Protocols; Humana New York: New York, NY, USA, 2020; pp. 189–206. [Google Scholar]
- Graw, S.; Chappell, K.; Washam, C.L.; Gies, A.; Bird, J.; Robeson, M.S.; Byrum, S.D. Multi-omics data integration considerations and study design for biological systems and disease. Mol. Omics 2021, 17, 170–185. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, M.; Wojtusciszyn, A.; Favre, L.; Boughorbel, S.; Shan, J.; Letaief, K.B.; Pitteloud, N.; Chouchane, L. Precision medicine in the era of artificial intelligence: Implications in chronic disease management. J. Transl. Med. 2020, 18, 472. [Google Scholar] [CrossRef] [PubMed]
- Denny, J.C.; Collins, F.S. Precision medicine in 2030—Seven ways to transform healthcare. Cell 2021, 184, 1415–1419. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Zhang, Z.; Zhou, L.; Han, C.; Huang, C.; Nice, E.C. Proteomics, personalized medicine and cancer. Cancers 2021, 13, 2512. [Google Scholar] [CrossRef] [PubMed]
- Park, D.I. Genomics, transcriptomics, proteomics and big data analysis in the discovery of new diagnostic markers and targets for therapy development. Prog. Mol. Biol. Transl. Sci. 2020, 173, 61–90. [Google Scholar] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).