Simple Summary
Cancer-associated glycosylation changes are widely used as biomarkers and strongly impact malignancy. However, the clinical significance of the deranged expression of glycosyltransferases observed in specimens is not always consistent with their role in experimental systems. We analyzed the overall survival curves of patients expressing high or low mRNA levels of 114 glycosyltransferases from the 21 cohorts of The Cancer Genome Atlas (TCGA). We identified 17 glycosyltransferases associated with poor prognosis and 4 associated with good prognosis in a large number of cohorts. In addition, we identified several glycosyltransferases with a very high prognostic value in only one or a few cohorts. Comparisons with published experimental works reveal partial consistency with TCGA clinical data. These data pave the way for the use of glycosyltransferases as prognostic markers and potential therapeutic targets and place experimental studies in an appropriate clinical context.
Abstract
Background: Glycosylation changes are a main feature of cancer. Some carbohydrate epitopes and expression levels of glycosyltransferases have been used or proposed as prognostic markers, while many experimental works have investigated the role of glycosyltransferases in malignancy. Using the transcriptomic data of the 21 TCGA cohorts, we correlated the expression level of 114 glycosyltransferases with the overall survival of patients. Methods: Using the Oncolnc website, we determined the Kaplan–Meier survival curves for the patients falling in the 15% upper or lower percentile of mRNA expression of each glycosyltransferase. Results: Seventeen glycosyltransferases involved in initial steps of N- or O-glycosylation and of glycolipid biosynthesis, in chain extension and sialylation were unequivocally associated with bad prognosis in a majority of cohorts. Four glycosyltransferases were associated with good prognosis. Other glycosyltransferases displayed an extremely high predictive value in only one or a few cohorts. The top were GALNT3, ALG6 and B3GNT7, which displayed a p < 1 × 10−9 in the low-grade glioma (LGG) cohort. Comparison with published experimental data points to ALG3, GALNT2, B4GALNT1, POFUT1, B4GALT5, B3GNT5 and ST3GAL2 as the most consistently malignancy-associated enzymes. Conclusions: We identified several cancer-associated glycosyltransferases as potential prognostic markers and therapeutic targets.
1. Introduction
Glycosylation is a widely occurring modification of proteins and lipids that plays a crucial role in the modulation of cellular and molecular interactions [1]. Glycosylation is profoundly altered in cancer [2,3] and a huge number of clinical and experimental studies support the role of specific carbohydrate structures in determining cancer malignancy. However, studies performed in different experimental systems do not always provide consistent and reliable conclusions about the role of sugar chains and their cognate glycosyltransferases in cancer. On the other hand, clinical studies are often performed on small cohorts, which do not allow us to reach reliable conclusions on the impact of the overexpression of a glycosyltransferase on patient survival. The cancer genome atlas (TCGA) contains transcriptomic and clinical data from hundreds of patients affected by 21 malignancies. In this study, we determined the association between the level of expression of 114 glycosyltransferases in the 21 TCGA cohorts with patients’ overall survival. We identified a few glycosyltransferases whose high expression was unambiguously associated with a better or poorer prognosis in different cohorts. In addition, we identified glycosyltransferase with a very high prognostic value in one or a few cohorts. The role of the glycosyltransferases emerging from TCGA data analysis was compared with data obtained from experimental studies through an extensive literature review.
2. Glycosyltransferase Genes Associated with Prognosis in TCGA Cohorts
We first determined the association with the prognosis of 114 glycosyltransferases in all 21 TCGA cohorts. Table S1 reports the p value for the association with overall survival of the 15% upper percentile vs. the 15% lower percentile of glycosyltransferase mRNA expression, as obtained from the Oncolnc website. A dark red code label or a dark blue code label was assigned to significant (p ≤ 0.05) associations with a bad (red) or a good (blue) prognosis. A light red or light blue code label was assigned to strong but not significant associations (0.1 ≥ p ≥ 0.05). The percentage of glycosyltransferases significantly associated with overall survival was strikingly different in the different cohorts (Table S1, penultimate row). Obviously, a low number of patients in a cohort would make it harder to reach statistical significance. However, this was not the reason for the discrepancy. In fact, in the BRCA cohort, which is the most numerous (1006 cases Table S1, third row), only 16 glycosyltransferases displayed an association with prognosis (14%) (Table S1), while in the LGG cohort, which contains about half of the BRCA patients (510 cases), 69 glycosyltransferases were associated with prognosis (60%) (Table S1). For each cohort, one or more enzymes showing the lowest p value were identified as “best predictors” of bad (red) or good (blue) prognosis. Notably, MGAT4A and B4GALNT1 were best predictors of bad prognosis in 3 (ESCA, UCEC and LUSC) and 2 (HNSC and KIRP) cohorts, respectively.
3. Glycosyltransferase Genes Playing a Consistent Association in a Large Number of Cohorts
Several glycosyltransferase genes presented a prevalent association with a bad prognosis in a large number of cohorts, while a few displayed a prevalent association with a good prognosis. The former are referred to hereafter as “Bad Prognosis-associated”, (BPA) genes, while the latter as “Good Prognosis-associated”, (GPA) glycosyltransferases. Inclusion in either category was based on the difference between the number of cohorts in which it was associated with a bad prognosis and the number of cohorts in which it was related with a good prognosis. When this “score” reached a value ≥ 5, the glycosyltransferase was referred to as BPA, while GPA was referred to a glycosyltransferase with a score value ≤ −5. For example, ALG3 was associated with a bad prognosis in 11 cohorts and with a good prognosis in 2. Consequently, its BPA score was 9. According to this analysis, we identified 17 BPA and 4 GPA enzymes (Table 1).

Table 1.
Prognostic value in TCGA cohorts of BPA and GPA glycosyltransferases.
Glycosyltransferases can be grouped as: initiating glycosyltransferases elaborating core structures of N- and O-linked chains and glycolipids; extending glycosyltransferases elongating sugar chains, which can be in common among N- and O-linked chains and glycolipids; and capping glycosyltransferases terminating sugar chains [4]. Table 2 reports the role of the 17 BPA and of the 4 GPA glycosyltransferases from Table 1 in glycan biosynthesis, as well as their score.

Table 2.
Glycosyltransferases show an association with prognosis in a large number of cohorts.
4. Glycosyltransferases with Very High Prognostic Value (VHPV)
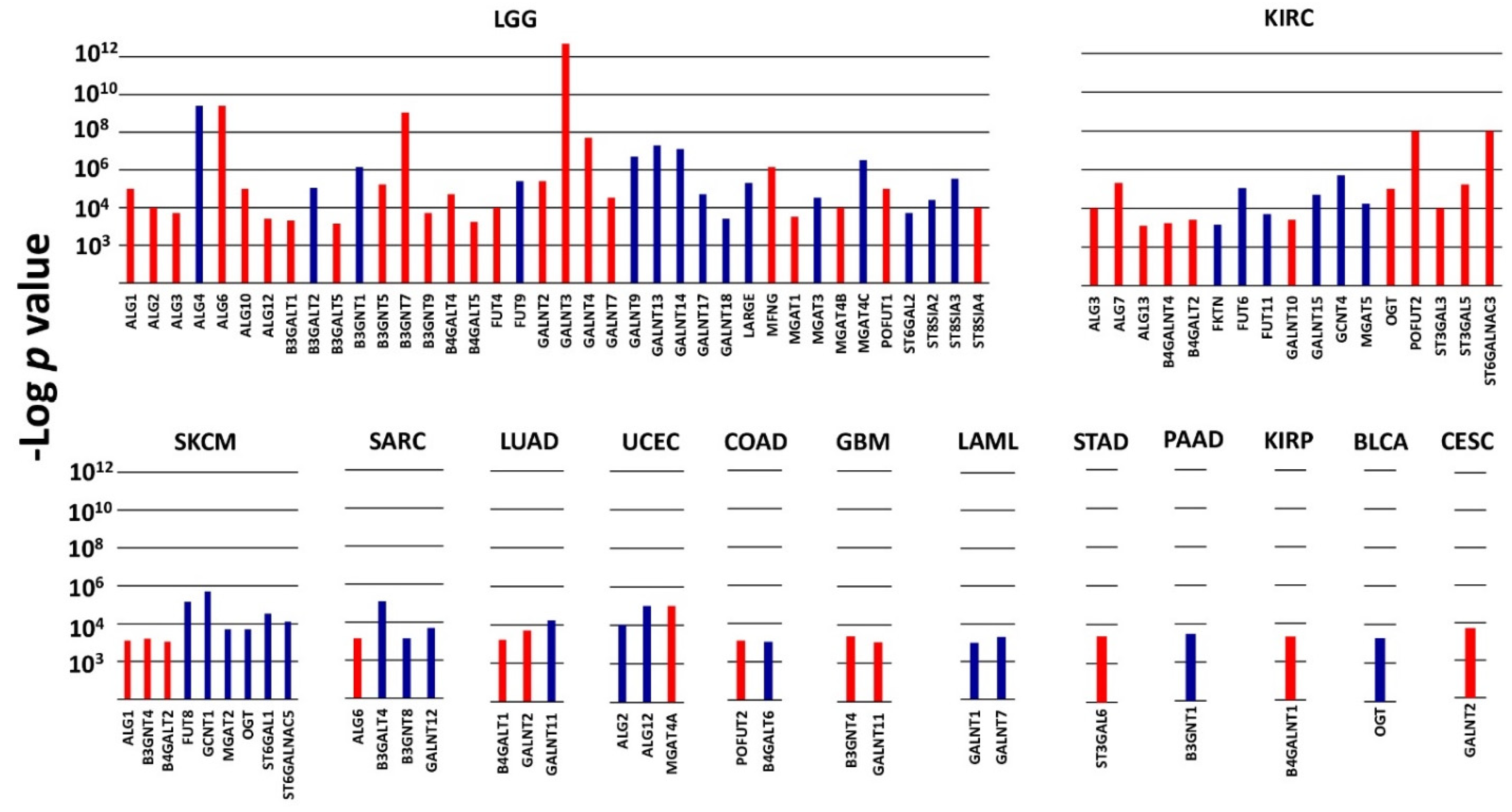
In the context of poor or good prognosis, several glycosyltransferases displayed a very high prognostic value (p ≤ 1 × 10−3) in a limited number of cohorts (Figure 1). These enzymes, which will be referred to as VHPV afterwards, were strikingly numerous in some cohorts. The cohort with the highest number of VHPV was LGG, followed by KIRC. Among the top 4 VHPV enzymes (p < 1 × 10−8), 3 were in LGG (GALNT3, ALG6 and B3GNT7), and 1 (POFUT2) was in KIRC (Figure 2). In LGG, a group of enzymes initiating N-glycosylation (ALG1 -2, -3, -6, -10, 12) or O-glycosylation (GALNT2, -3, -4, -7) displayed very strong association with poor prognosis. On the other hand, another group of GALNTs (9, 13, 14, 17, 18) showed a strong association with a good prognosis. Many VHPV glycosyltransferases displayed prognostic potential in only one cohort.
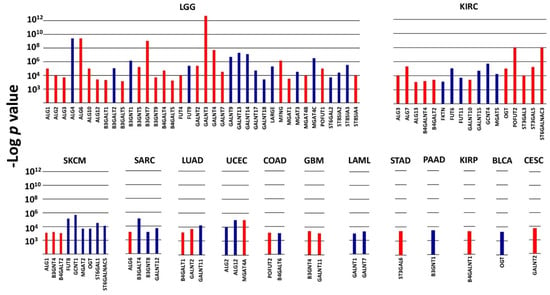
Figure 1.
VHPV in TCGA cohorts. Histograms represent the −Log of the p value for the comparison between the overall survival curves of the 15% higher expressers of each glycosyltransferase gene and the 15% lower expressers. Color labels indicate the association with a bad (red) or good (blue) prognosis. p < 1 × 10−3 was arbitrarily set as the threshold limit for inclusion. Cohorts not present in the figure did not contain any VHPV enzymes.
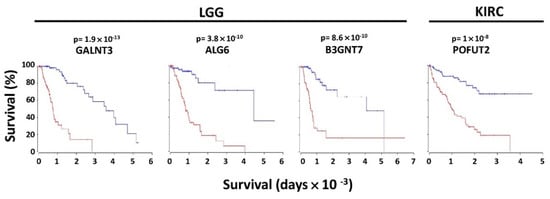
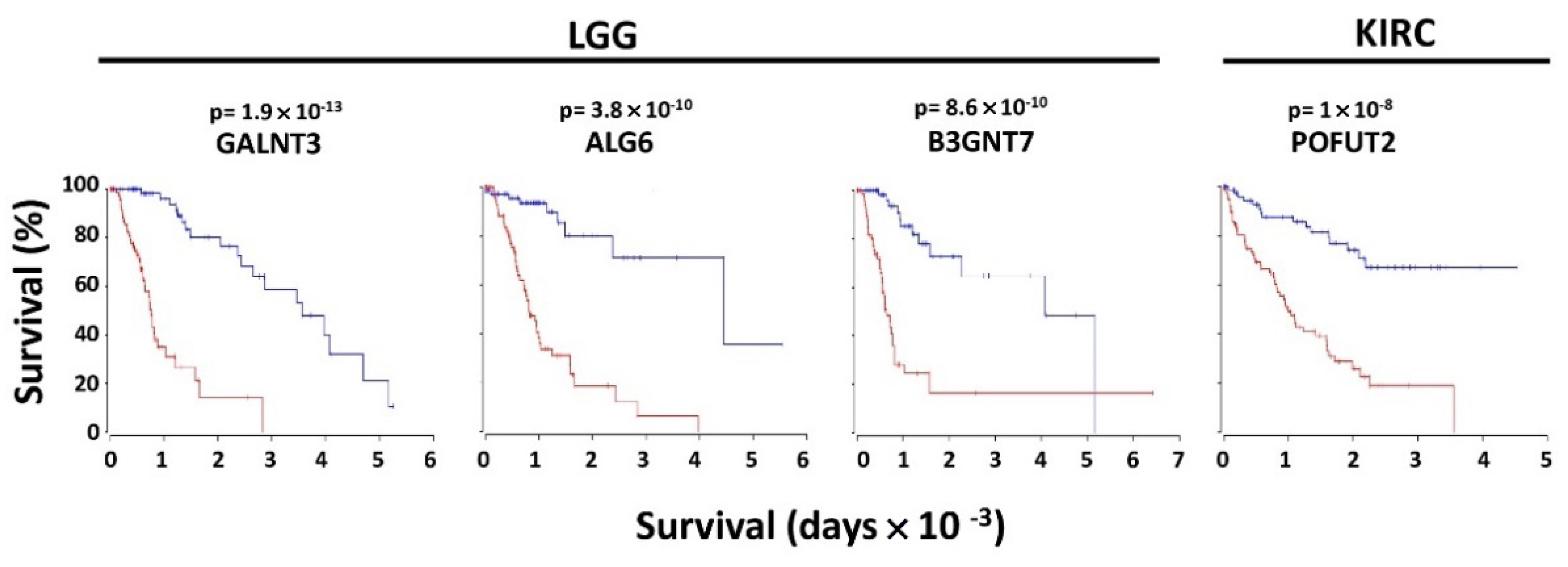
Figure 2.
Kaplan–Meier of overall survival curves of the top four VHPV glycosyltransferases. Curves were determined by the Oncolnc website for the 15% higher (red) and 15% lower (blue) expressers of the four glycosyltransferases. LGG and KIRC refer to brain lower grade glioma and kidney clear cell carcinoma, respectively.
5. Role of Glycosyltransferases in Experimental Systems
The role of relevant glycosyltransferases, including BPA and GPA, in experimental systems was assessed through an extensive literature search.
5.1. Initiating Glycosyltransferases
5.1.1. Glycosyltransferases Initiating N-Glycosylation
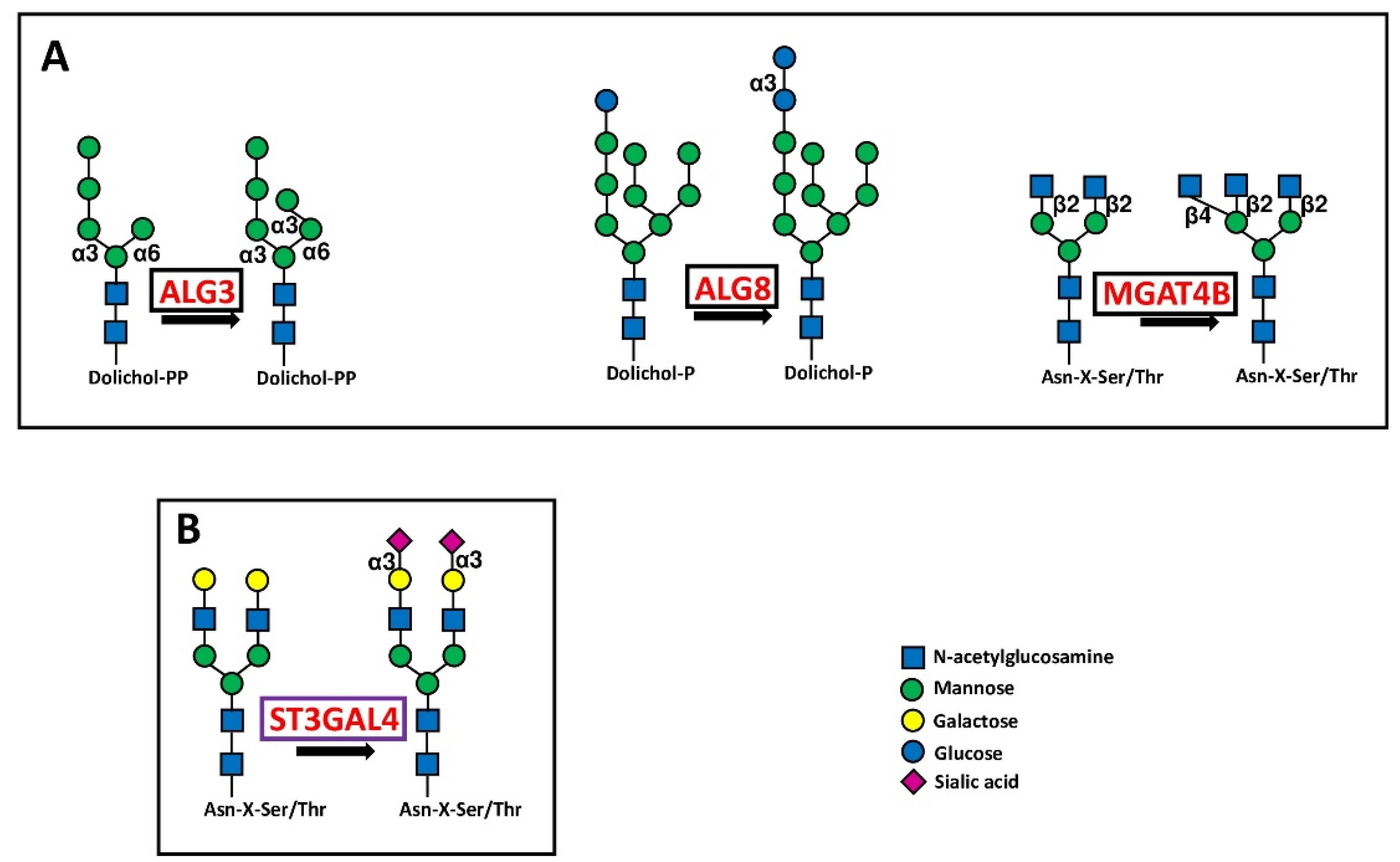
Glycosyltransferases ALG3, ALG8 and MGAT4B involved in the first steps of N-glycosylation behaved as BPA (Figure 3A).
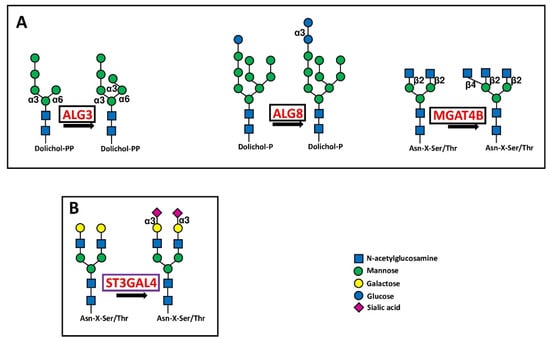
Figure 3.
Biosynthetic steps in N-glycosylation. (A) shows the core glycosylation steps catalyzed by three BPA glycosyltransferases in initiation of N-glycosylation. (B) shows the reaction catalyzed by the chain capping sialyltransferase ST3GAL4 in N-glycan biosynthesis. Enzymes catalyzing core glycosylation are boxed in black, while that catalyzing chain capping is boxed in violet.
Experimental data indicate that ALG3 contributes to malignancy of lung [5] and oral [6] cancer cell lines, in agreement with TCGA data reporting an association with a worse prognosis in LUAD and HNSC cohorts. However, the association with malignancy reported for esophageal [7] and cervical cancer [8] was not supported by TCGA data. ALG8 was reported to be associated with gastric [9] and colorectal [10] cancer. However, in the current study, we failed to observe any correlation with overall survival in these two malignancies. Very little or no information is available on the role of MGAT4B in experimental cancer systems.
5.1.2. Glycosyltransferases Initiating O-Glycosylation
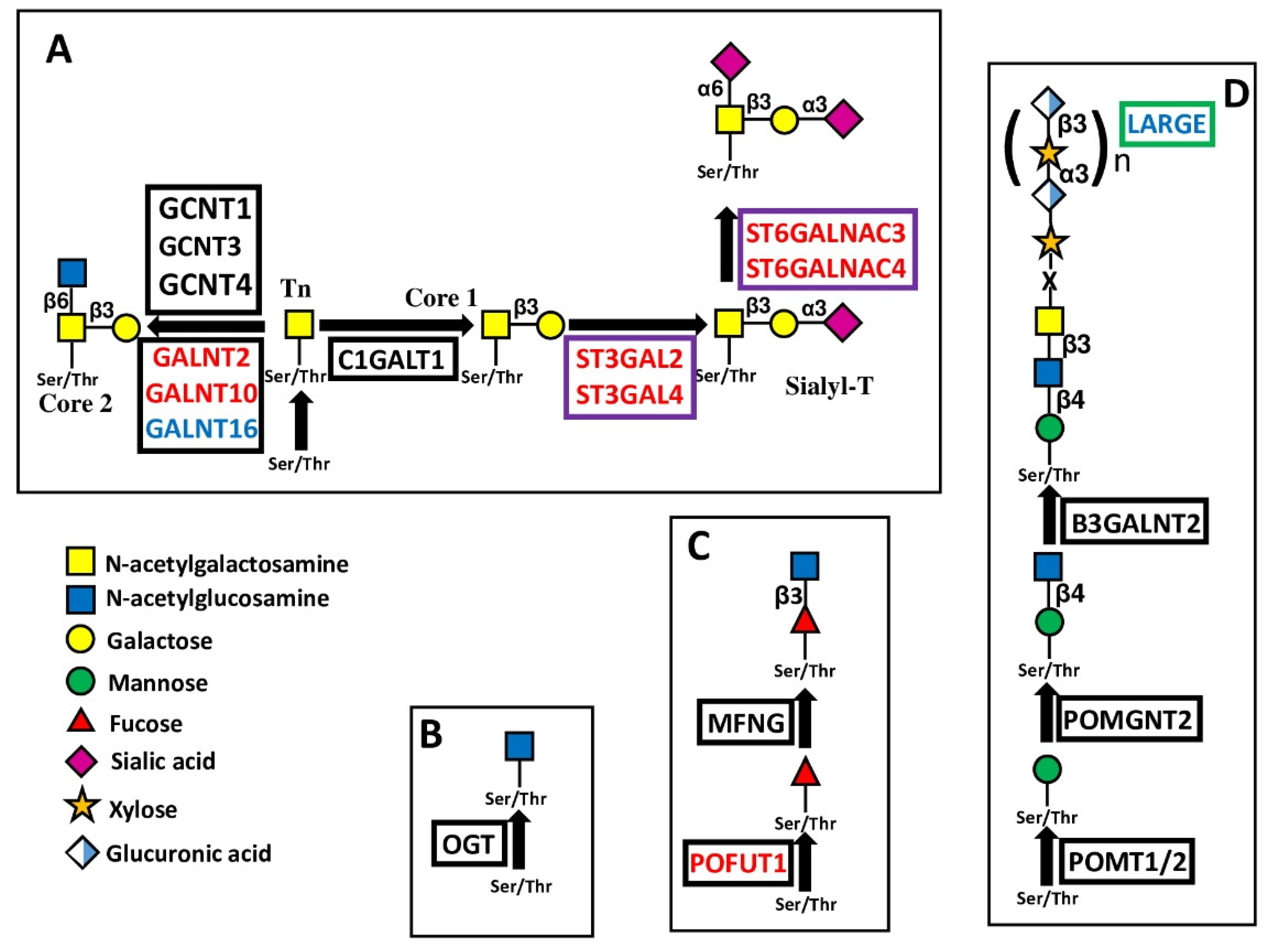
In the context of the 20 protein:O-GalNAc transferases mediating the addition of the first GalNAc of O-linked chains [11], GALNT2 and GALNT10 were identified as BPA. On the other hand, GALNT16 was GPA (Figure 4A). GALNT2, which is also the best predictor in CESC, provided a remarkable example of consistency between experimental data and prognosis. GALNT2 promoted malignancy through O-glycosylation of EGFR in oral cancer [12], glioma [13] and endometrial hyperplasia [14] cell lines, and by Notch signaling modulation [15] resulting in PD-L1 expression [16] in lung adenocarcinoma. Consistently, high GALNT2 expression was associated with poor overall survival in HNSC, LGG, UCEC and LUAD. On the other hand, increased malignancy related with high GALNT2 expression was also observed in hepatocellular carcinoma [17], while no relationship with overall survival was observed in the LIHC cohort. In gastric cancer cells, GALNT2 suppressed malignancy [18], but did not impact overall survival in STAD patients. Consistent with data of the OV cohort, in ovarian serous adenocarcinoma, high GALNT10 expression is related to an immunosuppressive microenvironment [19]. However, GALNT10 was causally associated with malignancy in cholangiocarcinoma [20] and hepatocellular carcinoma [21] but no relationship in the LIHC cohort was observed. Little or no information was available on GALNT16 in cancer.
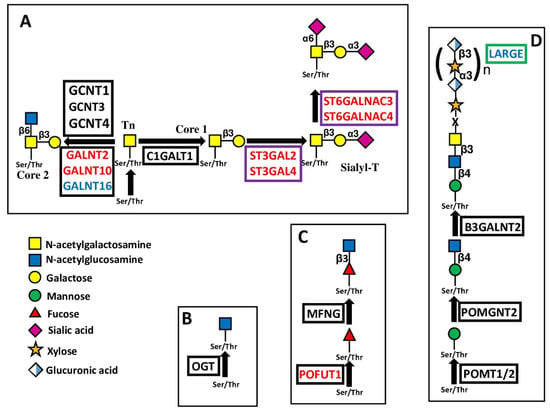
Figure 4.
Biosynthesis and structure of O-linked chains. The sugar O-glycosidically linked to Ser/Thr can be GalNAc (A), as in Mucin-type O-glycosylation; GlcNAc (B), as in many cytosolic and nuclear proteins; fucose (C), as in NOTCH receptors; or mannose (D), as in dystroglycan. Enzymes catalyzing core glycosylation are boxed in black, those catalyzing chain extensions are boxed in green, while those catalyzing chain cappings are boxed in violet.
Another type of O-glycosylation is O-GlcNAcylation (Figure 4B) [22]. The addition of a single O-GlcNAc residue to serine or threonine of cytosolic and nuclear proteins is mediated by a single enzyme, O-GlcNAc transferase (OGT). This enzyme is the best predictor of a good prognosis in BLCA. However, studies in bladder cancer cell lines have highlighted its tumor supporting activity [23,24].
A third type of O-glycosylation is O-fucosylation. POFUT1, which adds O-fucose to the NOTCH receptors (Figure 4C) [25], was found to be BPA. POFUT1, reported as a tumor-promoting glycosyltransferase in several studies, has also been proposed as a marker of colon cancer [26] and of high risk of tumor progression in adenomas [27]. Inhibition of POFUT1 decreased malignancy of CRC cell lines [28] by reducing stemness [29]. In a few cases, POFUT1 undergoes point mutation in CRC, resulting in enzyme hyperactivation and cancer progression [30]. POFUT2 is the best predictor of a poor prognosis in COAD. In hepatocarcinoma cells, POFUT1 promotes proliferation and invasion [31,32,33]. A high POFUT1 level correlates with glioblastoma [34] and lung [35], stomach [36], esophagus [37], breast [38], mouth [39] and bladder cancers [40]. However, only in the latter was an association with worse prognosis confirmed by TCGA data.
5.1.3. Glycosyltransferases Initiating Gangliosides
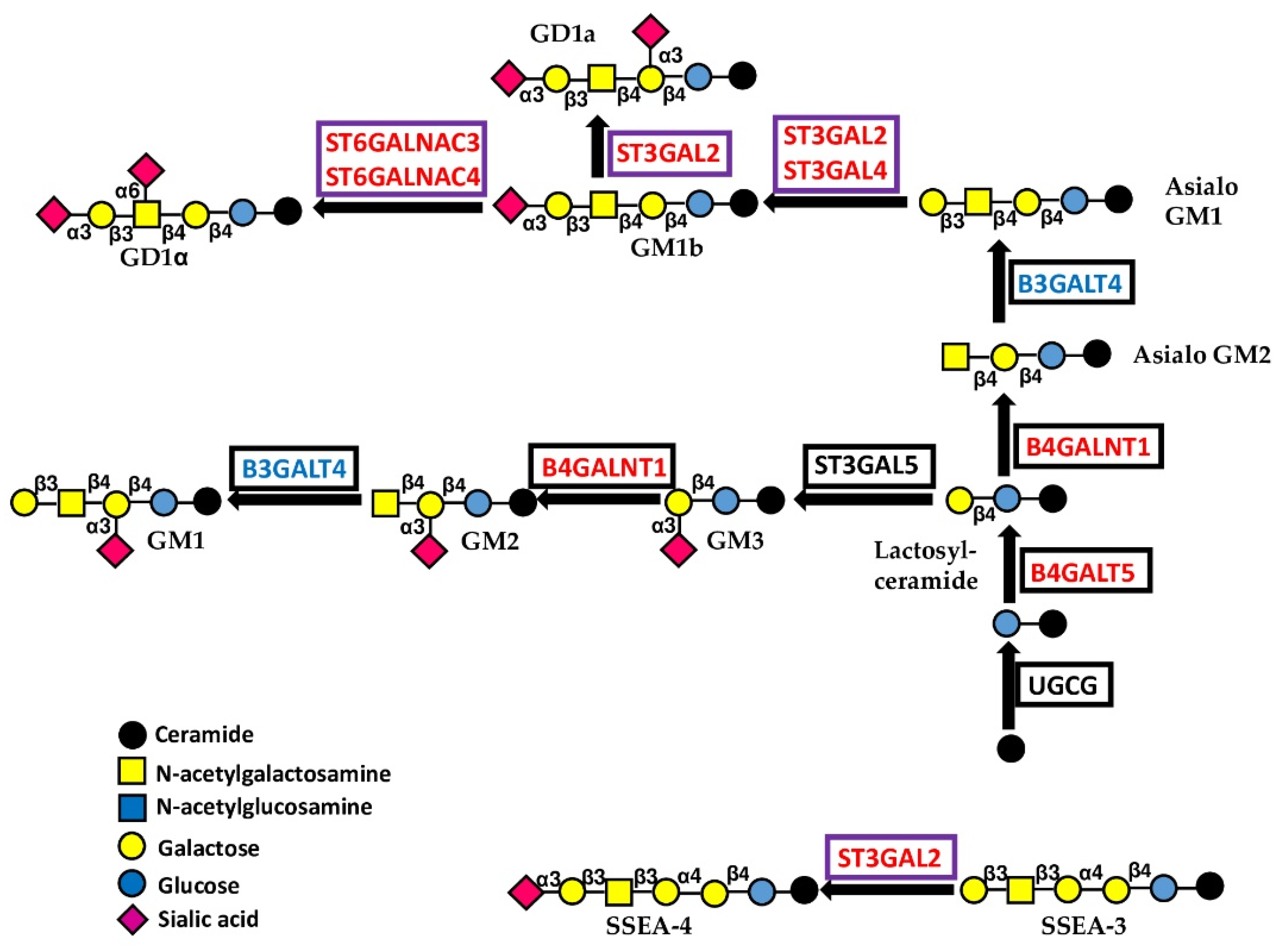
UGCG catalyzes the addition of glucose to ceramide (Figure 5). High expression of this enzyme increases malignancy in cervical cancer cells [41], consistent with the TCGA data of the CESC cohort (Table S1). B4GALT5 is the major enzyme involved in the biosynthesis of lactosyceramide, the root of all glycolipids [42,43], although it is also involved in other glycoconjugate formations. B4GALT5 increases the stemness and invasion of breast cancer cells [44] and multidrug resistance in leukemia cells [45]. No relationship with prognosis was evident in the BRCA cohort, while a tendency for a better prognosis was observed in the LAML cohort. B4GALNT1 catalyzes the synthesis of both GM2 and its asialo counterpart, asialo-GM2 (Figure 5). GM1, as well as GD2 and GD3, derive from GM2, while GD1a, GD1b and GD1α arise from asialo-GM2. Gangliosides GD2, GD3, GM2 and GD1a are greatly increased in breast cancer stem cells [46]. A causal correlation between high B4GALNT1 expression and malignancy has been noted in cell lines from lung, breast and kidney cancer, as well as in glioma and melanoma [46,47,48,49,50,51,52]. TCGA data are coherent with the B4GALNT1 role in kidney (KIRC) and lung (LUAD) cohorts. Phenotypically, the expression of B4GALNT1 has been associated with increased integrin signaling [52], reduced propensity to anoikis [49], stemness [46,50], augmented angiogenesis [51] and decreased immune surveillance [53]. B4GALNT1 is one of the most consistently and unambiguously glycosyltransferases associated with a bad prognosis (Table 1 and Table 2) in a large number of cohorts.
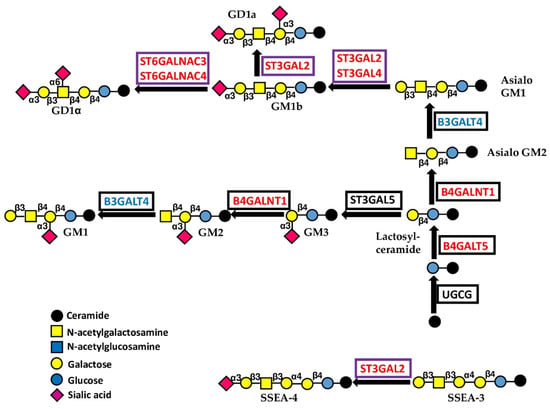
Figure 5.
Biosynthesis and structure of gangliosides. The sugar linked to ceramide is glucose. Enzymes catalyzing core glycosylation are boxed in black, while those catalyzing chain capping are boxed in violet.
5.2. Extending Glycosyltransferases
Among the extending glycosyltransferases, we will consider the enzymes involved in polylactosamine biosynthesis and LARGE.
5.2.1. Polylactosaminic Chains
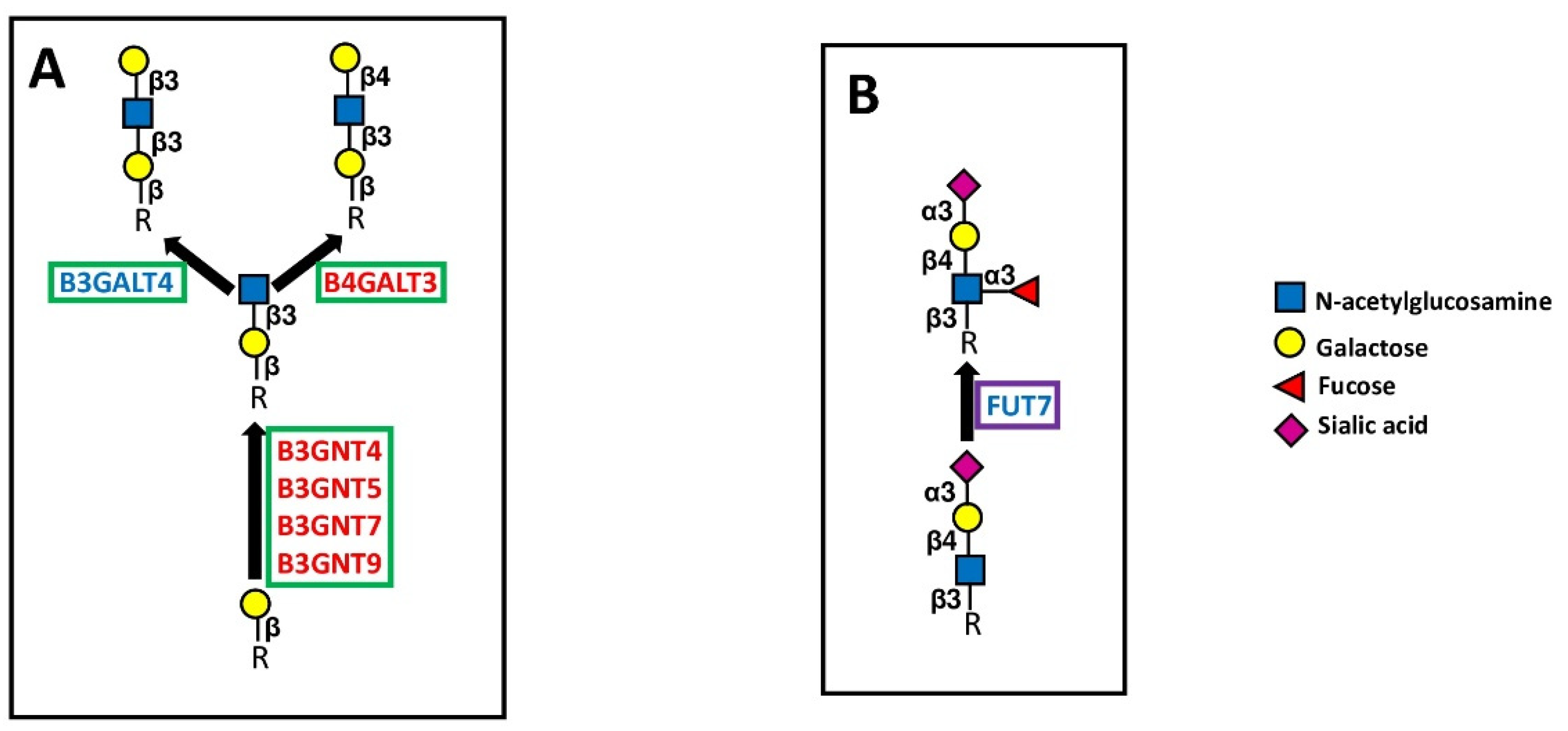
Polylactosamines constitute repeated Gal-GlcNAc (lactosamine) units. The two sugars can be linked either by a β1,3 bond (type 1 chains) or by a β1,4 bond (type 2 chains) (Figure 6A). N-linked chains, as well as O-linked chains and glycolipids, can be elongated by polylactosaminic chains. The first step of their biosynthesis consists of the addition of a GlcNAc residue in β1,3 linkage to an underlying galactose (Figure 6A).
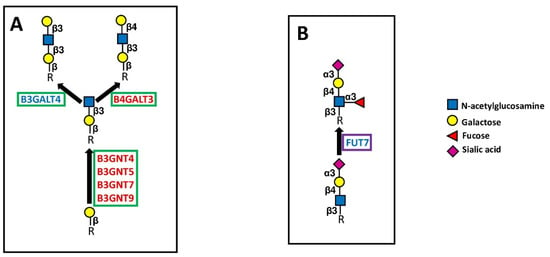
Figure 6.
Extension and capping of chains. (A) shows the elongation of polylactosaminic chains. (B) shows the structure and biosynthesis of the sialyl Lewis x antigen, catalyzed by FUT7, as an example of chain capping. Elongating enzymes are boxed in green, and capping enzymes are boxed in violet.
This reaction is mediated by different B3GNTs, specific to several types of sugar chains (e.g., type 2 chains for B3GNT4 and B3GNT7, glycolipids for B3GNT5, O-linked for B3GNT9). These four B3GNTs are BPA. B3GNT5 is a key enzyme for the biosynthesis of both type 1 and type 2 chains in glycolipids (Figure 6). Consistent with TCGA data, B3GNT5 enhances malignancy of glioma cells [54] and is stimulated by Helicobacter pylori infection in the stomach [55]. B3GNT7 promotes Lewis antigen expression [56] and suppresses malignancy in colon cancer cell lines [57], although in a large number of cohorts (but not in COAD), it is associated with worse prognosis. Within the B3GNTs group, B3GNT3 (which is neither a BPA nor a GPA) represents the subject of a larger number of studies. It mainly plays tumor-promoting activity in various types of tumors, including pancreatic [58,59], cervical [60], endometrial [61], and lung cancer [62,63]. In some instances, the expression of B3GNT3 inhibits the anti-cancer immune response, as in pancreatic [64], breast [65] and lung cancer [66]. In particular, in triple negative breast cancer, B3GNT3 promotes through EGFR the interaction between PD-1 and PD-L1, resulting in immune escape [65]. These insights are in good agreement with TCGA data, which report an association with poor prognosis in PAAD and LUAD cohorts. However, the tumor-suppressive role of B3GNT3 in pancreatic cancer [67] and neuroblastoma [68] has also been reported.
The second step in polylactosamine biosynthesis involves the addition of a galactose residue either through a β1,3 or a β1,4 linkage, generating type 1 or type 2 chains, respectively (Figure 6A). The enzyme B3GALT4, which both synthesizes type 1 chains and participates in ganglioside biosynthesis (Figure 5), is a GPA, although its association with poor survival in colon cancer has been reported [69]. On the other hand, the BPA B4GALT3 synthesizing type 2 chains behaves as a tumor-promoting gene in neuroblastoma [70,71], glioblastoma [72] and cervical carcinoma [73]. Consistently, B4GALT3 is a predictor of negative prognosis in the endometrial carcinoma (UCEC) cohort. However, B4GALT3 reduces malignancy in colon cancer [74].
5.2.2. LARGE
α-Dystroglycan is a plasma membrane glycoprotein that indirectly links the cytoskeleton with the laminin of the extracellular matrix. The laminin- α-dystroglycan interaction is mediated by its peculiar O-mannosyl glycans [75,76]. The addition of mannose to the peptide is catalyzed by POMT1 and POMT2 (Figure 4D). The chain starting with the first O-linked mannose is elongated by other sugars and terminated by repeated disaccharide units comprised of xylose and glucuronic acid. The glycosyltransferase LARGE is responsible for the biosynthesis of these repeated disaccharide units. TCGA data show that in 6 cohorts, LARGE expression is associated with better prognosis. Although little data have been published on the relationship between LARGE expression and cancer, it has been described that O-mannosylation as a whole exerts tumor-suppressing activity in gastric cancer [77].
5.3. Capping Glycosyltransferases
5.3.1. Sialyltransferases
The BPA sialyltransferases ST3GAL2, ST3GAL4, ST6GALNAC3 and ST6GALNAC4 are involved in the sialylation of both O-linked chains and glycolipids (Figure 4 and Figure 5), while ST3GAL4 sialylates also N-linked chains (Figure 3B). ST3GAL2 is differentially methylated in cancer [78] and is positively associated in oral cancer with advanced stages of the disease, lymph node involvement, and perineural invasion [79]. In addition to its involvement in sialylation of O-linked chains, ST3GAL2 is also a key player in ganglioside biosynthesis [80]. The ganglioside stage-specific embryonic antigen 4 (SSEA4), which is also a ST3GAL2 product, marks chemotherapy-resistant breast cancer cells with mesenchymal features [81]. Although not strictly associated with prognosis in BRCA and HNSC cohorts, the tumor-promoting activity of ST3GAL2 is supported by both experimental and clinical data. ST6GALNAC3 and ST6GALNAC4 are also involved in sialylation of both O-linked chains and glycolipids. ST6GALNAC3 was reduced in lung cancer tissues [82], while increased ST6GALNAC4 enhanced invasion of follicular thyroid carcinoma [83] and lung cancer [84]. Inconsistently, the latter is associated with a better prognosis in LUAD. ST3GAL6 is specific to type 2 chains and is the best predictor of poor survival in STAD. Experimental work has shown that its overexpression in gastric cancer cell lines protects against tyrosine kinase inhibitors [85].
5.3.2. Fucosyltransferases
Fucosyltransferase FUT7 is one of the major α1,3 FUTs involved in the biosynthesis of the cancer-associated sialyl Lewis x antigen (Figure 6B). In this work, we observed that in the LAML cohort, high FUT7 was associated with worse prognosis, confirming a previous study [86]. Although in a variety of other malignancies, including lung [87,88], liver [89], bladder [90], thyroid [91], and breast [92] cancers FUT7 behaves as a tumor-promoting enzyme, in 7 of the TCGA cohorts, including LUAD, it is associated with better overall survival. In addition, FUT7 is the best predictor of good prognosis in BRCA.
6. Mechanistic Aspects of Glycosyltransferase Expression
Like other genes, glycosyltransferases are regulated at multiple levels, including the activity of specific transcription factors, promoter methylation, and the network of non-coding RNAs, such as micro RNA (miRNA), long non-coding RNAs (lnRNA) and circular RNAs (circRNA). On the other hand, glycosylated cell surface molecules, such as growth factor receptors and cell adhesion molecules, trigger multiple signaling pathways, resulting in modulation of cell behavior [93]. Table 3 reports the mechanisms regulating the expression of relevant glycosyltransferases (upstream regulators) and their downstream pathways.

Table 3.
Mechanistic aspects of glycosyltransferase action and regulation.
From these data, it is evident that glycosyltransferases modulate different pathways in different cellular contexts. Sometimes, the activation of the same pathway induces opposite phenotypes in different tissues. For example, B4GALT3 activates β-integrin signaling in both neuroblastoma [71] and colon cancer [74], resulting in progression in the former and inhibition in the latter.
7. Discussion
The present work aims to combine the huge amount of clinical data from the public database TCGA with experimental studies on the glycosyltransferase role in cancer biology. Several key points emerged from the TCGA data analysis. First, some glycosyltransferases (BPA or GPA) are consistently associated with either poor or favorable prognosis in a large number of cohorts, while others (for example, ALG6 and GALNT12) displayed opposite associations in different cohorts. These findings support the notion that a few glycosyltransferases have a pleiotropic effect on several cell types and tissues, while the majority exert their effects in a tissue-specific manner. A paradigmatic example of this statement is provided by the B4GALNT2 gene, whose product synthesizes the carbohydrate antigen Sda. A high level of B4GALNT2 expression is associated with longer overall survival in the COAD cohort and attenuation of malignant phenotype in colon cancer cell lines [105,106]. However, high B4GALNT2 expression correlated with a worse prognosis in the BRCA cohort [107] and increased malignancy in breast cancer cell lines [108]. Some BPA genes are involved in the early steps of N-glycosylation (ALG3, ALG8 and MGAT4B) and of mucin-type O-glycosylation (GALNT2 and GALNT10). Intriguingly, GALNT16, another member of the protein:O-GalNAc transferases, behaves as a GPA, indicating that subtle variations in the first step of O-glycosylation can lead to opposite effects on malignancy. The very strong association of POFUT1 with poor prognosis is probably due to its effect on the first step of NOTCH receptor glycosylation. The BPA group also includes enzymes involved in the biosynthesis of the core portion of glycolipids, such as B4GALT5 and B4GALNT1. B3GNT4, -5, -7 and -9, participating in initiation/extension of polylactosaminic chains, are also BPA, consistent with the recognized role of extended polylatosaminic chains in promoting malignancy. However, of the two galactosyltransferases synthesizing polylactosamines, the one producing type 2 chains (B4GALT3) is a BPA, while that producing type 1 chains (B3GALT4) is a GPA. The gene LARGE, responsible for the elongation of α-dystroglycan sugar chains, represents one of the stronger GPA, probably because of the role of its product in promoting cell adhesion. Among the capping enzymes, we identified 4 sialyltranferases acting mainly on glycolipids and/or O-linked chains behaving as BPA. This finding is not surprising, considering the well-established association of sialyltransferases with malignant phenotype [109,110]. By contrast, fucosyltransferases, another major class of capping enzymes, displayed an opposite behavior. This was unexpected, considering that several members of this group (FUT3-7) are responsible for the biosynthesis of well-known cancer-associated Lewis type antigens and their sialylated counterparts sialyl Lewis x and sialyl Lewis a [111]. FUT7 was found to be a GPA and a best predictor of good prognosis in BRCA, despite experimental studies showing its tumor-promoting activity. Several glycosyltransferases, including MGAT5 [112], FUT8 [113], ST6GAL1 [110], ST6GALNAC1 [114,115] and ST8SIA1 [116] have an established reputation as tumor-promoting enzymes. On the other hand, MGAT3 is probably the best-recognized tumor-restraining glycosyltransferase [117,118]. However, no one of these enzymes displays a relevant association with prognosis in different cohorts. Comparison of TCGA data with literature indicates a consistent malignancy-oriented behavior by some glycosyltransferases, including ALG3, GALNT2, B4GALNT1, POFUT1, B4GALT5, B3GNT5 and ST3GAL2. On the other hand, the profile of other glycosyltransferases emerging from TCGA data analysis appears to be inconsistent with that emerging from experimental studies. This group includes B3GALT5, B3GNT7, B3GALT4 and FUT7. The limited consistency between the experimental and clinical data could be explained by the fact that cell lines derived from a single or a few cancer cases might not be representative of the many patients of the whole cohort. Moreover, transcriptomic data are not necessarily representative of enzyme activity and cancer antigen expression levels. In fact, the biosynthesis of a given carbohydrate antigen is the final effect of many factors, including the translational efficiency of glycosyltransferase mRNA, the half-life of enzyme protein, the effect of postranslational modifications on enzmatic activity, the availability of donor and acceptor substrates, the competition with other glycosyltransferases and probably many others. In addition to the identification of glycosyltransferases playing a pleiotropic effect in many cohorts, we also pursued the identification of glycosyltransferases with a very high prognostic value (VHPV), in which the overall survival of the top 15% expressers was statistically different from that of the bottom 15% expressers with a p < 1 × 10−3. There were no VHPV glycosyltransferases in some cohorts, such as BRCA, while in others, such as LGG and KIRC, they were numerous. These discrepancies suggest that several tumors display intrinsically different sensitivity to glycosylation changes. We have shown that glycosyltransferases involved in the biosynthesis of different sugar chains are able to activate relatively few signal transduction pathways. EGFR/AKT appears to be one of the most frequently involved. Among the mechanisms regulating glycosyltransferase expression, the contribution of non-coding RNAs is increasingly recognized. The complex network of interactions between lncRNA, circRNA and miRNAs is essential to ensure the fine-tuning of glycosyltransferase expression. Considering the huge therapeutic importance of immune checkpoint inhibitors targeting the PD-1/PD-L1 interaction, it is worth mentioning that such interaction is modulated by glycosylation and that glycosylation inhibitors are able to revert the cancer-induced inhibition of the immune system [65,119,120,121,122,123,124].
8. Conclusions
In conclusion, the wide analysis of TCGA data allows the identification of glycosyltransferases whose over- or under-expression impacts patients’ overall survival more dramatically. Even if the studies on experimental systems remain crucial to understanding the molecular mechanisms linking glycosyltransferase expression and malignancy, information from databases appears to be the best way to identify glycosyltransferases as potential biomarkers, either alone or in combination [125,126,127,128]. Owing to their very strict association with survival in specific malignancies, VHPV glycosyltransferases are ideal candidates as prognostic biomarkers and targets of therapeutic approaches.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/cancers14092128/s1, Table S1: Prognostic value of glycosyltransferase gene expression in TCGA cohorts.
Author Contributions
Conceptualization, F.D.; methodology, M.P.; validation, M.D. and N.M.; formal analysis, M.P. and M.D.; investigation, F.D. and M.P.; data curation, N.M.; writing—original draft preparation, F.D. and N.M.; writing—review and editing, F.D. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| BPA | bad prognosis-associated |
| circRNA | circular RNA |
| GPA | good prognosis-associated |
| lncRNA | long non-coding RNA |
| PD-1 | programmed death 1 |
| PD-L1 | programmed death ligand 1 |
| VHPV | very high prognostic value |
| BLCA | bladder urothelial carcinoma |
| BRCA | breast invasive carcinoma |
| CESC | cervical squamous cell carcinoma and endocervical adenocarcinoma |
| COAD | colon adenocarcinoma |
| ESCA | esophageal carcinoma |
| GBM | glioblastoma multiforme |
| HNSC | head and neck squamous cell carcinoma |
| KIRC | kidney renal clear cell carcinoma |
| KIRP | kidney renal papillary carcinoma |
| LAML | acute myeloid leukemia |
| LGG | brain lower grade glioma |
| LIHC | liver hepatocellular carcinoma |
| LUAD | lung adenocarcinoma |
| LUSC | lung squamous cell carcinoma |
| OV | ovary serous cystadenocarcinoma |
| PAAD | pancreatic adenocarcinoma |
| READ | rectum adenocarcinoma |
| SARC | sarcoma |
| SKCM | skin cutaneous melanoma |
| STAD | stomach adenocarcinoma |
| TCGA | the cancer genome atlas |
| UCEC | uterine corpus endometrial carcinoma |
References
- Varki, A. Biological roles of glycans. Glycobiology 2017, 27, 3–49. [Google Scholar] [CrossRef] [Green Version]
- Dall’Olio, F.; Malagolini, N.; Trinchera, M.; Chiricolo, M. Mechanisms of cancer-associated glycosylation changes. Front. Biosci. 2012, 17, 670–699. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pinho, S.S.; Reis, C.A. Glycosylation in cancer: Mechanisms and clinical implications. Nat. Rev. Cancer 2015, 15, 540–555. [Google Scholar] [CrossRef] [PubMed]
- Schjoldager, K.T.; Narimatsu, Y.; Joshi, H.J.; Clausen, H. Global view of human protein glycosylation pathways and functions. Nat. Rev. Mol. Cell Biol. 2020, 21, 729–749. [Google Scholar] [CrossRef] [PubMed]
- Ke, S.B.; Qiu, H.; Chen, J.M.; Shi, W.; Han, C.; Gong, Y.; Chen, Y.S. ALG3 contributes to the malignancy of non-small cell lung cancer and is negatively regulated by MiR-98-5p. Pathol.-Res. Pract. 2020, 216, 152761. [Google Scholar] [CrossRef] [PubMed]
- Shao, P.; Wei, C.; Wang, Y. ALG3 contributes to the malignant properties of OSCC cells by regulating CDK-Cyclin pathway. Oral Dis. 2021, 27, 1426–1434. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.Z.; Jiang, Y.Y.; Hao, J.J.; Zhang, Y.; Zhang, T.T.; Shang, L.; Liu, S.G.; Shi, F.; Wang, M.R. Identification of putative target genes for amplification within 11q13.2 and 3q27.1 in esophageal squamous cell carcinoma. Clin. Transl. Oncol. 2014, 16, 606–615. [Google Scholar] [CrossRef]
- Choi, Y.W.; Bae, S.M.; Kim, Y.W.; Lee, H.N.; Kim, Y.W.; Park, T.C.; Ro, D.Y.; Shin, J.C.; Shin, S.J.; Seo, J.S.; et al. Gene expression profiles in squamous cell cervical carcinoma using array-based comparative genomic hybridization analysis. Int. J. Gynecol. Cancer 2007, 17, 687–696. [Google Scholar] [CrossRef]
- Zhou, W.; Li, J.; Lu, X.; Liu, F.; An, T.; Xiao, X.; Kuo, Z.C.; Wu, W.; He, Y. Derivation and Validation of a Prognostic Model for Cancer Dependency Genes Based on CRISPR-Cas9 in Gastric Adenocarcinoma. Front. Oncol. 2021, 11, 617289. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, D.; Chen, Y.; Lu, Y.; Zhou, F.; Li, C.; Zeng, Z.; Cai, W.; Lin, L.; Li, Q.; et al. Robust Glycogene-Based Prognostic Signature for Proficient Mismatch Repair Colorectal Adenocarcinoma. Front. Oncol. 2021, 11, 727752. [Google Scholar] [CrossRef] [PubMed]
- Kato, K.; Hansen, L.; Clausen, H. Polypeptide N-acetylgalactosaminyltransferase-Associated Phenotypes in Mammals. Molecules 2021, 26, 5504. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.C.; Huang, M.J.; Liu, C.H.; Yang, T.L.; Huang, M.C. GALNT2 enhances migration and invasion of oral squamous cell carcinoma by regulating EGFR glycosylation and activity. Oral Oncol. 2014, 50, 478–484. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Xue, H.; Wei, Y.; Wang, C.; Yu, R.; Wang, C.; Wang, S.; Xu, J.; Qian, M.; Meng, Q.; et al. Mucin O-glycosylating enzyme GALNT2 facilitates the malignant character of glioma by activating the EGFR/PI3K/Akt/mTOR axis. Clin. Sci. 2019, 133, 1167–1184. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Xu, Y.; Yin, D.; Zhao, F.; Hao, Z.; Zhong, Y.; Zhang, J.; Zhang, B.; Yin, X. Type 2 diabetes mellitus facilitates endometrial hyperplasia progression by activating the proliferative function of mucin O-glycosylating enzyme GALNT2. Biomed. Pharmacother. 2020, 131, 110764. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Sun, R.; Zeng, L.; Chen, Y.; Zhang, N.; Cao, S.; Deng, S.; Meng, X.; Yang, S. GALNT2 promotes cell proliferation, migration, and invasion by activating the Notch/Hes1-PTEN-PI3K/Akt signaling pathway in lung adenocarcinoma. Life Sci. 2021, 276, 119439. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Wang, Z.; Zheng, Q.; Li, J. GALNT2/14 overexpression correlate with prognosis and methylation: Potential therapeutic targets for lung adenocarcinoma. Gene 2021, 790, 145689. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.M.; Liu, C.H.; Hu, R.H.; Huang, M.J.; Lee, J.J.; Chen, C.H.; Huang, J.; Lai, H.S.; Lee, P.H.; Hsu, W.M.; et al. Mucin Glycosylating Enzyme GALNT2 Regulates the Malignant Character of Hepatocellular Carcinoma by Modifying the EGF Receptor. Cancer Res. 2011, 71, 7270–7279. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.Y.; Shun, C.T.; Hung, K.Y.; Juan, H.F.; Hsu, C.L.; Huang, M.C.; Lai, I.R. Mucin glycosylating enzyme GALNT2 suppresses malignancy in gastric adenocarcinoma by reducing MET phosphorylation. Oncotarget 2016, 7, 11251–11262. [Google Scholar] [CrossRef]
- Zhang, G.; Lu, J.; Yang, M.; Wang, Y.; Liu, H.; Xu, C. Elevated GALNT10 expression identifies immunosuppressive microenvironment and dismal prognosis of patients with high grade serous ovarian cancer. Cancer Immunol. Immunother. 2020, 69, 175–187. [Google Scholar] [CrossRef]
- Liu, Z.; Pan, L.; Yan, X.; Duan, X. The long noncoding RNA DLGAP1-AS2 facilitates cholangiocarcinoma progression via miR-505 and GALNT10. FEBS Open Bio 2021, 11, 413–422. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Liu, H.O.; Liu, Y.D.; Liu, W.S.; Pan, D.; Zhang, W.J.; Yang, L.; Fu, Q.; Xu, J.J.; Gu, J.X. Decreased Expression of Hepatocyte Nuclear Factor 4α (Hnf4α)/MicroRNA-122 (miR-122) Axis in Hepatitis B Virus-associated Hepatocellular Carcinoma Enhances Potential Oncogenic GALNT10 Protein Activity. J. Biol. Chem. 2015, 290, 1170–1185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, J.B.; Pyo, K.H.; Kim, H.R. Role and Function of O-GlcNAcylation in Cancer. Cancers 2021, 13, 5365. [Google Scholar] [CrossRef] [PubMed]
- Rozanski, W.; Krzeslak, A.; Forma, E.; Brys, M.; Blewniewski, M.; Wozniak, P.; Lipinski, M. Prediction of bladder cancer based on urinary content of MGEA5 and OGT mRNA level. Clin. Lab. 2012, 58, 579–583. [Google Scholar] [PubMed]
- Wang, L.; Chen, S.; Zhang, Z.; Zhang, J.; Mao, S.; Zheng, J.; Xuan, Y.; Liu, M.; Cai, K.; Zhang, W.; et al. Suppressed OGT expression inhibits cell proliferation while inducing cell apoptosis in bladder cancer. BMC Cancer 2018, 18, 1141. [Google Scholar] [CrossRef] [Green Version]
- Holdener, B.C.; Haltiwanger, R.S. Protein O-fucosylation: Structure and function. Curr. Opin. Struct. Biol. 2019, 56, 78–86. [Google Scholar] [CrossRef]
- Chabanais, J.; Labrousse, F.; Chaunavel, A.; Germot, A.; Maftah, A. POFUT1 as a Promising Novel Biomarker of Colorectal Cancer. Cancers 2018, 10, 411. [Google Scholar] [CrossRef] [Green Version]
- Komor, M.A.; de Wit, M.; van den Berg, J.; Martens de Kemp, S.R.; Delis-van Diemen, P.M.; Bolijn, A.S.; Tijssen, M.; Schelfhorst, T.; Piersma, S.R.; Chiasserini, D.; et al. Molecular characterization of colorectal adenomas reveals POFUT1 as a candidate driver of tumor progression. Int. J. Cancer 2020, 146, 1979–1992. [Google Scholar] [CrossRef]
- Du, Y.; Li, D.; Li, N.; Su, C.; Yang, C.; Lin, C.; Chen, M.; Wu, R.; Li, X.; Hu, G. POFUT1 promotes colorectal cancer development through the activation of Notch1 signaling. Cell Death Dis. 2018, 9, 995. [Google Scholar] [CrossRef]
- Li, D.; Lin, C.; Li, N.; Du, Y.; Yang, C.; Bai, Y.; Feng, Z.; Su, C.; Wu, R.; Song, S.; et al. PLAGL2 and POFUT1 are regulated by an evolutionarily conserved bidirectional promoter and are collaboratively involved in colorectal cancer by maintaining stemness. EBioMedicine 2019, 45, 124–138. [Google Scholar] [CrossRef] [Green Version]
- Deschuyter, M.; Pennarubia, F.; Pinault, E.; Legardinier, S.; Maftah, A. Functional Characterization of POFUT1 Variants Associated with Colorectal Cancer. Cancers 2020, 12, 1430. [Google Scholar] [CrossRef]
- Annani-Akollor, M.E.; Wang, S.; Fan, J.; Liu, L.; Padhiar, A.A.; Zhang, J. Downregulated protein O-fucosyl transferase 1 (Pofut1) expression exerts antiproliferative and antiadhesive effects on hepatocytes by inhibiting Notch signalling. Biomed. Pharmacother. 2014, 68, 785–790. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Dong, P.; Liu, L.; Gao, Q.; Duan, M.; Zhang, S.; Chen, S.; Xue, R.; Wang, X. Overexpression of protein O-fucosyltransferase 1 accelerates hepatocellular carcinoma progression via the Notch signaling pathway. Biochem. Biophys. Res. Commun. 2016, 473, 503–510. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Huang, H.; Zhang, J.; Wu, Q.; Chen, X.; Huang, T.; Li, W.; Liu, Y.; Zhang, J. Caveolin-1 promotes invasion and metastasis by upregulating Pofut1 expression in mouse hepatocellular carcinoma. Cell Death Dis. 2019, 10, 477. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wang, J.; Ma, X.; Wang, M.; Zhou, L. POFUT1 acts as a tumor promoter in glioblastoma by enhancing the activation of Notch signaling. J. Bioenerg. Biomembr. 2021, 53, 621–632. [Google Scholar] [CrossRef]
- Leng, Q.; Tsou, J.H.; Zhan, M.; Jiang, F. Fucosylation genes as circulating biomarkers for lung cancer. J. Cancer Res. Clin. Oncol. 2018, 144, 2109–2115. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Wang, Z.; Huang, B.; Zhang, J.; Ge, Y.; Fan, Q.; Wang, Z. Bioinformatics insight into glycosyltransferase gene expression in gastric cancer: POFUT1 is a potential biomarker. Biochem. Biophys. Res. Commun. 2017, 483, 171–177. [Google Scholar] [CrossRef] [PubMed]
- Sadeghzadeh, Z.; Khosravi, A.; Jazi, M.S.; Asadi, J. Upregulation of Fucosyltransferase 3, 8 and protein O-Fucosyltransferase 1, 2 genes in esophageal cancer stem-like cells (CSLCs). Glycoconj. J. 2020, 37, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Wan, G.; Tian, L.; Yu, Y.; Li, F.; Wang, X.; Li, C.; Deng, S.; Yu, X.; Cai, X.; Zuo, Z.; et al. Overexpression of Pofut1 and activated Notch1 may be associated with poor prognosis in breast cancer. Biochem. Biophys. Res. Commun. 2017, 491, 104–111. [Google Scholar] [CrossRef]
- Yokota, S.; Ogawara, K.; Kimura, R.; Shimizu, F.; Baba, T.; Minakawa, Y.; Higo, M.; Kasamatsu, A.; Endo-Sakamoto, Y.; Shiiba, M.; et al. Protein O-fucosyltransferase 1: A potential diagnostic marker and therapeutic target for human oral cancer. Int. J. Oncol. 2013, 43, 1864–1870. [Google Scholar] [CrossRef] [Green Version]
- Wahby, S.; Jarczyk, J.; Fierek, A.; Heinkele, J.; Weis, C.A.; Eckstein, M.; Martini, T.; Porubsky, S.; Hafner, M.; Erben, P. POFUT1 mRNA expression as an independent prognostic parameter in muscle-invasive bladder cancer. Transl. Oncol. 2021, 14, 100900. [Google Scholar] [CrossRef]
- Zhang, F.; Zhang, H. UDP-Glucose Ceramide Glycosyltransferase Contributes to the Proliferation and Glycolysis of Cervical Cancer Cells by Regulating the PI3K/AKT Pathway. Ann. Clin. Lab. Sci. 2021, 51, 663–669. [Google Scholar] [PubMed]
- Tokuda, N.; Numata, S.; Li, X.; Nomura, T.; Takizawa, M.; Kondo, Y.; Yamashita, Y.; Hashimoto, N.; Kiyono, T.; Urano, T.; et al. β4GalT6 is involved in the synthesis of lactosylceramide with less intensity than β4GalT5. Glycobiology 2013, 23, 1175–1183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamaji, T.; Hanada, K. Establishment of HeLa cell mutants deficient in sphingolipid-related genes using TALENs. PLoS ONE 2014, 9, e88124. [Google Scholar] [CrossRef] [Green Version]
- Tang, W.; Li, M.; Qi, X.; Li, J. β1,4-Galactosyltransferase V Modulates Breast Cancer Stem Cells through Wnt/β-catenin Signaling Pathway. Cancer Res. Treat. 2020, 52, 1084–1102. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Ma, H.; Wei, W.; Ji, D.; Song, X.; Sun, J.; Zhang, J.; Jia, L. B4GALT family mediates the multidrug resistance of human leukemia cells by regulating the hedgehog pathway and the expression of p-glycoprotein and multidrug resistance-associated protein 1. Cell Death Dis. 2013, 4, e654. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liang, Y.J.; Ding, Y.; Levery, S.B.; Lobaton, M.; Handa, K.; Hakomori, S.I. Differential expression profiles of glycosphingolipids in human breast cancer stem cells vs. cancer non-stem cells. Proc. Natl. Acad. Sci. USA 2013, 110, 4968–4973. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, T.; Wu, H.; Lin, M.; Yin, J.; Tan, L.; Ruan, Y.; Feng, M. B4GALNT1 promotes progression and metastasis in lung adenocarcinoma through JNK/c-Jun/Slug pathway. Carcinogenesis 2021, 42, 621–630. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Mahata, B.; Dhir, A.; Mandal, T.K.; Biswas, K. Elevated histone H3 acetylation and loss of the Sp1-HDAC1 complex de-repress the GM2-synthase gene in renal cell carcinoma. J. Biol. Chem. 2019, 294, 1005–1018. [Google Scholar] [CrossRef] [Green Version]
- Mahata, B.; Banerjee, A.; Kundu, M.; Bandyopadhyay, U.; Biswas, K. TALEN mediated targeted editing of GM2/GD2-synthase gene modulates anchorage independent growth by reducing anoikis resistance in mouse tumor cells. Sci. Rep. 2015, 5, 9048. [Google Scholar] [CrossRef]
- Liang, Y.J.; Wang, C.Y.; Wang, I.A.; Chen, Y.W.; Li, L.T.; Lin, C.Y.; Ho, M.Y.; Chou, T.L.; Wang, Y.H.; Chiou, S.P.; et al. Interaction of glycosphingolipids GD3 and GD2 with growth factor receptors maintains breast cancer stem cell phenotype. Oncotarget 2017, 8, 47454–47473. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, H.; Koodie, L.; Jacobsen, K.; Hanzawa, K.; Miyamoto, Y.; Yamamoto, M. B4GALNT1 induces angiogenesis, anchorage independence growth and motility, and promotes tumorigenesis in melanoma by induction of ganglioside GM2/GD2. Sci. Rep. 2020, 10, 1199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kundu, M.; Mahata, B.; Banerjee, A.; Chakraborty, S.; Debnath, S.; Ray, S.S.; Ghosh, Z.; Biswas, K. Ganglioside GM2 mediates migration of tumor cells by interacting with integrin and modulating the downstream signaling pathway. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2016, 1863, 1472–1489. [Google Scholar] [CrossRef] [PubMed]
- Pereira, C.S.; Ribeiro, H.; Perez-Cabezas, B.; Cardoso, M.T.; Alegrete, N.; Gaspar, A.; Leao-Teles, E.; Macedo, M.F. The GM2 ganglioside inhibits iNKT cell responses in a CD1d-dependent manner. Mol. Genet. Metab. 2018, 125, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.Y.; Park, S.Y.; Kim, H.J.; Moon, S.; Lee, S.; Lee, S.H.; Kim, S.H. B3GNT5 is a novel marker correlated with stem-like phenotype and poor clinical outcome in human gliomas. CNS Neurosci. Ther. 2020, 26, 1147–1154. [Google Scholar] [CrossRef] [PubMed]
- Magalhaes, A.; Marcos-Pinto, R.; Nairn, A.V.; Dela, R.M.; Ferreira, R.M.; Junqueira-Neto, S.; Freitas, D.; Gomes, J.; Oliveira, P.; Santos, M.R.; et al. Helicobacter pylori chronic infection and mucosal inflammation switches the human gastric glycosylation pathways. Biochim. Biophys. Acta (BBA)-Mol. Basis Dis. 2015, 1852, 1928–1939. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carroll, D.J.; Burns, M.W.N.; Mottram, L.; Propheter, D.C.; Boucher, A.; Lessen, G.M.; Kumar, A.; Malaker, S.A.; Xing, C.; Hooper, L.V.; et al. Interleukin-22 regulates B3GNT7 expression to induce fucosylation of glycoproteins in intestinal epithelial cells. J. Biol. Chem. 2021, 298, 101463. [Google Scholar] [CrossRef]
- Lu, C.H.; Wu, W.Y.; Lai, Y.J.; Yang, C.M.; Yu, L.C. Suppression of B3GNT7 gene expression in colon adenocarcinoma and its potential effect in the metastasis of colon cancer cells. Glycobiology 2014, 24, 359–367. [Google Scholar] [CrossRef] [Green Version]
- Barkeer, S.; Chugh, S.; Karmakar, S.; Kaushik, G.; Rauth, S.; Rachagani, S.; Batra, S.K.; Ponnusamy, M.P. Novel role of O-glycosyltransferases GALNT3 and B3GNT3 in the self-renewal of pancreatic cancer stem cells. BMC Cancer 2018, 18, 1157. [Google Scholar] [CrossRef]
- Zhuang, H.; Zhou, Z.; Zhang, Z.; Chen, X.; Ma, Z.; Huang, S.; Gong, Y.; Zhang, C.; Hou, B. B3GNT3 overexpression promotes tumor progression and inhibits infiltration of CD8(+) T cells in pancreatic cancer. Aging 2021, 13, 2310–2329. [Google Scholar] [CrossRef]
- Zhang, W.; Hou, T.; Niu, C.; Song, L.; Zhang, Y. B3GNT3 Expression Is a Novel Marker Correlated with Pelvic Lymph Node Metastasis and Poor Clinical Outcome in Early-Stage Cervical Cancer. PLoS ONE 2015, 10, e0144360. [Google Scholar] [CrossRef]
- Wang, J.S.; Ruan, F.; Guo, L.Z.; Wang, F.G.; Wang, F.L.; An, H.M. B3GNT3 acts as a carcinogenic factor in endometrial cancer via facilitating cell growth, invasion and migration through regulating RhoA/RAC1 pathway-associated markers. Genes Genom. 2021, 43, 447–457. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Liu, T.; Xian, L.; Liu, W.; Liu, J.; Zhou, H. B3GNT3, a Direct Target of miR-149-5p, Promotes Lung Cancer Development and Indicates Poor Prognosis of Lung Cancer. Cancer Manag. Res. 2020, 12, 2381–2391. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, L.; Zhang, H.; Zhang, B.; Zhu, J.; Chen, C.; Liu, W. B3GNT3 overexpression is associated with unfavourable survival in non-small cell lung cancer. J. Clin. Pathol. 2018, 71, 642–647. [Google Scholar] [CrossRef] [PubMed]
- Kong, K.; Zhao, Y.; Xia, L.; Jiang, H.; Xu, M.; Zheng, J. B3GNT3: A prognostic biomarker associated with immune cell infiltration in pancreatic adenocarcinoma. Oncol. Lett. 2021, 21, 159. [Google Scholar] [CrossRef] [PubMed]
- Li, C.W.; Lim, S.O.; Chung, E.M.; Kim, Y.S.; Park, A.H.; Yao, J.; Cha, J.H.; Xia, W.; Chan, L.C.; Kim, T.; et al. Eradication of Triple-Negative Breast Cancer Cells by Targeting Glycosylated PD-L1. Cancer Cell 2018, 33, 187–201. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leng, X.; Wei, S.; Mei, J.; Deng, S.; Yang, Z.; Liu, Z.; Guo, C.; Deng, Y.; Xia, L.; Cheng, J.; et al. Identifying the prognostic significance of B3GNT3 with PD-L1 expression in lung adenocarcinoma. Transl. Lung Cancer Res. 2021, 10, 965–980. [Google Scholar] [CrossRef]
- Gupta, R.; Leon, F.; Thompson, C.M.; Nimmakayala, R.; Karmakar, S.; Nallasamy, P.; Chugh, S.; Prajapati, D.R.; Rachagani, S.; Kumar, S.; et al. Global analysis of human glycosyltransferases reveals novel targets for pancreatic cancer pathogenesis. Br. J. Cancer 2020, 122, 1661–1672. [Google Scholar] [CrossRef]
- Ho, W.L.; Che, M.I.; Chou, C.H.; Chang, H.H.; Jeng, Y.M.; Hsu, W.M.; Lin, K.H.; Huang, M.C. B3GNT3 expression suppresses cell migration and invasion and predicts favorable outcomes in neuroblastoma. Cancer Sci. 2013, 104, 1600–1608. [Google Scholar] [CrossRef]
- Zhang, T.; Wang, F.; Wu, J.Y.; Qiu, Z.C.; Wang, Y.; Liu, F.; Ge, X.S.; Qi, X.W.; Mao, Y.; Hua, D. Clinical correlation of B7-H3 and B3GALT4 with the prognosis of colorectal cancer. World J. Gastroenterol. 2018, 24, 3538–3546. [Google Scholar] [CrossRef]
- Bi, C.; Shan, J.; Li, M.; Zhang, Q.; Li, C.; Tong, J.; Huang, Q. Long noncoding RNA differentiation antagonizing nonprotein coding RNA promotes the proliferation, invasion and migration of neuroblastoma cells via targeting β-1, 4-galactosyltransferase III by sponging miR-338-3p. Neuroreport 2021, 32, 965–974. [Google Scholar] [CrossRef]
- Chang, H.H.; Chen, C.H.; Chou, C.H.; Liao, Y.F.; Huang, M.J.; Chen, Y.H.; Wang, W.J.; Huang, J.; Hung, J.S.; Ho, W.L.; et al. β-1,4-Galactosyltransferase III Enhances Invasive Phenotypes Via β1-Integrin and Predicts Poor Prognosis in Neuroblastoma. Clin. Cancer Res. 2013, 19, 1705–1716. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, T.; Li, Y.; Chen, B. B4GALT3 promotes cell proliferation and invasion in glioblastoma. Neurol. Res. 2020, 42, 463–470. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Yang, X.; Liu, M.; Tang, H. B4GALT3 up-regulation by miR-27a contributes to the oncogenic activity in human cervical cancer cells. Cancer Lett. 2016, 375, 284–292. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.H.; Wang, S.H.; Liu, C.H.; Wu, Y.L.; Wang, W.J.; Huang, J.; Hung, J.S.; Lai, I.R.; Liang, J.T.; Huang, M.C. β-1,4-Galactosyltransferase III suppresses β1 integrin-mediated invasive phenotypes and negatively correlates with metastasis in colorectal cancer. Carcinogenesis 2014, 35, 1258–1266. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sheikh, M.O.; Halmo, S.M.; Wells, L. Recent advancements in understanding mammalian O-mannosylation. Glycobiology 2017, 27, 806–819. [Google Scholar] [CrossRef] [Green Version]
- Wells, L. The O-mannosylation pathway: Glycosyltransferases and proteins implicated in congenital muscular dystrophy. J. Biol. Chem. 2013, 288, 6930–6935. [Google Scholar] [CrossRef] [Green Version]
- Carvalho, S.; Oliveira, T.; Bartels, M.F.; Miyoshi, E.; Pierce, M.; Taniguchi, N.; Carneiro, F.; Seruca, R.; Reis, C.A.; Strahl, S.; et al. O-mannosylation and N-glycosylation: Two coordinated mechanisms regulating the tumour suppressor functions of E-cadherin in cancer. Oncotarget 2016, 7, 65231–65246. [Google Scholar] [CrossRef] [Green Version]
- Vojta, A.; Samarzija, I.; Bockor, L.; Zoldos, V. Glyco-genes change expression in cancer through aberrant methylation. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2016, 1860, 1776–1785. [Google Scholar] [CrossRef]
- Mehta, K.A.; Patel, K.A.; Pandya, S.J.; Patel, P.S. Aberrant sialylation plays a significant role in oral squamous cell carcinoma progression. J. Oral Pathol. Med. 2020, 49, 253–259. [Google Scholar] [CrossRef]
- Sturgill, E.R.; Aoki, K.; Lopez, P.H.; Colacurcio, D.; Vajn, K.; Lorenzini, I.; Majic, S.; Yang, W.H.; Heffer, M.; Tiemeyer, M.; et al. Biosynthesis of the major brain gangliosides GD1a and GT1b. Glycobiology 2012, 22, 1289–1301. [Google Scholar] [CrossRef] [Green Version]
- Aloia, A.; Petrova, E.; Tomiuk, S.; Bissels, U.; Deas, O.; Saini, M.; Zickgraf, F.M.; Wagner, S.; Spaich, S.; Sutterlin, M.; et al. The sialyl-glycolipid stage-specific embryonic antigen 4 marks a subpopulation of chemotherapy-resistant breast cancer cells with mesenchymal features. Breast Cancer Res. 2015, 17, 146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, Q.; Chen, X.; Han, Y.; Lei, T.; Wu, Q.; Yu, X.; Wang, L.; Fan, Z.; Wang, S. Modification of α2,6-sialylation mediates the invasiveness and tumorigenicity of non-small cell lung cancer cells in vitro and in vivo via Notch1/Hes1/MMPs pathway. Int. J. Cancer 2018, 143, 2319–2330. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miao, X.; Jia, L.; Zhou, H.; Song, X.; Zhou, M.; Xu, J.; Zhao, L.; Feng, X.; Zhao, Y. miR-4299 mediates the invasive properties and tumorigenicity of human follicular thyroid carcinoma by targeting ST6GALNAC4. IUBMB Life 2016, 68, 136–144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reticker-Flynn, N.E.; Bhatia, S.N. Aberrant Glycosylation Promotes Lung Cancer Metastasis through Adhesion to Galectins in the Metastatic Niche. Cancer Discov. 2015, 5, 168–181. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Balmana, M.; Diniz, F.; Feijao, T.; Barrias, C.C.; Mereiter, S.; Reis, C.A. Analysis of the Effect of Increased α2,3-Sialylation on RTK Activation in MKN45 Gastric Cancer Spheroids Treated with Crizotinib. Int. J. Mol. Sci. 2020, 21, 722. [Google Scholar] [CrossRef] [Green Version]
- Dai, Y.; Cheng, Z.; Pang, Y.; Jiao, Y.; Qian, T.; Quan, L.; Cui, L.; Liu, Y.; Si, C.; Chen, J.; et al. Prognostic value of the FUT family in acute myeloid leukemia. Cancer Gene Ther. 2020, 27, 70–80. [Google Scholar] [CrossRef]
- Jassam, S.A.; Maherally, Z.; Ashkan, K.; Pilkington, G.J.; Fillmore, H.L. Fucosyltransferase 4 and 7 mediates adhesion of non-small cell lung cancer cells to brain-derived endothelial cells and results in modification of the blood-brain-barrier: In vitro investigation of CD15 and CD15s in lung-to-brain metastasis. J. Neuro-Oncol. 2019, 143, 405–415. [Google Scholar] [CrossRef] [Green Version]
- Liang, J.X.; Gao, W.; Cai, L. Fucosyltransferase VII promotes proliferation via the EGFR/AKT/mTOR pathway in A549 cells. OncoTargets Ther. 2017, 10, 3971–3978. [Google Scholar] [CrossRef] [Green Version]
- Li, D.; Sun, H.; Bai, G.; Wang, W.; Liu, M.; Bao, Z.; Li, J.; Liu, H. α-1,3-Fucosyltransferase-VII siRNA inhibits the expression of SLex and hepatocarcinoma cell proliferation. Int. J. Mol. Med. 2018, 42, 2700–2708. [Google Scholar] [CrossRef] [Green Version]
- Liu, M.; Zheng, Q.; Chen, S.; Liu, J.; Li, S. FUT7 Promotes the Epithelial-Mesenchymal Transition and Immune Infiltration in Bladder Urothelial Carcinoma. J. Inflamm. Res. 2021, 14, 1069–1084. [Google Scholar] [CrossRef]
- Qin, H.; Liu, J.; Yu, M.; Wang, H.; Thomas, A.M.; Li, S.; Yan, Q.; Wang, L. FUT7 promotes the malignant transformation of follicular thyroid carcinoma through alpha1,3-fucosylation of EGF receptor. Exp. Cell Res. 2020, 393, 112095. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Liu, J.; Xia, Y.; Wang, Z.; Li, X.; Gao, Q. Integrated analysis reveals the participation of IL4I1, ITGB7, and FUT7 in reshaping the TNBC immune microenvironment by targeting glycolysis. Ann. Med. 2021, 53, 916–928. [Google Scholar] [CrossRef] [PubMed]
- Gomes Ferreira, I.; Pucci, M.; Venturi, G.; Malagolini, N.; Chiricolo, M.; Dall’Olio, F. Glycosylation as a Main Regulator of Growth and Death Factor Receptors Signaling. Int. J. Mol. Sci. 2018, 19, 580. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.; He, Z.; Guo, L.; Wang, C.; Lin, C.; Ye, L.; Wang, X.; Li, Y.; Yang, M.; Liu, S.; et al. ALG3 contributes to stemness and radioresistance through regulating glycosylation of TGF-beta receptor II in breast cancer. J. Exp. Clin. Cancer Res. 2021, 40, 149. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhou, Y.; Xiong, X.; Huang, M.; Ying, X.; Wang, M. ALG3 Is Activated by Heat Shock Factor 2 and Promotes Breast Cancer Growth. Med. Sci. Monit. 2018, 24, 3479–3487. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Li, B.; Xu, G.; Han, C.; Xing, G. lncRNA MIR44352HG promotes the progression of liver cancer by upregulating B3GNT5 expression. Mol. Med. Rep. 2022, 25, 38. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhao, B.; Chen, X.; Geng, X.; Zhang, Z. Circ_0009910 sponges miR-491-5p to promote acute myeloid leukemia progression through modulating B4GALT5 expression and PI3K/AKT signaling pathway. Int. J. Lab. Hematol. 2022, 44, 320–332. [Google Scholar] [CrossRef] [PubMed]
- Ho, W.L.; Chou, C.H.; Jeng, Y.M.; Lu, M.Y.; Yang, Y.L.; Jou, S.T.; Lin, D.T.; Chang, H.H.; Lin, K.H.; Hsu, W.M.; et al. GALNT2 suppresses malignant phenotypes through IGF-1 receptor and predicts favorable prognosis in neuroblastoma. Oncotarget 2014, 5, 12247–12259. [Google Scholar] [CrossRef] [Green Version]
- Zhao, F.Y.; Zhang, Q.; Wang, J.M.; Jiang, J.Y.; Huyan, L.Y.; Liu, B.Q.; Yan, J.; Li, C.; Wang, H.Q. BAG3 epigenetically regulates GALNT10 expression via WDR5 and facilitates the stem cell-like properties of platin-resistant ovarian cancer cells. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2021, 1868, 119077. [Google Scholar] [CrossRef]
- Gomes, C.; Osorio, H.; Pinto, M.T.; Campos, D.; Oliveira, M.J.; Reis, C.A. Expression of ST3GAL4 leads to SLex expression and induces c-Met activation and an invasive phenotype in gastric carcinoma cells. PLoS ONE 2013, 8, e66737. [Google Scholar] [CrossRef]
- Mereiter, S.; Magalhaes, A.; Adamczyk, B.; Jin, C.; Almeida, A.; Drici, L.; Ibanez-Vea, M.; Gomes, C.; Ferreira, J.A.; Afonso, L.P.; et al. Glycomic analysis of gastric carcinoma cells discloses glycans as modulators of RON receptor tyrosine kinase activation in cancer. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2016, 1860, 1795–1808. [Google Scholar] [CrossRef] [PubMed]
- Greville, G.; Llop, E.; Huang, C.; Creagh-Flynn, J.; Pfister, S.; O’Flaherty, R.; Madden, S.F.; Peracaula, R.; Rudd, P.M.; McCann, A.; et al. Hypoxia Alters Epigenetic and N-Glycosylation Profiles of Ovarian and Breast Cancer Cell Lines in-vitro. Front. Oncol. 2020, 10, 1218. [Google Scholar] [CrossRef]
- Greville, G.; Llop, E.; Howard, J.; Madden, S.F.; Perry, A.S.; Peracaula, R.; Rudd, P.M.; McCann, A.; Saldova, R. 5-AZA-dC induces epigenetic changes associated with modified glycosylation of secreted glycoproteins and increased EMT and migration in chemo-sensitive cancer cells. Clin. Epigenetics 2021, 13, 34. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Shao, J.; Wang, Y.; Shen, H.; Yu, S.; Zhang, J.; Yin, L. Hsa-miR-370 inhibited P-selectin-induced cell adhesion in human colon adenocarcinoma cells. Mol. Cell. Biochem. 2019, 450, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Pucci, M.; Gomes Ferreira, I.; Orlandani, M.; Malagolini, N.; Ferracin, M.; Dall’Olio, F. High Expression of the Sda Synthase B4GALNT2 Associates with Good Prognosis and Attenuates Stemness in Colon Cancer. Cells 2020, 9, 948. [Google Scholar] [CrossRef] [Green Version]
- Pucci, M.; Ferreira, I.G.; Malagolini, N.; Ferracin, M.; Dall’Olio, F. The Sda Synthase B4GALNT2 Reduces Malignancy and Stemness in Colon Cancer Cell Lines Independently of Sialyl Lewis X Inhibition. Int. J. Mol. Sci. 2020, 21, 6558. [Google Scholar] [CrossRef]
- Pucci, M.; Malagolini, N.; Dall’Olio, F. Glycosyltransferase B4GALNT2 as a Predictor of Good Prognosis in Colon Cancer: Lessons from Databases. Int. J. Mol. Sci. 2021, 22, 4331. [Google Scholar] [CrossRef]
- Yu, P.; Zhu, L.; Cui, K.; Du, Y.; Zhang, C.; Ma, W.; Guo, J. B4GALNT2 Gene Promotes Proliferation, and Invasiveness and Migration Abilities of Model Triple Negative Breast Cancer (TNBC) Cells by Interacting With HLA-B Protein. Front. Oncol. 2021, 11, 722828. [Google Scholar] [CrossRef]
- Dall’Olio, F.; Chiricolo, M. Sialyltransferases in cancer. Glycoconj. J. 2001, 18, 841–850. [Google Scholar] [CrossRef]
- Dall’Olio, F.; Malagolini, N.; Trinchera, M.; Chiricolo, M. Sialosignaling: Sialyltransferases as engines of self-fueling loops in cancer progression. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2014, 1840, 2752–2764. [Google Scholar] [CrossRef] [Green Version]
- Shan, M.; Yang, D.; Dou, H.; Zhang, L. Fucosylation in cancer biology and its clinical applications. Prog. Mol. Biol. Transl. Sci. 2019, 162, 93–119. [Google Scholar] [CrossRef] [PubMed]
- Granovsky, M.; Fata, J.; Pawling, J.; Muller, W.J.; Khokha, R.; Dennis, J.W. Suppression of tumor growth and metastasis in Mgat5-deficient mice. Nat. Med. 2000, 6, 306–312. [Google Scholar] [CrossRef] [PubMed]
- Bastian, K.; Scott, E.; Elliott, D.J.; Munkley, J. FUT8 α-1,6-Fucosyltransferase in Cancer. Int. J. Mol. Sci. 2021, 22, 455. [Google Scholar] [CrossRef] [PubMed]
- Pinho, S.; Marcos, N.T.; Ferreira, B.; Carvalho, A.S.; Oliveira, M.J.; Santos-Silva, F.; Harduin-Lepers, A.; Reis, C.A. Biological significance of cancer-associated sialyl-Tn antigen: Modulation of malignant phenotype in gastric carcinoma cells. Cancer Lett. 2007, 249, 157–170. [Google Scholar] [CrossRef]
- Julien, S.; Adriaenssens, E.; Ottenberg, K.; Furlan, A.; Courtand, G.; Vercoutter-Edouart, A.-S.; Hanisch, F.-G.; Delannoy, P.; Le Bourhis, X. ST6GalNAc I expression in MDA-MB-231 breast cancer cells greatly modifies their O-glycosylation pattern and enhances their tumourigenicity. Glycobiology 2006, 16, 54–64. [Google Scholar] [CrossRef] [Green Version]
- Cazet, A.; Lefebvre, J.; Adriaenssens, E.; Julien, S.; Bobowski, M.; Grigoriadis, A.; Tutt, A.; Tulasne, D.; Le Bourhis, X.; Delannoy, P. GD3 synthase expression enhances proliferation and tumor growth of MDA-MB-231 breast cancer cells through c-Met activation. Mol. Cancer Res. 2010, 8, 1526–1535. [Google Scholar] [CrossRef] [Green Version]
- Pinho, S.S.; Oliveira, P.; Cabral, J.; Carvalho, S.; Huntsman, D.; Gartner, F.; Seruca, R.; Reis, C.A.; Oliveira, C. Loss and recovery of Mgat3 and GnT-III Mediated E-cadherin N-glycosylation is a mechanism involved in epithelial-mesenchymal-epithelial transitions. PLoS ONE 2012, 7, e33191. [Google Scholar] [CrossRef] [Green Version]
- Pinho, S.S.; Reis, C.A.; Paredes, J.; Magalhaes, A.M.; Ferreira, A.C.; Figueiredo, J.; Xiaogang, W.; Carneiro, F.; Gartner, F.; Seruca, R. The role of N-acetylglucosaminyltransferase III and V in the post-transcriptional modifications of E-cadherin. Hum. Mol. Genet. 2009, 18, 2599–2608. [Google Scholar] [CrossRef] [Green Version]
- Kim, B.; Sun, R.; Oh, W.; Kim, A.M.J.; Schwarz, J.R.; Lim, S.O. Saccharide analog, 2-deoxy-d-glucose enhances 4-1BB-mediated antitumor immunity via PD-L1 deglycosylation. Mol. Carcinog. 2020, 59, 691–700. [Google Scholar] [CrossRef]
- Lee, H.H.; Wang, Y.N.; Xia, W.; Chen, C.H.; Rau, K.M.; Ye, L.; Wei, Y.; Chou, C.K.; Wang, S.C.; Yan, M.; et al. Removal of N-Linked Glycosylation Enhances PD-L1 Detection and Predicts Anti-PD-1/PD-L1 Therapeutic Efficacy. Cancer Cell 2019, 36, 168–178. [Google Scholar] [CrossRef]
- Repas, J.; Zupin, M.; Vodlan, M.; Veranic, P.; Gole, B.; Potocnik, U.; Pavlin, M. Dual Effect of Combined Metformin and 2-Deoxy-D-Glucose Treatment on Mitochondrial Biogenesis and PD-L1 Expression in Triple-Negative Breast Cancer Cells. Cancers 2022, 14, 1343. [Google Scholar] [CrossRef] [PubMed]
- Shao, B.; Li, C.W.; Lim, S.O.; Sun, L.; Lai, Y.J.; Hou, J.; Liu, C.; Chang, C.W.; Qiu, Y.; Hsu, J.M.; et al. Deglycosylation of PD-L1 by 2-deoxyglucose reverses PARP inhibitor-induced immunosuppression in triple-negative breast cancer. Am. J. Cancer Res. 2018, 8, 1837–1846. [Google Scholar]
- Sun, L.; Li, C.W.; Chung, E.M.; Yang, R.; Kim, Y.S.; Park, A.H.; Lai, Y.J.; Yang, Y.; Wang, Y.H.; Liu, J.; et al. Targeting Glycosylated PD-1 Induces Potent Antitumor Immunity. Cancer Res. 2020, 80, 2298–2310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.N.; Lee, H.H.; Hsu, J.L.; Yu, D.; Hung, M.C. The impact of PD-L1 N-linked glycosylation on cancer therapy and clinical diagnosis. J. Biomed. Sci. 2020, 27, 77. [Google Scholar] [CrossRef] [PubMed]
- Ashkani, J.; Naidoo, K.J. Glycosyltransferase Gene Expression Profiles Classify Cancer Types and Propose Prognostic Subtypes. Sci. Rep. 2016, 6, 26451. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhat, S.A.; Mir, M.U.R.; Majid, S.; Hassan, T.; Rehman, M.U.; Kuchy, S. Diagnostic utility of glycosyltransferase mRNA expression in gastric cancer. Hematol./Oncol. Stem Cell Ther. 2018, 11, 158–168. [Google Scholar] [CrossRef] [PubMed]
- Mohamed Abd-El-Halim, Y.; El, K.A.; Silvy, F.; Rubis, M.; Bigonnet, M.; Roques, J.; Cros, J.; Nicolle, R.; Iovanna, J.; Dusetti, N.; et al. A glycosyltransferase gene signature to detect pancreatic ductal adenocarcinoma patients with poor prognosis. EBioMedicine 2021, 71, 103541. [Google Scholar] [CrossRef]
- Noda, M.; Okayama, H.; Tachibana, K.; Sakamoto, W.; Saito, K.; Thar Min, A.K.; Ashizawa, M.; Nakajima, T.; Aoto, K.; Momma, T.; et al. Glycosyltransferase gene expression identifies a poor prognostic colorectal cancer subtype associated with mismatch repair deficiency and incomplete glycan synthesis. Clin. Cancer Res. 2018, 24, 4468–4481. [Google Scholar] [CrossRef] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).