Generation of Chloroplast Molecular Markers to Differentiate Sophora toromiro and Its Hybrids as a First Approach to Its Reintroduction in Rapa Nui (Easter Island)
Abstract
:1. Introduction
2. Results and Discussion
2.1. Chloroplast Database Mapping
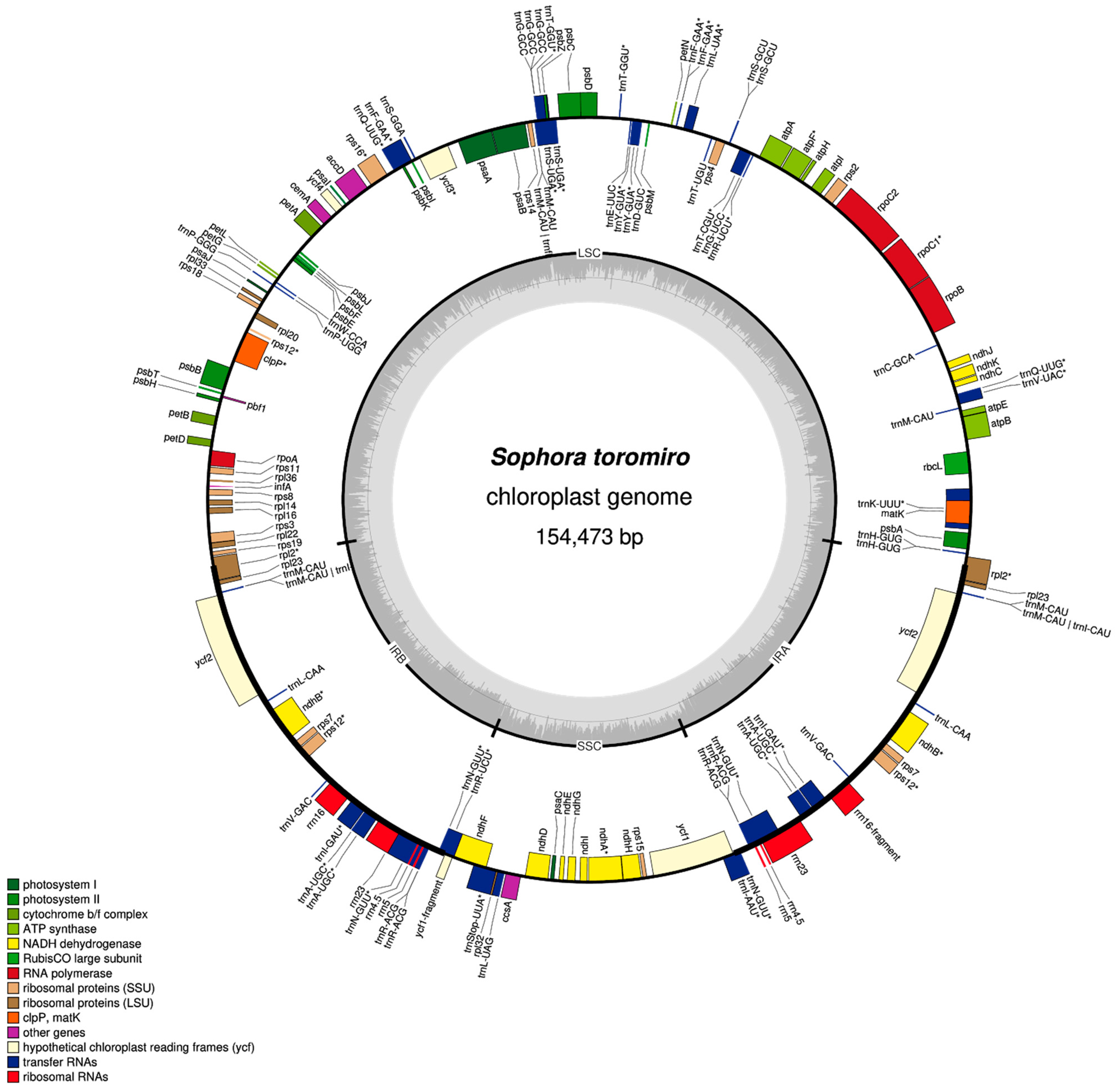
2.2. Chloroplast Assembly and Annotation
2.3. Phylogenomic Analysis
2.4. Chloroplast Genome Comparison
2.5. Phylogenetic Analysis
2.6. Simple Sequence Repeats (SSR) Analysis
2.7. Single Nucleotide Polymorphism (SNP) Analysis
3. Materials and Methods
3.1. Samples
3.2. Genomic DNA Extraction, Quantification and Quality Assessment
3.3. DNA Library Preparation and NGS
3.4. Chloroplast Assembly and Annotation
3.5. Phylogenomics, Phylogenetics and SSR Search
3.6. Single Nucleotide Polymorphism (SNP) Search and Characterization
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- World Conservation Monitoring Centre. Sophora toromiro. In The IUCN Red List of Threatened Species; IUCN: Gland, Switzerland, 1998. [Google Scholar] [CrossRef]
- Maunder, M. Conservation of the extinct Toromiro Tree, Sophora toromiro. Curtis’s Bot. Mag. 1997, 14, 226–231. [Google Scholar] [CrossRef]
- Palmer, J.L.; Linton Palmer, J. A Visit to Easter Island, or Rapa Nui, in 1868. J. R. Geogr. Soc. Lond. 1870, 40, 167. [Google Scholar] [CrossRef]
- Vargas, P.; Cristino, P.; Izaurieta, R. 1000 Años en Rapa nui: Arqueología del Asentamiento; Editorial Universitaria: Santiago, Chile, 2006. [Google Scholar]
- Toro, P. Isla de Pascua. Memoria del Ministerio del Culto y Colonización presentada al Congreso Nacional. 1892. Available online: https://obtienearchivo.bcn.cl/obtienearchivo?id=documentos/10221.1/40382/1/209432.pdf (accessed on 15 January 2021).
- Maunder, M.; Culham, A.; Alden, B.; Zizka, G.; Orliac, C.; Lobin, W.; Bordeu, A.; Ramirez, J.M.; Glissmann-Gough, S. Conservation of the Toromiro Tree: Case Study in the Management of a Plant Extinct in the Wild. Conserv. Biol. 2000, 14, 1341–1350. [Google Scholar] [CrossRef] [Green Version]
- Maunder, M.; Culham, A.; Bordeu, A.; Allainguillaume, J.; Wilkinson, M.J. Genetic diversity and pedigree for Sophora toromiro (Leguminosae): A tree extinct in the wild. Mol. Ecol. 1999, 8, 725–738. [Google Scholar] [CrossRef]
- Ricci, M.; Eaton, L. Do all existing Sophora toromiro descend from one individual? Biodivers. Conserv. 1997, 6, 1697–1702. [Google Scholar] [CrossRef]
- Heenan, P.B.; de Lange, P.J.; Wilton, A.D. Sophora (Fabaceae) in New Zealand: Taxonomy, distribution, and biogeography. N. Z. J. Bot. 2001, 39, 17–53. [Google Scholar] [CrossRef] [Green Version]
- Donoso, C. Un híbrido entre Sophora macrocarpa y Sophora microphylla. Boletın Técnico Facultad de Ciencias Forestales 1975, 30, 11–19. [Google Scholar]
- Donoso, C. Antecedentes de variación conocidos de otras latifoliadas de los bosques templados. In Variación Intraespecifica en Especies Arbóreas de los Bosques Templados de Chile y Argentina; Editorial Universitaria: Santiago, Chile, 2004; pp. 393–395. [Google Scholar]
- Püschel, T.A.; Espejo, J.; Sanzana, M.-J.; Benítez, H.A. Analysing the floral elements of the lost tree of Easter Island: A morphometric comparison between the remaining ex-situ lines of the endemic extinct species Sophora toromiro. PLoS ONE 2014, 9, e115548. [Google Scholar] [CrossRef] [Green Version]
- Peña, R.C.; Cassels, B.K. Phylogenetic relationships among Chilean Sophora species. Biochem. Syst. Ecol. 1996, 24, 725–733. [Google Scholar] [CrossRef]
- Stuessy, T.F.; Takayama, K.; López-Sepúlveda, P.; Crawford, D.J. Interpretation of patterns of genetic variation in endemic plant species of oceanic islands. Bot. J. Linn. Soc. 2014, 174, 276–288. [Google Scholar] [CrossRef] [Green Version]
- Shepherd, L.D.; Thiedemann, M.; Lehnebach, C. Genetic identification of historic Sophora (Fabaceae) specimens suggests toromiro (S. toromiro) from Rapa Nui/Easter Island may have been in cultivation in Europe in the 1700s. N. Z. J. Bot. 2020, 58, 255–267. [Google Scholar] [CrossRef]
- Hurr, K.A.; Lockhart, P.J.; Heenan, P.B.; Penny, D. Evidence for the recent dispersal of Sophora (Leguminosae) around the Southern Oceans: Molecular data. J. Biogeogr. 1999, 26, 565–577. [Google Scholar] [CrossRef]
- Shepherd, L.D.; Heenan, P.B. Evidence for both long-distance dispersal and isolation in the Southern Oceans: Molecular phylogeny of Sophora sect Edwardsia (Fabaceae). N. Z. J. Bot. 2017, 55, 334–346. [Google Scholar] [CrossRef]
- Mitchell, A.D.; Heenan, P.B. Sophora sect. Edwardsia (Fabaceae): Further evidence from nrDNA sequence data of a recent and rapid radiation around the Southern Oceans. Bot. J. Linn. Soc. 2002, 140, 435–441. [Google Scholar] [CrossRef] [Green Version]
- Shaw, J.; Lickey, E.B.; Schilling, E.E.; Small, R.L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: The tortoise and the hare III. Am. J. Bot. 2007, 94, 275–288. [Google Scholar] [CrossRef] [Green Version]
- Bell, K.L.; Loeffler, V.M.; Brosi, B.J. A reference library to aid in the identification of plant species mixtures by DNA metabarcoding. Appl. Plant Sci. 2017, 5. [Google Scholar] [CrossRef]
- Kress, W.J.; Erickson, D.L. A two-locus global DNA barcode for land plants: The coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE 2007, 2, e508. [Google Scholar] [CrossRef] [Green Version]
- Palmer, J.D.; Jansen, R.K.; Michaels, H.J.; Chase, M.W.; Manhart, J.R. Chloroplast DNA Variation and Plant Phylogeny. Ann. Mo. Bot. Gard. 1988, 75, 1180. [Google Scholar] [CrossRef]
- Roston, R.L.; Jouhet, J.; Yu, F.; Gao, H. Editorial: Structure and Function of Chloroplasts. Front. Plant Sci. 2018, 9, 1656. [Google Scholar] [CrossRef]
- Pogson, B.J.; Ganguly, D.; Albrecht-Borth, V. Insights into chloroplast biogenesis and development. Biochim. Biophys. Acta 2015, 1847, 1017–1024. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rogalski, M.; do Nascimento Vieira, L.; Fraga, H.P.; Guerra, M.P. Plastid genomics in horticultural species: Importance and applications for plant population genetics, evolution, and biotechnology. Front. Plant Sci. 2015, 6, 586. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.-B.; Tang, M.; Li, H.-T.; Zhang, Z.-R.; Li, D.-Z. Complete chloroplast genome of the genus Cymbidium: Lights into the species identification, phylogenetic implications and population genetic analyses. BMC Evol. Biol. 2013, 13, 84. [Google Scholar] [CrossRef] [Green Version]
- Zhang, T.; Fang, Y.; Wang, X.; Deng, X.; Zhang, X.; Hu, S.; Yu, J. The complete chloroplast and mitochondrial genome sequences of Boea hygrometrica: Insights into the evolution of plant organellar genomes. PLoS ONE 2012, 7, e30531. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, F.-H.; Chan, M.-T.; Liao, D.-C.; Hsu, C.-T.; Lee, Y.-W.; Daniell, H.; Duvall, M.R.; Lin, C.-S. Complete chloroplast genome of Oncidium Gower Ramsey and evaluation of molecular markers for identification and breeding in Oncidiinae. BMC Plant Biol. 2010, 10, 68. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Zhou, P.; Choi, Y.A.; Huang, S.; Gmitter, F.G., Jr. Mining and characterizing microsatellites from citrus ESTs. Appl. Genet. 2006, 112, 1248–1257. [Google Scholar] [CrossRef]
- Ellegren, H. Microsatellites: Simple sequences with complex evolution. Nat. Rev. Genet. 2004, 5, 435–445. [Google Scholar] [CrossRef]
- Mason, A.S. SSR genotyping. Methods Mol. Biol. 2015, 1245, 77–89. [Google Scholar] [CrossRef] [PubMed]
- Vieira, M.L.C.; Santini, L.; Diniz, A.L.; de Munhoz, C.F. Microsatellite markers: What they mean and why they are so useful. Genet. Mol. Biol. 2016, 39, 312–328. [Google Scholar] [CrossRef] [PubMed]
- Morgante, M.; Hanafey, M.; Powell, W. Microsatellites are preferentially associated with nonrepetitive DNA in plant genomes. Nat. Genet. 2002, 30, 194–200. [Google Scholar] [CrossRef]
- Mokhtar, M.M.; Atia, M.A.M. SSRome: An integrated database and pipelines for exploring microsatellites in all organisms. Nucleic Acids Res. 2019, 47, D244–D252. [Google Scholar] [CrossRef]
- Schroeder, H.; Fladung, M. Differentiation of Populus species by chloroplast SNP markers for barcoding and breeding approaches. iForest Biogeosci. For. 2015, 8, 544–546. [Google Scholar] [CrossRef] [Green Version]
- Singh, N.; Choudhury, D.R.; Singh, A.K.; Kumar, S.; Srinivasan, K.; Tyagi, R.K.; Singh, N.K.; Singh, R. Comparison of SSR and SNP markers in estimation of genetic diversity and population structure of Indian rice varieties. PLoS ONE 2013, 8, e84136. [Google Scholar] [CrossRef] [Green Version]
- Zuo, Y.; Chen, Z.; Kondo, K.; Funamoto, T.; Wen, J.; Zhou, S. DNA barcoding of Panax species. Planta Med. 2011, 77, 182–187. [Google Scholar] [CrossRef]
- Vignal, A.; Milan, D.; SanCristobal, M.; Eggen, A. A review on SNP and other types of molecular markers and their use in animal genetics. Genet. Sel. Evol. 2002, 34, 275–305. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, V.B.; Giang, V.N.L.; Waminal, N.E.; Park, H.-S.; Kim, N.-H.; Jang, W.; Lee, J.; Yang, T.-J. Comprehensive comparative analysis of chloroplast genomes from seven Panax species and development of an authentication system based on species-unique single nucleotide polymorphism markers. J. Ginseng Res. 2020, 44, 135–144. [Google Scholar] [CrossRef]
- Zhang, Y.; Iaffaldano, B.J.; Zhuang, X.; Cardina, J.; Cornish, K. Chloroplast genome resources and molecular markers differentiate rubber dandelion species from weedy relatives. BMC Plant Biol. 2017, 17, 34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Izan, S.; Esselink, D.; Visser, R.G.F.; Smulders, M.J.M.; Borm, T. Assembly of Complete Chloroplast Genomes from Non-model Species Based on a K-mer Frequency-Based Selection of Chloroplast Reads from Total DNA Sequences. Front. Plant Sci. 2017, 8, 1271. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Palmer, J.D. Plastid Chromosomes: Structure and Evolution. Mol. Biol. Plast. 1991, 7, 5–53. [Google Scholar]
- Jansen, R.K.; Raubeson, L.A.; Boore, J.L.; de Pamphilis, C.W.; Chumley, T.W.; Haberle, R.C.; Wyman, S.K.; Alverson, A.J.; Peery, R.; Herman, S.J.; et al. Methods for Obtaining and Analyzing Whole Chloroplast Genome Sequences. In Molecular Evolution: Producing the Biochemical Data; Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 2005; Volume 395, pp. 348–384. ISBN 9780121828004. [Google Scholar]
- Kikuchi, S.; Bédard, J.; Hirano, M.; Hirabayashi, Y.; Oishi, M.; Imai, M.; Takase, M.; Ide, T.; Nakai, M. Uncovering the protein translocon at the chloroplast inner envelope membrane. Science 2013, 339, 571–574. [Google Scholar] [CrossRef]
- Kikuchi, S.; Asakura, Y.; Imai, M.; Nakahira, Y.; Kotani, Y.; Hashiguchi, Y.; Nakai, Y.; Takafuji, K.; Bédard, J.; Hirabayashi-Ishioka, Y.; et al. A Ycf2-FtsHi Heteromeric AAA-ATPase Complex Is Required for Chloroplast Protein Import. Plant Cell 2018, 30, 2677–2703. [Google Scholar] [CrossRef] [Green Version]
- de Vries, J.; Sousa, F.L.; Bölter, B.; Soll, J.; Gould, S.B. YCF1: A Green TIC? Plant Cell 2015, 27, 1827–1833. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, W.; Li, L.; Li, G. Characterization of the complete chloroplast genome of shrubby sophora (Sophora flavescens Ait.). Mitochondrial DNA Part B 2018, 3, 1282–1283. [Google Scholar] [CrossRef] [Green Version]
- Shi, Y.; Liu, B. Complete chloroplast genome sequence of Sophora japonica “JinhuaiJ2” (Papilionaceae), an important traditional chinese herb. Mitochondrial DNA Part B 2020, 5, 319–320. [Google Scholar] [CrossRef] [Green Version]
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Chevreux, B.; Pfisterer, T.; Drescher, B.; Driesel, A.J.; Müller, W.E.G.; Wetter, T.; Suhai, S. Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res. 2004, 14, 1147–1159. [Google Scholar] [CrossRef] [Green Version]
- Tillich, M.; Lehwark, P.; Pellizzer, T.; Ulbricht-Jones, E.S.; Fischer, A.; Bock, R.; Greiner, S. GeSeq—Versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar] [CrossRef] [PubMed]
- Amiryousefi, A.; Hyvönen, J.; Poczai, P. IRscope: An online program to visualize the junction sites of chloroplast genomes. Bioinformatics 2018, 34, 3030–3031. [Google Scholar] [CrossRef] [PubMed]
- Greiner, S.; Lehwark, P.; Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 2019, 47, W59–W64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sievers, F.; Higgins, D.G. Clustal Omega, accurate alignment of very large numbers of sequences. Methods Mol. Biol. 2014, 1079, 105–116. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Peterson, D.; Tamura, K. MEGA-CC: Computing core of molecular evolutionary genetics analysis program for automated and iterative data analysis. Bioinformatics 2012, 28, 2685–2686. [Google Scholar] [CrossRef] [Green Version]
- Grant, J.R.; Arantes, A.S.; Stothard, P. Comparing thousands of circular genomes using the CGView Comparison Tool. BMC Genom. 2012, 13, 202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Page, A.J.; Taylor, B.; Delaney, A.J.; Soares, J.; Seemann, T.; Keane, J.A.; Harris, S.R. SNP-sites: Rapid efficient extraction of SNPs from multi-FASTA alignments. Microb. Genom. 2016, 2, e000056. [Google Scholar] [CrossRef] [Green Version]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef] [Green Version]





| Species | Raw Data Reads | Alignment Percentage | Aligned Reads | Average Coverage | GC Content | Size (bp) |
|---|---|---|---|---|---|---|
| ST1 | 48,875,452 | 4.28% | 2,093,045 | 3549.43 | 36.48% | 154,450 |
| ST2 | 65,161,387 | 5.26% | 3,424,884 | 5881.42 | 36.46% | 154,473 |
| ST-hyb1 | 47,916,209 | 5.39% | 2,581,423 | 4440.65 | 36.48% | 154,239 |
| ST-hyb2 | 42,444,815 | 7.67% | 3,253,301 | 5613.72 | 36.49% | 154,325 |
| SM | 46,355,472 | 4.97% | 2,301,647 | 3920.12 | 36.43% | 154,426 |
| Specie | rRNA | tRNA | CDS | Duplicated Genes | Total Genes | LSC (bp) | IR (bp) | SSC (bp) |
|---|---|---|---|---|---|---|---|---|
| ST1 | 4 | 32 | 78 | 17 | 131 | 85,769 | 24,773 | 19,135 |
| ST2 | 4 | 32 | 78 | 17 | 131 | 85,735 | 24,804 | 19,130 |
| ST-hyb1 | 4 | 32 | 78 | 17 | 131 | 85,600 | 24,774 | 19,091 |
| ST-hyb2 | 4 | 32 | 78 | 17 | 131 | 85,590 | 24,767 | 19,201 |
| SM | 4 | 32 | 78 | 17 | 131 | 85,690 | 24,800 | 19,136 |
| Gene Category | Gene Functional Group | Name |
|---|---|---|
| Photosynthesis related | ATP synthase | atpA, atpB, atpE, atpF ★, atpH, atpI |
| Cytochrome b/f complex | petA, petB, petD, petG, petL, petN | |
| Cytochrome c synthesis | ccsA | |
| NADPH dehydrogenase | ndhA ★, ndhB ★, ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, ndhK | |
| Photosystem I | psaA, psaB, psaC, psaI, psaJ | |
| Photosystem I stability | ycf3 ★, ycf4 | |
| Photosystem II | psbA, psbB (x2), psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbN (pbf1), psbT, psbZ | |
| Rubisco | rbcL | |
| Transcription and translation related | Ribosomal proteins | rps2, rps3, rps4, rps7 (x2), rps8, rps11, rps12 ★ (x2), rps14, rps15, rps16 ★, rps18, rps19, rpl2 ★ (x2), rpl14, rpl16, rpl20, rpl22, rpl23 (x2), rpl32, rpl33, rpl36 |
| Transcription | rpoA, rpoB, rpoC1 ★, rpoC2 | |
| RNA | Ribosomal RNA | rrn4.5 (x2), rrn5 (x2), rrn16 (x2), rrn23 (x2) |
| Transfer RNA | trnA-UGC ★ (x2), trnC-GCA, trnD-GUC, trnE-UUC ★, trnF-GAA, trnfM-CAU, trnG-GCC, trnG-UCC, trnH-GUG, trnI-CAU (x2), trnI-GAU (x2), trnK-UUU ★, trnL-CAA (x2), trnL-UAA ★, trnL-UAG, trnM-CAU, trnN-GUU (x2), trnP-GGG, trnP-UGG, trnQ-UUG, trnR-ACG (x2), trnR-UCU, trnS-GCU, trnS-GGA, trnS-UGA, trnT-CGU ★, trnT-GGU, trnT-UGU, trnV-GAC (x2), trnV-UAC ★, trnW-CCA, trnY-GUA | |
| Others | Carbon metabolism | cemA |
| Fatty acid synthesis | accD | |
| Proteolysis | clpP ★ | |
| RNA processing | matK | |
| Unknown function | Conserved reading frames | yfc1, ycf2 (x2) |
| Species | 1x | 2x | 3x | 4x | 5x | 6x | Total | CDS | IGS | Intron | IRa | IRb | SSC | LSC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ST1 | 100 | 11 | 11 | 7 | 1 | 1 | 131 | 21 | 89 | 21 | 6 | 5 | 24 | 96 |
| ST2 | 100 | 11 | 12 | 8 | 2 | 1 | 134 | 23 | 90 | 21 | 6 | 6 | 26 | 96 |
| ST-hyb1 | 101 | 10 | 11 | 8 | 1 | 1 | 132 | 21 | 92 | 19 | 6 | 5 | 25 | 96 |
| ST-hyb2 | 98 | 12 | 11 | 7 | 1 | 1 | 130 | 20 | 91 | 19 | 6 | 6 | 25 | 93 |
| SM | 113 | 10 | 11 | 7 | 1 | 1 | 143 | 22 | 103 | 18 | 6 | 6 | 29 | 102 |
| SNP Shared | ST1 | ST2 | ST-hyb1 | ST-hyb2 | SM |
|---|---|---|---|---|---|
| ST1 | - | 9 | 18 | 2 | 8 |
| ST2 | 9 | - | 8 | 2 | 4 |
| ST-hyb1 | 18 | 8 | - | 4 | 5 |
| ST-hyb2 | 2 | 2 | 4 | - | 29 |
| SM | 8 | 4 | 5 | 29 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pezoa, I.; Villacreses, J.; Rubilar, M.; Pizarro, C.; Galleguillos, M.J.; Ejsmentewicz, T.; Fonseca, B.; Espejo, J.; Polanco, V.; Sánchez, C. Generation of Chloroplast Molecular Markers to Differentiate Sophora toromiro and Its Hybrids as a First Approach to Its Reintroduction in Rapa Nui (Easter Island). Plants 2021, 10, 342. https://doi.org/10.3390/plants10020342
Pezoa I, Villacreses J, Rubilar M, Pizarro C, Galleguillos MJ, Ejsmentewicz T, Fonseca B, Espejo J, Polanco V, Sánchez C. Generation of Chloroplast Molecular Markers to Differentiate Sophora toromiro and Its Hybrids as a First Approach to Its Reintroduction in Rapa Nui (Easter Island). Plants. 2021; 10(2):342. https://doi.org/10.3390/plants10020342
Chicago/Turabian StylePezoa, Ignacio, Javier Villacreses, Miguel Rubilar, Carolina Pizarro, María Jesús Galleguillos, Troy Ejsmentewicz, Beatriz Fonseca, Jaime Espejo, Víctor Polanco, and Carolina Sánchez. 2021. "Generation of Chloroplast Molecular Markers to Differentiate Sophora toromiro and Its Hybrids as a First Approach to Its Reintroduction in Rapa Nui (Easter Island)" Plants 10, no. 2: 342. https://doi.org/10.3390/plants10020342







