Cytological and Transcriptomic Analysis Provide Insights into the Formation of Variegated Leaves in Ilex × altaclerensis ‘Belgica Aurea’
Abstract
:1. Introduction
2. Results
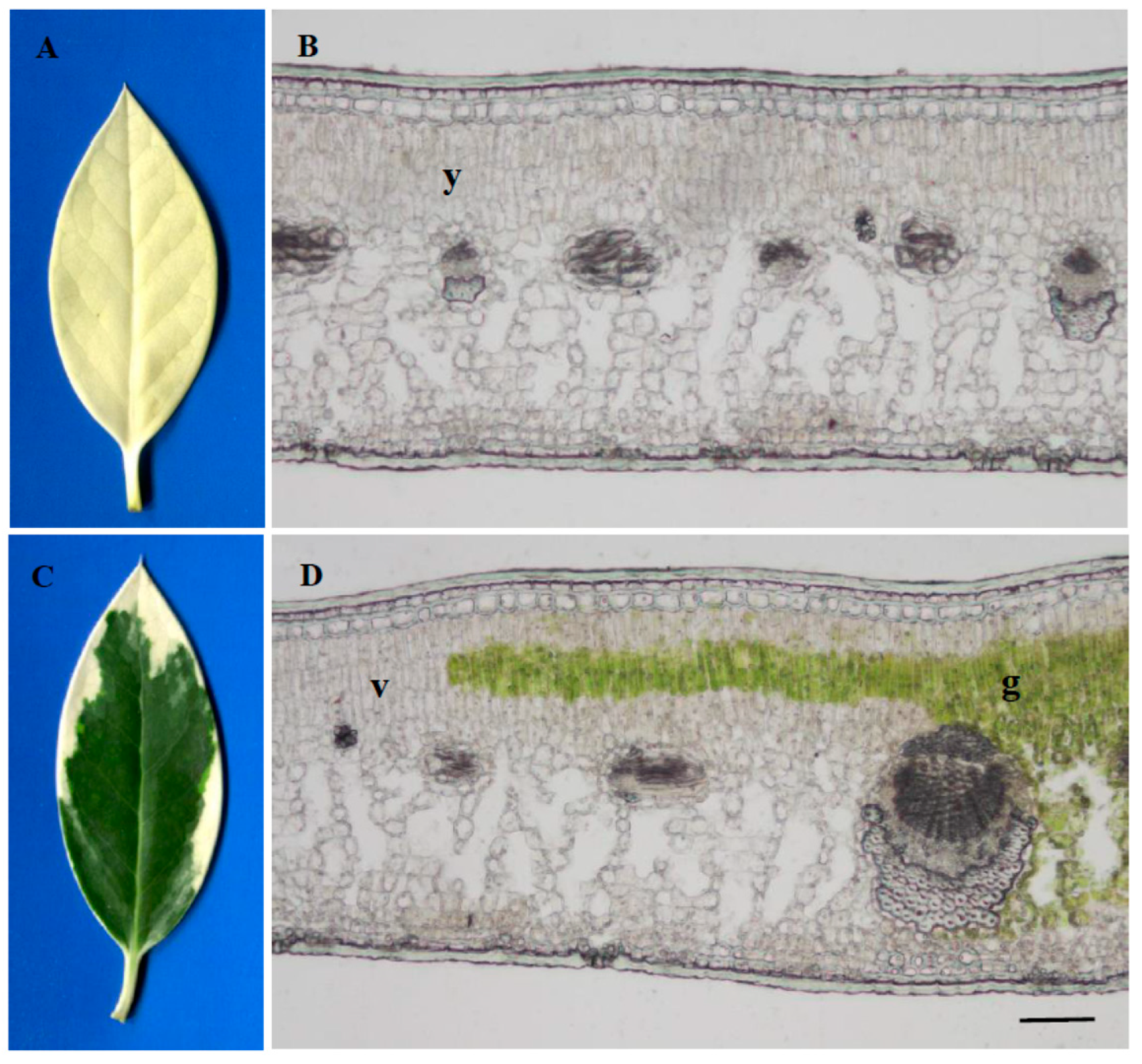
2.1. Leaf Anatomical Characteristics and Ultrastructure
2.2. Pigment Content in Yellow and Variegated Leaves
2.3. Localization of Pigments in the Leaves
2.4. RNA-Seq Analysis
2.5. Gene Annotation and Function Classification
2.6. Gene Expression Analysis
2.7. GO Enrichment Analysis of DEGs
2.8. KEGG Enrichment Analysis of DEGs
2.9. Analysis of the Genes Involved in Chlorophyll Metabolic Pathway and Chloroplast Integrity and Function
2.10. Validation of RNA Sequencing Data by qRT-PCR
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Measurements of Contents of Chlorophyll, Carotenoids and Anthocyanins
4.3. Transmission Electron Microscopic Observation
4.4. Leaf Histology
4.5. Laser Scanning Confocal Microscopy
4.6. RNA Extraction and cDNA Library Preparation
4.7. Transcriptome Assembly and Functional Annotation
4.8. Analysis of Differentially Expressed Genes (DEGs)
4.9. Quantitative Real-Time PCR
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, J.H.; Zeng, J.C.; Wang, X.M.; Chen, S.F.; Albach, D.C.; Li, H.Q. A revised classification of leaf variegation types. Flora 2020, 272, 151703. [Google Scholar] [CrossRef]
- Lev-Yadun, S.; Niemelä, P. Leaf pseudo-variegation: Definition, common types, and probably the defended models for real defensive leaf variegation mimicking them? Flora 2017, 22, 682–688. [Google Scholar] [CrossRef]
- Campitelli, B.E.; Stehlik, I.; Stinchcombe, J.R. Leaf variegation is associated with reduced herbivore damage in Hydrophyllum virginianum. Botany 2008, 86, 306–313. [Google Scholar] [CrossRef] [Green Version]
- Shelef, O.; Summerfield, L.; Lev-Yadun, S.; Villamarin-Cortez, S.; Sadeh, R.; Herrmann, I.; Rachmilevitch, S. Thermal benefits from white variegation of Silybum marianum leaves. Front. Plant Sci. 2019, 10, 688. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.L.; Liu, Z.H.; Song, H.N.; Chen, M.H.; Cheng, S.P. Protective role of leaf variegation in Pittosporum tobira under low temperature: Insights into the physio-biochemical and molecular mechanisms. Int. J. Mol. Sci. 2019, 20, 4857. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, M.H.; Li, X.; Zhang, X.X.; Zhang, H.; Zhao, X.Y. Mutation mechanism of leaf color in plants: A review. Forests 2020, 11, 851. [Google Scholar] [CrossRef]
- Yu, F.; Fu, A.G.; Aluru, M.; Park, S.; Xu, Y.; Liu, H.Y.; Liu, X.Y.; Foudree, A.; Nambogga, M.; Rodermel, S. Variegation mutants and mechanisms of chloroplast biogenesis. Plant Cell Environ. 2007, 30, 350–365. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, W.; Tamura, T.; Hanba-Tomita, Y.; Sodmergen Murata, M. The VAR1 locus of Arabidopsis encodes a chloroplastic FtsH and is responsible for leaf variegation in the mutant alleles. Genes Cells 2002, 7, 769–780. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.J.; Zhao, J.; Jia, M.; Xu, N.; Liang, S.; Shao, J.X.; Qi, Y.F.; Liu, X.Y.; An, L.J.; Yu, F. Balance between cytosolic and chloroplast translation affects leaf variegation. Plant Physiol. 2018, 176, 804–818. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Albrecht, V.; Ingenfeld, A.; Apel, K. Snowy cotyledon 2: The identification of a zinc finger domain protein essential for chloroplast development in cotyledons but not in true leaves. Plant Mol. Biol. 2008, 66, 599–608. [Google Scholar] [CrossRef] [Green Version]
- Zagari, N.; Sandoval-Ibañez, O.; Sandal, N.; Su, J.; Rodriguez-Concepcion, M.; Stougaard, J.; Pribil, M.; Leister, D.; Pulido, P. SNOWY COTYLEDON 2 promotes chloroplast development and has a role in leaf variegation in both Lotus japonicus and Arabidopsis thaliana. Mol. Plant 2017, 10, 721–734. [Google Scholar] [CrossRef] [Green Version]
- Azarin, K.; Usatov, A.; Makarenko, M.; Kozel, N.; Kovalevich, A.; Dremuk, I.; Yemelyanova, A.; Logacheva, M.; Fedorenko, A.; Averina, N. A point mutation in the photosystem I P700 chlorophyll a apoprotein A1 gene confers variegation in Helianthus annuus L. Plant Mol. Biol. 2020, 103, 373–389. [Google Scholar] [CrossRef]
- Gao, J.; Liang, D.; Xu, Q.Q.; Yang, F.X.; Zhu, G.F. Involvement of CsERF2 in leaf variegation of Cymbidium sinense ‘Dharma’. Planta 2020, 252, 29. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.B.; Hu, K.N.; Yan, M.J.; Yi, B.; Wen, J.; Ma, C.Z.; Shen, J.X.; Fu, T.D.; Tu, J.X. Disruption of carotene biosynthesis leads to abnormal plastids and variegated leaves in Brassica napus. Mol. Genet. Genom. 2020, 295, 981–999. [Google Scholar] [CrossRef] [PubMed]
- Cuénoud, P.; Martinez, M.A.D.P.; Loizeau, P.A.; Spichiger, R.; Andrews, S.; Manen, J.F. Molecular phylogeny and biogeography of the genus Ilex L. (Aquifoliaceae). Ann. Bot. 2000, 85, 111–122. [Google Scholar] [CrossRef] [Green Version]
- Yao, X.; Tan, Y.H.; Liu, Y.Y.; Song, Y.; Yang, J.B.; Corlett, R.T. Chloroplast genome structure in Ilex (Aquifoliaceae). Sci. Rep. 2016, 6, 28559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andrews, S. The clones of Ilex × Altaclerensis. Acta Hortic. 1986, 182, 377–380. [Google Scholar] [CrossRef]
- Tanaka, A.; Tanaka, R. Chlorophyll metabolism. Curr. Opin. Plant Biol. 2006, 9, 248–255. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, W. Leaf-variegated mutations and their responsible genes in Arabidopsis thaliana. Genes Genet. Syst. 2003, 78, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benning, C. A role for lipid trafficking in chloroplast biogenesis. Prog. Lipid Res. 2008, 47, 381–389. [Google Scholar] [CrossRef]
- Tsai, C.C.; Wu, Y.J.; Sheue, C.R.; Liao, P.C.; Chen, Y.H.; Li, S.J.; Liu, J.W.; Chang, H.T.; Liu, W.L.; Ko, Y.Z.; et al. Molecular basis underlying leaf variegation of a moth orchid mutant (Phalaenopsis aphrodite subsp. formosana). Front. Plant Sci. 2017, 8, 1333. [Google Scholar] [CrossRef] [Green Version]
- Li, W.X.; Yang, S.B.; Lu, Z.G.; He, Z.C.; Ye, Y.L.; Zhao, B.B.; Wang, L.; Jin, B. Cytological, physiological, and transcriptomic analyses of golden leaf coloration in Ginkgo biloba L. Hortic. Res. 2018, 5, 12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, X.Y.; Huang, L.B.; Chen, Q.S.; Lv, Y.Z.; Sun, H.N.; Liang, Z.H. Physiological and anatomical differences and differentially expressed genes reveal yellow leaf coloration in shumard oak. Plants 2020, 9, 169. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maekawa, S.; Takabayashi, A.; Reyes, T.H.; Yamamoto, H.; Tanaka, A.; Sato, T.; Yamaguchi, J. Pale-green phenotype of atl31 atl6 double mutant leaves is caused by disruption of 5-aminolevulinic acid biosynthesis in Arabidopsis thaliana. PLoS ONE 2015, 10, e0117662. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Kanakala, S.; He, Y.H.; Zhong, X.L.; Yu, S.M.; Li, R.X.; Sun, L.X.; Ma, J. Physiological characterization and comparative transcriptome analysis of white and green leaves of Ananas comosus var. bracteatus. PLoS ONE 2017, 12, e0169838. [Google Scholar] [CrossRef] [PubMed]
- Miura, E.; Kato, Y.; Sakamoto, W. Comparative transcriptome analysis of green/white variegated sectors in Arabidopsis yellow variegated2: Responses to oxidative and other stresses in white sectors. J. Exp. Bot. 2010, 61, 2433–2445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kong, W.Y.; Yu, X.W.; Chen, H.Y.; Liu, L.L.; Xiao, Y.J.; Wang, Y.L.; Wang, C.L.; Lin, Y.; Yu, Y.; Wang, C.M.; et al. The catalytic subunit of magnesium-protoporphyrin IX monomethyl ester cyclase forms a chloroplast complex to regulate chlorophyll biosynthesis in rice. Plant Mol. Biol. 2016, 92, 177–191. [Google Scholar] [CrossRef] [PubMed]
- Dörmann, P.; Benning, C. Galactolipids rule in seed plants. Trends Plant Sci. 2002, 7, 112–118. [Google Scholar] [CrossRef]
- Klaus, D.; Härtel, H.; Fitzpatrick, L.M.; Froehlich, J.E.; Hubert, J.; Benning, C.; Dörmann, P. Digalactosyldiacylglycerol synthesis in chloroplasts of the Arabidopsis dgd1 mutant. Plant Physiol. 2002, 128, 885–895. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, K.; Wada, H. Role of lipids in chloroplast biogenesis. In Lipids in Plant and Algae Development; Nakamura, Y., Li-Beisson, Y., Eds.; Springer International Publishing: Basel, Switzerland, 2016; pp. 103–125. [Google Scholar]
- Kobayashi, K.; Narise, T.; Sonoike, K.; Hashimoto, H.; Sato, N.; Kondo, M.; Nishimura, M.; Sato, M.; Toyooka, K.; Sugimoto, K.; et al. Role of galactolipid biosynthesis in coordinated development of photosynthetic complexes and thylakoid membranes during chloroplast biogenesis in Arabidopsis. Plant J. 2013, 73, 250–261. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, H.K. Chlorophylls and carotenoids: Pigments of photosynthetic biomembranes. Method. Enzymol. 1987, 148, 350–382. [Google Scholar]
- Hughes, N.M.; Morley, C.B.; Smith, W.K. Coordination of anthocyanin decline and photosynthetic maturation in juvenile leaves of three deciduous tree species. New Phytol. 2007, 175, 675–685. [Google Scholar] [CrossRef]
- Egea, I.; Bian, W.P.; Barsan, C.; Jauneau, A.; Pech, J.C.; Latché, A.; Li, Z.G.; Chervin, C. Chloroplast to chromoplast transition in tomato fruit: Spectral confocal microscopy analyses of carotenoids and chlorophylls in isolated plastids and time-lapse recording on intact live tissue. Ann. Bot. 2011, 108, 291–297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gomez, C.; Conejero, G.; Torregrosa, L.; Cheynier, V.; Terrier, N.; Ageorges, A. In vivo grapevine anthocyanin transport involves vesicle-mediated trafficking and the contribution of anthoMATE transporters and GST. Plant J. 2011, 67, 960–970. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.X.; Chen, Y.S.; Shi, C.M.; Huang, Z.B.; Zhang, Y.; Li, S.K.; Li, Y.; Ye, J.; Yu, C.; Li, Z.; et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience 2018, 7, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Feng, Z.; Wang, X.; Wang, X.W.; Zhang, X.G. DEGseq: An R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010, 26, 136–138. [Google Scholar] [CrossRef] [PubMed]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Alicia, O. Gene ontology analysis for RNA-Seq: Accounting for selection bias. Gen. Biol. 2010, 11, R14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mao, X.; Cai, T.; Olyarchuk, J.G.; Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 2005, 21, 3787–3793. [Google Scholar] [CrossRef] [PubMed]
- Koressaar, T.; Remm, M. Enhancements and modifications of primer design program Primer3. Bioinformatics 2007, 23, 1289–1291. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3- new capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed] [Green Version]









| Yellow | Green | Variegation | |
|---|---|---|---|
| Chl a (mg·g−1 FW) | 0.09 ± 0.01 b | 1.28 ± 0.09 a | 0.11 ± 0.01 b |
| Chl b (mg·g−1 FW) | 0.05 ± 0.01 c | 0.38 ± 0.01 a | 0.10 ± 0.01 b |
| Chl a+b (mg·g−1 FW) | 0.14 ± 0.01 b | 1.66 ± 0.10 a | 0.21 ± 0.01 b |
| Chl a/b | 1.63 ± 0.07 b | 3.33 ± 0.12 a | 1.14 ± 0.03 c |
| Car (mg·g−1 FW) | 0.05 ± 0.01 b | 0.46 ± 0.03 a | 0.06 ± 0.01 b |
| Chl a+b/Car | 2.91 ± 0.12 c | 3.65 ± 0.04 a | 3.31 ± 0.09 b |
| Ant (mg·g−1 FW) | 0.10 ± 0.01 b | 0.145 ± 0.01 a | 0.10 ± 0.01 b |
| Database | Number of Annotated Unigenes | Percentage of Annotated Unigenes |
|---|---|---|
| NR | 59,752 | 75.98% |
| NT | 52,105 | 66.26% |
| Swiss-Prot | 45,066 | 57.31% |
| KEGG | 47,967 | 61.00% |
| KOG | 47,369 | 60.24% |
| Pfam | 46,101 | 58.63% |
| GO | 34,793 | 44.25% |
| Intersection | 20,462 | 26.02% |
| Overall | 62,275 | 79.19% |
| Gene ID | Log2 Fold Change | Annotation | ||
|---|---|---|---|---|
| Variegation vs. Green | Yellow vs. Green | Yellow vs. Variegation | ||
| CL5332.Contig4_All | 3.20 | 2.16 | −1.04 | magnesium-protoporphyrin IX monomethyl ester [oxidative] cyclase |
| CL374.Contig9_All | 8.64 | 3.97 | −4.66 | chlorophyll(ide) b reductase |
| CL8272.Contig2_All | 7.34 | 2.73 | −4.61 | chlorophyll(ide) b reductase |
| CL8272.Contig3_All | −3.41 | −2.15 | 1.26 | chlorophyll(ide) b reductase |
| Unigene31820_All | 8.64 | 2.79 | −5.85 | monogalactosyldiacylglycerol synthase |
| CL3869.Contig6_All | 2.09 | 8.83 | 6.73 | digalactosyldiacylglycerol synthase |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Q.; Huang, J.; Zhou, P.; Hao, M.; Zhang, M. Cytological and Transcriptomic Analysis Provide Insights into the Formation of Variegated Leaves in Ilex × altaclerensis ‘Belgica Aurea’. Plants 2021, 10, 552. https://doi.org/10.3390/plants10030552
Zhang Q, Huang J, Zhou P, Hao M, Zhang M. Cytological and Transcriptomic Analysis Provide Insights into the Formation of Variegated Leaves in Ilex × altaclerensis ‘Belgica Aurea’. Plants. 2021; 10(3):552. https://doi.org/10.3390/plants10030552
Chicago/Turabian StyleZhang, Qiang, Jing Huang, Peng Zhou, Mingzhuo Hao, and Min Zhang. 2021. "Cytological and Transcriptomic Analysis Provide Insights into the Formation of Variegated Leaves in Ilex × altaclerensis ‘Belgica Aurea’" Plants 10, no. 3: 552. https://doi.org/10.3390/plants10030552







